DNA Nanostructures for siRNA DeliveryClick to copy article linkArticle link copied!
- Bharath Raj Madhanagopal*Bharath Raj Madhanagopal*Email: [email protected]Department of Nanoscale Science and Engineering, University at Albany, State University of New York, Albany, New York 12222, United StatesMore by Bharath Raj Madhanagopal
- Sarah YoussefSarah YoussefDepartment of Nanoscale Science and Engineering, University at Albany, State University of New York, Albany, New York 12222, United StatesMore by Sarah Youssef
- Arun Richard Chandrasekaran*Arun Richard Chandrasekaran*Email: [email protected]Department of Nanoscale Science and Engineering, University at Albany, State University of New York, Albany, New York 12222, United StatesThe RNA Institute, University at Albany, State University of New York, Albany, New York 12222, United StatesMore by Arun Richard Chandrasekaran
Abstract
Small interfering RNAs (siRNAs) represent an emerging class of versatile nucleic acid drugs for a broad spectrum of genetic and metabolic disorders. Since siRNAs can be developed to silence any target gene with relative ease compared to conventional drugs, there is enormous potential in this therapeutic modality for combating a variety of illnesses. However, its application is limited by low biostability, rapid clearance, and poor biodistribution of naked RNA. This is overcome by employing backbone modifications, conjugation of cell-targeting ligands, and the use of nanocarriers. DNA-based nanostructures are well suited to carry siRNA drugs since the use of DNA as a construction material provides the ability to tune the size, shape, and other morphological features of the nanostructure. DNA nanostructures also allow easy loading of multiple siRNA drugs with stoichiometric precision, enable functionalization with various targeting and tracking agents, and can be designed to deliver siRNA cargo in response to various stimuli. In this review, we provide an overview of recent reports on the use of DNA-based nanostructures to achieve targeted delivery of siRNA in vitro and in vivo. We discuss aspects of nanostructure design for various drug-loading and drug-release strategies and pharmacodynamic and pharmacokinetic properties of DNA nanocarriers and provide a survey of various diseases that have been targeted by siRNA-carrying DNA nanostructures. We also highlight the challenges facing these new-generation nanocarriers in achieving their therapeutic potential and clinical applications.
This publication is licensed under
License Summary*
You are free to share(copy and redistribute) this article in any medium or format and to adapt(remix, transform, and build upon) the material for any purpose, even commercially within the parameters below:
Creative Commons (CC): This is a Creative Commons license.
Attribution (BY): Credit must be given to the creator.
*Disclaimer
This summary highlights only some of the key features and terms of the actual license. It is not a license and has no legal value. Carefully review the actual license before using these materials.
License Summary*
You are free to share(copy and redistribute) this article in any medium or format and to adapt(remix, transform, and build upon) the material for any purpose, even commercially within the parameters below:
Creative Commons (CC): This is a Creative Commons license.
Attribution (BY): Credit must be given to the creator.
*Disclaimer
This summary highlights only some of the key features and terms of the actual license. It is not a license and has no legal value. Carefully review the actual license before using these materials.
License Summary*
You are free to share(copy and redistribute) this article in any medium or format and to adapt(remix, transform, and build upon) the material for any purpose, even commercially within the parameters below:
Creative Commons (CC): This is a Creative Commons license.
Attribution (BY): Credit must be given to the creator.
*Disclaimer
This summary highlights only some of the key features and terms of the actual license. It is not a license and has no legal value. Carefully review the actual license before using these materials.
Introduction
Figure 1
Figure 1. Overview of siRNA-mediated gene silencing by RNA interference (RNAi).
DNA Nanocarriers
Figure 2
Figure 2. DNA nanostructures. (a) DNA nanostructures assembled by cooperative assembly of short DNA single strands (e.g., a tetrahedron). (b) Modular assembly of DNA motifs into polyhedral structures (e.g., an icosahedron). (c) Hierarchical assembly of DNA motifs into 2D arrays. (d) Folding of a long single-stranded DNA scaffold into arbitrary shapes using the DNA origami strategy. (e) Single-stranded DNA built into larger structures using the DNA brick strategy.
Pharmacokinetics and Pharmacodynamics of DNA Nanostructures
Figure 3
Figure 3. Overview of drug delivery pathway using DNA nanostructures. DNA nanostructures can be functionalized to be (a) biostable against nuclease activity, (b) survive protein corona formation, and (c) used as imaging modalities. (d) DNA nanostructures with siRNA cargos can be targeted to specific cells for uptake, tracking, and disassembly or degradation in response to external stimuli to release the cargo.
Biostability
Protein Corona Formation
Cell Uptake
Endosomal Escape
Immunogenicity
Biodistribution
Targeted Delivery
Pharmacokinetic Techniques for Studying DNA Nanostructures
Loading of siRNA on DNA Nanocarriers and Release Strategies
Figure 4
Figure 4. Loading and release of siRNAs. (a) Attachment to single-stranded overhangs on DNA nanostructures. (b) Antisense strand is part of the DNA nanostructure that can bind to the sense strand of the siRNA. (c) siRNA linkers can connect DNA nanostructures to create a nanogel. (d) Release of siRNA by RNase H activity, (e) Dicer activity, and (f) reduction of disulfide linkages by glutathione.
DNA Nanocarriers for Therapeutic siRNA Delivery
| disease | model organism | nanocarrier | siRNA/small-molecule drug | response | administration | references |
|---|---|---|---|---|---|---|
| Breast cancer | MDA-MB-231 tumor-bearing mice | DNA nanogel | Anti-PLK1 | Reduction in Ki67-positive tumor cells | I.V. | (44) |
| Tumor-bearing humanized nude mice | DNA nanowire | Anti-PLK1 and doxorubicin | Inhibition of tumor growth | I.V. | (109) | |
| MCF-7R tumor-bearing mice | DNA origami | Anti-Bcl2, anti-P-gp and doxorubicin | Reduction of tumor growth | I.V. | (43) | |
| Non-small-cell lung cancer | Immunodeficient mouse model | DNA nanotube | Anti-KRASG12C and metformin | Reduction of tumor growth | I.V. | (52) |
| Glioblastoma | CB17 SCID mice | Spherical nucleic acids | Bcl2L12 | Impaired tumor growth | I.C. | (110) |
| Aging | Chemotherapy-induced senescent mice model | DNA tetrahedra | Anti-Raptor | Increase in the mean life span | I.P. | (48) |
| Acute lung injury | Acute lung injury mice model | DNA tetrahedra | Anti-mTOR and spermidine | Shift in the macrophage polarization | I.V. | (113) |
| Acute kidney injury | BALB/c mice | l-DNA tetrahedra | Anti-p53 | p53 knockdown | I.V. | (49) |
| C57BL/6 mice | DNA tetrahedra | I.V. | (114) | |||
| Hypercholesterolemia | BALB/c mice | DNA tetrahedra | Anti-ApoB1 | Reduction in the lipid levels in the blood | I.P. | (104) |
| Inflammation | C57BL/6 | DNA tetrahedra | Anti-TNFα | Reduction in anti-TNFα mRNA level | I.P. | (42) |
| Ulcerative colitis | C57BL/6 J | DNA nanotube | Anti-TNFα and anti-integrin α4 | reduced intestinal macrophage recruitment and T-cell homing | I.V. | (106) |
Abbreviations: PLK1, Polo-like kinase 1; Bcl2, B-cell lymphoma 2; P-gp, P-glycoprotein; KRAS, Kirsten rat sarcoma viral oncogene; Raptor, Regulatory associated protein of mTOR complex 1; mTOR, Mechanistic target of rapamycin; ApoB1, Apolipoprotein B; TNFα, Tumor necrosis factor alpha; I.V., intravenous; I.P., intraperitoneal, I.C., intracranial.
Cancer
Aging
Acute Lung Injury
Acute Kidney Injury
Hypercholesterolemia
Inflammation
Ulcerative Colitis
Swine Fever
Outlook
Acknowledgments
Research reported in this publication was supported by the National Institutes of Health (NIH) through National Institute of General Medical Sciences (NIGMS) under award number R35GM150672 to A.R.C. This manuscript is the result of funding in whole or in part by the National Institutes of Health (NIH). It is subject to the NIH Public Access Policy. Through acceptance of this federal funding, NIH has been given a right to make this manuscript publicly available in PubMed Central upon the Official Date of Publication, as defined by NIH.
References
This article references 132 other publications.
- 1Khvorova, A.; Watts, J. K. The Chemical Evolution of Oligonucleotide Therapies of Clinical Utility. Nat. Biotechnol. 2017, 35 (3), 238– 248, DOI: 10.1038/nbt.3765Google ScholarThere is no corresponding record for this reference.
- 2Egli, M.; Manoharan, M. Chemistry, Structure and Function of Approved Oligonucleotide Therapeutics. Nucleic Acids Res. 2023, 51 (6), 2529– 2573, DOI: 10.1093/nar/gkad067Google ScholarThere is no corresponding record for this reference.
- 3Paunovska, K.; Loughrey, D.; Dahlman, J. E. Drug Delivery Systems for RNA Therapeutics. Nat. Rev. Genet. 2022, 23 (5), 265– 280, DOI: 10.1038/s41576-021-00439-4Google ScholarThere is no corresponding record for this reference.
- 4Elbashir, S. M.; Harborth, J.; Lendeckel, W.; Yalcin, A.; Weber, K.; Tuschl, T. Duplexes of 21-Nucleotide RNAs Mediate RNA Interference in Cultured Mammalian Cells. Nature 2001, 411 (6836), 494– 498, DOI: 10.1038/35078107Google ScholarThere is no corresponding record for this reference.
- 5Jadhav, V.; Vaishnaw, A.; Fitzgerald, K.; Maier, M. A. RNA Interference in the Era of Nucleic Acid Therapeutics. Nat. Biotechnol. 2024, 42 (3), 394– 405, DOI: 10.1038/s41587-023-02105-yGoogle ScholarThere is no corresponding record for this reference.
- 6Xiao, B.; Wang, S.; Pan, Y.; Zhi, W.; Gu, C.; Guo, T.; Zhai, J.; Li, C.; Chen, Y. Q.; Wang, R. Development, Opportunities, and Challenges of siRNA Nucleic Acid Drugs. Mol. Ther. Nucleic Acids 2025, 36 (1), 102437 DOI: 10.1016/j.omtn.2024.102437Google ScholarThere is no corresponding record for this reference.
- 7Zamore, P. D.; Tuschl, T.; Sharp, P. A.; Bartel, D. P. RNAi: Double-Stranded RNA Directs the ATP-Dependent Cleavage of mRNA at 21 to 23 Nucleotide Intervals. Cell 2000, 101 (1), 25– 33, DOI: 10.1016/S0092-8674(00)80620-0Google ScholarThere is no corresponding record for this reference.
- 8Semizarov, D.; Frost, L.; Sarthy, A.; Kroeger, P.; Halbert, D. N.; Fesik, S. W. Specificity of Short Interfering RNA Determined through Gene Expression Signatures. Proc. Natl. Acad. Sci. U.S.A. 2003, 100 (11), 6347– 6352, DOI: 10.1073/pnas.1131959100Google ScholarThere is no corresponding record for this reference.
- 9Hammond, S. M.; Bernstein, E.; Beach, D.; Hannon, G. J. An RNA-Directed Nuclease Mediates Post-Transcriptional Gene Silencing in Drosophila Cells. Nature 2000, 404 (6775), 293– 296, DOI: 10.1038/35005107Google ScholarThere is no corresponding record for this reference.
- 10Wittrup, A.; Ai, A.; Liu, X.; Hamar, P.; Trifonova, R.; Charisse, K.; Manoharan, M.; Kirchhausen, T.; Lieberman, J. Visualizing Lipid-Formulated siRNA Release from Endosomes and Target Gene Knockdown. Nat. Biotechnol. 2015, 33 (8), 870– 876, DOI: 10.1038/nbt.3298Google ScholarThere is no corresponding record for this reference.
- 11Tabernero, J.; Shapiro, G. I.; LoRusso, P. M.; Cervantes, A.; Schwartz, G. K.; Weiss, G. J.; Paz-Ares, L.; Cho, D. C.; Infante, J. R.; Alsina, M.; Gounder, M. M.; Falzone, R.; Harrop, J.; White, A. C. S.; Toudjarska, I.; Bumcrot, D.; Meyers, R. E.; Hinkle, G.; Svrzikapa, N.; Hutabarat, R. M.; Clausen, V. A.; Cehelsky, J.; Nochur, S. V.; Gamba-Vitalo, C.; Vaishnaw, A. K.; Sah, D. W. Y.; Gollob, J. A.; Burris, H. A. III. First-in-Humans Trial of an RNA Interference Therapeutic Targeting VEGF and KSP in Cancer Patients with Liver Involvement. Cancer Discovery 2013, 3 (4), 406– 417, DOI: 10.1158/2159-8290.CD-12-0429Google ScholarThere is no corresponding record for this reference.
- 12Kato, H.; Takeuchi, O.; Mikamo-Satoh, E.; Hirai, R.; Kawai, T.; Matsushita, K.; Hiiragi, A.; Dermody, T. S.; Fujita, T.; Akira, S. Length-Dependent Recognition of Double-Stranded Ribonucleic Acids by Retinoic Acid–Inducible Gene-I and Melanoma Differentiation–Associated Gene 5. J. Exp. Med. 2008, 205 (7), 1601– 1610, DOI: 10.1084/jem.20080091Google ScholarThere is no corresponding record for this reference.
- 13Bereczki, Z.; Benczik, B.; Balogh, O. M.; Marton, S.; Puhl, E.; Pétervári, M.; Váczy-Földi, M.; Papp, Z. T.; Makkos, A.; Glass, K.; Locquet, F.; Euler, G.; Schulz, R.; Ferdinandy, P.; Ágg, B. Mitigating Off-Target Effects of Small RNAs: Conventional Approaches, Network Theory and Artificial Intelligence. Br. J. Pharmacol. 2025, 182 (2), 340– 379, DOI: 10.1111/bph.17302Google ScholarThere is no corresponding record for this reference.
- 14Janas, M. M.; Schlegel, M. K.; Harbison, C. E.; Yilmaz, V. O.; Jiang, Y.; Parmar, R.; Zlatev, I.; Castoreno, A.; Xu, H.; Shulga-Morskaya, S.; Rajeev, K. G.; Manoharan, M.; Keirstead, N. D.; Maier, M. A.; Jadhav, V. Selection of GalNAc-Conjugated siRNAs with Limited off-Target-Driven Rat Hepatotoxicity. Nat. Commun. 2018, 9, 723 DOI: 10.1038/s41467-018-02989-4Google ScholarThere is no corresponding record for this reference.
- 15Jackson, A. L.; Burchard, J.; Leake, D.; Reynolds, A.; Schelter, J.; Guo, J.; Johnson, J. M.; Lim, L.; Karpilow, J.; Nichols, K.; Marshall, W.; Khvorova, A.; Linsley, P. S. Position-Specific Chemical Modification of siRNAs Reduces “off-Target” Transcript Silencing. RNA 2006, 12 (7), 1197– 1205, DOI: 10.1261/rna.30706Google ScholarThere is no corresponding record for this reference.
- 16Corey, D. R.; Damha, M. J.; Manoharan, M. Challenges and Opportunities for Nucleic Acid Therapeutics. Nucleic Acid Ther. 2022, 32 (1), 8– 13, DOI: 10.1089/nat.2021.0085Google ScholarThere is no corresponding record for this reference.
- 17Couto, L. B.; High, K. A. Viral Vector-Mediated RNA Interference. Curr. Opin. Pharmacol. 2010, 10 (5), 534– 542, DOI: 10.1016/j.coph.2010.06.007Google ScholarThere is no corresponding record for this reference.
- 18Pérez-Martínez, F. C.; Guerra, J.; Posadas, I.; Ceña, V. Barriers to Non-Viral Vector-Mediated Gene Delivery in the Nervous System. Pharm. Res. 2011, 28 (8), 1843– 1858, DOI: 10.1007/s11095-010-0364-7Google ScholarThere is no corresponding record for this reference.
- 19Gregoriadis, G. Engineering Liposomes for Drug Delivery: Progress and Problems. Trends Biotechnol. 1995, 13 (12), 527– 537, DOI: 10.1016/S0167-7799(00)89017-4Google ScholarThere is no corresponding record for this reference.
- 20Wijagkanalan, W.; Kawakami, S.; Hashida, M. Designing Dendrimers for Drug Delivery and Imaging: Pharmacokinetic Considerations. Pharm. Res. 2011, 28 (7), 1500– 1519, DOI: 10.1007/s11095-010-0339-8Google ScholarThere is no corresponding record for this reference.
- 21Pack, D. W.; Hoffman, A. S.; Pun, S.; Stayton, P. S. Design and Development of Polymers for Gene Delivery. Nat. Rev. Drug Discovery 2005, 4 (7), 581– 593, DOI: 10.1038/nrd1775Google ScholarThere is no corresponding record for this reference.
- 22Duncan, B.; Kim, C.; Rotello, V. M. Gold Nanoparticle Platforms as Drug and Biomacromolecule Delivery Systems. J. Controlled Release 2010, 148 (1), 122– 127, DOI: 10.1016/j.jconrel.2010.06.004Google ScholarThere is no corresponding record for this reference.
- 23Kakkar, A.; Traverso, G.; Farokhzad, O. C.; Weissleder, R.; Langer, R. Evolution of Macromolecular Complexity in Drug Delivery Systems. Nat. Rev. Chem. 2017, 1 (8), 0063 DOI: 10.1038/s41570-017-0063Google ScholarThere is no corresponding record for this reference.
- 24Zhang, S.; Zhao, Y.; Zhi, D.; Zhang, S. Non-Viral Vectors for the Mediation of RNAi. Bioorg. Chem. 2012, 40, 10– 18, DOI: 10.1016/j.bioorg.2011.07.005Google ScholarThere is no corresponding record for this reference.
- 25Seow, Y.; Wood, M. J. Biological Gene Delivery Vehicles: Beyond Viral Vectors. Mol. Ther. 2009, 17 (5), 767– 777, DOI: 10.1038/mt.2009.41Google ScholarThere is no corresponding record for this reference.
- 26Madhanagopal, B. R.; Zhang, S.; Demirel, E.; Wady, H.; Chandrasekaran, A. R. DNA Nanocarriers: Programmed to Deliver. Trends Biochem. Sci. 2018, 43 (12), 997– 1013, DOI: 10.1016/j.tibs.2018.09.010Google ScholarThere is no corresponding record for this reference.
- 27Xavier, P. L.; Chandrasekaran, A. R. DNA-Based Construction at the Nanoscale: Emerging Trends and Applications. Nanotechnology 2018, 29 (6), 062001 DOI: 10.1088/1361-6528/aaa120Google ScholarThere is no corresponding record for this reference.
- 28Chandrasekaran, A. R.; Levchenko, O. DNA Nanocages. Chem. Mater. 2016, 28 (16), 5569– 5581, DOI: 10.1021/acs.chemmater.6b02546Google ScholarThere is no corresponding record for this reference.
- 29Goodman, R. P.; Schaap, I. A. T.; Tardin, C. F.; Erben, C. M.; Berry, R. M.; Schmidt, C. F.; Turberfield, A. J. Rapid Chiral Assembly of Rigid DNA Building Blocks for Molecular Nanofabrication. Science 2005, 310 (5754), 1661– 1665, DOI: 10.1126/science.1120367Google ScholarThere is no corresponding record for this reference.
- 30Bhatia, D.; Mehtab, S.; Krishnan, R.; Indi, S. S.; Basu, A.; Krishnan, Y. Icosahedral DNA Nanocapsules by Modular Assembly. Angew. Chem., Int. Ed. 2009, 48 (23), 4134– 4137, DOI: 10.1002/anie.200806000Google ScholarThere is no corresponding record for this reference.
- 31Winfree, E.; Liu, F.; Wenzler, L. A.; Seeman, N. C. Design and Self-Assembly of Two-Dimensional DNA Crystals. Nature 1998, 394 (6693), 539– 544, DOI: 10.1038/28998Google ScholarThere is no corresponding record for this reference.
- 32Rothemund, P. W. K. Folding DNA to Create Nanoscale Shapes and Patterns. Nature 2006, 440 (7082), 297– 302, DOI: 10.1038/nature04586Google ScholarThere is no corresponding record for this reference.
- 33Ke, Y.; Ong, L. L.; Shih, W. M.; Yin, P. Three-Dimensional Structures Self-Assembled from DNA Bricks. Science 2012, 338 (6111), 1177– 1183, DOI: 10.1126/science.1227268Google ScholarThere is no corresponding record for this reference.
- 34Madsen, M.; Gothelf, K. V. Chemistries for DNA Nanotechnology. Chem. Rev. 2019, 119 (10), 6384– 6458, DOI: 10.1021/acs.chemrev.8b00570Google ScholarThere is no corresponding record for this reference.
- 35Pinheiro, V. B.; Holliger, P. Towards XNA Nanotechnology: New Materials from Synthetic Genetic Polymers. Trends Biotechnol. 2014, 32 (6), 321– 328, DOI: 10.1016/j.tibtech.2014.03.010Google ScholarThere is no corresponding record for this reference.
- 36Goodman, R. P.; Berry, R. M.; Turberfield, A. J. The Single-Step Synthesis of a DNA Tetrahedron. Chem. Commun. 2004, (No. 12), 1372– 1373, DOI: 10.1039/B402293AGoogle ScholarThere is no corresponding record for this reference.
- 37Huang, J.; Chakraborty, A.; Tadepalli, L. S.; Paul, A. Adoption of a Tetrahedral DNA Nanostructure as a Multifunctional Biomaterial for Drug Delivery. ACS Pharmacol. Transl. Sci. 2024, 7 (8), 2204– 2214, DOI: 10.1021/acsptsci.4c00308Google ScholarThere is no corresponding record for this reference.
- 38Walsh, A. S.; Yin, H.; Erben, C. M.; Wood, M. J. A.; Turberfield, A. J. DNA Cage Delivery to Mammalian Cells. ACS Nano 2011, 5 (7), 5427– 5432, DOI: 10.1021/nn2005574Google ScholarThere is no corresponding record for this reference.
- 39Vilcapoma, J.; Patel, A.; Chandrasekaran, A. R.; Halvorsen, K. The Role of Size in Biostability of DNA Tetrahedra. Chem. Commun. 2023, 59 (34), 5083– 5085, DOI: 10.1039/D3CC01123BGoogle ScholarThere is no corresponding record for this reference.
- 40Rajwar, A.; Shetty, S. R.; Vaswani, P.; Morya, V.; Barai, A.; Sen, S.; Sonawane, M.; Bhatia, D. Geometry of a DNA Nanostructure Influences Its Endocytosis: Cellular Study on 2D, 3D, and in Vivo Systems. ACS Nano 2022, 16 (7), 10496– 10508, DOI: 10.1021/acsnano.2c01382Google ScholarThere is no corresponding record for this reference.
- 41Xia, K.; Kong, H.; Cui, Y.; Ren, N.; Li, Q.; Ma, J.; Cui, R.; Zhang, Y.; Shi, J.; Li, Q.; Lv, M.; Sun, Y.; Wang, L.; Li, J.; Zhu, Y. Systematic Study in Mammalian Cells Showing No Adverse Response to Tetrahedral DNA Nanostructure. ACS Appl. Mater. Interfaces 2018, 10 (18), 15442– 15448, DOI: 10.1021/acsami.8b02626Google ScholarThere is no corresponding record for this reference.
- 42Gao, Y.; Chen, X.; Tian, T.; Zhang, T.; Gao, S.; Zhang, X.; Yao, Y.; Lin, Y.; Cai, X. A Lysosome-Activated Tetrahedral Nanobox for Encapsulated siRNA Delivery. Adv. Mater. 2022, 34 (46), 2201731 DOI: 10.1002/adma.202201731Google ScholarThere is no corresponding record for this reference.
- 43Wang, Z.; Song, L.; Liu, Q.; Tian, R.; Shang, Y.; Liu, F.; Liu, S.; Zhao, S.; Han, Z.; Sun, J.; Jiang, Q.; Ding, B. A Tubular DNA Nanodevice as a siRNA/Chemo-Drug Co-delivery Vehicle for Combined Cancer Therapy. Angew. Chem. Int. Ed. 2021, 60 (5), 2594– 2598, DOI: 10.1002/anie.202009842Google ScholarThere is no corresponding record for this reference.
- 44Ding, F.; Mou, Q.; Ma, Y.; Pan, G.; Guo, Y.; Tong, G.; Choi, C. H. J.; Zhu, X.; Zhang, C. A Crosslinked Nucleic Acid Nanogel for Effective siRNA Delivery and Antitumor Therapy. Angew. Chem., Int. Ed. 2018, 57 (12), 3064– 3068, DOI: 10.1002/anie.201711242Google ScholarThere is no corresponding record for this reference.
- 45Frtús, A.; Smolková, B.; Uzhytchak, M.; Lunova, M.; Jirsa, M.; Henry, S. J. W.; Dejneka, A.; Stephanopoulos, N.; Lunov, O. The Interactions between DNA Nanostructures and Cells: A Critical Overview from a Cell Biology Perspective. Acta Biomater. 2022, 146, 10– 22, DOI: 10.1016/j.actbio.2022.04.046Google ScholarThere is no corresponding record for this reference.
- 46Chandrasekaran, A. R. Nuclease Resistance of DNA Nanostructures. Nat. Rev. Chem. 2021, 5 (4), 225– 239, DOI: 10.1038/s41570-021-00251-yGoogle ScholarThere is no corresponding record for this reference.
- 47Keum, J.-W.; Bermudez, H. Enhanced Resistance of DNA Nanostructures to Enzymatic Digestion. Chem. Commun. 2009, 2009, 7036– 7038, DOI: 10.1039/b917661fGoogle ScholarThere is no corresponding record for this reference.
- 48Yue, Z.; Yang, Y.; Nie, L.; Sun, Y.; Wang, Q.; Lin, Y.; Gao, Y.; Cai, X. A Binary siRNA-Loaded Tetrahedral DNA Nanobox for Synergetic Anti-Aging Therapy. Small 2025, 21 (18), 2408323 DOI: 10.1002/smll.202408323Google ScholarThere is no corresponding record for this reference.
- 49Thai, H. B. D.; Kim, K.-R.; Hong, K. T.; Voitsitskyi, T.; Lee, J.-S.; Mao, C.; Ahn, D.-R. Kidney-Targeted Cytosolic Delivery of siRNA Using a Small-Sized Mirror DNA Tetrahedron for Enhanced Potency. ACS Cent. Sci. 2020, 6 (12), 2250– 2258, DOI: 10.1021/acscentsci.0c00763Google ScholarThere is no corresponding record for this reference.
- 50Skaanning, M. K.; Bønnelykke, J.; Nijenhuis, M. A. D.; Samanta, A.; Smidt, J. M.; Gothelf, K. V. Self-Assembly of Ultrasmall 3D Architectures of (l)-Acyclic Threoninol Nucleic Acids with High Thermal and Serum Stability. J. Am. Chem. Soc. 2024, 146 (29), 20141– 20146, DOI: 10.1021/jacs.4c04919Google ScholarThere is no corresponding record for this reference.
- 51Madhanagopal, B. R.; Patel, A.; Talbot, H.; Geary, J. A.; Ganesh, K. N.; Chandrasekaran, A. R. Peptide Nucleic Acid (PNA) and DNA Hybrid Three-Way Junctions and Mesojunctions bioRxiv 2025 DOI: 10.1101/2025.05.15.653195 .Google ScholarThere is no corresponding record for this reference.
- 52Qian, H.; Wang, D.; He, B.; Liu, Q.; Xu, Y.; Wu, D.; Chen, C.; Zhang, W.; Leong, D. T.; Wang, G. Assembling Defined DNA Nanostructures with Anticancer Drugs: A Metformin/DNA Complex Nanoplatform with a Synergistic Antitumor Effect for KRAS-Mutated Lung Cancer Therapy. NPG Asia Mater. 2022, 14 (1), 81 DOI: 10.1038/s41427-022-00427-yGoogle ScholarThere is no corresponding record for this reference.
- 53Wamhoff, E.-C.; Romanov, A.; Huang, H.; Read, B. J.; Ginsburg, E.; Knappe, G. A.; Kim, H. M.; Farrell, N. P.; Irvine, D. J.; Bathe, M. Controlling Nuclease Degradation of Wireframe DNA Origami with Minor Groove Binders. ACS Nano 2022, 16 (6), 8954– 8966, DOI: 10.1021/acsnano.1c11575Google ScholarThere is no corresponding record for this reference.
- 54Rodriguez, A.; Gandavadi, D.; Mathivanan, J.; Song, T.; Madhanagopal, B. R.; Talbot, H.; Sheng, J.; Wang, X.; Chandrasekaran, A. R. Self-Assembly of DNA Nanostructures in Different Cations. Small 2023, 19 (39), 2300040 DOI: 10.1002/smll.202300040Google ScholarThere is no corresponding record for this reference.
- 55Gandavadi, D.; Talbot, H.; Dwivedy, A.; Umrao, S.; Rodriguez, A.; Cho, H.; Zheng, M.; Chandrasekaran, A. R.; Wang, X. DNA Nanostructure Self-Assembly in an Aqueous Ionic Liquid Solution with Enhanced Stability and Target Binding Affinity. J. Am. Chem. Soc. 2025, 147 (46), 42635– 42646, DOI: 10.1021/jacs.5c13969Google ScholarThere is no corresponding record for this reference.
- 56Chandrasekaran, A. R.; Vilcapoma, J.; Dey, P.; Wong-Deyrup, S. W.; Dey, B. K.; Halvorsen, K. Exceptional Nuclease Resistance of Paranemic Crossover (PX) DNA and Crossover-Dependent Biostability of DNA Motifs. J. Am. Chem. Soc. 2020, 142 (14), 6814– 6821, DOI: 10.1021/jacs.0c02211Google ScholarThere is no corresponding record for this reference.
- 57Madhanagopal, B. R.; Talbot, H.; Rodriguez, A.; Louis, J. M.; Zeghal, H.; Vangaveti, S.; Reddy, K.; Chandrasekaran, A. R. The Unusual Structural Properties and Potential Biological Relevance of Switchback DNA. Nat. Commun. 2024, 15 (1), 6636 DOI: 10.1038/s41467-024-50348-3Google ScholarThere is no corresponding record for this reference.
- 58Perrault, S. D.; Shih, W. M. Virus-Inspired Membrane Encapsulation of DNA Nanostructures To Achieve In Vivo Stability. ACS Nano 2014, 8 (5), 5132– 5140, DOI: 10.1021/nn5011914Google ScholarThere is no corresponding record for this reference.
- 59Auvinen, H.; Zhang, H.; Nonappa; Kopilow, A.; Niemelä, E. H.; Nummelin, S.; Correia, A.; Santos, H. A.; Linko, V.; Kostiainen, M. A. Protein Coating of DNA Nanostructures for Enhanced Stability and Immunocompatibility. Adv. Healthcare Mater. 2017, 6 (18), 1700692 DOI: 10.1002/adhm.201700692Google ScholarThere is no corresponding record for this reference.
- 60Seitz, I.; McNeale, D.; Sainsbury, F.; Linko, V.; Kostiainen, M. A. Modular Virus Capsid Coatings for Biocatalytic DNA Origami Nanoreactors. ACS Nano 2025, 19 (41), 36465– 36477, DOI: 10.1021/acsnano.5c10734Google ScholarThere is no corresponding record for this reference.
- 61Ponnuswamy, N.; Bastings, M. M. C.; Nathwani, B.; Ryu, J. H.; Chou, L. Y. T.; Vinther, M.; Li, W. A.; Anastassacos, F. M.; Mooney, D. J.; Shih, W. M. Oligolysine-Based Coating Protects DNA Nanostructures from Low-Salt Denaturation and Nuclease Degradation. Nat. Commun. 2017, 8 (1), 15654 DOI: 10.1038/ncomms15654Google ScholarThere is no corresponding record for this reference.
- 62Agarwal, N. P.; Matthies, M.; Gür, F. N.; Osada, K.; Schmidt, T. L. Block Copolymer Micellization as a Protection Strategy for DNA Origami. Angew. Chem., Int. Ed. 2017, 56 (20), 5460– 5464, DOI: 10.1002/anie.201608873Google ScholarThere is no corresponding record for this reference.
- 63Youssef, S.; Tsang, E.; Samanta, A.; Kumar, V.; Gothelf, K. V. Reversible Protection and Targeted Delivery of DNA Origami with a Disulfide-Containing Cationic Polymer. Small 2024, 20 (10), 2301058 DOI: 10.1002/smll.202301058Google ScholarThere is no corresponding record for this reference.
- 64Schöttler, S.; Landfester, K.; Mailänder, V. Controlling the Stealth Effect of Nanocarriers through Understanding the Protein Corona. Angew. Chem., Int. Ed. 2016, 55 (31), 8806– 8815, DOI: 10.1002/anie.201602233Google ScholarThere is no corresponding record for this reference.
- 65Huzar, J.; Coreas, R.; Landry, M. P.; Tikhomirov, G. AI-Based Prediction of Protein Corona Composition on DNA Nanostructures. ACS Nano 2025, 19 (4), 4333– 4345, DOI: 10.1021/acsnano.4c12259Google ScholarThere is no corresponding record for this reference.
- 66Kim, K.-R.; Kim, J.; Back, J. H.; Lee, J. E.; Ahn, D.-R. Cholesterol-Mediated Seeding of Protein Corona on DNA Nanostructures for Targeted Delivery of Oligonucleotide Therapeutics to Treat Liver Fibrosis. ACS Nano 2022, 16 (5), 7331– 7343, DOI: 10.1021/acsnano.1c08508Google ScholarThere is no corresponding record for this reference.
- 67Rodríguez-Franco, H. J.; Hendrickx, P. B. M.; Bastings, M. M. C. Tailoring DNA Origami Protection: A Study of Oligolysine-PEG Coatings for Improved Colloidal, Structural, and Functional Integrity. ACS Polym. Au 2025, 5 (1), 35– 44, DOI: 10.1021/acspolymersau.4c00085Google ScholarThere is no corresponding record for this reference.
- 68Rodríguez-Franco, H. J.; Weiden, J.; Bastings, M. M. C. Stabilizing Polymer Coatings Alter the Protein Corona of DNA Origami and Can Be Engineered to Bias the Cellular Uptake. ACS Polym. Au 2023, 3 (4), 344– 353, DOI: 10.1021/acspolymersau.3c00009Google ScholarThere is no corresponding record for this reference.
- 69Tang, X.; Zhai, T.; Li, T.; Jin, Y.; Lei, D.; Zhu, C.; Qu, L.; Li, Y.; Wang, Y.; Gu, H.; Fang, B. Dimensional Control of DNA Nanostructures Enhances Cellular Uptake and Guides Tissue-Regenerative Responses. J. Nanobiotechnol. 2025, 23 (1), 615 DOI: 10.1186/s12951-025-03707-1Google ScholarThere is no corresponding record for this reference.
- 70Wang, P.; Rahman, M. A.; Zhao, Z.; Weiss, K.; Zhang, C.; Chen, Z.; Hurwitz, S. J.; Chen, Z. G.; Shin, D. M.; Ke, Y. Visualization of the Cellular Uptake and Trafficking of DNA Origami Nanostructures in Cancer Cells. J. Am. Chem. Soc. 2018, 140 (7), 2478– 2484, DOI: 10.1021/jacs.7b09024Google ScholarThere is no corresponding record for this reference.
- 71Gopinath, S. C. B.; Lakshmipriya, T.; Chen, Y.; Arshad, M. K. M.; Kerishnan, J. P.; Ruslinda, A. R.; Al-Douri, Y.; Voon, C. H.; Hashim, U. Cell-Targeting Aptamers Act as Intracellular Delivery Vehicles. Appl. Microbiol. Biotechnol. 2016, 100 (16), 6955– 6969, DOI: 10.1007/s00253-016-7686-2Google ScholarThere is no corresponding record for this reference.
- 72Xia, Z.; Wang, P.; Liu, X.; Liu, T.; Yan, Y.; Yan, J.; Zhong, J.; Sun, G.; He, D. Tumor-Penetrating Peptide-Modified DNA Tetrahedron for Targeting Drug Delivery. Biochemistry 2016, 55 (9), 1326– 1331, DOI: 10.1021/acs.biochem.5b01181Google ScholarThere is no corresponding record for this reference.
- 73Nagaraj, H.; Lehot, V.; Nasim, N.; Cicek, Y. A.; Goswami, R.; Jeon, T.; Rotello, V. M. Breaking the Cellular Delivery Bottleneck: Recent Developments in Direct Cytosolic Delivery of Biologics. RSC Pharm. 2025, 2 (5), 850– 864, DOI: 10.1039/D5PM00129CGoogle ScholarThere is no corresponding record for this reference.
- 74Dowdy, S. F.; Setten, R. L.; Cui, X.-S.; Jadhav, S. G. Delivery of RNA Therapeutics: The Great Endosomal Escape!. Nucleic Acid Ther. 2022, 32 (5), 361– 368, DOI: 10.1089/nat.2022.0004Google ScholarThere is no corresponding record for this reference.
- 75Lönn, P.; Kacsinta, A. D.; Cui, X.-S.; Hamil, A. S.; Kaulich, M.; Gogoi, K.; Dowdy, S. F. Enhancing Endosomal Escape for Intracellular Delivery of Macromolecular Biologic Therapeutics. Sci. Rep. 2016, 6 (1), 32301 DOI: 10.1038/srep32301Google ScholarThere is no corresponding record for this reference.
- 76Liang, X.-h.; Sun, H.; Hsu, C.-W.; Nichols, J. G.; Vickers, T. A.; De Hoyos, C. L.; Crooke, S. T. Golgi-Endosome Transport Mediated by M6PR Facilitates Release of Antisense Oligonucleotides from Endosomes. Nucleic Acids Res. 2020, 48 (3), 1372– 1391, DOI: 10.1093/nar/gkz1171Google ScholarThere is no corresponding record for this reference.
- 77Hwang, H. S.; Hu, J.; Na, K.; Bae, Y. H. Role of Polymeric Endosomolytic Agents in Gene Transfection: A Comparative Study of Poly(l-Lysine) Grafted with Monomeric l-Histidine Analogue and Poly(l-Histidine). Biomacromolecules 2014, 15 (10), 3577– 3586, DOI: 10.1021/bm500843rGoogle ScholarThere is no corresponding record for this reference.
- 78Sun, W.; Ji, W.; Hall, J. M.; Hu, Q.; Wang, C.; Beisel, C. L.; Gu, Z. Self-Assembled DNA Nanoclews for the Efficient Delivery of CRISPR–Cas9 for Genome Editing. Angew. Chem., Int. Ed. 2015, 54 (41), 12029– 12033, DOI: 10.1002/anie.201506030Google ScholarThere is no corresponding record for this reference.
- 79Liang, L.; Li, J.; Li, Q.; Huang, Q.; Shi, J.; Yan, H.; Fan, C. Single-Particle Tracking and Modulation of Cell Entry Pathways of a Tetrahedral DNA Nanostructure in Live Cells. Angew. Chem., Int. Ed. 2014, 53 (30), 7745– 7750, DOI: 10.1002/anie.201403236Google ScholarThere is no corresponding record for this reference.
- 80Wang, D.; Liu, Q.; Wu, D.; He, B.; Li, J.; Mao, C.; Wang, G.; Qian, H. Isothermal Self-Assembly of Spermidine–DNA Nanostructure Complex as a Functional Platform for Cancer Therapy. ACS Appl. Mater. Interfaces 2018, 10 (18), 15504– 15516, DOI: 10.1021/acsami.8b03464Google ScholarThere is no corresponding record for this reference.
- 81Smolková, B.; MacCulloch, T.; Rockwood, T. F.; Liu, M.; Henry, S. J. W.; Frtús, A.; Uzhytchak, M.; Lunova, M.; Hof, M.; Jurkiewicz, P.; Dejneka, A.; Stephanopoulos, N.; Lunov, O. Protein Corona Inhibits Endosomal Escape of Functionalized DNA Nanostructures in Living Cells. ACS Appl. Mater. Interfaces 2021, 13 (39), 46375– 46390, DOI: 10.1021/acsami.1c14401Google ScholarThere is no corresponding record for this reference.
- 82Han, X.; Zhang, H.; Butowska, K.; Swingle, K. L.; Alameh, M.-G.; Weissman, D.; Mitchell, M. J. An Ionizable Lipid Toolbox for RNA Delivery. Nat. Commun. 2021, 12 (1), 7233 DOI: 10.1038/s41467-021-27493-0Google ScholarThere is no corresponding record for this reference.
- 83Huang, P.; Qi, M.; Chen, C.; Xu, F.; Li, S.; Xu, Q.; Pan, H.; Wang, Y.; Yu, C.; Zhang, S.; Zhou, Y. Asymmetric Vesicles Self-Assembled by Amphiphilic Sequence-Controlled Polymers. ACS Macro Lett. 2021, 10 (7), 894– 900, DOI: 10.1021/acsmacrolett.1c00301Google ScholarThere is no corresponding record for this reference.
- 84Yu, L.; Xu, Y.; Al-Amin, M.; Jiang, S.; Sample, M.; Prasad, A.; Stephanopoulos, N.; Šulc, P.; Yan, H. CytoDirect: A Nucleic Acid Nanodevice for Specific and Efficient Delivery of Functional Payloads to the Cytoplasm. J. Am. Chem. Soc. 2023, 145 (50), 27336– 27347, DOI: 10.1021/jacs.3c07491Google ScholarThere is no corresponding record for this reference.
- 85Cognet, M.; Renno, G.; Rose, N.; Zhang, Y.; Josso, P.; Moreno, J.; Ren, X.; Bouffard, J.; Saidjalolov, S.; Sakai, N.; Matile, S. Cell-Penetrating Poly(Disulfide)s. Helv. Chim. Acta 2025, 108 (12), e00129 DOI: 10.1002/hlca.202500129Google ScholarThere is no corresponding record for this reference.
- 86Wang, W.; Chopra, B.; Walawalkar, V.; Liang, Z.; Adams, R.; Deserno, M.; Ren, X.; Taylor, R. E. Cell–Surface Binding of DNA Nanostructures for Enhanced Intracellular and Intranuclear Delivery. ACS Appl. Mater. Interfaces 2024, 16 (13), 15783– 15797, DOI: 10.1021/acsami.3c18068Google ScholarThere is no corresponding record for this reference.
- 87Samanta, A.; Malle, M. G.; Tsang, E.; Omer, M.; Skaanning, M. K.; Youssef, S.; Kjems, J.; Gothelf, K. V. Bacteriophage-Mimetic DNA Origami Needle for Targeted Membrane Penetration and Cytosolic Cargo Delivery. Adv. Sci. 2026, 13 (10), e12844 DOI: 10.1002/advs.202512844Google ScholarThere is no corresponding record for this reference.
- 88Du, R. R.; Cedrone, E.; Romanov, A.; Falkovich, R.; Dobrovolskaia, M. A.; Bathe, M. Innate Immune Stimulation Using 3D Wireframe DNA Origami. ACS Nano 2022, 16 (12), 20340– 20352, DOI: 10.1021/acsnano.2c06275Google ScholarThere is no corresponding record for this reference.
- 89Lucas, C. R.; Halley, P. D.; Chowdury, A. A.; Harrington, B. K.; Beaver, L.; Lapalombella, R.; Johnson, A. J.; Hertlein, E. K.; Phelps, M. A.; Byrd, J. C.; Castro, C. E. DNA Origami Nanostructures Elicit Dose-Dependent Immunogenicity and Are Nontoxic up to High Doses In Vivo. Small 2022, 18 (26), 2108063 DOI: 10.1002/smll.202108063Google ScholarThere is no corresponding record for this reference.
- 90Arulkumaran, N.; Lanphere, C.; Gaupp, C.; Burns, J. R.; Singer, M.; Howorka, S. DNA Nanodevices with Selective Immune Cell Interaction and Function. ACS Nano 2021, 15 (3), 4394– 4404, DOI: 10.1021/acsnano.0c07915Google ScholarThere is no corresponding record for this reference.
- 91Rodriguez, A.; Madhanagopal, B. R.; Sarkar, K.; Nowzari, Z.; Mathivanan, J.; Talbot, H.; Patel, A.; Morya, V.; Halvorsen, K.; Vangaveti, S.; Berglund, J. A.; Chandrasekaran, A. R. Counterions Influence the Isothermal Self-Assembly of DNA Nanostructures. Sc. Adv. 2025, 11 (11), eadu7366 DOI: 10.1126/sciadv.adu7366Google ScholarThere is no corresponding record for this reference.
- 92Guo, Y.; Huang, Y.; Liu, M.; Liu, J.; Liu, J.; Yang, D. Evaluation of Pharmacokinetics, Immunogenicity, and Immunotoxicity of DNA Tetrahedral and DNA Polymeric Nanostructures. Small Methods 2025, 9 (6), 2401007 DOI: 10.1002/smtd.202401007Google ScholarThere is no corresponding record for this reference.
- 93Jiang, D.; Ge, Z.; Im, H.-J.; England, C. G.; Ni, D.; Hou, J.; Zhang, L.; Kutyreff, C. J.; Yan, Y.; Liu, Y.; Cho, S. Y.; Engle, J. W.; Shi, J.; Huang, P.; Fan, C.; Yan, H.; Cai, W. DNA Origami Nanostructures Can Exhibit Preferential Renal Uptake and Alleviate Acute Kidney Injury. Nat. Biomed. Eng. 2018, 2 (11), 865– 877, DOI: 10.1038/s41551-018-0317-8Google ScholarThere is no corresponding record for this reference.
- 94Wang, Y.; Baars, I.; Berzina, I.; Rocamonde-Lago, I.; Shen, B.; Yang, Y.; Lolaico, M.; Waldvogel, J.; Smyrlaki, I.; Zhu, K.; Harris, R. A.; Högberg, B. A DNA Robotic Switch with Regulated Autonomous Display of Cytotoxic Ligand Nanopatterns. Nat. Nanotechnol. 2024, 19 (9), 1366– 1374, DOI: 10.1038/s41565-024-01676-4Google ScholarThere is no corresponding record for this reference.
- 95Li, L.; Yin, J.; Ma, W.; Tang, L.; Zou, J.; Yang, L.; Du, T.; Zhao, Y.; Wang, L.; Yang, Z.; Fan, C.; Chao, J.; Chen, X. A DNA Origami Device Spatially Controls CD95 Signalling to Induce Immune Tolerance in Rheumatoid Arthritis. Nat. Mater. 2024, 23 (7), 993– 1001, DOI: 10.1038/s41563-024-01865-5Google ScholarThere is no corresponding record for this reference.
- 96Zeng, Y. C.; Young, O. J.; Wintersinger, C. M.; Anastassacos, F. M.; MacDonald, J. I.; Isinelli, G.; Dellacherie, M. O.; Sobral, M.; Bai, H.; Graveline, A. R.; Vernet, A.; Sanchez, M.; Mulligan, K.; Choi, Y.; Ferrante, T. C.; Keskin, D. B.; Fell, G. G.; Neuberg, D.; Wu, C. J.; Mooney, D. J.; Kwon, I. C.; Ryu, J. H.; Shih, W. M. Fine Tuning of CpG Spatial Distribution with DNA Origami for Improved Cancer Vaccination. Nat. Nanotechnol. 2024, 19 (7), 1055– 1065, DOI: 10.1038/s41565-024-01615-3Google ScholarThere is no corresponding record for this reference.
- 97Zeng, Y. C.; Young, O. J.; Xiong, Q.; Si, L.; Ku, M. W.; Bernier, S. G.; Dembele, H.; Isinelli, G.; Gilboa, T.; Swank, Z.; Seok, S. H.; Rajwar, A.; Jiang, A.; Zhai, Y.; Williams, L. D.; Hellman, C. A.; Wintersinger, C. M.; Graveline, A. R.; Vernet, A.; Sanchez, M.; Bardales, S.; Tomaras, G. D.; Ryu, J. H.; Kwon, I. C.; Goyal, G.; Ingber, D. E.; Shih, W. M. DNA Origami Vaccine Nanoparticles Improve Humoral and Cellular Immune Responses to Infectious Diseases. Nat. Biomed. Eng. 2026, 1– 18, DOI: 10.1038/s41551-026-01614-wGoogle ScholarThere is no corresponding record for this reference.
- 98Glassman, P. M.; Muzykantov, V. R. Pharmacokinetic and Pharmacodynamic Properties of Drug Delivery Systems. J. Pharmacol. Exp. Ther. 2019, 370 (3), 570– 580, DOI: 10.1124/jpet.119.257113Google ScholarThere is no corresponding record for this reference.
- 99Chen, L.; Bosmajian, C.; Woo, S. Mechanistic Intracellular PK/PD Modeling to Inform Development Strategies for Small Interfering RNA Therapeutics. Mol. Ther. Nucleic Acids 2025, 36 (2), 102516 DOI: 10.1016/j.omtn.2025.102516Google ScholarThere is no corresponding record for this reference.
- 100Wang, Y.; Rocamonde-Lago, I.; Waldvogel, J.; Shen, B.; Wu, Y.-C.; Zhu, J.; Zang, S.; Jia, Y.; Baars, I.; Kloosterman, A.; Hoffecker, I. T.; Wu, M.-R.; He, Q.; Högberg, B. Resolving DNA Origami Structural Integrity and Pharmacokinetics in Vivo. Nat. Nanotechnol. 2026, 21 (2), 268– 276, DOI: 10.1038/s41565-025-02091-zGoogle ScholarThere is no corresponding record for this reference.
- 101Li, S.; Sun, Y.; Tian, T.; Qin, X.; Lin, S.; Zhang, T.; Zhang, Q.; Zhou, M.; Zhang, X.; Zhou, Y.; Zhao, H.; Zhu, B.; Cai, X. MicroRNA-214–3p Modified Tetrahedral Framework Nucleic Acids Target Survivin to Induce Tumour Cell Apoptosis. Cell Proliferation 2020, 53 (1), e12708 DOI: 10.1111/cpr.12708Google ScholarThere is no corresponding record for this reference.
- 102Wei, M.; Li, S.; Yang, Z.; Cheng, C.; Li, T.; Le, W. Tetrahedral DNA Nanostructures Functionalized by Multivalent microRNA132 Antisense Oligonucleotides Promote the Differentiation of Mouse Embryonic Stem Cells into Dopaminergic Neurons. Nanomed.: Nanotechnol., Biol. Med. 2021, 34, 102375 DOI: 10.1016/j.nano.2021.102375Google ScholarThere is no corresponding record for this reference.
- 103Su, J.; Wu, F.; Xia, H.; Wu, Y.; Liu, S. Accurate Cancer Cell Identification and microRNA Silencing Induced Therapy Using Tailored DNA Tetrahedron Nanostructures. Chem. Sci. 2020, 11 (1), 80– 86, DOI: 10.1039/C9SC04823EGoogle ScholarThere is no corresponding record for this reference.
- 104Kim, K.-R.; Jegal, H.; Kim, J.; Ahn, D.-R. A Self-Assembled DNA Tetrahedron as a Carrier for in Vivo Liver-Specific Delivery of siRNA. Biomater. Sci. 2020, 8 (2), 586– 590, DOI: 10.1039/C9BM01769KGoogle ScholarThere is no corresponding record for this reference.
- 105Moreno, P. M. D.; Cortinhas, J.; Martins, A. S.; Pêgo, A. P. Engineering a Novel Self-Assembled Multi-siRNA Nanocaged Architecture with Controlled Enzyme-Mediated siRNA Release. ACS Appl. Mater. Interfaces 2022, 14 (51), 56483– 56497, DOI: 10.1021/acsami.2c15086Google ScholarThere is no corresponding record for this reference.
- 106Zhang, T.; Li, R.; Wang, Z.; Zhou, Y.; Zhou, Y.; Chen, X.; Peng, C.; Jiang, Y.; Tong, N.; Li, W. Inflammation-Specific DNA Origami Nanodevice for Delivery of siRNAs to Treat Ulcerative Colitis. Nat. Commun. 2026, 17 (1), 495 DOI: 10.1038/s41467-025-67183-9Google ScholarThere is no corresponding record for this reference.
- 107Bujold, K. E.; Hsu, J. C. C.; Sleiman, H. F. Optimized DNA “Nanosuitcases” for Encapsulation and Conditional Release of siRNA. J. Am. Chem. Soc. 2016, 138 (42), 14030– 14038, DOI: 10.1021/jacs.6b08369Google ScholarThere is no corresponding record for this reference.
- 108Zhang, H.; Demirer, G. S.; Zhang, H.; Ye, T.; Goh, N. S.; Aditham, A. J.; Cunningham, F. J.; Fan, C.; Landry, M. P. DNA Nanostructures Coordinate Gene Silencing in Mature Plants. Proc. Natl. Acad. Sci. U.S.A. 2019, 116 (15), 7543– 7548, DOI: 10.1073/pnas.1818290116Google ScholarThere is no corresponding record for this reference.
- 109Li, C.; Lin, W.; Wang, W.; Wu, J.; Luo, S.; Chen, L.; Wu, R.; Shen, Z.; Wu, Z.-S. Folding an RCA Scaffold into an Intelligent Coiled Nanosnake for Precise/Synergistic RNAi-/Chemotherapy of Cancer. Anal. Chem. 2025, 97 (2), 1107– 1116, DOI: 10.1021/acs.analchem.4c03437Google ScholarThere is no corresponding record for this reference.
- 110Jensen, S. A.; Day, E. S.; Ko, C. H.; Hurley, L. A.; Luciano, J. P.; Kouri, F. M.; Merkel, T. J.; Luthi, A. J.; Patel, P. C.; Cutler, J. I.; Daniel, W. L.; Scott, A. W.; Rotz, M. W.; Meade, T. J.; Giljohann, D. A.; Mirkin, C. A.; Stegh, A. H. Spherical Nucleic Acid Nanoparticle Conjugates as an RNAi-Based Therapy for Glioblastoma. Sci. Transl. Med. 2013, 5 (209), 209ra152 DOI: 10.1126/scitranslmed.3006839Google ScholarThere is no corresponding record for this reference.
- 111Kumthekar, P.; Ko, C. H.; Paunesku, T.; Dixit, K.; Sonabend, A. M.; Bloch, O.; Tate, M.; Schwartz, M.; Zuckerman, L.; Lezon, R.; Lukas, R. V.; Jovanovic, B.; McCortney, K.; Colman, H.; Chen, S.; Lai, B.; Antipova, O.; Deng, J.; Li, L.; Tommasini-Ghelfi, S.; Hurley, L. A.; Unruh, D.; Sharma, N. V.; Kandpal, M.; Kouri, F. M.; Davuluri, R. V.; Brat, D. J.; Muzzio, M.; Glass, M.; Vijayakumar, V.; Heidel, J.; Giles, F. J.; Adams, A. K.; James, C. D.; Woloschak, G. E.; Horbinski, C.; Stegh, A. H. A First-in-Human Phase 0 Clinical Study of RNA Interference–Based Spherical Nucleic Acids in Patients with Recurrent Glioblastoma. Sci. Transl. Med. 2021, 13 (584), eabb3945 DOI: 10.1126/scitranslmed.abb3945Google ScholarThere is no corresponding record for this reference.
- 112Chen, X.; Tang, J.; Shuai, W.; Meng, J.; Feng, J.; Han, Z. Macrophage Polarization and Its Role in the Pathogenesis of Acute Lung Injury/Acute Respiratory Distress Syndrome. Inflamm. Res. 2020, 69 (9), 883– 895, DOI: 10.1007/s00011-020-01378-2Google ScholarThere is no corresponding record for this reference.
- 113Huang, C.; You, Q.; Xu, J.; Wu, D.; Chen, H.; Guo, Y.; Xu, J.; Hu, M.; Qian, H. An mTOR siRNA-Loaded Spermidine/DNA Tetrahedron Nanoplatform with a Synergistic Anti-Inflammatory Effect on Acute Lung Injury. Adv. Healthcare Mater. 2022, 11 (11), 2200008 DOI: 10.1002/adhm.202200008Google ScholarThere is no corresponding record for this reference.
- 114Li, C.; Zhao, W.; Hu, Z.; Yu, H. Cholesterol-Modified DNA Nanostructures Serve as Effective Non-Viral Carriers for Delivering siRNA to the Kidneys to Prevent Acute Kidney Injury. Small 2024, 20 (30), 2311690 DOI: 10.1002/smll.202311690Google ScholarThere is no corresponding record for this reference.
- 115Kim, K.-R.; Kim, H. Y.; Lee, Y.-D.; Ha, J. S.; Kang, J. H.; Jeong, H.; Bang, D.; Ko, Y. T.; Kim, S.; Lee, H.; Ahn, D.-R. Self-Assembled Mirror DNA Nanostructures for Tumor-Specific Delivery of Anticancer Drugs. J. Controlled Release 2016, 243, 121– 131, DOI: 10.1016/j.jconrel.2016.10.015Google ScholarThere is no corresponding record for this reference.
- 116Jiang, D.; Sun, Y.; Li, J.; Li, Q.; Lv, M.; Zhu, B.; Tian, T.; Cheng, D.; Xia, J.; Zhang, L.; Wang, L.; Huang, Q.; Shi, J.; Fan, C. Multiple-Armed Tetrahedral DNA Nanostructures for Tumor-Targeting, Dual-Modality in Vivo Imaging. ACS Appl. Mater. Interfaces 2016, 8 (7), 4378– 4384, DOI: 10.1021/acsami.5b10792Google ScholarThere is no corresponding record for this reference.
- 117Zhou, X.; Yu, M.; Ma, L.; Fu, J.; Guo, J.; Lei, J.; Fu, Z.; Fu, Y.; Zhang, Q.; Zhang, C.-Y.; Chen, X. In Vivo Self-Assembled siRNA as a Modality for Combination Therapy of Ulcerative Colitis. Nat. Commun. 2022, 13 (1), 5700 DOI: 10.1038/s41467-022-33436-0Google ScholarThere is no corresponding record for this reference.
- 118Dong, H.; Song, G.; Ma, D.; Wang, T.; Jing, S.; Yang, H.; Tao, Y.; Tang, Y.; Shi, Y.; Dai, Z.; Zhu, J.; Liu, T.; Wang, B.; Leng, X.; Shen, X.; Zhu, C.; Zhao, Y. Improved Antiviral Activity of Classical Swine Fever Virus-Targeted siRNA by Tetrahedral Framework Nucleic Acid-Enhanced Delivery. ACS Appl. Mater. Interfaces 2021, 13 (25), 29416– 29423, DOI: 10.1021/acsami.1c08143Google ScholarThere is no corresponding record for this reference.
- 119Fang, Y.; Xue, J.; Gao, S.; Lu, A.; Yang, D.; Jiang, H.; He, Y.; Shi, K. Cleavable PEGylation: A Strategy for Overcoming the “PEG Dilemma” in Efficient Drug Delivery. Drug Delivery 2017, 24 (2), 22– 32, DOI: 10.1080/10717544.2017.1388451Google ScholarThere is no corresponding record for this reference.
- 120Ou, M.; Wang, X.-L.; Xu, R.; Chang, C.-W.; Bull, D. A.; Kim, S. W. Novel Biodegradable Poly(Disulfide Amine)s for Gene Delivery with High Efficiency and Low Cytotoxicity. Bioconjugate Chem. 2008, 19 (3), 626– 633, DOI: 10.1021/bc700397xGoogle ScholarThere is no corresponding record for this reference.
- 121Shi, D.; Beasock, D.; Fessler, A.; Szebeni, J.; Ljubimova, J. Y.; Afonin, K. A.; Dobrovolskaia, M. A. To PEGylate or Not to PEGylate: Immunological Properties of Nanomedicine’s Most Popular Component, Polyethylene Glycol and Its Alternatives. Adv. Drug Delivery Rev. 2022, 180, 114079 DOI: 10.1016/j.addr.2021.114079Google ScholarThere is no corresponding record for this reference.
- 122Ju, Y.; Carreño, J. M.; Simon, V.; Dawson, K.; Krammer, F.; Kent, S. J. Impact of Anti-PEG Antibodies Induced by SARS-CoV-2 mRNA Vaccines. Nat. Rev. Immunol. 2023, 23 (3), 135– 136, DOI: 10.1038/s41577-022-00825-xGoogle ScholarThere is no corresponding record for this reference.
- 123Klipp, A.; Burger, M.; Leroux, J.-C. Get out or Die Trying: Peptide- and Protein-Based Endosomal Escape of RNA Therapeutics. Adv. Drug Delivery Rev. 2023, 200, 115047 DOI: 10.1016/j.addr.2023.115047Google ScholarThere is no corresponding record for this reference.
- 124Rhee, W. J.; Bao, G. Slow Non-Specific Accumulation of 2′-Deoxy and 2′-O-Methyl Oligonucleotide Probes at Mitochondria in Live Cells. Nucleic Acids Res. 2010, 38 (9), e109 DOI: 10.1093/nar/gkq050Google ScholarThere is no corresponding record for this reference.
- 125Høiberg, H. C.; Sparvath, S. M.; Andersen, V. L.; Kjems, J.; Andersen, E. S. An RNA Origami Octahedron with Intrinsic siRNAs for Potent Gene Knockdown. Biotechnol. J. 2019, 14 (1), 1700634 DOI: 10.1002/biot.201700634Google ScholarThere is no corresponding record for this reference.
- 126Wu, X. A.; Choi, C. H. J.; Zhang, C.; Hao, L.; Mirkin, C. A. Intracellular Fate of Spherical Nucleic Acid Nanoparticle Conjugates. J. Am. Chem. Soc. 2014, 136 (21), 7726– 7733, DOI: 10.1021/ja503010aGoogle ScholarThere is no corresponding record for this reference.
- 127Lacroix, A.; Vengut-Climent, E.; de Rochambeau, D.; Sleiman, H. F. Uptake and Fate of Fluorescently Labeled DNA Nanostructures in Cellular Environments: A Cautionary Tale. ACS Cent. Sci. 2019, 5 (5), 882– 891, DOI: 10.1021/acscentsci.9b00174Google ScholarThere is no corresponding record for this reference.
- 128Kozlov, S.; Panigaj, M.; Rebolledo, L.; Bhaskaran, H.; Afonin, K. A. Nucleic Acid Nanoparticles Redefine Traditional Regulatory Terminology: The Blurred Line between Active Pharmaceutical Ingredients and Excipients. ACS Nano Med. 2026, 1 (1), 5– 17, DOI: 10.1021/acsnanomed.5c00070Google ScholarThere is no corresponding record for this reference.
- 129Praetorius, F.; Kick, B.; Behler, K. L.; Honemann, M. N.; Weuster-Botz, D.; Dietz, H. Biotechnological Mass Production of DNA Origami. Nature 2017, 552 (7683), 84– 87, DOI: 10.1038/nature24650Google ScholarThere is no corresponding record for this reference.
- 130Langlois, N. I.; Clark, H. A. Characterization of DNA Nanostructure Stability by Size Exclusion Chromatography. Anal. Methods 2022, 14 (10), 1006– 1014, DOI: 10.1039/D1AY02146JGoogle ScholarThere is no corresponding record for this reference.
- 131Halvorsen, K.; Kizer, M. E.; Wang, X.; Chandrasekaran, A. R.; Basanta-Sanchez, M. Shear Dependent LC Purification of an Engineered DNA Nanoswitch and Implications for DNA Origami. Anal. Chem. 2017, 89 (11), 5673– 5677, DOI: 10.1021/acs.analchem.7b00791Google ScholarThere is no corresponding record for this reference.
- 132Douglas, S. M.; Bachelet, I.; Church, G. M. A Logic-Gated Nanorobot for Targeted Transport of Molecular Payloads. Science 2012, 335 (6070), 831– 834, DOI: 10.1126/science.1214081Google ScholarThere is no corresponding record for this reference.
Cited By
This article has not yet been cited by other publications.
Article Views
Altmetric
Citations
Article Views are the COUNTER-compliant sum of full text article downloads since November 2008 (both PDF and HTML) across all institutions and individuals. These metrics are regularly updated to reflect usage leading up to the last few days.
Citations are the number of other articles citing this article, calculated by Crossref and updated daily. Find more information about Crossref citation counts.
The Altmetric Attention Score is a quantitative measure of the attention that a research article has received online. Clicking on the donut icon will load a page at altmetric.com with additional details about the score and the social media presence for the given article. Find more information on the Altmetric Attention Score and how the score is calculated.
Recommended Articles
Abstract
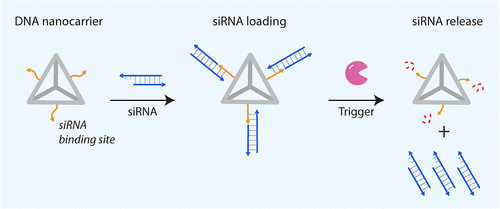
Figure 1

Figure 1. Overview of siRNA-mediated gene silencing by RNA interference (RNAi).
Figure 2

Figure 2. DNA nanostructures. (a) DNA nanostructures assembled by cooperative assembly of short DNA single strands (e.g., a tetrahedron). (b) Modular assembly of DNA motifs into polyhedral structures (e.g., an icosahedron). (c) Hierarchical assembly of DNA motifs into 2D arrays. (d) Folding of a long single-stranded DNA scaffold into arbitrary shapes using the DNA origami strategy. (e) Single-stranded DNA built into larger structures using the DNA brick strategy.
Figure 3

Figure 3. Overview of drug delivery pathway using DNA nanostructures. DNA nanostructures can be functionalized to be (a) biostable against nuclease activity, (b) survive protein corona formation, and (c) used as imaging modalities. (d) DNA nanostructures with siRNA cargos can be targeted to specific cells for uptake, tracking, and disassembly or degradation in response to external stimuli to release the cargo.
Figure 4
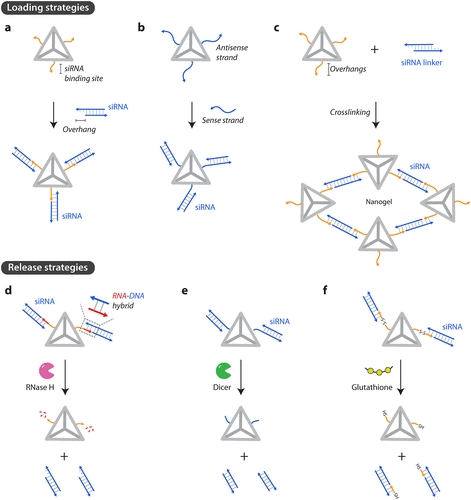
Figure 4. Loading and release of siRNAs. (a) Attachment to single-stranded overhangs on DNA nanostructures. (b) Antisense strand is part of the DNA nanostructure that can bind to the sense strand of the siRNA. (c) siRNA linkers can connect DNA nanostructures to create a nanogel. (d) Release of siRNA by RNase H activity, (e) Dicer activity, and (f) reduction of disulfide linkages by glutathione.
References
This article references 132 other publications.
- 1Khvorova, A.; Watts, J. K. The Chemical Evolution of Oligonucleotide Therapies of Clinical Utility. Nat. Biotechnol. 2017, 35 (3), 238– 248, DOI: 10.1038/nbt.3765There is no corresponding record for this reference.
- 2Egli, M.; Manoharan, M. Chemistry, Structure and Function of Approved Oligonucleotide Therapeutics. Nucleic Acids Res. 2023, 51 (6), 2529– 2573, DOI: 10.1093/nar/gkad067There is no corresponding record for this reference.
- 3Paunovska, K.; Loughrey, D.; Dahlman, J. E. Drug Delivery Systems for RNA Therapeutics. Nat. Rev. Genet. 2022, 23 (5), 265– 280, DOI: 10.1038/s41576-021-00439-4There is no corresponding record for this reference.
- 4Elbashir, S. M.; Harborth, J.; Lendeckel, W.; Yalcin, A.; Weber, K.; Tuschl, T. Duplexes of 21-Nucleotide RNAs Mediate RNA Interference in Cultured Mammalian Cells. Nature 2001, 411 (6836), 494– 498, DOI: 10.1038/35078107There is no corresponding record for this reference.
- 5Jadhav, V.; Vaishnaw, A.; Fitzgerald, K.; Maier, M. A. RNA Interference in the Era of Nucleic Acid Therapeutics. Nat. Biotechnol. 2024, 42 (3), 394– 405, DOI: 10.1038/s41587-023-02105-yThere is no corresponding record for this reference.
- 6Xiao, B.; Wang, S.; Pan, Y.; Zhi, W.; Gu, C.; Guo, T.; Zhai, J.; Li, C.; Chen, Y. Q.; Wang, R. Development, Opportunities, and Challenges of siRNA Nucleic Acid Drugs. Mol. Ther. Nucleic Acids 2025, 36 (1), 102437 DOI: 10.1016/j.omtn.2024.102437There is no corresponding record for this reference.
- 7Zamore, P. D.; Tuschl, T.; Sharp, P. A.; Bartel, D. P. RNAi: Double-Stranded RNA Directs the ATP-Dependent Cleavage of mRNA at 21 to 23 Nucleotide Intervals. Cell 2000, 101 (1), 25– 33, DOI: 10.1016/S0092-8674(00)80620-0There is no corresponding record for this reference.
- 8Semizarov, D.; Frost, L.; Sarthy, A.; Kroeger, P.; Halbert, D. N.; Fesik, S. W. Specificity of Short Interfering RNA Determined through Gene Expression Signatures. Proc. Natl. Acad. Sci. U.S.A. 2003, 100 (11), 6347– 6352, DOI: 10.1073/pnas.1131959100There is no corresponding record for this reference.
- 9Hammond, S. M.; Bernstein, E.; Beach, D.; Hannon, G. J. An RNA-Directed Nuclease Mediates Post-Transcriptional Gene Silencing in Drosophila Cells. Nature 2000, 404 (6775), 293– 296, DOI: 10.1038/35005107There is no corresponding record for this reference.
- 10Wittrup, A.; Ai, A.; Liu, X.; Hamar, P.; Trifonova, R.; Charisse, K.; Manoharan, M.; Kirchhausen, T.; Lieberman, J. Visualizing Lipid-Formulated siRNA Release from Endosomes and Target Gene Knockdown. Nat. Biotechnol. 2015, 33 (8), 870– 876, DOI: 10.1038/nbt.3298There is no corresponding record for this reference.
- 11Tabernero, J.; Shapiro, G. I.; LoRusso, P. M.; Cervantes, A.; Schwartz, G. K.; Weiss, G. J.; Paz-Ares, L.; Cho, D. C.; Infante, J. R.; Alsina, M.; Gounder, M. M.; Falzone, R.; Harrop, J.; White, A. C. S.; Toudjarska, I.; Bumcrot, D.; Meyers, R. E.; Hinkle, G.; Svrzikapa, N.; Hutabarat, R. M.; Clausen, V. A.; Cehelsky, J.; Nochur, S. V.; Gamba-Vitalo, C.; Vaishnaw, A. K.; Sah, D. W. Y.; Gollob, J. A.; Burris, H. A. III. First-in-Humans Trial of an RNA Interference Therapeutic Targeting VEGF and KSP in Cancer Patients with Liver Involvement. Cancer Discovery 2013, 3 (4), 406– 417, DOI: 10.1158/2159-8290.CD-12-0429There is no corresponding record for this reference.
- 12Kato, H.; Takeuchi, O.; Mikamo-Satoh, E.; Hirai, R.; Kawai, T.; Matsushita, K.; Hiiragi, A.; Dermody, T. S.; Fujita, T.; Akira, S. Length-Dependent Recognition of Double-Stranded Ribonucleic Acids by Retinoic Acid–Inducible Gene-I and Melanoma Differentiation–Associated Gene 5. J. Exp. Med. 2008, 205 (7), 1601– 1610, DOI: 10.1084/jem.20080091There is no corresponding record for this reference.
- 13Bereczki, Z.; Benczik, B.; Balogh, O. M.; Marton, S.; Puhl, E.; Pétervári, M.; Váczy-Földi, M.; Papp, Z. T.; Makkos, A.; Glass, K.; Locquet, F.; Euler, G.; Schulz, R.; Ferdinandy, P.; Ágg, B. Mitigating Off-Target Effects of Small RNAs: Conventional Approaches, Network Theory and Artificial Intelligence. Br. J. Pharmacol. 2025, 182 (2), 340– 379, DOI: 10.1111/bph.17302There is no corresponding record for this reference.
- 14Janas, M. M.; Schlegel, M. K.; Harbison, C. E.; Yilmaz, V. O.; Jiang, Y.; Parmar, R.; Zlatev, I.; Castoreno, A.; Xu, H.; Shulga-Morskaya, S.; Rajeev, K. G.; Manoharan, M.; Keirstead, N. D.; Maier, M. A.; Jadhav, V. Selection of GalNAc-Conjugated siRNAs with Limited off-Target-Driven Rat Hepatotoxicity. Nat. Commun. 2018, 9, 723 DOI: 10.1038/s41467-018-02989-4There is no corresponding record for this reference.
- 15Jackson, A. L.; Burchard, J.; Leake, D.; Reynolds, A.; Schelter, J.; Guo, J.; Johnson, J. M.; Lim, L.; Karpilow, J.; Nichols, K.; Marshall, W.; Khvorova, A.; Linsley, P. S. Position-Specific Chemical Modification of siRNAs Reduces “off-Target” Transcript Silencing. RNA 2006, 12 (7), 1197– 1205, DOI: 10.1261/rna.30706There is no corresponding record for this reference.
- 16Corey, D. R.; Damha, M. J.; Manoharan, M. Challenges and Opportunities for Nucleic Acid Therapeutics. Nucleic Acid Ther. 2022, 32 (1), 8– 13, DOI: 10.1089/nat.2021.0085There is no corresponding record for this reference.
- 17Couto, L. B.; High, K. A. Viral Vector-Mediated RNA Interference. Curr. Opin. Pharmacol. 2010, 10 (5), 534– 542, DOI: 10.1016/j.coph.2010.06.007There is no corresponding record for this reference.
- 18Pérez-Martínez, F. C.; Guerra, J.; Posadas, I.; Ceña, V. Barriers to Non-Viral Vector-Mediated Gene Delivery in the Nervous System. Pharm. Res. 2011, 28 (8), 1843– 1858, DOI: 10.1007/s11095-010-0364-7There is no corresponding record for this reference.
- 19Gregoriadis, G. Engineering Liposomes for Drug Delivery: Progress and Problems. Trends Biotechnol. 1995, 13 (12), 527– 537, DOI: 10.1016/S0167-7799(00)89017-4There is no corresponding record for this reference.
- 20Wijagkanalan, W.; Kawakami, S.; Hashida, M. Designing Dendrimers for Drug Delivery and Imaging: Pharmacokinetic Considerations. Pharm. Res. 2011, 28 (7), 1500– 1519, DOI: 10.1007/s11095-010-0339-8There is no corresponding record for this reference.
- 21Pack, D. W.; Hoffman, A. S.; Pun, S.; Stayton, P. S. Design and Development of Polymers for Gene Delivery. Nat. Rev. Drug Discovery 2005, 4 (7), 581– 593, DOI: 10.1038/nrd1775There is no corresponding record for this reference.
- 22Duncan, B.; Kim, C.; Rotello, V. M. Gold Nanoparticle Platforms as Drug and Biomacromolecule Delivery Systems. J. Controlled Release 2010, 148 (1), 122– 127, DOI: 10.1016/j.jconrel.2010.06.004There is no corresponding record for this reference.
- 23Kakkar, A.; Traverso, G.; Farokhzad, O. C.; Weissleder, R.; Langer, R. Evolution of Macromolecular Complexity in Drug Delivery Systems. Nat. Rev. Chem. 2017, 1 (8), 0063 DOI: 10.1038/s41570-017-0063There is no corresponding record for this reference.
- 24Zhang, S.; Zhao, Y.; Zhi, D.; Zhang, S. Non-Viral Vectors for the Mediation of RNAi. Bioorg. Chem. 2012, 40, 10– 18, DOI: 10.1016/j.bioorg.2011.07.005There is no corresponding record for this reference.
- 25Seow, Y.; Wood, M. J. Biological Gene Delivery Vehicles: Beyond Viral Vectors. Mol. Ther. 2009, 17 (5), 767– 777, DOI: 10.1038/mt.2009.41There is no corresponding record for this reference.
- 26Madhanagopal, B. R.; Zhang, S.; Demirel, E.; Wady, H.; Chandrasekaran, A. R. DNA Nanocarriers: Programmed to Deliver. Trends Biochem. Sci. 2018, 43 (12), 997– 1013, DOI: 10.1016/j.tibs.2018.09.010There is no corresponding record for this reference.
- 27Xavier, P. L.; Chandrasekaran, A. R. DNA-Based Construction at the Nanoscale: Emerging Trends and Applications. Nanotechnology 2018, 29 (6), 062001 DOI: 10.1088/1361-6528/aaa120There is no corresponding record for this reference.
- 28Chandrasekaran, A. R.; Levchenko, O. DNA Nanocages. Chem. Mater. 2016, 28 (16), 5569– 5581, DOI: 10.1021/acs.chemmater.6b02546There is no corresponding record for this reference.
- 29Goodman, R. P.; Schaap, I. A. T.; Tardin, C. F.; Erben, C. M.; Berry, R. M.; Schmidt, C. F.; Turberfield, A. J. Rapid Chiral Assembly of Rigid DNA Building Blocks for Molecular Nanofabrication. Science 2005, 310 (5754), 1661– 1665, DOI: 10.1126/science.1120367There is no corresponding record for this reference.
- 30Bhatia, D.; Mehtab, S.; Krishnan, R.; Indi, S. S.; Basu, A.; Krishnan, Y. Icosahedral DNA Nanocapsules by Modular Assembly. Angew. Chem., Int. Ed. 2009, 48 (23), 4134– 4137, DOI: 10.1002/anie.200806000There is no corresponding record for this reference.
- 31Winfree, E.; Liu, F.; Wenzler, L. A.; Seeman, N. C. Design and Self-Assembly of Two-Dimensional DNA Crystals. Nature 1998, 394 (6693), 539– 544, DOI: 10.1038/28998There is no corresponding record for this reference.
- 32Rothemund, P. W. K. Folding DNA to Create Nanoscale Shapes and Patterns. Nature 2006, 440 (7082), 297– 302, DOI: 10.1038/nature04586There is no corresponding record for this reference.
- 33Ke, Y.; Ong, L. L.; Shih, W. M.; Yin, P. Three-Dimensional Structures Self-Assembled from DNA Bricks. Science 2012, 338 (6111), 1177– 1183, DOI: 10.1126/science.1227268There is no corresponding record for this reference.
- 34Madsen, M.; Gothelf, K. V. Chemistries for DNA Nanotechnology. Chem. Rev. 2019, 119 (10), 6384– 6458, DOI: 10.1021/acs.chemrev.8b00570There is no corresponding record for this reference.
- 35Pinheiro, V. B.; Holliger, P. Towards XNA Nanotechnology: New Materials from Synthetic Genetic Polymers. Trends Biotechnol. 2014, 32 (6), 321– 328, DOI: 10.1016/j.tibtech.2014.03.010There is no corresponding record for this reference.
- 36Goodman, R. P.; Berry, R. M.; Turberfield, A. J. The Single-Step Synthesis of a DNA Tetrahedron. Chem. Commun. 2004, (No. 12), 1372– 1373, DOI: 10.1039/B402293AThere is no corresponding record for this reference.
- 37Huang, J.; Chakraborty, A.; Tadepalli, L. S.; Paul, A. Adoption of a Tetrahedral DNA Nanostructure as a Multifunctional Biomaterial for Drug Delivery. ACS Pharmacol. Transl. Sci. 2024, 7 (8), 2204– 2214, DOI: 10.1021/acsptsci.4c00308There is no corresponding record for this reference.
- 38Walsh, A. S.; Yin, H.; Erben, C. M.; Wood, M. J. A.; Turberfield, A. J. DNA Cage Delivery to Mammalian Cells. ACS Nano 2011, 5 (7), 5427– 5432, DOI: 10.1021/nn2005574There is no corresponding record for this reference.
- 39Vilcapoma, J.; Patel, A.; Chandrasekaran, A. R.; Halvorsen, K. The Role of Size in Biostability of DNA Tetrahedra. Chem. Commun. 2023, 59 (34), 5083– 5085, DOI: 10.1039/D3CC01123BThere is no corresponding record for this reference.
- 40Rajwar, A.; Shetty, S. R.; Vaswani, P.; Morya, V.; Barai, A.; Sen, S.; Sonawane, M.; Bhatia, D. Geometry of a DNA Nanostructure Influences Its Endocytosis: Cellular Study on 2D, 3D, and in Vivo Systems. ACS Nano 2022, 16 (7), 10496– 10508, DOI: 10.1021/acsnano.2c01382There is no corresponding record for this reference.
- 41Xia, K.; Kong, H.; Cui, Y.; Ren, N.; Li, Q.; Ma, J.; Cui, R.; Zhang, Y.; Shi, J.; Li, Q.; Lv, M.; Sun, Y.; Wang, L.; Li, J.; Zhu, Y. Systematic Study in Mammalian Cells Showing No Adverse Response to Tetrahedral DNA Nanostructure. ACS Appl. Mater. Interfaces 2018, 10 (18), 15442– 15448, DOI: 10.1021/acsami.8b02626There is no corresponding record for this reference.
- 42Gao, Y.; Chen, X.; Tian, T.; Zhang, T.; Gao, S.; Zhang, X.; Yao, Y.; Lin, Y.; Cai, X. A Lysosome-Activated Tetrahedral Nanobox for Encapsulated siRNA Delivery. Adv. Mater. 2022, 34 (46), 2201731 DOI: 10.1002/adma.202201731There is no corresponding record for this reference.
- 43Wang, Z.; Song, L.; Liu, Q.; Tian, R.; Shang, Y.; Liu, F.; Liu, S.; Zhao, S.; Han, Z.; Sun, J.; Jiang, Q.; Ding, B. A Tubular DNA Nanodevice as a siRNA/Chemo-Drug Co-delivery Vehicle for Combined Cancer Therapy. Angew. Chem. Int. Ed. 2021, 60 (5), 2594– 2598, DOI: 10.1002/anie.202009842There is no corresponding record for this reference.
- 44Ding, F.; Mou, Q.; Ma, Y.; Pan, G.; Guo, Y.; Tong, G.; Choi, C. H. J.; Zhu, X.; Zhang, C. A Crosslinked Nucleic Acid Nanogel for Effective siRNA Delivery and Antitumor Therapy. Angew. Chem., Int. Ed. 2018, 57 (12), 3064– 3068, DOI: 10.1002/anie.201711242There is no corresponding record for this reference.
- 45Frtús, A.; Smolková, B.; Uzhytchak, M.; Lunova, M.; Jirsa, M.; Henry, S. J. W.; Dejneka, A.; Stephanopoulos, N.; Lunov, O. The Interactions between DNA Nanostructures and Cells: A Critical Overview from a Cell Biology Perspective. Acta Biomater. 2022, 146, 10– 22, DOI: 10.1016/j.actbio.2022.04.046There is no corresponding record for this reference.
- 46Chandrasekaran, A. R. Nuclease Resistance of DNA Nanostructures. Nat. Rev. Chem. 2021, 5 (4), 225– 239, DOI: 10.1038/s41570-021-00251-yThere is no corresponding record for this reference.
- 47Keum, J.-W.; Bermudez, H. Enhanced Resistance of DNA Nanostructures to Enzymatic Digestion. Chem. Commun. 2009, 2009, 7036– 7038, DOI: 10.1039/b917661fThere is no corresponding record for this reference.
- 48Yue, Z.; Yang, Y.; Nie, L.; Sun, Y.; Wang, Q.; Lin, Y.; Gao, Y.; Cai, X. A Binary siRNA-Loaded Tetrahedral DNA Nanobox for Synergetic Anti-Aging Therapy. Small 2025, 21 (18), 2408323 DOI: 10.1002/smll.202408323There is no corresponding record for this reference.
- 49Thai, H. B. D.; Kim, K.-R.; Hong, K. T.; Voitsitskyi, T.; Lee, J.-S.; Mao, C.; Ahn, D.-R. Kidney-Targeted Cytosolic Delivery of siRNA Using a Small-Sized Mirror DNA Tetrahedron for Enhanced Potency. ACS Cent. Sci. 2020, 6 (12), 2250– 2258, DOI: 10.1021/acscentsci.0c00763There is no corresponding record for this reference.
- 50Skaanning, M. K.; Bønnelykke, J.; Nijenhuis, M. A. D.; Samanta, A.; Smidt, J. M.; Gothelf, K. V. Self-Assembly of Ultrasmall 3D Architectures of (l)-Acyclic Threoninol Nucleic Acids with High Thermal and Serum Stability. J. Am. Chem. Soc. 2024, 146 (29), 20141– 20146, DOI: 10.1021/jacs.4c04919There is no corresponding record for this reference.
- 51Madhanagopal, B. R.; Patel, A.; Talbot, H.; Geary, J. A.; Ganesh, K. N.; Chandrasekaran, A. R. Peptide Nucleic Acid (PNA) and DNA Hybrid Three-Way Junctions and Mesojunctions bioRxiv 2025 DOI: 10.1101/2025.05.15.653195 .There is no corresponding record for this reference.
- 52Qian, H.; Wang, D.; He, B.; Liu, Q.; Xu, Y.; Wu, D.; Chen, C.; Zhang, W.; Leong, D. T.; Wang, G. Assembling Defined DNA Nanostructures with Anticancer Drugs: A Metformin/DNA Complex Nanoplatform with a Synergistic Antitumor Effect for KRAS-Mutated Lung Cancer Therapy. NPG Asia Mater. 2022, 14 (1), 81 DOI: 10.1038/s41427-022-00427-yThere is no corresponding record for this reference.
- 53Wamhoff, E.-C.; Romanov, A.; Huang, H.; Read, B. J.; Ginsburg, E.; Knappe, G. A.; Kim, H. M.; Farrell, N. P.; Irvine, D. J.; Bathe, M. Controlling Nuclease Degradation of Wireframe DNA Origami with Minor Groove Binders. ACS Nano 2022, 16 (6), 8954– 8966, DOI: 10.1021/acsnano.1c11575There is no corresponding record for this reference.
- 54Rodriguez, A.; Gandavadi, D.; Mathivanan, J.; Song, T.; Madhanagopal, B. R.; Talbot, H.; Sheng, J.; Wang, X.; Chandrasekaran, A. R. Self-Assembly of DNA Nanostructures in Different Cations. Small 2023, 19 (39), 2300040 DOI: 10.1002/smll.202300040There is no corresponding record for this reference.
- 55Gandavadi, D.; Talbot, H.; Dwivedy, A.; Umrao, S.; Rodriguez, A.; Cho, H.; Zheng, M.; Chandrasekaran, A. R.; Wang, X. DNA Nanostructure Self-Assembly in an Aqueous Ionic Liquid Solution with Enhanced Stability and Target Binding Affinity. J. Am. Chem. Soc. 2025, 147 (46), 42635– 42646, DOI: 10.1021/jacs.5c13969There is no corresponding record for this reference.
- 56Chandrasekaran, A. R.; Vilcapoma, J.; Dey, P.; Wong-Deyrup, S. W.; Dey, B. K.; Halvorsen, K. Exceptional Nuclease Resistance of Paranemic Crossover (PX) DNA and Crossover-Dependent Biostability of DNA Motifs. J. Am. Chem. Soc. 2020, 142 (14), 6814– 6821, DOI: 10.1021/jacs.0c02211There is no corresponding record for this reference.
- 57Madhanagopal, B. R.; Talbot, H.; Rodriguez, A.; Louis, J. M.; Zeghal, H.; Vangaveti, S.; Reddy, K.; Chandrasekaran, A. R. The Unusual Structural Properties and Potential Biological Relevance of Switchback DNA. Nat. Commun. 2024, 15 (1), 6636 DOI: 10.1038/s41467-024-50348-3There is no corresponding record for this reference.
- 58Perrault, S. D.; Shih, W. M. Virus-Inspired Membrane Encapsulation of DNA Nanostructures To Achieve In Vivo Stability. ACS Nano 2014, 8 (5), 5132– 5140, DOI: 10.1021/nn5011914There is no corresponding record for this reference.
- 59Auvinen, H.; Zhang, H.; Nonappa; Kopilow, A.; Niemelä, E. H.; Nummelin, S.; Correia, A.; Santos, H. A.; Linko, V.; Kostiainen, M. A. Protein Coating of DNA Nanostructures for Enhanced Stability and Immunocompatibility. Adv. Healthcare Mater. 2017, 6 (18), 1700692 DOI: 10.1002/adhm.201700692There is no corresponding record for this reference.
- 60Seitz, I.; McNeale, D.; Sainsbury, F.; Linko, V.; Kostiainen, M. A. Modular Virus Capsid Coatings for Biocatalytic DNA Origami Nanoreactors. ACS Nano 2025, 19 (41), 36465– 36477, DOI: 10.1021/acsnano.5c10734There is no corresponding record for this reference.
- 61Ponnuswamy, N.; Bastings, M. M. C.; Nathwani, B.; Ryu, J. H.; Chou, L. Y. T.; Vinther, M.; Li, W. A.; Anastassacos, F. M.; Mooney, D. J.; Shih, W. M. Oligolysine-Based Coating Protects DNA Nanostructures from Low-Salt Denaturation and Nuclease Degradation. Nat. Commun. 2017, 8 (1), 15654 DOI: 10.1038/ncomms15654There is no corresponding record for this reference.
- 62Agarwal, N. P.; Matthies, M.; Gür, F. N.; Osada, K.; Schmidt, T. L. Block Copolymer Micellization as a Protection Strategy for DNA Origami. Angew. Chem., Int. Ed. 2017, 56 (20), 5460– 5464, DOI: 10.1002/anie.201608873There is no corresponding record for this reference.
- 63Youssef, S.; Tsang, E.; Samanta, A.; Kumar, V.; Gothelf, K. V. Reversible Protection and Targeted Delivery of DNA Origami with a Disulfide-Containing Cationic Polymer. Small 2024, 20 (10), 2301058 DOI: 10.1002/smll.202301058There is no corresponding record for this reference.
- 64Schöttler, S.; Landfester, K.; Mailänder, V. Controlling the Stealth Effect of Nanocarriers through Understanding the Protein Corona. Angew. Chem., Int. Ed. 2016, 55 (31), 8806– 8815, DOI: 10.1002/anie.201602233There is no corresponding record for this reference.
- 65Huzar, J.; Coreas, R.; Landry, M. P.; Tikhomirov, G. AI-Based Prediction of Protein Corona Composition on DNA Nanostructures. ACS Nano 2025, 19 (4), 4333– 4345, DOI: 10.1021/acsnano.4c12259There is no corresponding record for this reference.
- 66Kim, K.-R.; Kim, J.; Back, J. H.; Lee, J. E.; Ahn, D.-R. Cholesterol-Mediated Seeding of Protein Corona on DNA Nanostructures for Targeted Delivery of Oligonucleotide Therapeutics to Treat Liver Fibrosis. ACS Nano 2022, 16 (5), 7331– 7343, DOI: 10.1021/acsnano.1c08508There is no corresponding record for this reference.
- 67Rodríguez-Franco, H. J.; Hendrickx, P. B. M.; Bastings, M. M. C. Tailoring DNA Origami Protection: A Study of Oligolysine-PEG Coatings for Improved Colloidal, Structural, and Functional Integrity. ACS Polym. Au 2025, 5 (1), 35– 44, DOI: 10.1021/acspolymersau.4c00085There is no corresponding record for this reference.
- 68Rodríguez-Franco, H. J.; Weiden, J.; Bastings, M. M. C. Stabilizing Polymer Coatings Alter the Protein Corona of DNA Origami and Can Be Engineered to Bias the Cellular Uptake. ACS Polym. Au 2023, 3 (4), 344– 353, DOI: 10.1021/acspolymersau.3c00009There is no corresponding record for this reference.
- 69Tang, X.; Zhai, T.; Li, T.; Jin, Y.; Lei, D.; Zhu, C.; Qu, L.; Li, Y.; Wang, Y.; Gu, H.; Fang, B. Dimensional Control of DNA Nanostructures Enhances Cellular Uptake and Guides Tissue-Regenerative Responses. J. Nanobiotechnol. 2025, 23 (1), 615 DOI: 10.1186/s12951-025-03707-1There is no corresponding record for this reference.
- 70Wang, P.; Rahman, M. A.; Zhao, Z.; Weiss, K.; Zhang, C.; Chen, Z.; Hurwitz, S. J.; Chen, Z. G.; Shin, D. M.; Ke, Y. Visualization of the Cellular Uptake and Trafficking of DNA Origami Nanostructures in Cancer Cells. J. Am. Chem. Soc. 2018, 140 (7), 2478– 2484, DOI: 10.1021/jacs.7b09024There is no corresponding record for this reference.
- 71Gopinath, S. C. B.; Lakshmipriya, T.; Chen, Y.; Arshad, M. K. M.; Kerishnan, J. P.; Ruslinda, A. R.; Al-Douri, Y.; Voon, C. H.; Hashim, U. Cell-Targeting Aptamers Act as Intracellular Delivery Vehicles. Appl. Microbiol. Biotechnol. 2016, 100 (16), 6955– 6969, DOI: 10.1007/s00253-016-7686-2There is no corresponding record for this reference.
- 72Xia, Z.; Wang, P.; Liu, X.; Liu, T.; Yan, Y.; Yan, J.; Zhong, J.; Sun, G.; He, D. Tumor-Penetrating Peptide-Modified DNA Tetrahedron for Targeting Drug Delivery. Biochemistry 2016, 55 (9), 1326– 1331, DOI: 10.1021/acs.biochem.5b01181There is no corresponding record for this reference.
- 73Nagaraj, H.; Lehot, V.; Nasim, N.; Cicek, Y. A.; Goswami, R.; Jeon, T.; Rotello, V. M. Breaking the Cellular Delivery Bottleneck: Recent Developments in Direct Cytosolic Delivery of Biologics. RSC Pharm. 2025, 2 (5), 850– 864, DOI: 10.1039/D5PM00129CThere is no corresponding record for this reference.
- 74Dowdy, S. F.; Setten, R. L.; Cui, X.-S.; Jadhav, S. G. Delivery of RNA Therapeutics: The Great Endosomal Escape!. Nucleic Acid Ther. 2022, 32 (5), 361– 368, DOI: 10.1089/nat.2022.0004There is no corresponding record for this reference.
- 75Lönn, P.; Kacsinta, A. D.; Cui, X.-S.; Hamil, A. S.; Kaulich, M.; Gogoi, K.; Dowdy, S. F. Enhancing Endosomal Escape for Intracellular Delivery of Macromolecular Biologic Therapeutics. Sci. Rep. 2016, 6 (1), 32301 DOI: 10.1038/srep32301There is no corresponding record for this reference.
- 76Liang, X.-h.; Sun, H.; Hsu, C.-W.; Nichols, J. G.; Vickers, T. A.; De Hoyos, C. L.; Crooke, S. T. Golgi-Endosome Transport Mediated by M6PR Facilitates Release of Antisense Oligonucleotides from Endosomes. Nucleic Acids Res. 2020, 48 (3), 1372– 1391, DOI: 10.1093/nar/gkz1171There is no corresponding record for this reference.
- 77Hwang, H. S.; Hu, J.; Na, K.; Bae, Y. H. Role of Polymeric Endosomolytic Agents in Gene Transfection: A Comparative Study of Poly(l-Lysine) Grafted with Monomeric l-Histidine Analogue and Poly(l-Histidine). Biomacromolecules 2014, 15 (10), 3577– 3586, DOI: 10.1021/bm500843rThere is no corresponding record for this reference.
- 78Sun, W.; Ji, W.; Hall, J. M.; Hu, Q.; Wang, C.; Beisel, C. L.; Gu, Z. Self-Assembled DNA Nanoclews for the Efficient Delivery of CRISPR–Cas9 for Genome Editing. Angew. Chem., Int. Ed. 2015, 54 (41), 12029– 12033, DOI: 10.1002/anie.201506030There is no corresponding record for this reference.
- 79Liang, L.; Li, J.; Li, Q.; Huang, Q.; Shi, J.; Yan, H.; Fan, C. Single-Particle Tracking and Modulation of Cell Entry Pathways of a Tetrahedral DNA Nanostructure in Live Cells. Angew. Chem., Int. Ed. 2014, 53 (30), 7745– 7750, DOI: 10.1002/anie.201403236There is no corresponding record for this reference.
- 80Wang, D.; Liu, Q.; Wu, D.; He, B.; Li, J.; Mao, C.; Wang, G.; Qian, H. Isothermal Self-Assembly of Spermidine–DNA Nanostructure Complex as a Functional Platform for Cancer Therapy. ACS Appl. Mater. Interfaces 2018, 10 (18), 15504– 15516, DOI: 10.1021/acsami.8b03464There is no corresponding record for this reference.
- 81Smolková, B.; MacCulloch, T.; Rockwood, T. F.; Liu, M.; Henry, S. J. W.; Frtús, A.; Uzhytchak, M.; Lunova, M.; Hof, M.; Jurkiewicz, P.; Dejneka, A.; Stephanopoulos, N.; Lunov, O. Protein Corona Inhibits Endosomal Escape of Functionalized DNA Nanostructures in Living Cells. ACS Appl. Mater. Interfaces 2021, 13 (39), 46375– 46390, DOI: 10.1021/acsami.1c14401There is no corresponding record for this reference.
- 82Han, X.; Zhang, H.; Butowska, K.; Swingle, K. L.; Alameh, M.-G.; Weissman, D.; Mitchell, M. J. An Ionizable Lipid Toolbox for RNA Delivery. Nat. Commun. 2021, 12 (1), 7233 DOI: 10.1038/s41467-021-27493-0There is no corresponding record for this reference.
- 83Huang, P.; Qi, M.; Chen, C.; Xu, F.; Li, S.; Xu, Q.; Pan, H.; Wang, Y.; Yu, C.; Zhang, S.; Zhou, Y. Asymmetric Vesicles Self-Assembled by Amphiphilic Sequence-Controlled Polymers. ACS Macro Lett. 2021, 10 (7), 894– 900, DOI: 10.1021/acsmacrolett.1c00301There is no corresponding record for this reference.
- 84Yu, L.; Xu, Y.; Al-Amin, M.; Jiang, S.; Sample, M.; Prasad, A.; Stephanopoulos, N.; Šulc, P.; Yan, H. CytoDirect: A Nucleic Acid Nanodevice for Specific and Efficient Delivery of Functional Payloads to the Cytoplasm. J. Am. Chem. Soc. 2023, 145 (50), 27336– 27347, DOI: 10.1021/jacs.3c07491There is no corresponding record for this reference.
- 85Cognet, M.; Renno, G.; Rose, N.; Zhang, Y.; Josso, P.; Moreno, J.; Ren, X.; Bouffard, J.; Saidjalolov, S.; Sakai, N.; Matile, S. Cell-Penetrating Poly(Disulfide)s. Helv. Chim. Acta 2025, 108 (12), e00129 DOI: 10.1002/hlca.202500129There is no corresponding record for this reference.
- 86Wang, W.; Chopra, B.; Walawalkar, V.; Liang, Z.; Adams, R.; Deserno, M.; Ren, X.; Taylor, R. E. Cell–Surface Binding of DNA Nanostructures for Enhanced Intracellular and Intranuclear Delivery. ACS Appl. Mater. Interfaces 2024, 16 (13), 15783– 15797, DOI: 10.1021/acsami.3c18068There is no corresponding record for this reference.
- 87Samanta, A.; Malle, M. G.; Tsang, E.; Omer, M.; Skaanning, M. K.; Youssef, S.; Kjems, J.; Gothelf, K. V. Bacteriophage-Mimetic DNA Origami Needle for Targeted Membrane Penetration and Cytosolic Cargo Delivery. Adv. Sci. 2026, 13 (10), e12844 DOI: 10.1002/advs.202512844There is no corresponding record for this reference.
- 88Du, R. R.; Cedrone, E.; Romanov, A.; Falkovich, R.; Dobrovolskaia, M. A.; Bathe, M. Innate Immune Stimulation Using 3D Wireframe DNA Origami. ACS Nano 2022, 16 (12), 20340– 20352, DOI: 10.1021/acsnano.2c06275There is no corresponding record for this reference.
- 89Lucas, C. R.; Halley, P. D.; Chowdury, A. A.; Harrington, B. K.; Beaver, L.; Lapalombella, R.; Johnson, A. J.; Hertlein, E. K.; Phelps, M. A.; Byrd, J. C.; Castro, C. E. DNA Origami Nanostructures Elicit Dose-Dependent Immunogenicity and Are Nontoxic up to High Doses In Vivo. Small 2022, 18 (26), 2108063 DOI: 10.1002/smll.202108063There is no corresponding record for this reference.
- 90Arulkumaran, N.; Lanphere, C.; Gaupp, C.; Burns, J. R.; Singer, M.; Howorka, S. DNA Nanodevices with Selective Immune Cell Interaction and Function. ACS Nano 2021, 15 (3), 4394– 4404, DOI: 10.1021/acsnano.0c07915There is no corresponding record for this reference.
- 91Rodriguez, A.; Madhanagopal, B. R.; Sarkar, K.; Nowzari, Z.; Mathivanan, J.; Talbot, H.; Patel, A.; Morya, V.; Halvorsen, K.; Vangaveti, S.; Berglund, J. A.; Chandrasekaran, A. R. Counterions Influence the Isothermal Self-Assembly of DNA Nanostructures. Sc. Adv. 2025, 11 (11), eadu7366 DOI: 10.1126/sciadv.adu7366There is no corresponding record for this reference.
- 92Guo, Y.; Huang, Y.; Liu, M.; Liu, J.; Liu, J.; Yang, D. Evaluation of Pharmacokinetics, Immunogenicity, and Immunotoxicity of DNA Tetrahedral and DNA Polymeric Nanostructures. Small Methods 2025, 9 (6), 2401007 DOI: 10.1002/smtd.202401007There is no corresponding record for this reference.
- 93Jiang, D.; Ge, Z.; Im, H.-J.; England, C. G.; Ni, D.; Hou, J.; Zhang, L.; Kutyreff, C. J.; Yan, Y.; Liu, Y.; Cho, S. Y.; Engle, J. W.; Shi, J.; Huang, P.; Fan, C.; Yan, H.; Cai, W. DNA Origami Nanostructures Can Exhibit Preferential Renal Uptake and Alleviate Acute Kidney Injury. Nat. Biomed. Eng. 2018, 2 (11), 865– 877, DOI: 10.1038/s41551-018-0317-8There is no corresponding record for this reference.
- 94Wang, Y.; Baars, I.; Berzina, I.; Rocamonde-Lago, I.; Shen, B.; Yang, Y.; Lolaico, M.; Waldvogel, J.; Smyrlaki, I.; Zhu, K.; Harris, R. A.; Högberg, B. A DNA Robotic Switch with Regulated Autonomous Display of Cytotoxic Ligand Nanopatterns. Nat. Nanotechnol. 2024, 19 (9), 1366– 1374, DOI: 10.1038/s41565-024-01676-4There is no corresponding record for this reference.
- 95Li, L.; Yin, J.; Ma, W.; Tang, L.; Zou, J.; Yang, L.; Du, T.; Zhao, Y.; Wang, L.; Yang, Z.; Fan, C.; Chao, J.; Chen, X. A DNA Origami Device Spatially Controls CD95 Signalling to Induce Immune Tolerance in Rheumatoid Arthritis. Nat. Mater. 2024, 23 (7), 993– 1001, DOI: 10.1038/s41563-024-01865-5There is no corresponding record for this reference.
- 96Zeng, Y. C.; Young, O. J.; Wintersinger, C. M.; Anastassacos, F. M.; MacDonald, J. I.; Isinelli, G.; Dellacherie, M. O.; Sobral, M.; Bai, H.; Graveline, A. R.; Vernet, A.; Sanchez, M.; Mulligan, K.; Choi, Y.; Ferrante, T. C.; Keskin, D. B.; Fell, G. G.; Neuberg, D.; Wu, C. J.; Mooney, D. J.; Kwon, I. C.; Ryu, J. H.; Shih, W. M. Fine Tuning of CpG Spatial Distribution with DNA Origami for Improved Cancer Vaccination. Nat. Nanotechnol. 2024, 19 (7), 1055– 1065, DOI: 10.1038/s41565-024-01615-3There is no corresponding record for this reference.
- 97Zeng, Y. C.; Young, O. J.; Xiong, Q.; Si, L.; Ku, M. W.; Bernier, S. G.; Dembele, H.; Isinelli, G.; Gilboa, T.; Swank, Z.; Seok, S. H.; Rajwar, A.; Jiang, A.; Zhai, Y.; Williams, L. D.; Hellman, C. A.; Wintersinger, C. M.; Graveline, A. R.; Vernet, A.; Sanchez, M.; Bardales, S.; Tomaras, G. D.; Ryu, J. H.; Kwon, I. C.; Goyal, G.; Ingber, D. E.; Shih, W. M. DNA Origami Vaccine Nanoparticles Improve Humoral and Cellular Immune Responses to Infectious Diseases. Nat. Biomed. Eng. 2026, 1– 18, DOI: 10.1038/s41551-026-01614-wThere is no corresponding record for this reference.
- 98Glassman, P. M.; Muzykantov, V. R. Pharmacokinetic and Pharmacodynamic Properties of Drug Delivery Systems. J. Pharmacol. Exp. Ther. 2019, 370 (3), 570– 580, DOI: 10.1124/jpet.119.257113There is no corresponding record for this reference.
- 99Chen, L.; Bosmajian, C.; Woo, S. Mechanistic Intracellular PK/PD Modeling to Inform Development Strategies for Small Interfering RNA Therapeutics. Mol. Ther. Nucleic Acids 2025, 36 (2), 102516 DOI: 10.1016/j.omtn.2025.102516There is no corresponding record for this reference.
- 100Wang, Y.; Rocamonde-Lago, I.; Waldvogel, J.; Shen, B.; Wu, Y.-C.; Zhu, J.; Zang, S.; Jia, Y.; Baars, I.; Kloosterman, A.; Hoffecker, I. T.; Wu, M.-R.; He, Q.; Högberg, B. Resolving DNA Origami Structural Integrity and Pharmacokinetics in Vivo. Nat. Nanotechnol. 2026, 21 (2), 268– 276, DOI: 10.1038/s41565-025-02091-zThere is no corresponding record for this reference.
- 101Li, S.; Sun, Y.; Tian, T.; Qin, X.; Lin, S.; Zhang, T.; Zhang, Q.; Zhou, M.; Zhang, X.; Zhou, Y.; Zhao, H.; Zhu, B.; Cai, X. MicroRNA-214–3p Modified Tetrahedral Framework Nucleic Acids Target Survivin to Induce Tumour Cell Apoptosis. Cell Proliferation 2020, 53 (1), e12708 DOI: 10.1111/cpr.12708There is no corresponding record for this reference.
- 102Wei, M.; Li, S.; Yang, Z.; Cheng, C.; Li, T.; Le, W. Tetrahedral DNA Nanostructures Functionalized by Multivalent microRNA132 Antisense Oligonucleotides Promote the Differentiation of Mouse Embryonic Stem Cells into Dopaminergic Neurons. Nanomed.: Nanotechnol., Biol. Med. 2021, 34, 102375 DOI: 10.1016/j.nano.2021.102375There is no corresponding record for this reference.
- 103Su, J.; Wu, F.; Xia, H.; Wu, Y.; Liu, S. Accurate Cancer Cell Identification and microRNA Silencing Induced Therapy Using Tailored DNA Tetrahedron Nanostructures. Chem. Sci. 2020, 11 (1), 80– 86, DOI: 10.1039/C9SC04823EThere is no corresponding record for this reference.
- 104Kim, K.-R.; Jegal, H.; Kim, J.; Ahn, D.-R. A Self-Assembled DNA Tetrahedron as a Carrier for in Vivo Liver-Specific Delivery of siRNA. Biomater. Sci. 2020, 8 (2), 586– 590, DOI: 10.1039/C9BM01769KThere is no corresponding record for this reference.
- 105Moreno, P. M. D.; Cortinhas, J.; Martins, A. S.; Pêgo, A. P. Engineering a Novel Self-Assembled Multi-siRNA Nanocaged Architecture with Controlled Enzyme-Mediated siRNA Release. ACS Appl. Mater. Interfaces 2022, 14 (51), 56483– 56497, DOI: 10.1021/acsami.2c15086There is no corresponding record for this reference.
- 106Zhang, T.; Li, R.; Wang, Z.; Zhou, Y.; Zhou, Y.; Chen, X.; Peng, C.; Jiang, Y.; Tong, N.; Li, W. Inflammation-Specific DNA Origami Nanodevice for Delivery of siRNAs to Treat Ulcerative Colitis. Nat. Commun. 2026, 17 (1), 495 DOI: 10.1038/s41467-025-67183-9There is no corresponding record for this reference.
- 107Bujold, K. E.; Hsu, J. C. C.; Sleiman, H. F. Optimized DNA “Nanosuitcases” for Encapsulation and Conditional Release of siRNA. J. Am. Chem. Soc. 2016, 138 (42), 14030– 14038, DOI: 10.1021/jacs.6b08369There is no corresponding record for this reference.
- 108Zhang, H.; Demirer, G. S.; Zhang, H.; Ye, T.; Goh, N. S.; Aditham, A. J.; Cunningham, F. J.; Fan, C.; Landry, M. P. DNA Nanostructures Coordinate Gene Silencing in Mature Plants. Proc. Natl. Acad. Sci. U.S.A. 2019, 116 (15), 7543– 7548, DOI: 10.1073/pnas.1818290116There is no corresponding record for this reference.
- 109Li, C.; Lin, W.; Wang, W.; Wu, J.; Luo, S.; Chen, L.; Wu, R.; Shen, Z.; Wu, Z.-S. Folding an RCA Scaffold into an Intelligent Coiled Nanosnake for Precise/Synergistic RNAi-/Chemotherapy of Cancer. Anal. Chem. 2025, 97 (2), 1107– 1116, DOI: 10.1021/acs.analchem.4c03437There is no corresponding record for this reference.
- 110Jensen, S. A.; Day, E. S.; Ko, C. H.; Hurley, L. A.; Luciano, J. P.; Kouri, F. M.; Merkel, T. J.; Luthi, A. J.; Patel, P. C.; Cutler, J. I.; Daniel, W. L.; Scott, A. W.; Rotz, M. W.; Meade, T. J.; Giljohann, D. A.; Mirkin, C. A.; Stegh, A. H. Spherical Nucleic Acid Nanoparticle Conjugates as an RNAi-Based Therapy for Glioblastoma. Sci. Transl. Med. 2013, 5 (209), 209ra152 DOI: 10.1126/scitranslmed.3006839There is no corresponding record for this reference.
- 111Kumthekar, P.; Ko, C. H.; Paunesku, T.; Dixit, K.; Sonabend, A. M.; Bloch, O.; Tate, M.; Schwartz, M.; Zuckerman, L.; Lezon, R.; Lukas, R. V.; Jovanovic, B.; McCortney, K.; Colman, H.; Chen, S.; Lai, B.; Antipova, O.; Deng, J.; Li, L.; Tommasini-Ghelfi, S.; Hurley, L. A.; Unruh, D.; Sharma, N. V.; Kandpal, M.; Kouri, F. M.; Davuluri, R. V.; Brat, D. J.; Muzzio, M.; Glass, M.; Vijayakumar, V.; Heidel, J.; Giles, F. J.; Adams, A. K.; James, C. D.; Woloschak, G. E.; Horbinski, C.; Stegh, A. H. A First-in-Human Phase 0 Clinical Study of RNA Interference–Based Spherical Nucleic Acids in Patients with Recurrent Glioblastoma. Sci. Transl. Med. 2021, 13 (584), eabb3945 DOI: 10.1126/scitranslmed.abb3945There is no corresponding record for this reference.
- 112Chen, X.; Tang, J.; Shuai, W.; Meng, J.; Feng, J.; Han, Z. Macrophage Polarization and Its Role in the Pathogenesis of Acute Lung Injury/Acute Respiratory Distress Syndrome. Inflamm. Res. 2020, 69 (9), 883– 895, DOI: 10.1007/s00011-020-01378-2There is no corresponding record for this reference.
- 113Huang, C.; You, Q.; Xu, J.; Wu, D.; Chen, H.; Guo, Y.; Xu, J.; Hu, M.; Qian, H. An mTOR siRNA-Loaded Spermidine/DNA Tetrahedron Nanoplatform with a Synergistic Anti-Inflammatory Effect on Acute Lung Injury. Adv. Healthcare Mater. 2022, 11 (11), 2200008 DOI: 10.1002/adhm.202200008There is no corresponding record for this reference.
- 114Li, C.; Zhao, W.; Hu, Z.; Yu, H. Cholesterol-Modified DNA Nanostructures Serve as Effective Non-Viral Carriers for Delivering siRNA to the Kidneys to Prevent Acute Kidney Injury. Small 2024, 20 (30), 2311690 DOI: 10.1002/smll.202311690There is no corresponding record for this reference.
- 115Kim, K.-R.; Kim, H. Y.; Lee, Y.-D.; Ha, J. S.; Kang, J. H.; Jeong, H.; Bang, D.; Ko, Y. T.; Kim, S.; Lee, H.; Ahn, D.-R. Self-Assembled Mirror DNA Nanostructures for Tumor-Specific Delivery of Anticancer Drugs. J. Controlled Release 2016, 243, 121– 131, DOI: 10.1016/j.jconrel.2016.10.015There is no corresponding record for this reference.
- 116Jiang, D.; Sun, Y.; Li, J.; Li, Q.; Lv, M.; Zhu, B.; Tian, T.; Cheng, D.; Xia, J.; Zhang, L.; Wang, L.; Huang, Q.; Shi, J.; Fan, C. Multiple-Armed Tetrahedral DNA Nanostructures for Tumor-Targeting, Dual-Modality in Vivo Imaging. ACS Appl. Mater. Interfaces 2016, 8 (7), 4378– 4384, DOI: 10.1021/acsami.5b10792There is no corresponding record for this reference.
- 117Zhou, X.; Yu, M.; Ma, L.; Fu, J.; Guo, J.; Lei, J.; Fu, Z.; Fu, Y.; Zhang, Q.; Zhang, C.-Y.; Chen, X. In Vivo Self-Assembled siRNA as a Modality for Combination Therapy of Ulcerative Colitis. Nat. Commun. 2022, 13 (1), 5700 DOI: 10.1038/s41467-022-33436-0There is no corresponding record for this reference.
- 118Dong, H.; Song, G.; Ma, D.; Wang, T.; Jing, S.; Yang, H.; Tao, Y.; Tang, Y.; Shi, Y.; Dai, Z.; Zhu, J.; Liu, T.; Wang, B.; Leng, X.; Shen, X.; Zhu, C.; Zhao, Y. Improved Antiviral Activity of Classical Swine Fever Virus-Targeted siRNA by Tetrahedral Framework Nucleic Acid-Enhanced Delivery. ACS Appl. Mater. Interfaces 2021, 13 (25), 29416– 29423, DOI: 10.1021/acsami.1c08143There is no corresponding record for this reference.
- 119Fang, Y.; Xue, J.; Gao, S.; Lu, A.; Yang, D.; Jiang, H.; He, Y.; Shi, K. Cleavable PEGylation: A Strategy for Overcoming the “PEG Dilemma” in Efficient Drug Delivery. Drug Delivery 2017, 24 (2), 22– 32, DOI: 10.1080/10717544.2017.1388451There is no corresponding record for this reference.
- 120Ou, M.; Wang, X.-L.; Xu, R.; Chang, C.-W.; Bull, D. A.; Kim, S. W. Novel Biodegradable Poly(Disulfide Amine)s for Gene Delivery with High Efficiency and Low Cytotoxicity. Bioconjugate Chem. 2008, 19 (3), 626– 633, DOI: 10.1021/bc700397xThere is no corresponding record for this reference.
- 121Shi, D.; Beasock, D.; Fessler, A.; Szebeni, J.; Ljubimova, J. Y.; Afonin, K. A.; Dobrovolskaia, M. A. To PEGylate or Not to PEGylate: Immunological Properties of Nanomedicine’s Most Popular Component, Polyethylene Glycol and Its Alternatives. Adv. Drug Delivery Rev. 2022, 180, 114079 DOI: 10.1016/j.addr.2021.114079There is no corresponding record for this reference.
- 122Ju, Y.; Carreño, J. M.; Simon, V.; Dawson, K.; Krammer, F.; Kent, S. J. Impact of Anti-PEG Antibodies Induced by SARS-CoV-2 mRNA Vaccines. Nat. Rev. Immunol. 2023, 23 (3), 135– 136, DOI: 10.1038/s41577-022-00825-xThere is no corresponding record for this reference.
- 123Klipp, A.; Burger, M.; Leroux, J.-C. Get out or Die Trying: Peptide- and Protein-Based Endosomal Escape of RNA Therapeutics. Adv. Drug Delivery Rev. 2023, 200, 115047 DOI: 10.1016/j.addr.2023.115047There is no corresponding record for this reference.
- 124Rhee, W. J.; Bao, G. Slow Non-Specific Accumulation of 2′-Deoxy and 2′-O-Methyl Oligonucleotide Probes at Mitochondria in Live Cells. Nucleic Acids Res. 2010, 38 (9), e109 DOI: 10.1093/nar/gkq050There is no corresponding record for this reference.
- 125Høiberg, H. C.; Sparvath, S. M.; Andersen, V. L.; Kjems, J.; Andersen, E. S. An RNA Origami Octahedron with Intrinsic siRNAs for Potent Gene Knockdown. Biotechnol. J. 2019, 14 (1), 1700634 DOI: 10.1002/biot.201700634There is no corresponding record for this reference.
- 126Wu, X. A.; Choi, C. H. J.; Zhang, C.; Hao, L.; Mirkin, C. A. Intracellular Fate of Spherical Nucleic Acid Nanoparticle Conjugates. J. Am. Chem. Soc. 2014, 136 (21), 7726– 7733, DOI: 10.1021/ja503010aThere is no corresponding record for this reference.
- 127Lacroix, A.; Vengut-Climent, E.; de Rochambeau, D.; Sleiman, H. F. Uptake and Fate of Fluorescently Labeled DNA Nanostructures in Cellular Environments: A Cautionary Tale. ACS Cent. Sci. 2019, 5 (5), 882– 891, DOI: 10.1021/acscentsci.9b00174There is no corresponding record for this reference.
- 128Kozlov, S.; Panigaj, M.; Rebolledo, L.; Bhaskaran, H.; Afonin, K. A. Nucleic Acid Nanoparticles Redefine Traditional Regulatory Terminology: The Blurred Line between Active Pharmaceutical Ingredients and Excipients. ACS Nano Med. 2026, 1 (1), 5– 17, DOI: 10.1021/acsnanomed.5c00070There is no corresponding record for this reference.
- 129Praetorius, F.; Kick, B.; Behler, K. L.; Honemann, M. N.; Weuster-Botz, D.; Dietz, H. Biotechnological Mass Production of DNA Origami. Nature 2017, 552 (7683), 84– 87, DOI: 10.1038/nature24650There is no corresponding record for this reference.
- 130Langlois, N. I.; Clark, H. A. Characterization of DNA Nanostructure Stability by Size Exclusion Chromatography. Anal. Methods 2022, 14 (10), 1006– 1014, DOI: 10.1039/D1AY02146JThere is no corresponding record for this reference.
- 131Halvorsen, K.; Kizer, M. E.; Wang, X.; Chandrasekaran, A. R.; Basanta-Sanchez, M. Shear Dependent LC Purification of an Engineered DNA Nanoswitch and Implications for DNA Origami. Anal. Chem. 2017, 89 (11), 5673– 5677, DOI: 10.1021/acs.analchem.7b00791There is no corresponding record for this reference.
- 132Douglas, S. M.; Bachelet, I.; Church, G. M. A Logic-Gated Nanorobot for Targeted Transport of Molecular Payloads. Science 2012, 335 (6070), 831– 834, DOI: 10.1126/science.1214081There is no corresponding record for this reference.



