Mitochondrial Ubiquitination as a Signaling Hub: Balancing Mitophagy, Inflammation, and Cell DeathClick to copy article linkArticle link copied!
- Ashwini KumarAshwini KumarJames Tarpo Jr. and Margaret Tarpo Department of Chemistry, Purdue University, West Lafayette, Indiana 47907, United StatesPurdue Institute for Drug Discovery, Purdue University, West Lafayette, Indiana 47907, United StatesPurdue Institute for Cancer Research, Purdue University, West Lafayette, Indiana 47907, United StatesMore by Ashwini Kumar
- Emmanouil Zacharioudakis*Emmanouil Zacharioudakis*Email: [email protected]James Tarpo Jr. and Margaret Tarpo Department of Chemistry, Purdue University, West Lafayette, Indiana 47907, United StatesPurdue Institute for Drug Discovery, Purdue University, West Lafayette, Indiana 47907, United StatesPurdue Institute for Cancer Research, Purdue University, West Lafayette, Indiana 47907, United StatesMore by Emmanouil Zacharioudakis
Abstract
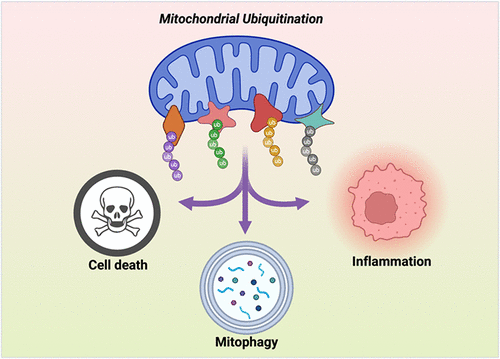
Mitochondria are increasingly recognized as signaling organelles that coordinate cell-fate decisions during stress. Because outer mitochondrial membrane (OMM) proteins are exposed to the cytosol, they are prominent substrates for ubiquitination, a dynamic post-translational modification that encodes information through diverse chain architectures and linkage types. In this review, we examine how ubiquitination of OMM proteins functions as a molecular switch that integrates mitochondrial stress signals and engages three major, often antagonistic, stress-response mechanisms: mitophagy, cell death, and innate immune signaling. We highlight an emerging concept that a stress-responsive “ubiquitin code” is written on OMM substrates, in which pathway selection is coordinated by the identity of ubiquitinated OMM proteins together with the linkage type and branching of attached polyubiquitin chains. We provide an updated overview of the E3 ubiquitin ligases and deubiquitinases (DUBs) that write and erase this code and summarize ubiquitin linkage types reported on key OMM substrates across these pathways. For mitophagy, we cover both PARKIN-dependent and PARKIN-independent mechanisms mediated by other E3 ligases and counteracted by DUBs. For innate immunity, we discuss how ubiquitination of OMM proteins regulates the MDA5/RIG-I-MAVS axis and NF-κB signaling. For cell death, we describe how ubiquitination of anti- and pro-apoptotic BCL-2 family proteins can either lower or increase the threshold for the induction of apoptosis. We also highlight the newfound role of PARKIN to drive apoptosis through a BAX/BAK-independent mechanism. Finally, we discuss therapeutic opportunities to reprogram OMM ubiquitination by targeting E3 ligases or DUBs directly, or by using PROTAC- and DUBTAC-based strategies.
Cited By
This article has not yet been cited by other publications.
Article Views
Altmetric
Citations
Article Views are the COUNTER-compliant sum of full text article downloads since November 2008 (both PDF and HTML) across all institutions and individuals. These metrics are regularly updated to reflect usage leading up to the last few days.
Citations are the number of other articles citing this article, calculated by Crossref and updated daily. Find more information about Crossref citation counts.
The Altmetric Attention Score is a quantitative measure of the attention that a research article has received online. Clicking on the donut icon will load a page at altmetric.com with additional details about the score and the social media presence for the given article. Find more information on the Altmetric Attention Score and how the score is calculated.


