Modern RNA Quantification Methods: From RT-qPCR to Advanced MicroscopyClick to copy article linkArticle link copied!
- Tyrese BoddieTyrese BoddieDepartment of Chemistry, University of Alabama at Birmingham, 901 14th Street South, Birmingham, Alabama 35294, United StatesMore by Tyrese Boddie
- Arianna LacenArianna LacenDepartment of Chemistry, University of Alabama at Birmingham, 901 14th Street South, Birmingham, Alabama 35294, United StatesMore by Arianna Lacen
- Hui-Ting Lee*Hui-Ting Lee*Email: [email protected]Department of Chemistry, University of Alabama at Birmingham, 901 14th Street South, Birmingham, Alabama 35294, United StatesMore by Hui-Ting Lee
Abstract
RNA plays a crucial role in gene expression, regulation, protein synthesis, and other cellular functions. The diversity that exists between different RNAs makes information beyond their expression level necessary for understanding more about their complex functions in a cell. Conventional ensemble approaches to RNA quantification have been used extensively to measure the quantity of RNA but lack cellular-level spatial information. This review highlights important contributions that high resolution microscopy has made to RNA quantification and cellular biophysics. Using advanced microscopy for precise localization, real-time tracking, and quantitative measurements of RNA increases our understanding of different disease states, cell- and tissue-specific gene regulation, and cellular architecture.
This publication is licensed under
License Summary*
You are free to share(copy and redistribute) this article in any medium or format and to adapt(remix, transform, and build upon) the material for any purpose, even commercially within the parameters below:
Creative Commons (CC): This is a Creative Commons license.
Attribution (BY): Credit must be given to the creator.
*Disclaimer
This summary highlights only some of the key features and terms of the actual license. It is not a license and has no legal value. Carefully review the actual license before using these materials.
License Summary*
You are free to share(copy and redistribute) this article in any medium or format and to adapt(remix, transform, and build upon) the material for any purpose, even commercially within the parameters below:
Creative Commons (CC): This is a Creative Commons license.
Attribution (BY): Credit must be given to the creator.
*Disclaimer
This summary highlights only some of the key features and terms of the actual license. It is not a license and has no legal value. Carefully review the actual license before using these materials.
License Summary*
You are free to share(copy and redistribute) this article in any medium or format and to adapt(remix, transform, and build upon) the material for any purpose, even commercially within the parameters below:
Creative Commons (CC): This is a Creative Commons license.
Attribution (BY): Credit must be given to the creator.
*Disclaimer
This summary highlights only some of the key features and terms of the actual license. It is not a license and has no legal value. Carefully review the actual license before using these materials.
1. Introduction
2. Ensemble RNA Quantification
2.1. RT-qPCR
2.1.1. Microarray
2.1.2. RNA Sequencing
2.2. Flow Cytometry and Nano Biopsy
3. Image-Based RNA Quantification
3.1. Fixed-Cell Imaging of RNA
3.1.1. Fluorescence in Situ Hybridization (FISH)
3.1.2. smFISH
Figure 1
Figure 1. Common fixed-cell imaging techniques used for RNA quantification. (A) Illustration of smFISH (top) to label GFP RNA in CHO cells. Reproduced from ref (39) under Creative Commons License CC-BY-NC-ND. Copyright 2008 Raj A; et al. Published by Springer Nature. (B) Illustration of RNAscope (right) and an example of RNAscope being used for multicolor detection of β-actin, PLP0 (60S acidic ribosomal protein P0), PPIB (peptidylprolyl isomerase B), and HPRT-1 (hypoxanthine phosphoribosyltransferase 1) (left). Reproduced from ref (44) under Creative Commons License CC-BY-NC-ND. Copyright 2012 Wang, F; et al. Published by Elsevier. (C) Illustration of RCAFISH with the target padlock probe (bottom) to image TK1 mRNA in MCF-7 cells. Adapted from ref (50) under Creative Commons License CC BY 3.0. Copyright 2017 Deng, R.; et al. Published by Royal Society of Chemistry. (D) Illustration of HCR FISH (top) and validation of the technique by detecting EGFP mRNA in wild-type Arabidopsis. Adapted from ref (46) under Creative Commons License CC BY 4.0. Copyright 2023 Huang, T.; et al. Published by Springer Nature. (E) Principle of seqFISH and example images. Reproduced from ref (54) under Creative Commons License CC-BY-NC-ND. Copyright 2014 Lubeck, E.; et al. Published by Springer Nature. (F) MERFISH workflow (left) and images of RNA molecules in an IMR90 cell after each hybridization round. Adapted with permission from ref (59). Copyright 2015 AAAS.
3.1.3. RNAscope
3.1.4. HCR-FISH
3.1.5. RCA-FISH
3.1.6. SeqFISH
3.1.7. MERFISH
3.2. Live-Cell Imaging of RNA
3.2.1. MS2-MCP System
Figure 2
Figure 2. Common live-cell imaging techniques used for RNA quantification. (A) Schematic of MS2-MCP system and MS2-based signal amplification with the suntag system (top) and representative live-cell images of β-actin (bottom). Reporduced from ref (67) under Creative Commons License CC BY 4.0. Copyright 2023 Hu Y.; et al. Published by eLife; (B) Illustration of Molecular beacons for live-cell imaging being used to visualize the transport of native oskar mRNA from a nurse cell to the posterior cortex of the oocyte. Adapted with permission from ref (82). Copyright (2003) National Academy of Sciences, U.S.A. (C) Example of fluorogenic RNA being used to target CXCL1 mRNA after 5 ng/mL TNF-α treatment. Adapted with permission from ref (68). Copyright 2023 American Chemical Society. (D) Example of different dCas12a mutants fused with GFP in the presence of a PAMmer sequence targeting β-actin mRNA in HeLa cells. Reporduced with permission from ref (77). Copyright 2024 American Chemical Society. (E) dCas 13b with different RNA sgRNA aptamers for multicolor imaging of MUC4 and SatIII RNA. Reproduced from ref (79) under the Creative Commons License CC BY-NC 3.0. Copyright 2022 Tang, H.; et al. Published by Royal Society of Chemistry.
3.2.2. Fluorogenic RNAs
3.2.3. CRISPR/Cas Systems
3.2.4. Molecular Beacons
3.3. Image Analysis
3.3.1. Intensity-Based Analysis
3.3.2. Point Pattern Analysis
Figure 3
Figure 3. RNA quantification using STORM, PAINT, and ExM. (A) Fluorophore localization for SMLM reconstruction. Reproduced with permission from ref (117). Copyright 2020 Elsevier. (B) Nearest Neighbor distances to count the number of Xist molecules and their distance to a histone marker, respectively. Reproduced with permission from ref (147). Copyright 2015 PNAS. (C) Localization of different sRNAs using sRNA-PAINT and their reported expression levels compared to RNA-seq Reproduced from ref (99) under Creative Commons License CC BY 4.0. Copyright 2020 Published by Oxford Academic Huang, K.; et al. (D) Bivariate pair correlation to measure the correlation between Sec61β with vgRNA and dsRNA and Sec61β with nsp3. Reproduced from ref (115) under Creative Commons License CC BY 4.0. Copyright 2024 Published by Springer Nature. Andronov, L.; et al. (E) Super-resolution time trace of Pol II cluster colocalizing with the active gene locus of β-actin (top) and real-time monitoring of mRNA output of ACTB following serum stimulation (bottom). Reproduced from ref (141) under Creative Commons License CC BY 4.0. Copyright 2016 Published by elife. Cho, W.-K.; et al. (F) Detection of miRNA using DNA PAINT. Expression reported by counts and each peak is a different miRNA. Reproduced from ref (13) under Creative Commons License CC-BY-NC-ND. Copyright 2023 Published by Elsevier Kocabey, S.; et al. (G) Voronoi Tessellation of RNA nanodomains clustering to different RNAP II using STORM and DNA-PAINT. Reproduced from ref (149) under Creative Commons License CC BY 4.0. Copyright 2022 Published by Oxford Academic. Castells-Garcia, A et al. (H) Spatial transcriptome wide analysis using expansion microscopy and MERFISH. Reproduced from ref (167) under Creative Commons License CC BY-NC-ND. Copyright 2019 Published by National Academy of Sciences Xia, C.; et al.
Figure 4
Figure 4. RNA quantification using SIM, STED, MINFLUX, and SHaSM. (A) SIM imaging of stress granules using a small molecule fluorescent probe (scale bar 5 μm). Reproduced from ref (170). Copyright 2023 American Chemical Society. (B) Schematic of using RNA-SPLIT to monitor Xist Turnover and representative 3D SIM images of Xist turnover during expansion. Reproduced with permission from ref (169). Copyright 2021 AAAS. (C) Single particle tracking of the comovement of TOI1-B and tdMCP-mCherry labeled trajectories. Reproduced from ref (171) under Creative Commons License CC BY 4.0. Copyright 2020 Cawte, A. D. et al. Published by Springer Nature. (D) Subcellular characterization of mtRNA using STED and MINFLUX. Reproduced from ref (168) under Creative Commons License CC BY 4.0. Copyright 2025 Stoldt, S.; et al. Published by Springer Nature. (E) Detection of Her2 mRNA in three different cell lines using SHaSM. Reproduced from ref (174). Copyright 2014 American Chemical Society.
3.3.3. Cluster Analysis
4. Super-Resolution Microscopy for RNA Quantification
4.1. Single Molecule Localization Microscopy
| Technique | General Principle | Advantages | Disadvantages | ref |
|---|---|---|---|---|
| SMLM | Uses blinking principle to locate the center of a dye. | •Similar labeling and sample preparation as most conventional fluorescent microscopy | •Requires thin samples | (99), (115), (124), (126), (127), (141), (147) |
| •Multicolor imaging | •Mainly fixed samples | |||
| •Mainly 2D images | ||||
| •Large data storage and long data processing times | ||||
| ExM | Increases distance between molecules through expansion in a hydrogel matrix. | •Thick cell and tissue samples | •Only fixed cells | (62), (118) |
| •Multicolor imaging | ||||
| •3D imaging | ||||
| •Easy to combine with another microscopy | ||||
| STED | Uses depletion laser | •1–10 nm resolution | •Not many reports on RNA imaging | (168), (184) |
| •3D imaging | •Expensive high-power lasers | |||
| •Strong lasers cause fast photobleaching | ||||
| MINFLUX | Uses differential emission intensity of dyes with doughnut shape laser and unique scanning pattern. | •Low photon budget (compared to SMLM) | •Complex instrument alignment | (168) |
| •3D imaging | •Long acquisition time for static images | |||
| •Multicolor imaging | •Small field of view | |||
| •1–10 nm resolution | ||||
| SIM | Uses patterned illumination and reconstruction from images at different phases shifts and rotations | •Small raw data size (9–15 images). | •Complex instrument alignment | (121), (169), (170), (185) |
| •Least invasive for live-cell imaging | •Lowest resolution | |||
| •3D imaging |
4.1.1. PALM/STORM
Gene Regulation
Genome Architecture
Cellular Environment During Viral Infection
4.1.2. DNA-PAINT
DNA PAINT as a Biosensor
Localization of small RNAs
4.2. Expansion Microscopy (ExM)
4.2.1. Transcriptomic Profiling of RNA
4.3. Stimulated Emission Depletion (STED)
4.4. MINFLUX
4.4.1. STED and MINFLUX to Visualize Mitochondrial mRNA
4.5. Structured Illumination Microscopy (SIM)
4.5.1. RNA Spreading and Turnover
4.5.2. Stress Granule Formation
4.5.3. Single-Particle Tracking
4.6. Super-Resolved Second Harmonic Microscopy (SHaSM)
5. Nonoptical Microscopy
6. Conclusion
References
This article references 185 other publications.
- 1Cao, X.; Zhang, Y.; Ding, Y.; Wan, Y. Identification of RNA structures and their roles in RNA functions. Nat. Rev. Mol. Cell Biol. 2024, 25 (10), 784– 801, DOI: 10.1038/s41580-024-00748-6Google ScholarThere is no corresponding record for this reference.
- 2Singh, K. P.; Miaskowski, C.; Dhruva, A. A.; Flowers, E.; Kober, K. M. Mechanisms and Measurement of Changes in Gene Expression. Biol. Res. Nurs 2018, 20 (4), 369– 382, DOI: 10.1177/1099800418772161Google ScholarThere is no corresponding record for this reference.
From NLM.
- 3Chen, Y.; Davidson, N. M.; Wan, Y. K.; Yao, F.; Su, Y.; Gamaarachchi, H.; Sim, A.; Patel, H.; Low, H. M.; Hendra, C.; Wratten, L.; Hakkaart, C.; Sawyer, C.; Iakovleva, V.; Lee, P. L.; Xin, L.; Ng, H. E. V.; Loo, J. M.; Ong, X.; Ng, H. Q. A.; Wang, J.; Koh, W. Q. C.; Poon, S. Y. P.; Stanojevic, D.; Tran, H.-D.; Lim, K. H. E.; Toh, S. Y.; Ewels, P. A.; Ng, H.-H.; Iyer, N. G.; Thiery, A.; Chng, W. J.; Chen, L.; DasGupta, R.; Sikic, M.; Chan, Y.-S.; Tan, B. O. P.; Wan, Y.; Tam, W. L.; Yu, Q.; Khor, C. C.; Wustefeld, T.; Lezhava, A.; Pratanwanich, P. N.; Love, M. I.; Goh, W. S. S.; Ng, S. B.; Oshlack, A.; Iyer, N. G.; Yu, Q.; Goke, J. A systematic benchmark of Nanopore long-read RNA sequencing for transcript-level analysis in human cell lines. Nat. Methods 2025, 22 (4), 801– 812, DOI: 10.1038/s41592-025-02623-4Google ScholarThere is no corresponding record for this reference.
- 4Chen, L.-L.; Kim, V. N. Small and long non-coding RNAs: Past, present, and future. Cell 2024, 187 (23), 6451– 6485, DOI: 10.1016/j.cell.2024.10.024Google ScholarThere is no corresponding record for this reference.
- 5Mattick, J. S.; Amaral, P. P.; Carninci, P.; Carpenter, S.; Chang, H. Y.; Chen, L.-L.; Chen, R.; Dean, C.; Dinger, M. E.; Fitzgerald, K. A.; Gingeras, T. R.; Guttman, M.; Hirose, T.; Huarte, M.; Johnson, R.; Kanduri, C.; Kapranov, P.; Lawrence, J. B.; Lee, J. T.; Mendell, J. T.; Mercer, T. R.; Moore, K. J.; Nakagawa, S.; Rinn, J. L.; Spector, D. L.; Ulitsky, I.; Wan, Y.; Wilusz, J. E.; Wu, M. Long non-coding RNAs: definitions, functions, challenges and recommendations. Nat. Rev. Mol. Cell Biol. 2023, 24 (6), 430– 447, DOI: 10.1038/s41580-022-00566-8Google ScholarThere is no corresponding record for this reference.
- 6Le, P.; Ahmed, N.; Yeo, G. W. Illuminating RNA biology through imaging. Nat. Cell Biol. 2022, 24 (6), 815– 824, DOI: 10.1038/s41556-022-00933-9Google ScholarThere is no corresponding record for this reference.
- 7Xia, Y.; Zhang, R.; Wang, Z.; Tian, J.; Chen, X. Recent advances in high-performance fluorescent and bioluminescent RNA imaging probes. Chem. Soc. Rev. 2017, 46 (10), 2824– 2843, DOI: 10.1039/C6CS00675BGoogle ScholarThere is no corresponding record for this reference.
From NLM.
- 8Boutorine, A. S.; Novopashina, D. S.; Krasheninina, O. A.; Nozeret, K.; Venyaminova, A. G. Fluorescent probes for nucleic Acid visualization in fixed and live cells. Molecules 2013, 18 (12), 15357– 15397, DOI: 10.3390/molecules181215357Google ScholarThere is no corresponding record for this reference.
From NLM.
- 9Schermelleh, L.; Ferrand, A.; Huser, T.; Eggeling, C.; Sauer, M.; Biehlmaier, O.; Drummen, G. P. C. Super-resolution microscopy demystified. Nat. Cell Biol. 2019, 21 (1), 72– 84, DOI: 10.1038/s41556-018-0251-8Google ScholarThere is no corresponding record for this reference.
- 10Sigal, Y. M.; Zhou, R.; Zhuang, X. Visualizing and discovering cellular structures with super-resolution microscopy. Science 2018, 361 (6405), 880– 887, DOI: 10.1126/science.aau1044Google ScholarThere is no corresponding record for this reference.
From NLM.
- 11Huang, B.; Babcock, H.; Zhuang, X. Breaking the diffraction barrier: super-resolution imaging of cells. Cell 2010, 143 (7), 1047– 1058, DOI: 10.1016/j.cell.2010.12.002Google ScholarThere is no corresponding record for this reference.
From NLM.
- 12Zhong, J.; Xu, Z.; Peng, J.; Guan, L.; Li, J.; Zhou, Z.; Zhang, Y.; Zhang, J.; Liu, S.; Yang, Y.; Hao, X. A CRISPR/Cas13a system based on a dumbbell-shaped hairpin combined with DNA-PAINT to establish the DCP-platform for highly sensitive detection of Hantaan virus RNA. Talanta 2025, 291, 127852, DOI: 10.1016/j.talanta.2025.127852Google ScholarThere is no corresponding record for this reference.
- 13Kocabey, S.; Chiarelli, G.; Acuna, G. P.; Ruegg, C. Ultrasensitive and multiplexed miRNA detection system with DNA-PAINT. Biosens. Bioelectron. 2023, 224, 115053, DOI: 10.1016/j.bios.2022.115053Google ScholarThere is no corresponding record for this reference.
- 14Higuchi, R.; Fockler, C.; Dollinger, G.; Watson, R. Kinetic PCR analysis: real-time monitoring of DNA amplification reactions. Biotechnology (N Y) 1993, 11 (9), 1026– 1030, DOI: 10.1038/nbt0993-1026Google ScholarThere is no corresponding record for this reference.
From NLM.
- 15Salimi, A.; Rahmani, S.; Sharifi-Zarchi, A. InterOpt: Improved gene expression quantification in qPCR experiments using weighted aggregation of reference genes. iScience 2023, 26 (10), 107945, DOI: 10.1016/j.isci.2023.107945Google ScholarThere is no corresponding record for this reference.
- 16Różański, M.; Walczak-Drzewiecka, A.; Witaszewska, J.; Wójcik, E.; Guziński, A.; Zimoń, B.; Matusiak, R.; Kazimierczak, J.; Borowiec, M.; Kania, K.; Paradowska, E.; Pawełczyk, J.; Dziadek, J.; Dastych, J. RT-qPCR-based tests for SARS-CoV-2 detection in pooled saliva samples for massive population screening to monitor epidemics. Sci. Rep. 2022, 12 (1), 8082, DOI: 10.1038/s41598-022-12179-4Google ScholarThere is no corresponding record for this reference.
- 17Bustin, S. A.; Nolan, T. Pitfalls of quantitative real-time reverse-transcription polymerase chain reaction. J. Biomol. Tech. 2004, 15 (3), 155– 166Google ScholarThere is no corresponding record for this reference.
From NLM.
- 18Gerbi, S. A.; Borovjagin, A. V. Pre-ribosomal RNA processing in multicellular organisms. Madame Curie Bioscience Database [Internet]; Landes Bioscience , 2004; pp 170– 198.Google ScholarThere is no corresponding record for this reference.
- 19Wu, J.; Xiao, J.; Zhang, Z.; Wang, X.; Hu, S.; Yu, J. Ribogenomics: the Science and Knowledge of RNA. Genomics, Proteomics & Bioinformatics 2014, 12 (2), 57– 63, DOI: 10.1016/j.gpb.2014.04.002Google ScholarThere is no corresponding record for this reference.
- 20Sanders, R.; Mason, D. J.; Foy, C. A.; Huggett, J. F. Considerations for accurate gene expression measurement by reverse transcription quantitative PCR when analysing clinical samples. Anal. Bioanal. Chem. 2014, 406 (26), 6471– 6483, DOI: 10.1007/s00216-014-7857-xGoogle ScholarThere is no corresponding record for this reference.
- 21Forster, T.; Roy, D.; Ghazal, P. Experiments using microarray technology: limitations and standard operating procedures. J. Endocrinol 2003, 178 (2), 195– 204, DOI: 10.1677/joe.0.1780195Google ScholarThere is no corresponding record for this reference.
From NLM.
- 22Coppola, G. Designing, performing, and interpreting a microarray-based gene expression study. Methods Mol. Biol. 2011, 793, 417– 439, DOI: 10.1007/978-1-61779-328-8_28Google ScholarThere is no corresponding record for this reference.
From NLM.
- 23Uhlen, M.; Quake, S. R. Sequential sequencing by synthesis and the next-generation sequencing revolution. Trends Biotechnol. 2023, 41 (12), 1565– 1572, DOI: 10.1016/j.tibtech.2023.06.007Google ScholarThere is no corresponding record for this reference.
(accessed 2025/10/30).
- 24Satam, H.; Joshi, K.; Mangrolia, U.; Waghoo, S.; Zaidi, G.; Rawool, S.; Thakare, R. P.; Banday, S.; Mishra, A. K.; Das, G.; Malonia, S. K. Next-Generation Sequencing Technology: Current Trends and Advancements. Biology (Basel) 2023, 12 (7), 997, DOI: 10.3390/biology12070997Google ScholarThere is no corresponding record for this reference.
- 25Ament, I. H.; DeBruyne, N.; Wang, F.; Lin, L. Long-read RNA sequencing: A transformative technology for exploring transcriptome complexity in human diseases. Molecular Therapy 2025, 33 (3), 883– 894, DOI: 10.1016/j.ymthe.2024.11.025Google ScholarThere is no corresponding record for this reference.
(accessed 2025/10/30).
- 26McKinnon, K. M. Flow Cytometry: An Overview. Curr. Protoc Immunol 2018, 120, 5.1.1– 5.1.11, DOI: 10.1002/cpim.40Google ScholarThere is no corresponding record for this reference.
From NLM.
- 27Zhao, Y.; Lin, C.; Wu, P.; Chen, X.; Zhao, Y.; Li, Y.; Chen, L.; Nilsson, M.; Ke, R. Single Cell RNA Expression Analysis Using Flow Cytometry Based on Specific Probe Ligation and Rolling Circle Amplification. ACS Sensors 2020, 5 (10), 3031– 3036, DOI: 10.1021/acssensors.0c01569Google ScholarThere is no corresponding record for this reference.
- 28Hanley, M. B.; Lomas, W.; Mittar, D.; Maino, V.; Park, E. Detection of low abundance RNA molecules in individual cells by flow cytometry. PLoS One 2013, 8 (2), e57002 DOI: 10.1371/journal.pone.0057002Google ScholarThere is no corresponding record for this reference.
From NLM.
- 29Warren, C. J.; Barbachano-Guerrero, A.; Huey, D.; Yang, Q.; Worden-Sapper, E. R.; Kuhn, J. H.; Sawyer, S. L. Quantification of virus-infected cells using RNA FISH-Flow. STAR Protoc 2023, 4 (2), 102291, DOI: 10.1016/j.xpro.2023.102291Google ScholarThere is no corresponding record for this reference.
From NLM.
- 30Gaudin, R.; Barteneva, N. S. Sorting of small infectious virus particles by flow virometry reveals distinct infectivity profiles. Nat. Commun. 2015, 6 (1), 6022, DOI: 10.1038/ncomms7022Google ScholarThere is no corresponding record for this reference.
- 31Actis, P.; Maalouf, M. M.; Kim, H. J.; Lohith, A.; Vilozny, B.; Seger, R. A.; Pourmand, N. Compartmental Genomics in Living Cells Revealed by Single-Cell Nanobiopsy. ACS Nano 2014, 8 (1), 546– 553, DOI: 10.1021/nn405097uGoogle ScholarThere is no corresponding record for this reference.
- 32Sahota, A.; Monteza Cabrejos, A.; Kwan, Z.; Paulose Nadappuram, B.; Ivanov, A. P.; Edel, J. B. Recent advances in single-cell subcellular sampling. Chem. Commun. (Camb) 2023, 59 (36), 5312– 5328, DOI: 10.1039/D3CC00573AGoogle ScholarThere is no corresponding record for this reference.
From NLM.
- 33Sahl, S. J.; Hell, S. W.; Jakobs, S. Fluorescence nanoscopy in cell biology. Nat. Rev. Mol. Cell Biol. 2017, 18 (11), 685– 701, DOI: 10.1038/nrm.2017.71Google ScholarThere is no corresponding record for this reference.
- 34McIsaac, R. S.; Silverman, S. J.; Parsons, L.; Xu, P.; Briehof, R.; McClean, M. N.; Botstein, D. Visualization and analysis of mRNA molecules using fluorescence in situ hybridization in Saccharomyces cerevisiae. J. Vis Exp 2013, (76), e50382 DOI: 10.3791/50382Google ScholarThere is no corresponding record for this reference.
From NLM.
- 35Elliott, A. D. Confocal Microscopy: Principles and Modern Practices. Curr. Protoc Cytom 2020, 92 (1), e68 DOI: 10.1002/cpcy.68Google ScholarThere is no corresponding record for this reference.
From NLM.
- 36Singer, R. H.; Ward, D. C. Actin gene expression visualized in chicken muscle tissue culture by using in situ hybridization with a biotinated nucleotide analog. Proc. Natl. Acad. Sci. U. S. A. 1982, 79 (23), 7331– 7335, DOI: 10.1073/pnas.79.23.7331Google ScholarThere is no corresponding record for this reference.
From NLM.
- 37Barakat, T. S.; Gribnau, J. Combined DNA-RNA fluorescent in situ hybridization (FISH) to study X chromosome inactivation in differentiated female mouse embryonic stem cells. J. Vis Exp 2014, 88, 51628, DOI: 10.3791/51628Google ScholarThere is no corresponding record for this reference.
- 38Young, A. P.; Jackson, D. J.; Wyeth, R. C. A technical review and guide to RNA fluorescence in situ hybridization. PeerJ. 2020, 8, e8806 DOI: 10.7717/peerj.8806Google ScholarThere is no corresponding record for this reference.
From NLM.
- 39Raj, A.; van den Bogaard, P.; Rifkin, S. A.; van Oudenaarden, A.; Tyagi, S. Imaging individual mRNA molecules using multiple singly labeled probes. Nat. Methods 2008, 5 (10), 877– 879, DOI: 10.1038/nmeth.1253Google ScholarThere is no corresponding record for this reference.
From NLM.
- 40Haimovich, G.; Gerst, J. E. Single-molecule Fluorescence in situ Hybridization (smFISH) for RNA Detection in Adherent Animal Cells. Bio-protocol 2018, 8 (21), e3070 DOI: 10.21769/BioProtoc.3070Google ScholarThere is no corresponding record for this reference.
- 41Femino, A. M.; Fay, F. S.; Fogarty, K.; Singer, R. H. Visualization of Single RNA Transcripts in Situ. Science 1998, 280 (5363), 585– 590, DOI: 10.1126/science.280.5363.585Google ScholarThere is no corresponding record for this reference.
- 42Levesque, M. J.; Ginart, P.; Wei, Y.; Raj, A. Visualizing SNVs to quantify allele-specific expression in single cells. Nat. Methods 2013, 10 (9), 865– 867, DOI: 10.1038/nmeth.2589Google ScholarThere is no corresponding record for this reference.
- 43Levesque, M. J.; Raj, A. Single-chromosome transcriptional profiling reveals chromosomal gene expression regulation. Nat. Methods 2013, 10 (3), 246– 248, DOI: 10.1038/nmeth.2372Google ScholarThere is no corresponding record for this reference.
- 44Wang, F.; Flanagan, J.; Su, N.; Wang, L. C.; Bui, S.; Nielson, A.; Wu, X.; Vo, H. T.; Ma, X. J.; Luo, Y. RNAscope: a novel in situ RNA analysis platform for formalin-fixed, paraffin-embedded tissues. J. Mol. Diagn 2012, 14 (1), 22– 29, DOI: 10.1016/j.jmoldx.2011.08.002Google ScholarThere is no corresponding record for this reference.
From NLM.
- 45Atout, S.; Shurrab, S.; Loveridge, C. Evaluation of the Suitability of RNAscope as a Technique to Measure Gene Expression in Clinical Diagnostics: A Systematic Review. Mol. Diagn Ther 2022, 26 (1), 19– 37, DOI: 10.1007/s40291-021-00570-2Google ScholarThere is no corresponding record for this reference.
From NLM.
- 46Huang, T.; Guillotin, B.; Rahni, R.; Birnbaum, K. D.; Wagner, D. A rapid and sensitive, multiplex, whole mount RNA fluorescence in situ hybridization and immunohistochemistry protocol. Plant Methods 2023, 19 (1), 131, DOI: 10.1186/s13007-023-01108-9Google ScholarThere is no corresponding record for this reference.
- 47Choi, H. M. T.; Beck, V. A.; Pierce, N. A. Next-Generation in Situ Hybridization Chain Reaction: Higher Gain, Lower Cost, Greater Durability. ACS Nano 2014, 8 (5), 4284– 4294, DOI: 10.1021/nn405717pGoogle ScholarThere is no corresponding record for this reference.
- 48Lovely, A. M.; Duerr, T. J.; Stein, D. F.; Mun, E. T.; Monaghan, J. R. Hybridization Chain Reaction Fluorescence In Situ Hybridization (HCR-FISH) in Ambystoma mexicanum Tissue. Methods Mol. Biol. 2023, 2562, 109– 122, DOI: 10.1007/978-1-0716-2659-7_6Google ScholarThere is no corresponding record for this reference.
From NLM.
- 49Choi, H. M. T.; Schwarzkopf, M.; Fornace, M. E.; Acharya, A.; Artavanis, G.; Stegmaier, J.; Cunha, A.; Pierce, N. A. Third-generation in situ hybridization chain reaction: multiplexed, quantitative, sensitive, versatile, robust. Development 2018, 145 (12), dev165753, DOI: 10.1242/dev.165753Google ScholarThere is no corresponding record for this reference.
- 50Deng, R.; Zhang, K.; Sun, Y.; Ren, X.; Li, J. Highly specific imaging of mRNA in single cells by target RNA-initiated rolling circle amplification. Chem. Sci. 2017, 8 (5), 3668– 3675, DOI: 10.1039/C7SC00292KGoogle ScholarThere is no corresponding record for this reference.
From NLM.
- 51Kurreck, J.; Wyszko, E.; Gillen, C.; Erdmann, V. A. Design of antisense oligonucleotides stabilized by locked nucleic acids. Nucleic Acids Res. 2002, 30 (9), 1911– 1918, DOI: 10.1093/nar/30.9.1911Google ScholarThere is no corresponding record for this reference.
From NLM.
- 52Wu, C.; Simonetti, M.; Rossell, C.; Mignardi, M.; Mirzazadeh, R.; Annaratone, L.; Marchiò, C.; Sapino, A.; Bienko, M.; Crosetto, N.; Nilsson, M. RollFISH achieves robust quantification of single-molecule RNA biomarkers in paraffin-embedded tumor tissue samples. Communications Biology 2018, 1 (1), 209, DOI: 10.1038/s42003-018-0218-0Google ScholarThere is no corresponding record for this reference.
- 53Lubeck, E.; Cai, L. Single-cell systems biology by super-resolution imaging and combinatorial labeling. Nat. Methods 2012, 9 (7), 743– 748, DOI: 10.1038/nmeth.2069Google ScholarThere is no corresponding record for this reference.
- 54Lubeck, E.; Coskun, A. F.; Zhiyentayev, T.; Ahmad, M.; Cai, L. Single-cell in situ RNA profiling by sequential hybridization. Nat. Methods 2014, 11 (4), 360– 361, DOI: 10.1038/nmeth.2892Google ScholarThere is no corresponding record for this reference.
- 55Shah, S.; Lubeck, E.; Zhou, W.; Cai, L. In Situ Transcription Profiling of Single Cells Reveals Spatial Organization of Cells in the Mouse Hippocampus. Neuron 2016, 92 (2), 342– 357, DOI: 10.1016/j.neuron.2016.10.001Google ScholarThere is no corresponding record for this reference.
accessed 2026/02/05).
- 56Shah, S.; Takei, Y.; Zhou, W.; Lubeck, E.; Yun, J.; Eng, C.-H. L.; Koulena, N.; Cronin, C.; Karp, C.; Liaw, E. J.; Amin, M.; Cai, L. Dynamics and Spatial Genomics of the Nascent Transcriptome by Intron seqFISH. Cell 2018, 174 (2), 363– 376.e316, DOI: 10.1016/j.cell.2018.05.035Google ScholarThere is no corresponding record for this reference.
accessed 2026/02/05).
- 57Eng, C.-H. L.; Shah, S.; Thomassie, J.; Cai, L. Profiling the transcriptome with RNA SPOTs. Nat. Methods 2017, 14 (12), 1153– 1155, DOI: 10.1038/nmeth.4500Google ScholarThere is no corresponding record for this reference.
- 58Eng, C.-H. L.; Lawson, M.; Zhu, Q.; Dries, R.; Koulena, N.; Takei, Y.; Yun, J.; Cronin, C.; Karp, C.; Yuan, G.-C.; Cai, L. Transcriptome-scale super-resolved imaging in tissues by RNA seqFISH+. Nature 2019, 568 (7751), 235– 239, DOI: 10.1038/s41586-019-1049-yGoogle ScholarThere is no corresponding record for this reference.
- 59Chen, K. H.; Boettiger, A. N.; Moffitt, J. R.; Wang, S.; Zhuang, X. Spatially resolved, highly multiplexed RNA profiling in single cells. Science 2015, 348 (6233), aaa6090, DOI: 10.1126/science.aaa6090Google ScholarThere is no corresponding record for this reference.
- 60Moffitt, J. R.; Hao, J.; Wang, G.; Chen, K. H.; Babcock, H. P.; Zhuang, X. High-throughput single-cell gene-expression profiling with multiplexed error-robust fluorescence in situ hybridization. Proc. Natl. Acad. Sci. U. S. A. 2016, 113 (39), 11046– 11051, DOI: 10.1073/pnas.1612826113Google ScholarThere is no corresponding record for this reference.
- 61Moffitt, J. R.; Zhuang, X. Chapter One - RNA Imaging with Multiplexed Error-Robust Fluorescence In Situ Hybridization (MERFISH). In Methods in Enzymology, Filonov, G. S., Jaffrey, S. R., Eds.; Academic Press, 2016; Vol. 572, pp 1– 49.Google ScholarThere is no corresponding record for this reference.
- 62Chen, F.; Tillberg, P. W.; Boyden, E. S. Expansion microscopy. Science 2015, 347 (6221), 543– 548, DOI: 10.1126/science.1260088Google ScholarThere is no corresponding record for this reference.
- 63Liu, J.; Tran, V.; Vemuri, V. N. P.; Byrne, A.; Borja, M.; Kim, Y. J.; Agarwal, S.; Wang, R.; Awayan, K.; Murti, A.; Taychameekiatchai, A.; Wang, B.; Emanuel, G.; He, J.; Haliburton, J.; Oliveira Pisco, A.; Neff, N. F. Concordance of MERFISH spatial transcriptomics with bulk and single-cell RNA sequencing. Life Sci. Alliance 2023, 6 (1), e202201701, DOI: 10.26508/lsa.202201701Google ScholarThere is no corresponding record for this reference.
- 64Fang, R.; Halpern, A.; Rahman, M. M.; Huang, Z.; Lei, Z.; Hell, S. J.; Dulac, C.; Zhuang, X. Three-dimensional single-cell transcriptome imaging of thick tissues. eLife 2024, 12, RP90029, DOI: 10.7554/eLife.90029Google ScholarThere is no corresponding record for this reference.
- 65Braselmann, E.; Rathbun, C.; Richards, E. M.; Palmer, A. E. Illuminating RNA Biology: Tools for Imaging RNA in Live Mammalian Cells. Cell Chemical Biology 2020, 27 (8), 891– 903, DOI: 10.1016/j.chembiol.2020.06.010Google ScholarThere is no corresponding record for this reference.
- 66Chiu, Y. F.; Huang, Y. W.; Chen, C. Y.; Chen, Y. C.; Gong, Y. N.; Kuo, R. L.; Huang, C. G.; Shih, S. R. Visualizing Influenza A Virus vRNA Replication. Front Microbiol 2022, 13, 812711, DOI: 10.3389/fmicb.2022.812711Google ScholarThere is no corresponding record for this reference.
From NLM.
- 67Hu, Y.; Xu, J.; Gao, E.; Fan, X.; Wei, J.; Ye, B.; Xu, S.; Ma, W. Enhanced single RNA imaging reveals dynamic gene expression in live animals. eLife 2023, 12, e82178 DOI: 10.7554/eLife.82178Google ScholarThere is no corresponding record for this reference.
- 68Peng, Y.; Shu, L.; Deng, X.; Huang, X.; Mo, X.; Du, F.; Tang, Z. Live-Cell Imaging of Endogenous RNA with a Genetically Encoded Fluorogenic Allosteric Aptamer. Anal. Chem. 2023, 95 (37), 13762– 13768, DOI: 10.1021/acs.analchem.2c05724Google ScholarThere is no corresponding record for this reference.
- 69Constantin, T. P.; Silva, G. L.; Robertson, K. L.; Hamilton, T. P.; Fague, K.; Waggoner, A. S.; Armitage, B. A. Synthesis of New Fluorogenic Cyanine Dyes and Incorporation into RNA Fluoromodules. Org. Lett. 2008, 10 (8), 1561– 1564, DOI: 10.1021/ol702920eGoogle ScholarThere is no corresponding record for this reference.
- 70Huang, K.; Chen, X.; Li, C.; Song, Q.; Li, H.; Zhu, L.; Yang, Y.; Ren, A. Structure-based investigation of fluorogenic Pepper aptamer. Nat. Chem. Biol. 2021, 17 (12), 1289– 1295, DOI: 10.1038/s41589-021-00884-6Google ScholarThere is no corresponding record for this reference.
- 71Zheng, H.; Liu, X.; Liu, L.; Hu, J.; Chen, X. Imaging of endogenous RNA in live cells using sequence-activated fluorescent RNA probes. Nucleic Acids Res. 2025, 53 (2), gkae1209, DOI: 10.1093/nar/gkae1209Google ScholarThere is no corresponding record for this reference.
- 72Jansen, R.; Embden, J. D.; Gaastra, W.; Schouls, L. M. Identification of genes that are associated with DNA repeats in prokaryotes. Mol. Microbiol. 2002, 43 (6), 1565– 1575, DOI: 10.1046/j.1365-2958.2002.02839.xGoogle ScholarThere is no corresponding record for this reference.
From NLM.
- 73Wang, S. W.; Gao, C.; Zheng, Y. M.; Yi, L.; Lu, J. C.; Huang, X. Y.; Cai, J. B.; Zhang, P. F.; Cui, Y. H.; Ke, A. W. Current applications and future perspective of CRISPR/Cas9 gene editing in cancer. Mol. Cancer 2022, 21 (1), 57, DOI: 10.1186/s12943-022-01518-8Google ScholarThere is no corresponding record for this reference.
From NLM.
- 74Chen, K.; Wang, Y. CRISPR/Cas systems for in situ imaging of intracellular nucleic acids: Concepts and applications. Biotechnol. Bioeng. 2023, 120 (12), 3446– 3464, DOI: 10.1002/bit.28543Google ScholarThere is no corresponding record for this reference.
From NLM.
- 75Hillary, V. E.; Ceasar, S. A. A Review on the Mechanism and Applications of CRISPR/Cas9/Cas12/Cas13/Cas14 Proteins Utilized for Genome Engineering. Mol. Biotechnol 2023, 65 (3), 311– 325, DOI: 10.1007/s12033-022-00567-0Google ScholarThere is no corresponding record for this reference.
From NLM.
- 76Cao, H.; Wang, Y.; Zhang, N.; Xia, S.; Tian, P.; Lu, L.; Du, J.; Du, Y. Progress of CRISPR-Cas13 Mediated Live-Cell RNA Imaging and Detection of RNA-Protein Interactions. Front Cell Dev Biol. 2022, 10, 866820, DOI: 10.3389/fcell.2022.866820Google ScholarThere is no corresponding record for this reference.
From NLM.
- 77Jia, H.-Y.; Zhang, X.-Y.; Ye, B.-C.; Yin, B.-C. An Orthogonal CRISPR/dCas12a System for RNA Imaging in Live Cells. Anal. Chem. 2024, 96 (15), 5913– 5921, DOI: 10.1021/acs.analchem.3c05975Google ScholarThere is no corresponding record for this reference.
- 78O’Connell, M. R.; Oakes, B. L.; Sternberg, S. H.; East-Seletsky, A.; Kaplan, M.; Doudna, J. A. Programmable RNA recognition and cleavage by CRISPR/Cas9. Nature 2014, 516 (7530), 263– 266, DOI: 10.1038/nature13769Google ScholarThere is no corresponding record for this reference.
From NLM.
- 79Tang, H.; Peng, J.; Peng, S.; Wang, Q.; Jiang, X.; Xue, X.; Tao, Y.; Xiang, L.; Ji, Q.; Liu, S. M.; Weng, X.; Zhou, X. Live-cell RNA imaging using the CRISPR-dCas13 system with modified sgRNAs appended with fluorescent RNA aptamers. Chem. Sci. 2022, 13 (47), 14032– 14040, DOI: 10.1039/D2SC04656CGoogle ScholarThere is no corresponding record for this reference.
From NLM.
- 80Tyagi, S.; Kramer, F. R. Molecular Beacons: Probes that Fluoresce upon Hybridization. Nat. Biotechnol. 1996, 14 (3), 303– 308, DOI: 10.1038/nbt0396-303Google ScholarThere is no corresponding record for this reference.
- 81Xia, C.; Colognori, D.; Jiang, X. S.; Xu, K.; Doudna, J. A. Single-molecule live-cell RNA imaging with CRISPR–Csm. Nat. Biotechnol. 2025, 43, 2023, DOI: 10.1038/s41587-024-02540-5Google ScholarThere is no corresponding record for this reference.
- 82Bratu, D. P.; Cha, B.-J.; Mhlanga, M. M.; Kramer, F. R.; Tyagi, S. Visualizing the distribution and transport of mRNAs in living cells. Proc. Natl. Acad. Sci. U. S. A. 2003, 100 (23), 13308– 13313, DOI: 10.1073/pnas.2233244100Google ScholarThere is no corresponding record for this reference.
- 83Turner-Bridger, B.; Jakobs, M.; Muresan, L.; Wong, H. H.-W.; Franze, K.; Harris, W. A.; Holt, C. E. Single-molecule analysis of endogenous β-actin mRNA trafficking reveals a mechanism for compartmentalized mRNA localization in axons. Proc. Natl. Acad. Sci. U. S. A. 2018, 115 (41), E9697– E9706, DOI: 10.1073/pnas.1806189115Google ScholarThere is no corresponding record for this reference.
- 84Culley, S.; Caballero, A. C.; Burden, J. J.; Uhlmann, V. Made to measure: An introduction to quantifying microscopy data in the life sciences. J. Microsc 2024, 295 (1), 61– 82, DOI: 10.1111/jmi.13208Google ScholarThere is no corresponding record for this reference.
From NLM.
- 85Schindelin, J.; Arganda-Carreras, I.; Frise, E.; Kaynig, V.; Longair, M.; Pietzsch, T.; Preibisch, S.; Rueden, C.; Saalfeld, S.; Schmid, B.; Tinevez, J. Y.; White, D. J.; Hartenstein, V.; Eliceiri, K.; Tomancak, P.; Cardona, A. Fiji: an open-source platform for biological-image analysis. Nat. Methods 2012, 9 (7), 676– 682, DOI: 10.1038/nmeth.2019Google ScholarThere is no corresponding record for this reference.
From NLM.
- 86Carpenter, A. E.; Jones, T. R.; Lamprecht, M. R.; Clarke, C.; Kang, I. H.; Friman, O.; Guertin, D. A.; Chang, J. H.; Lindquist, R. A.; Moffat, J.; Golland, P.; Sabatini, D. M. CellProfiler: image analysis software for identifying and quantifying cell phenotypes. Genome Biology 2006, 7 (10), R100, DOI: 10.1186/gb-2006-7-10-r100Google ScholarThere is no corresponding record for this reference.
- 87Waters, J. C. Accuracy and precision in quantitative fluorescence microscopy. J. Cell Biol. 2009, 185 (7), 1135– 1148, DOI: 10.1083/jcb.200903097Google ScholarThere is no corresponding record for this reference.
From NLM.
- 88Sasaki, A. Recent advances in the standardization of fluorescence microscopy for quantitative image analysis. Biophys Rev. 2022, 14 (1), 33– 39, DOI: 10.1007/s12551-021-00871-0Google ScholarThere is no corresponding record for this reference.
From NLM.
- 89Baskin, D. S.; Widmayer, M. A.; Sharpe, M. A. Quantification and calibration of images in fluorescence microscopy. Anal. Biochem. 2010, 404 (2), 118– 126, DOI: 10.1016/j.ab.2010.05.029Google ScholarThere is no corresponding record for this reference.
- 90Barbini, P.; Cevenini, G.; Massai, M. R. Nearest-Neighbor Analysis of Spatial Point Patterns: Application to Biomedical Image Interpretation. Computers and Biomedical Research 1996, 29 (6), 482– 493, DOI: 10.1006/cbmr.1996.0035Google ScholarThere is no corresponding record for this reference.
- 91Soltisz, A. M.; Craigmile, P. F.; Veeraraghavan, R. Spatial Pattern Analysis using Closest Events (SPACE)-A Nearest Neighbor Point Pattern Analysis Framework for Assessing Spatial Relationships from Digital Images. Microsc Microanal 2024, 30 (2), 306– 317, DOI: 10.1093/mam/ozae022Google ScholarThere is no corresponding record for this reference.
From NLM.
- 92Kiskowski, M. A.; Hancock, J. F.; Kenworthy, A. K. On the use of Ripley’s K-function and its derivatives to analyze domain size. Biophys. J. 2009, 97 (4), 1095– 1103, DOI: 10.1016/j.bpj.2009.05.039Google ScholarThere is no corresponding record for this reference.
From NLM.
- 93Ruan, Y.; Yin, P.; Li, F.; Li, D.; Lin, Q.; Li, K. The Accuracy of Determining Cluster Size by Analyzing Ripley’s K Function in Single Molecule Localization Microscopy. Applied Sciences 2019, 9 (16), 3271, DOI: 10.3390/app9163271Google ScholarThere is no corresponding record for this reference.
- 94Peters, R.; Benthem Muñiz, M.; Griffié, J.; Williamson, D. J.; Ashdown, G. W.; Lorenz, C. D.; Owen, D. M. Quantification of fibrous spatial point patterns from single-molecule localization microscopy (SMLM) data. Bioinformatics 2017, 33 (11), 1703– 1711, DOI: 10.1093/bioinformatics/btx026Google ScholarThere is no corresponding record for this reference.
(accessed 10/26/2025).
- 95Lee, C.; Roberts, S. E.; Gladfelter, A. S. Quantitative spatial analysis of transcripts in multinucleate cells using single-molecule FISH. Methods 2016, 98, 124– 133, DOI: 10.1016/j.ymeth.2015.12.007Google ScholarThere is no corresponding record for this reference.
- 96Mancebo, A.; Mehra, D.; Banerjee, C.; Kim, D. H.; Puchner, E. M. Efficient Cross-Correlation Filtering of One- and Two-Color Single Molecule Localization Microscopy Data. Front Bioinform 2021, 1, 739769, DOI: 10.3389/fbinf.2021.739769Google ScholarThere is no corresponding record for this reference.
From NLM.
- 97Schnitzbauer, J.; Wang, Y.; Zhao, S.; Bakalar, M.; Nuwal, T.; Chen, B.; Huang, B. Correlation analysis framework for localization-based superresolution microscopy. Proc. Natl. Acad. Sci. U. S. A. 2018, 115 (13), 3219– 3224, DOI: 10.1073/pnas.1711314115Google ScholarThere is no corresponding record for this reference.
From NLM.
- 98Ester, M.; Kriegel, H.-P.; Sander, J.; Xu, X. A density-based algorithm for discovering clusters in large spatial databases with noise. kdd Proceedings 1996, 96, 226– 231Google ScholarThere is no corresponding record for this reference.
- 99Huang, K.; Demirci, F.; Batish, M.; Treible, W.; Meyers, B. C.; Caplan, J. L. Quantitative, super-resolution localization of small RNAs with sRNA-PAINT. Nucleic Acids Res. 2020, 48 (16), e96– e96, DOI: 10.1093/nar/gkaa623Google ScholarThere is no corresponding record for this reference.
(accessed 10/26/2025).
- 100Pageon, S. V.; Nicovich, P. R.; Mollazade, M.; Tabarin, T.; Gaus, K. Clus-DoC: a combined cluster detection and colocalization analysis for single-molecule localization microscopy data. Mol. Biol. Cell 2016, 27 (22), 3627– 3636, DOI: 10.1091/mbc.e16-07-0478Google ScholarThere is no corresponding record for this reference.
From NLM.
- 101Bushra, A. A.; Kim, D.; Kan, Y.; Yi, G. AutoSCAN: automatic detection of DBSCAN parameters and efficient clustering of data in overlapping density regions. PeerJ. Comput. Sci. 2024, 10, e1921 DOI: 10.7717/peerj-cs.1921Google ScholarThere is no corresponding record for this reference.
From NLM.
- 102Yin, L.; Hu, H.; Li, K.; Zheng, G.; Qu, Y.; Chen, H. Improvement of DBSCAN Algorithm Based on K-Dist Graph for Adaptive Determining Parameters. Electronics 2023, 12 (15), 3213, DOI: 10.3390/electronics12153213Google ScholarThere is no corresponding record for this reference.
- 103Bushra, A. A.; Yi, G. Comparative Analysis Review of Pioneering DBSCAN and Successive Density-Based Clustering Algorithms. IEEE Access 2021, 9, 87918– 87935, DOI: 10.1109/ACCESS.2021.3089036Google ScholarThere is no corresponding record for this reference.
- 104Marenda, M.; Lazarova, E.; van de Linde, S.; Gilbert, N.; Michieletto, D. Parameter-free molecular super-structures quantification in single-molecule localization microscopy. J. Cell Biol. 2021, 220 (5), e202010003, DOI: 10.1083/jcb.202010003Google ScholarThere is no corresponding record for this reference.
- 105Mazouchi, A.; Milstein, J. N. Fast Optimized Cluster Algorithm for Localizations (FOCAL): a spatial cluster analysis for super-resolved microscopy. Bioinformatics 2016, 32 (5), 747– 754, DOI: 10.1093/bioinformatics/btv630Google ScholarThere is no corresponding record for this reference.
(accessed 10/26/2025).
- 106Bertin, E.; Parazza, F.; Chassery, J. M. Segmentation and measurement based on 3D Voronoi diagram: Application to confocal microscopy. Computerized Medical Imaging and Graphics 1993, 17 (3), 175– 182, DOI: 10.1016/0895-6111(93)90041-KGoogle ScholarThere is no corresponding record for this reference.
- 107Eils, R.; Bertin, E.; Saracoglu, K.; Rinke, B.; Schröck, E.; Parazza, F.; Usson, Y.; Robert-Nicoud, M.; Stelzer, E. H.; Chassery, J. M. Application of confocal laser microscopy and three-dimensional Voronoi diagrams for volume and surface estimates of interphase chromosomes. J. Microsc 1995, 177 (Pt 2), 150– 161, DOI: 10.1111/j.1365-2818.1995.tb03545.xGoogle ScholarThere is no corresponding record for this reference.
From NLM.
- 108Andronov, L.; Orlov, I.; Lutz, Y.; Vonesch, J.-L.; Klaholz, B. P. ClusterViSu, a method for clustering of protein complexes by Voronoi tessellation in super-resolution microscopy. Sci. Rep. 2016, 6 (1), 24084, DOI: 10.1038/srep24084Google ScholarThere is no corresponding record for this reference.
- 109Levet, F.; Hosy, E.; Kechkar, A.; Butler, C.; Beghin, A.; Choquet, D.; Sibarita, J.-B. SR-Tesseler: a method to segment and quantify localization-based super-resolution microscopy data. Nat. Methods 2015, 12 (11), 1065– 1071, DOI: 10.1038/nmeth.3579Google ScholarThere is no corresponding record for this reference.
- 110Andronov, L.; Lutz, Y.; Vonesch, J. L.; Klaholz, B. P. SharpViSu: integrated analysis and segmentation of super-resolution microscopy data. Bioinformatics 2016, 32 (14), 2239– 2241, DOI: 10.1093/bioinformatics/btw123Google ScholarThere is no corresponding record for this reference.
From NLM.
- 111Rubin-Delanchy, P.; Burn, G. L.; Griffié, J.; Williamson, D. J.; Heard, N. A.; Cope, A. P.; Owen, D. M. Bayesian cluster identification in single-molecule localization microscopy data. Nat. Methods 2015, 12 (11), 1072– 1076, DOI: 10.1038/nmeth.3612Google ScholarThere is no corresponding record for this reference.
- 112Tang, Y.; Hendriks, J.; Gensch, T.; Dai, L.; Li, J. Automatic Bayesian single molecule identification for localization microscopy. Sci. Rep. 2016, 6 (1), 33521, DOI: 10.1038/srep33521Google ScholarThere is no corresponding record for this reference.
- 113Griffié, J.; Shannon, M.; Bromley, C. L.; Boelen, L.; Burn, G. L.; Williamson, D. J.; Heard, N. A.; Cope, A. P.; Owen, D. M.; Rubin-Delanchy, P. A Bayesian cluster analysis method for single-molecule localization microscopy data. Nat. Protoc. 2016, 11 (12), 2499– 2514, DOI: 10.1038/nprot.2016.149Google ScholarThere is no corresponding record for this reference.
- 114Hammer, J. L.; Devanny, A. J.; Kaufman, L. J. Bayesian optimized parameter selection for density-based clustering applied to single molecule localization microscopy. Communications Biology 2025, 8 (1), 902, DOI: 10.1038/s42003-025-08332-0Google ScholarThere is no corresponding record for this reference.
- 115Andronov, L.; Han, M.; Zhu, Y.; Balaji, A.; Roy, A. R.; Barentine, A. E. S.; Patel, P.; Garhyan, J.; Qi, L. S.; Moerner, W. E. Nanoscale cellular organization of viral RNA and proteins in SARS-CoV-2 replication organelles. Nat. Commun. 2024, 15 (1), 4644, DOI: 10.1038/s41467-024-48991-xGoogle ScholarThere is no corresponding record for this reference.
- 116Mayerich, D.; Sun, R. Superresolution Image Processing. In Microscope Image Processing, Second ed.; Merchant, F. A., Castleman, K. R., Eds.; Academic Press, 2023; Chapter Twelve, pp 319– 334.Google ScholarThere is no corresponding record for this reference.
- 117Jimenez, A.; Friedl, K.; Leterrier, C. About samples, giving examples: Optimized Single Molecule Localization Microscopy. Methods 2020, 174, 100– 114, DOI: 10.1016/j.ymeth.2019.05.008Google ScholarThere is no corresponding record for this reference.
- 118Chen, F.; Wassie, A. T.; Cote, A. J.; Sinha, A.; Alon, S.; Asano, S.; Daugharthy, E. R.; Chang, J.-B.; Marblestone, A.; Church, G. M.; Raj, A.; Boyden, E. S. Nanoscale imaging of RNA with expansion microscopy. Nat. Methods 2016, 13 (8), 679– 684, DOI: 10.1038/nmeth.3899Google ScholarThere is no corresponding record for this reference.
- 119Klar, T. A.; Jakobs, S.; Dyba, M.; Egner, A.; Hell, S. W. Fluorescence microscopy with diffraction resolution barrier broken by stimulated emission. Proc. Natl. Acad. Sci. U. S. A. 2000, 97 (15), 8206– 8210, DOI: 10.1073/pnas.97.15.8206Google ScholarThere is no corresponding record for this reference.
- 120Balzarotti, F.; Eilers, Y.; Gwosch, K. C.; Gynnå, A. H.; Westphal, V.; Stefani, F. D.; Elf, J.; Hell, S. W. Nanometer resolution imaging and tracking of fluorescent molecules with minimal photon fluxes. Science 2017, 355 (6325), 606– 612, DOI: 10.1126/science.aak9913Google ScholarThere is no corresponding record for this reference.
- 121Gustafsson, M. G. Surpassing the lateral resolution limit by a factor of two using structured illumination microscopy. J. Microsc. 2000, 198 (2), 82– 87, DOI: 10.1046/j.1365-2818.2000.00710.xGoogle ScholarThere is no corresponding record for this reference.
From NLM.
- 122Betzig, E.; Patterson, G. H.; Sougrat, R.; Lindwasser, O. W.; Olenych, S.; Bonifacino, J. S.; Davidson, M. W.; Lippincott-Schwartz, J.; Hess, H. F. Imaging intracellular fluorescent proteins at nanometer resolution. Science 2006, 313 (5793), 1642– 1645, DOI: 10.1126/science.1127344Google ScholarThere is no corresponding record for this reference.
From NLM.
- 123Rust, M. J.; Bates, M.; Zhuang, X. Sub-diffraction-limit imaging by stochastic optical reconstruction microscopy (STORM). Nat. Methods 2006, 3 (10), 793– 796, DOI: 10.1038/nmeth929Google ScholarThere is no corresponding record for this reference.
- 124Schnitzbauer, J.; Strauss, M. T.; Schlichthaerle, T.; Schueder, F.; Jungmann, R. Super-resolution microscopy with DNA-PAINT. Nat. Protoc. 2017, 12 (6), 1198– 1228, DOI: 10.1038/nprot.2017.024Google ScholarThere is no corresponding record for this reference.
- 125Arnould, B.; Quillin, A. L.; Heemstra, J. M. Tracking the Message: Applying Single Molecule Localization Microscopy to Cellular RNA Imaging. Chembiochem 2023, 24 (10), e202300049 DOI: 10.1002/cbic.202300049Google ScholarThere is no corresponding record for this reference.
From NLM.
- 126Lelek, M.; Gyparaki, M. T.; Beliu, G.; Schueder, F.; Griffié, J.; Manley, S.; Jungmann, R.; Sauer, M.; Lakadamyali, M.; Zimmer, C. Single-molecule localization microscopy. Nature Reviews Methods Primers 2021, 1 (1), 39, DOI: 10.1038/s43586-021-00038-xGoogle ScholarThere is no corresponding record for this reference.
- 127Hugelier, S.; Colosi, P. L.; Lakadamyali, M. Quantitative Single-Molecule Localization Microscopy. Annu. Rev. Biophys 2023, 52, 139– 160, DOI: 10.1146/annurev-biophys-111622-091212Google ScholarThere is no corresponding record for this reference.
From NLM.
- 128Hyun, Y.; Kim, D. Recent development of computational cluster analysis methods for single-molecule localization microscopy images. Comput. Struct Biotechnol J. 2023, 21, 879– 888, DOI: 10.1016/j.csbj.2023.01.006Google ScholarThere is no corresponding record for this reference.
From NLM.
- 129Khater, I. M.; Nabi, I. R.; Hamarneh, G. A Review of Super-Resolution Single-Molecule Localization Microscopy Cluster Analysis and Quantification Methods. Patterns (N Y) 2020, 1 (3), 100038, DOI: 10.1016/j.patter.2020.100038Google ScholarThere is no corresponding record for this reference.
From NLM.
- 130Nieves, D. J.; Pike, J. A.; Levet, F.; Williamson, D. J.; Baragilly, M.; Oloketuyi, S.; de Marco, A.; Griffié, J.; Sage, D.; Cohen, E. A. K.; Sibarita, J.-B.; Heilemann, M.; Owen, D. M. A framework for evaluating the performance of SMLM cluster analysis algorithms. Nat. Methods 2023, 20 (2), 259– 267, DOI: 10.1038/s41592-022-01750-6Google ScholarThere is no corresponding record for this reference.
- 131Ejdrup, A. L.; Lycas, M. D.; Lorenzen, N.; Konomi, A.; Herborg, F.; Madsen, K. L.; Gether, U. A density-based enrichment measure for assessing colocalization in single-molecule localization microscopy data. Nat. Commun. 2022, 13 (1), 4388, DOI: 10.1038/s41467-022-32064-yGoogle ScholarThere is no corresponding record for this reference.
- 132Ovesný, M.; Křížek, P.; Borkovec, J.; Svindrych, Z.; Hagen, G. M. ThunderSTORM: a comprehensive ImageJ plug-in for PALM and STORM data analysis and super-resolution imaging. Bioinformatics 2014, 30 (16), 2389– 2390, DOI: 10.1093/bioinformatics/btu202Google ScholarThere is no corresponding record for this reference.
From NLM.
- 133Wolter, S.; Löschberger, A.; Holm, T.; Aufmkolk, S.; Dabauvalle, M.-C.; van de Linde, S.; Sauer, M. rapidSTORM: accurate, fast open-source software for localization microscopy. Nat. Methods 2012, 9 (11), 1040– 1041, DOI: 10.1038/nmeth.2224Google ScholarThere is no corresponding record for this reference.
- 134Sage, D.; Kirshner, H.; Pengo, T.; Stuurman, N.; Min, J.; Manley, S.; Unser, M. Quantitative evaluation of software packages for single-molecule localization microscopy. Nat. Methods 2015, 12 (8), 717– 724, DOI: 10.1038/nmeth.3442Google ScholarThere is no corresponding record for this reference.
- 135Liu, S.; Wang, C. Y.; Zheng, P.; Jia, B. B.; Zemke, N. R.; Ren, P.; Park, H. L.; Ren, B.; Zhuang, X. Cell type–specific 3D-genome organization and transcription regulation in the brain. Science Advances 2025, 11 (9), eadv2067 DOI: 10.1126/sciadv.adv2067Google ScholarThere is no corresponding record for this reference.
- 136Shivanandan, A.; Deschout, H.; Scarselli, M.; Radenovic, A. Challenges in quantitative single molecule localization microscopy. FEBS letters 2014, 588 (19), 3595– 3602, DOI: 10.1016/j.febslet.2014.06.014Google ScholarThere is no corresponding record for this reference.
- 137Olivier, N.; Keller, D.; Rajan, V. S.; Gönczy, P.; Manley, S. Simple buffers for 3D STORM microscopy. Biomed Opt Express 2013, 4 (6), 885– 899, DOI: 10.1364/BOE.4.000885Google ScholarThere is no corresponding record for this reference.
From NLM.
- 138Shcherbakova, D. M.; Sengupta, P.; Lippincott-Schwartz, J.; Verkhusha, V. V. Photocontrollable fluorescent proteins for superresolution imaging. Annu. Rev. Biophys 2014, 43, 303– 329, DOI: 10.1146/annurev-biophys-051013-022836Google ScholarThere is no corresponding record for this reference.
From NLM.
- 139Godin, A. G.; Lounis, B.; Cognet, L. Super-resolution microscopy approaches for live cell imaging. Biophys. J. 2014, 107 (8), 1777– 1784, DOI: 10.1016/j.bpj.2014.08.028Google ScholarThere is no corresponding record for this reference.
From NLM.
- 140Xu, J.; Ma, H.; Liu, Y. Stochastic Optical Reconstruction Microscopy (STORM). Curr. Protoc Cytom 2017, 81, 12.46.11– 12.46.27, DOI: 10.1002/cpcy.23Google ScholarThere is no corresponding record for this reference.
From NLM.
- 141Cho, W.-K.; Jayanth, N.; English, B. P.; Inoue, T.; Andrews, J. O.; Conway, W.; Grimm, J. B.; Spille, J.-H.; Lavis, L. D.; Lionnet, T.; Cisse, I. I. RNA Polymerase II cluster dynamics predict mRNA output in living cells. eLife 2016, 5, e13617 DOI: 10.7554/eLife.13617Google ScholarThere is no corresponding record for this reference.
- 142Dankovich, T. M.; Rizzoli, S. O. Challenges facing quantitative large-scale optical super-resolution, and some simple solutions. iScience 2021, 24 (3), 102134, DOI: 10.1016/j.isci.2021.102134Google ScholarThere is no corresponding record for this reference.
- 143Cho, W.-K.; Spille, J.-H.; Hecht, M.; Lee, C.; Li, C.; Grube, V.; Cisse, I. I. Mediator and RNA polymerase II clusters associate in transcription-dependent condensates. Science 2018, 361 (6400), 412– 415, DOI: 10.1126/science.aar4199Google ScholarThere is no corresponding record for this reference.
- 144Cisse, I. I.; Izeddin, I.; Causse, S. Z.; Boudarene, L.; Senecal, A.; Muresan, L.; Dugast-Darzacq, C.; Hajj, B.; Dahan, M.; Darzacq, X. Real-Time Dynamics of RNA Polymerase II Clustering in Live Human Cells. Science 2013, 341 (6146), 664– 667, DOI: 10.1126/science.1239053Google ScholarThere is no corresponding record for this reference.
- 145Yang, Y.; Qu, N.; Tan, J.; Rushdi, M. N.; Krueger, C. J.; Chen, A. K. Roles of Gag-RNA interactions in HIV-1 virus assembly deciphered by single-molecule localization microscopy. Proc. Natl. Acad. Sci. U. S. A. 2018, 115 (26), 6721– 6726, DOI: 10.1073/pnas.1805728115Google ScholarThere is no corresponding record for this reference.
From NLM.
- 146Xie, L.; Dong, P.; Chen, X.; Hsieh, T. S.; Banala, S.; De Marzio, M.; English, B. P.; Qi, Y.; Jung, S. K.; Kieffer-Kwon, K. R.; Legant, W. R.; Hansen, A. S.; Schulmann, A.; Casellas, R.; Zhang, B.; Betzig, E.; Lavis, L. D.; Chang, H. Y.; Tjian, R.; Liu, Z. 3D ATAC-PALM: super-resolution imaging of the accessible genome. Nat. Methods 2020, 17 (4), 430– 436, DOI: 10.1038/s41592-020-0775-2Google ScholarThere is no corresponding record for this reference.
From NLM.
- 147Sunwoo, H.; Wu, J. Y.; Lee, J. T. The Xist RNA-PRC2 complex at 20-nm resolution reveals a low Xist stoichiometry and suggests a hit-and-run mechanism in mouse cells. Proc. Natl. Acad. Sci. U. S. A. 2015, 112 (31), E4216– 4225, DOI: 10.1073/pnas.1503690112Google ScholarThere is no corresponding record for this reference.
From NLM.
- 148Maclary, E.; Hinten, M.; Harris, C.; Sethuraman, S.; Gayen, S.; Kalantry, S. PRC2 represses transcribed genes on the imprinted inactive X chromosome in mice. Genome Biology 2017, 18 (1), 82, DOI: 10.1186/s13059-017-1211-5Google ScholarThere is no corresponding record for this reference.
- 149Castells-Garcia, A.; Ed-daoui, I.; González-Almela, E.; Vicario, C.; Ottestrom, J.; Lakadamyali, M.; Neguembor, M. V.; Cosma, Maria P. Super resolution microscopy reveals how elongating RNA polymerase II and nascent RNA interact with nucleosome clutches. Nucleic Acids Res. 2022, 50 (1), 175– 190, DOI: 10.1093/nar/gkab1215Google ScholarThere is no corresponding record for this reference.
(accessed 10/26/2025).
- 150Hilbert, L.; Sato, Y.; Kuznetsova, K.; Bianucci, T.; Kimura, H.; Jülicher, F.; Honigmann, A.; Zaburdaev, V.; Vastenhouw, N. L. Transcription organizes euchromatin via microphase separation. Nat. Commun. 2021, 12 (1), 1360, DOI: 10.1038/s41467-021-21589-3Google ScholarThere is no corresponding record for this reference.
From NLM.
- 151Snijder, E. J.; Limpens, R.; de Wilde, A. H.; de Jong, A. W. M.; Zevenhoven-Dobbe, J. C.; Maier, H. J.; Faas, F.; Koster, A. J.; Bárcena, M. A unifying structural and functional model of the coronavirus replication organelle: Tracking down RNA synthesis. PLoS Biol. 2020, 18 (6), e3000715 DOI: 10.1371/journal.pbio.3000715Google ScholarThere is no corresponding record for this reference.
From NLM.
- 152Jungmann, R.; Steinhauer, C.; Scheible, M.; Kuzyk, A.; Tinnefeld, P.; Simmel, F. C. Single-molecule kinetics and super-resolution microscopy by fluorescence imaging of transient binding on DNA origami. Nano Lett. 2010, 10 (11), 4756– 4761, DOI: 10.1021/nl103427wGoogle ScholarThere is no corresponding record for this reference.
From NLM.
- 153Jungmann, R.; Avendaño, M. S.; Woehrstein, J. B.; Dai, M.; Shih, W. M.; Yin, P. Multiplexed 3D cellular super-resolution imaging with DNA-PAINT and Exchange-PAINT. Nat. Methods 2014, 11 (3), 313– 318, DOI: 10.1038/nmeth.2835Google ScholarThere is no corresponding record for this reference.
From NLM.
- 154Iinuma, R.; Ke, Y.; Jungmann, R.; Schlichthaerle, T.; Woehrstein, J. B.; Yin, P. Polyhedra self-assembled from DNA tripods and characterized with 3D DNA-PAINT. Science 2014, 344 (6179), 65– 69, DOI: 10.1126/science.1250944Google ScholarThere is no corresponding record for this reference.
From NLM.
- 155Jungmann, R.; Avendaño, M. S.; Dai, M.; Woehrstein, J. B.; Agasti, S. S.; Feiger, Z.; Rodal, A.; Yin, P. Quantitative super-resolution imaging with qPAINT. Nat. Methods 2016, 13 (5), 439– 442, DOI: 10.1038/nmeth.3804Google ScholarThere is no corresponding record for this reference.
From NLM.
- 156Schäfer, P.; Tenschert, W.; Schröter, M.; Gutensohn, K.; Laufs, R. False-Positive Results of Plasma PCR for Cytomegalovirus DNA due to Delayed Sample Preparation. Journal of Clinical Microbiology 2000, 38 (9), 3249– 3253, DOI: 10.1128/JCM.38.9.3249-3253.2000Google ScholarThere is no corresponding record for this reference.
(accessed 2025/10/26).
- 157Faubert, A.; Bohsina, R.; You, E.; Hébert, M.; Griffith, M.; Aubin, M. J. Diagnostic Accuracy of Conjunctival Rt-Pcr in Sars-Cov-2: A Systematic Review and Diagnostic Accuracy Meta-Analysis. Ocul Immunol Inflamm 2024, 32 (8), 1621– 1632, DOI: 10.1080/09273948.2023.2272200Google ScholarThere is no corresponding record for this reference.
From NLM.
- 158Ruijter, J. M.; Barnewall, R. J.; Marsh, I. B.; Szentirmay, A. N.; Quinn, J. C.; van Houdt, R.; Gunst, Q. D.; van den Hoff, M. J. B. Efficiency Correction Is Required for Accurate Quantitative PCR Analysis and Reporting. Clinical Chemistry 2021, 67 (6), 829– 842, DOI: 10.1093/clinchem/hvab052Google ScholarThere is no corresponding record for this reference.
(accessed 10/26/2025).
- 159Louw, T. M.; Booth, C. S.; Pienaar, E.; TerMaat, J. R.; Whitney, S. E.; Viljoen, H. J. Experimental validation of a fundamental model for PCR efficiency. Chem. Eng. Sci. 2011, 66 (8), 1783– 1789, DOI: 10.1016/j.ces.2011.01.029Google ScholarThere is no corresponding record for this reference.
- 160Zhao, L.; Qiu, M.; Li, X.; Yang, J.; Li, J. CRISPR-Cas13a system: A novel tool for molecular diagnostics. Front Microbiol 2022, 13, 1060947, DOI: 10.3389/fmicb.2022.1060947Google ScholarThere is no corresponding record for this reference.
From NLM.
- 161Metcalf, G. A. D. MicroRNAs: circulating biomarkers for the early detection of imperceptible cancers via biosensor and machine-learning advances. Oncogene 2024, 43 (28), 2135– 2142, DOI: 10.1038/s41388-024-03076-3Google ScholarThere is no corresponding record for this reference.
- 162Schofield, A. L.; Brown, J. P.; Brown, J.; Wilczynska, A.; Bell, C.; Glaab, W. E.; Hackl, M.; Howell, L.; Lee, S.; Dear, J. W.; Remes, M.; Reeves, P.; Zhang, E.; Allmer, J.; Norris, A.; Falciani, F.; Takeshita, L. Y.; Seyed Forootan, S.; Sutton, R.; Park, B. K.; Goldring, C. Systems analysis of miRNA biomarkers to inform drug safety. Arch. Toxicol. 2021, 95 (11), 3475– 3495, DOI: 10.1007/s00204-021-03150-9Google ScholarThere is no corresponding record for this reference.
From NLM.
- 163Visone, R.; Croce, C. M. MiRNAs and cancer. Am. J. Pathol. 2009, 174 (4), 1131– 1138, DOI: 10.2353/ajpath.2009.080794Google ScholarThere is no corresponding record for this reference.
From NLM.
- 164Kroh, E. M.; Parkin, R. K.; Mitchell, P. S.; Tewari, M. Analysis of circulating microRNA biomarkers in plasma and serum using quantitative reverse transcription-PCR (qRT-PCR). Methods 2010, 50 (4), 298– 301, DOI: 10.1016/j.ymeth.2010.01.032Google ScholarThere is no corresponding record for this reference.
From NLM.
- 165Vester, B.; Wengel, J. LNA (locked nucleic acid): high-affinity targeting of complementary RNA and DNA. Biochemistry 2004, 43 (42), 13233– 13241, DOI: 10.1021/bi0485732Google ScholarThere is no corresponding record for this reference.
From NLM.
- 166Woo, N.; Brown, C. M. Review of expansion microscopy combined with advanced imaging modalities. J. Microsc. 2026, 301, 335, DOI: 10.1111/jmi.70048Google ScholarThere is no corresponding record for this reference.
- 167Xia, C.; Fan, J.; Emanuel, G.; Hao, J.; Zhuang, X. Spatial transcriptome profiling by MERFISH reveals subcellular RNA compartmentalization and cell cycle-dependent gene expression. Proc. Natl. Acad. Sci. U. S. A. 2019, 116 (39), 19490– 19499, DOI: 10.1073/pnas.1912459116Google ScholarThere is no corresponding record for this reference.
- 168Stoldt, S.; Maass, F.; Weber, M.; Dennerlein, S.; Ilgen, P.; Gärtner, J.; Canfes, A.; Schweighofer, S. V.; Jans, D. C.; Rehling, P.; Jakobs, S. Super-resolution microscopy of mitochondrial mRNAs. Nat. Commun. 2025, 16 (1), 6391, DOI: 10.1038/s41467-025-61577-5Google ScholarThere is no corresponding record for this reference.
- 169Rodermund, L.; Coker, H.; Oldenkamp, R.; Wei, G.; Bowness, J.; Rajkumar, B.; Nesterova, T.; Susano Pinto, D. M.; Schermelleh, L.; Brockdorff, N. Time-resolved structured illumination microscopy reveals key principles of Xist RNA spreading. Science 2021, 372 (6547), eabe7500 DOI: 10.1126/science.abe7500Google ScholarThere is no corresponding record for this reference.
- 170Shao, W.; Zeng, S.-T.; Yu, Z.-Y.; Tang, G.-X.; Chen, S.-B.; Huang, Z.-S.; Chen, X.-C.; Tan, J.-H. Tracking Stress Granule Dynamics in Live Cells and In Vivo with a Small Molecule. Anal. Chem. 2021, 93 (49), 16297– 16301, DOI: 10.1021/acs.analchem.1c03577Google ScholarThere is no corresponding record for this reference.
- 171Cawte, A. D.; Unrau, P. J.; Rueda, D. S. Live cell imaging of single RNA molecules with fluorogenic Mango II arrays. Nat. Commun. 2020, 11 (1), 1283, DOI: 10.1038/s41467-020-14932-7Google ScholarThere is no corresponding record for this reference.
- 172Park, H. Y.; Trcek, T.; Wells, A. L.; Chao, J. A.; Singer, R. H. An Unbiased Analysis Method to Quantify mRNA Localization Reveals Its Correlation with Cell Motility. Cell Reports 2012, 1 (2), 179– 184, DOI: 10.1016/j.celrep.2011.12.009Google ScholarThere is no corresponding record for this reference.
(accessed 2025/10/26).
- 173Tinevez, J. Y.; Perry, N.; Schindelin, J.; Hoopes, G. M.; Reynolds, G. D.; Laplantine, E.; Bednarek, S. Y.; Shorte, S. L.; Eliceiri, K. W. TrackMate: An open and extensible platform for single-particle tracking. Methods 2017, 115, 80– 90, DOI: 10.1016/j.ymeth.2016.09.016Google ScholarThere is no corresponding record for this reference.
From NLM.
- 174Liu, J.; Cho, I.-H.; Cui, Y.; Irudayaraj, J. Second Harmonic Super-resolution Microscopy for Quantification of mRNA at Single Copy Sensitivity. ACS Nano 2014, 8 (12), 12418– 12427, DOI: 10.1021/nn505096tGoogle ScholarThere is no corresponding record for this reference.
- 175Campagnola, P. J.; Clark, H. A.; Mohler, W. A.; Lewis, A.; Loew, L. M. Second-harmonic imaging microscopy of living cells. J. Biomed Opt 2001, 6 (3), 277– 286, DOI: 10.1117/1.1383294Google ScholarThere is no corresponding record for this reference.
From NLM.
- 176Aghigh, A.; Bancelin, S.; Rivard, M.; Pinsard, M.; Ibrahim, H.; Légaré, F. Second harmonic generation microscopy: a powerful tool for bio-imaging. Biophys Rev. 2023, 15 (1), 43– 70, DOI: 10.1007/s12551-022-01041-6Google ScholarThere is no corresponding record for this reference.
From NLM.
- 177Chen, X.; Nadiarynkh, O.; Plotnikov, S.; Campagnola, P. J. Second harmonic generation microscopy for quantitative analysis of collagen fibrillar structure. Nat. Protoc 2012, 7 (4), 654– 669, DOI: 10.1038/nprot.2012.009Google ScholarThere is no corresponding record for this reference.
From NLM.
- 178Stanciu, S. G.; Hristu, R.; Stanciu, G. A.; Tranca, D. E.; Eftimie, L.; Dumitru, A.; Costache, M.; Stenmark, H. A.; Manders, H.; Cherian, A.; Tark-Dame, M.; Manders, E. M. M. Super-resolution re-scan second harmonic generation microscopy. Proc. Natl. Acad. Sci. U. S. A. 2022, 119 (47), e2214662119 DOI: 10.1073/pnas.2214662119Google ScholarThere is no corresponding record for this reference.
- 179Ding, J.; Lee, Y.-T.; Bhandari, Y.; Schwieters, C. D.; Fan, L.; Yu, P.; Tarosov, S. G.; Stagno, J. R.; Ma, B.; Nussinov, R.; Rein, A.; Zhang, J.; Wang, Y.-X. Visualizing RNA conformational and architectural heterogeneity in solution. Nat. Commun. 2023, 14 (1), 714, DOI: 10.1038/s41467-023-36184-xGoogle ScholarThere is no corresponding record for this reference.
- 180Degenhardt, M. F. S.; Degenhardt, H. F.; Bhandari, Y. R.; Lee, Y.-T.; Ding, J.; Yu, P.; Heinz, W. F.; Stagno, J. R.; Schwieters, C. D.; Watts, N. R.; Wingfield, P. T.; Rein, A.; Zhang, J.; Wang, Y.-X. Determining structures of RNA conformers using AFM and deep neural networks. Nature 2025, 637 (8048), 1234– 1243, DOI: 10.1038/s41586-024-07559-xGoogle ScholarThere is no corresponding record for this reference.
- 181Luo, B.; Zhang, C.; Ling, X.; Mukherjee, S.; Jia, G.; Xie, J.; Jia, X.; Liu, L.; Baulin, E. F.; Luo, Y.; Jiang, L.; Dong, H.; Wei, X.; Bujnicki, J. M.; Su, Z. Cryo-EM reveals dynamics of Tetrahymena group I intron self-splicing. Nature Catalysis 2023, 6 (4), 298– 309, DOI: 10.1038/s41929-023-00934-3Google ScholarThere is no corresponding record for this reference.
- 182Liu, J.; McRae, E. K. S.; Zhang, M.; Geary, C.; Andersen, E. S.; Ren, G. Non-averaged single-molecule tertiary structures reveal RNA self-folding through individual-particle cryo-electron tomography. Nat. Commun. 2024, 15 (1), 9084, DOI: 10.1038/s41467-024-52914-1Google ScholarThere is no corresponding record for this reference.
- 183Krieg, M.; Fläschner, G.; Alsteens, D.; Gaub, B. M.; Roos, W. H.; Wuite, G. J. L.; Gaub, H. E.; Gerber, C.; Dufrêne, Y. F.; Müller, D. J. Atomic force microscopy-based mechanobiology. Nature Reviews Physics 2019, 1 (1), 41– 57, DOI: 10.1038/s42254-018-0001-7Google ScholarThere is no corresponding record for this reference.
- 184Berrevoets, E. S.; Kessler, L. F.; Balakrishnan, A.; Okuda, E. K.; Müller-McNicoll, M.; Rieger, B.; Stallinga, S.; Heilemann, M. Smart 3D super-resolution microscopy reveals the architecture of the RNA scaffold in a nuclear body. Nat. Commun. 2025, 16 (1), 10689, DOI: 10.1038/s41467-025-65723-xGoogle ScholarThere is no corresponding record for this reference.
- 185Autour, A.; C. Y. Jeng, S.; D. Cawte, A.; Abdolahzadeh, A.; Galli, A.; Panchapakesan, S. S. S.; Rueda, D.; Ryckelynck, M.; Unrau, P. J. Fluorogenic RNA Mango aptamers for imaging small non-coding RNAs in mammalian cells. Nat. Commun. 2018, 9 (1), 656, DOI: 10.1038/s41467-018-02993-8Google ScholarThere is no corresponding record for this reference.
From NLM.
Cited By
This article has not yet been cited by other publications.
Article Views
Altmetric
Citations
Article Views are the COUNTER-compliant sum of full text article downloads since November 2008 (both PDF and HTML) across all institutions and individuals. These metrics are regularly updated to reflect usage leading up to the last few days.
Citations are the number of other articles citing this article, calculated by Crossref and updated daily. Find more information about Crossref citation counts.
The Altmetric Attention Score is a quantitative measure of the attention that a research article has received online. Clicking on the donut icon will load a page at altmetric.com with additional details about the score and the social media presence for the given article. Find more information on the Altmetric Attention Score and how the score is calculated.
Recommended Articles
Abstract
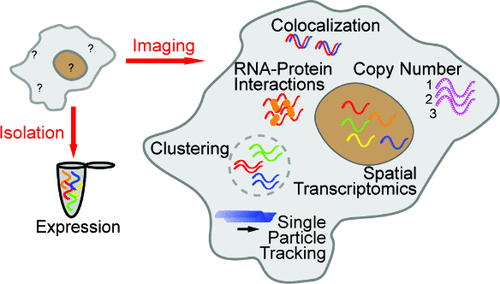
Figure 1

Figure 1. Common fixed-cell imaging techniques used for RNA quantification. (A) Illustration of smFISH (top) to label GFP RNA in CHO cells. Reproduced from ref (39) under Creative Commons License CC-BY-NC-ND. Copyright 2008 Raj A; et al. Published by Springer Nature. (B) Illustration of RNAscope (right) and an example of RNAscope being used for multicolor detection of β-actin, PLP0 (60S acidic ribosomal protein P0), PPIB (peptidylprolyl isomerase B), and HPRT-1 (hypoxanthine phosphoribosyltransferase 1) (left). Reproduced from ref (44) under Creative Commons License CC-BY-NC-ND. Copyright 2012 Wang, F; et al. Published by Elsevier. (C) Illustration of RCAFISH with the target padlock probe (bottom) to image TK1 mRNA in MCF-7 cells. Adapted from ref (50) under Creative Commons License CC BY 3.0. Copyright 2017 Deng, R.; et al. Published by Royal Society of Chemistry. (D) Illustration of HCR FISH (top) and validation of the technique by detecting EGFP mRNA in wild-type Arabidopsis. Adapted from ref (46) under Creative Commons License CC BY 4.0. Copyright 2023 Huang, T.; et al. Published by Springer Nature. (E) Principle of seqFISH and example images. Reproduced from ref (54) under Creative Commons License CC-BY-NC-ND. Copyright 2014 Lubeck, E.; et al. Published by Springer Nature. (F) MERFISH workflow (left) and images of RNA molecules in an IMR90 cell after each hybridization round. Adapted with permission from ref (59). Copyright 2015 AAAS.
Figure 2
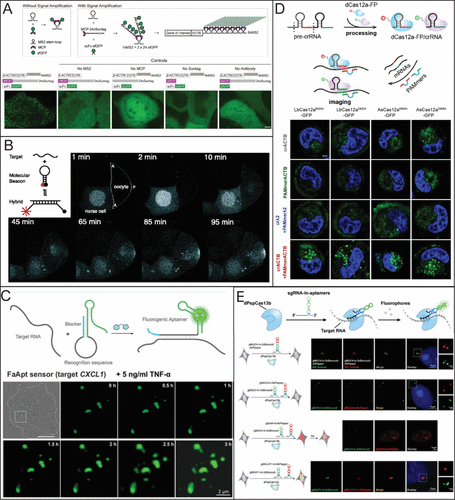
Figure 2. Common live-cell imaging techniques used for RNA quantification. (A) Schematic of MS2-MCP system and MS2-based signal amplification with the suntag system (top) and representative live-cell images of β-actin (bottom). Reporduced from ref (67) under Creative Commons License CC BY 4.0. Copyright 2023 Hu Y.; et al. Published by eLife; (B) Illustration of Molecular beacons for live-cell imaging being used to visualize the transport of native oskar mRNA from a nurse cell to the posterior cortex of the oocyte. Adapted with permission from ref (82). Copyright (2003) National Academy of Sciences, U.S.A. (C) Example of fluorogenic RNA being used to target CXCL1 mRNA after 5 ng/mL TNF-α treatment. Adapted with permission from ref (68). Copyright 2023 American Chemical Society. (D) Example of different dCas12a mutants fused with GFP in the presence of a PAMmer sequence targeting β-actin mRNA in HeLa cells. Reporduced with permission from ref (77). Copyright 2024 American Chemical Society. (E) dCas 13b with different RNA sgRNA aptamers for multicolor imaging of MUC4 and SatIII RNA. Reproduced from ref (79) under the Creative Commons License CC BY-NC 3.0. Copyright 2022 Tang, H.; et al. Published by Royal Society of Chemistry.
Figure 3
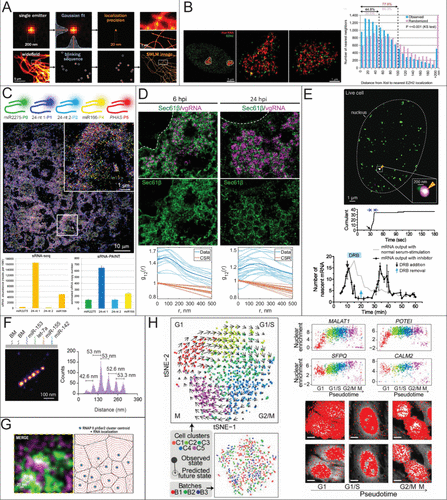
Figure 3. RNA quantification using STORM, PAINT, and ExM. (A) Fluorophore localization for SMLM reconstruction. Reproduced with permission from ref (117). Copyright 2020 Elsevier. (B) Nearest Neighbor distances to count the number of Xist molecules and their distance to a histone marker, respectively. Reproduced with permission from ref (147). Copyright 2015 PNAS. (C) Localization of different sRNAs using sRNA-PAINT and their reported expression levels compared to RNA-seq Reproduced from ref (99) under Creative Commons License CC BY 4.0. Copyright 2020 Published by Oxford Academic Huang, K.; et al. (D) Bivariate pair correlation to measure the correlation between Sec61β with vgRNA and dsRNA and Sec61β with nsp3. Reproduced from ref (115) under Creative Commons License CC BY 4.0. Copyright 2024 Published by Springer Nature. Andronov, L.; et al. (E) Super-resolution time trace of Pol II cluster colocalizing with the active gene locus of β-actin (top) and real-time monitoring of mRNA output of ACTB following serum stimulation (bottom). Reproduced from ref (141) under Creative Commons License CC BY 4.0. Copyright 2016 Published by elife. Cho, W.-K.; et al. (F) Detection of miRNA using DNA PAINT. Expression reported by counts and each peak is a different miRNA. Reproduced from ref (13) under Creative Commons License CC-BY-NC-ND. Copyright 2023 Published by Elsevier Kocabey, S.; et al. (G) Voronoi Tessellation of RNA nanodomains clustering to different RNAP II using STORM and DNA-PAINT. Reproduced from ref (149) under Creative Commons License CC BY 4.0. Copyright 2022 Published by Oxford Academic. Castells-Garcia, A et al. (H) Spatial transcriptome wide analysis using expansion microscopy and MERFISH. Reproduced from ref (167) under Creative Commons License CC BY-NC-ND. Copyright 2019 Published by National Academy of Sciences Xia, C.; et al.
Figure 4
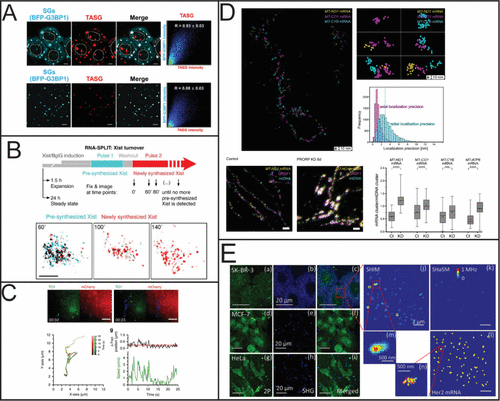
Figure 4. RNA quantification using SIM, STED, MINFLUX, and SHaSM. (A) SIM imaging of stress granules using a small molecule fluorescent probe (scale bar 5 μm). Reproduced from ref (170). Copyright 2023 American Chemical Society. (B) Schematic of using RNA-SPLIT to monitor Xist Turnover and representative 3D SIM images of Xist turnover during expansion. Reproduced with permission from ref (169). Copyright 2021 AAAS. (C) Single particle tracking of the comovement of TOI1-B and tdMCP-mCherry labeled trajectories. Reproduced from ref (171) under Creative Commons License CC BY 4.0. Copyright 2020 Cawte, A. D. et al. Published by Springer Nature. (D) Subcellular characterization of mtRNA using STED and MINFLUX. Reproduced from ref (168) under Creative Commons License CC BY 4.0. Copyright 2025 Stoldt, S.; et al. Published by Springer Nature. (E) Detection of Her2 mRNA in three different cell lines using SHaSM. Reproduced from ref (174). Copyright 2014 American Chemical Society.
References
This article references 185 other publications.
- 1Cao, X.; Zhang, Y.; Ding, Y.; Wan, Y. Identification of RNA structures and their roles in RNA functions. Nat. Rev. Mol. Cell Biol. 2024, 25 (10), 784– 801, DOI: 10.1038/s41580-024-00748-6There is no corresponding record for this reference.
- 2Singh, K. P.; Miaskowski, C.; Dhruva, A. A.; Flowers, E.; Kober, K. M. Mechanisms and Measurement of Changes in Gene Expression. Biol. Res. Nurs 2018, 20 (4), 369– 382, DOI: 10.1177/1099800418772161There is no corresponding record for this reference.
From NLM.
- 3Chen, Y.; Davidson, N. M.; Wan, Y. K.; Yao, F.; Su, Y.; Gamaarachchi, H.; Sim, A.; Patel, H.; Low, H. M.; Hendra, C.; Wratten, L.; Hakkaart, C.; Sawyer, C.; Iakovleva, V.; Lee, P. L.; Xin, L.; Ng, H. E. V.; Loo, J. M.; Ong, X.; Ng, H. Q. A.; Wang, J.; Koh, W. Q. C.; Poon, S. Y. P.; Stanojevic, D.; Tran, H.-D.; Lim, K. H. E.; Toh, S. Y.; Ewels, P. A.; Ng, H.-H.; Iyer, N. G.; Thiery, A.; Chng, W. J.; Chen, L.; DasGupta, R.; Sikic, M.; Chan, Y.-S.; Tan, B. O. P.; Wan, Y.; Tam, W. L.; Yu, Q.; Khor, C. C.; Wustefeld, T.; Lezhava, A.; Pratanwanich, P. N.; Love, M. I.; Goh, W. S. S.; Ng, S. B.; Oshlack, A.; Iyer, N. G.; Yu, Q.; Goke, J. A systematic benchmark of Nanopore long-read RNA sequencing for transcript-level analysis in human cell lines. Nat. Methods 2025, 22 (4), 801– 812, DOI: 10.1038/s41592-025-02623-4There is no corresponding record for this reference.
- 4Chen, L.-L.; Kim, V. N. Small and long non-coding RNAs: Past, present, and future. Cell 2024, 187 (23), 6451– 6485, DOI: 10.1016/j.cell.2024.10.024There is no corresponding record for this reference.
- 5Mattick, J. S.; Amaral, P. P.; Carninci, P.; Carpenter, S.; Chang, H. Y.; Chen, L.-L.; Chen, R.; Dean, C.; Dinger, M. E.; Fitzgerald, K. A.; Gingeras, T. R.; Guttman, M.; Hirose, T.; Huarte, M.; Johnson, R.; Kanduri, C.; Kapranov, P.; Lawrence, J. B.; Lee, J. T.; Mendell, J. T.; Mercer, T. R.; Moore, K. J.; Nakagawa, S.; Rinn, J. L.; Spector, D. L.; Ulitsky, I.; Wan, Y.; Wilusz, J. E.; Wu, M. Long non-coding RNAs: definitions, functions, challenges and recommendations. Nat. Rev. Mol. Cell Biol. 2023, 24 (6), 430– 447, DOI: 10.1038/s41580-022-00566-8There is no corresponding record for this reference.
- 6Le, P.; Ahmed, N.; Yeo, G. W. Illuminating RNA biology through imaging. Nat. Cell Biol. 2022, 24 (6), 815– 824, DOI: 10.1038/s41556-022-00933-9There is no corresponding record for this reference.
- 7Xia, Y.; Zhang, R.; Wang, Z.; Tian, J.; Chen, X. Recent advances in high-performance fluorescent and bioluminescent RNA imaging probes. Chem. Soc. Rev. 2017, 46 (10), 2824– 2843, DOI: 10.1039/C6CS00675BThere is no corresponding record for this reference.
From NLM.
- 8Boutorine, A. S.; Novopashina, D. S.; Krasheninina, O. A.; Nozeret, K.; Venyaminova, A. G. Fluorescent probes for nucleic Acid visualization in fixed and live cells. Molecules 2013, 18 (12), 15357– 15397, DOI: 10.3390/molecules181215357There is no corresponding record for this reference.
From NLM.
- 9Schermelleh, L.; Ferrand, A.; Huser, T.; Eggeling, C.; Sauer, M.; Biehlmaier, O.; Drummen, G. P. C. Super-resolution microscopy demystified. Nat. Cell Biol. 2019, 21 (1), 72– 84, DOI: 10.1038/s41556-018-0251-8There is no corresponding record for this reference.
- 10Sigal, Y. M.; Zhou, R.; Zhuang, X. Visualizing and discovering cellular structures with super-resolution microscopy. Science 2018, 361 (6405), 880– 887, DOI: 10.1126/science.aau1044There is no corresponding record for this reference.
From NLM.
- 11Huang, B.; Babcock, H.; Zhuang, X. Breaking the diffraction barrier: super-resolution imaging of cells. Cell 2010, 143 (7), 1047– 1058, DOI: 10.1016/j.cell.2010.12.002There is no corresponding record for this reference.
From NLM.
- 12Zhong, J.; Xu, Z.; Peng, J.; Guan, L.; Li, J.; Zhou, Z.; Zhang, Y.; Zhang, J.; Liu, S.; Yang, Y.; Hao, X. A CRISPR/Cas13a system based on a dumbbell-shaped hairpin combined with DNA-PAINT to establish the DCP-platform for highly sensitive detection of Hantaan virus RNA. Talanta 2025, 291, 127852, DOI: 10.1016/j.talanta.2025.127852There is no corresponding record for this reference.
- 13Kocabey, S.; Chiarelli, G.; Acuna, G. P.; Ruegg, C. Ultrasensitive and multiplexed miRNA detection system with DNA-PAINT. Biosens. Bioelectron. 2023, 224, 115053, DOI: 10.1016/j.bios.2022.115053There is no corresponding record for this reference.
- 14Higuchi, R.; Fockler, C.; Dollinger, G.; Watson, R. Kinetic PCR analysis: real-time monitoring of DNA amplification reactions. Biotechnology (N Y) 1993, 11 (9), 1026– 1030, DOI: 10.1038/nbt0993-1026There is no corresponding record for this reference.
From NLM.
- 15Salimi, A.; Rahmani, S.; Sharifi-Zarchi, A. InterOpt: Improved gene expression quantification in qPCR experiments using weighted aggregation of reference genes. iScience 2023, 26 (10), 107945, DOI: 10.1016/j.isci.2023.107945There is no corresponding record for this reference.
- 16Różański, M.; Walczak-Drzewiecka, A.; Witaszewska, J.; Wójcik, E.; Guziński, A.; Zimoń, B.; Matusiak, R.; Kazimierczak, J.; Borowiec, M.; Kania, K.; Paradowska, E.; Pawełczyk, J.; Dziadek, J.; Dastych, J. RT-qPCR-based tests for SARS-CoV-2 detection in pooled saliva samples for massive population screening to monitor epidemics. Sci. Rep. 2022, 12 (1), 8082, DOI: 10.1038/s41598-022-12179-4There is no corresponding record for this reference.
- 17Bustin, S. A.; Nolan, T. Pitfalls of quantitative real-time reverse-transcription polymerase chain reaction. J. Biomol. Tech. 2004, 15 (3), 155– 166There is no corresponding record for this reference.
From NLM.
- 18Gerbi, S. A.; Borovjagin, A. V. Pre-ribosomal RNA processing in multicellular organisms. Madame Curie Bioscience Database [Internet]; Landes Bioscience , 2004; pp 170– 198.There is no corresponding record for this reference.
- 19Wu, J.; Xiao, J.; Zhang, Z.; Wang, X.; Hu, S.; Yu, J. Ribogenomics: the Science and Knowledge of RNA. Genomics, Proteomics & Bioinformatics 2014, 12 (2), 57– 63, DOI: 10.1016/j.gpb.2014.04.002There is no corresponding record for this reference.
- 20Sanders, R.; Mason, D. J.; Foy, C. A.; Huggett, J. F. Considerations for accurate gene expression measurement by reverse transcription quantitative PCR when analysing clinical samples. Anal. Bioanal. Chem. 2014, 406 (26), 6471– 6483, DOI: 10.1007/s00216-014-7857-xThere is no corresponding record for this reference.
- 21Forster, T.; Roy, D.; Ghazal, P. Experiments using microarray technology: limitations and standard operating procedures. J. Endocrinol 2003, 178 (2), 195– 204, DOI: 10.1677/joe.0.1780195There is no corresponding record for this reference.
From NLM.
- 22Coppola, G. Designing, performing, and interpreting a microarray-based gene expression study. Methods Mol. Biol. 2011, 793, 417– 439, DOI: 10.1007/978-1-61779-328-8_28There is no corresponding record for this reference.
From NLM.
- 23Uhlen, M.; Quake, S. R. Sequential sequencing by synthesis and the next-generation sequencing revolution. Trends Biotechnol. 2023, 41 (12), 1565– 1572, DOI: 10.1016/j.tibtech.2023.06.007There is no corresponding record for this reference.
(accessed 2025/10/30).
- 24Satam, H.; Joshi, K.; Mangrolia, U.; Waghoo, S.; Zaidi, G.; Rawool, S.; Thakare, R. P.; Banday, S.; Mishra, A. K.; Das, G.; Malonia, S. K. Next-Generation Sequencing Technology: Current Trends and Advancements. Biology (Basel) 2023, 12 (7), 997, DOI: 10.3390/biology12070997There is no corresponding record for this reference.
- 25Ament, I. H.; DeBruyne, N.; Wang, F.; Lin, L. Long-read RNA sequencing: A transformative technology for exploring transcriptome complexity in human diseases. Molecular Therapy 2025, 33 (3), 883– 894, DOI: 10.1016/j.ymthe.2024.11.025There is no corresponding record for this reference.
(accessed 2025/10/30).
- 26McKinnon, K. M. Flow Cytometry: An Overview. Curr. Protoc Immunol 2018, 120, 5.1.1– 5.1.11, DOI: 10.1002/cpim.40There is no corresponding record for this reference.
From NLM.
- 27Zhao, Y.; Lin, C.; Wu, P.; Chen, X.; Zhao, Y.; Li, Y.; Chen, L.; Nilsson, M.; Ke, R. Single Cell RNA Expression Analysis Using Flow Cytometry Based on Specific Probe Ligation and Rolling Circle Amplification. ACS Sensors 2020, 5 (10), 3031– 3036, DOI: 10.1021/acssensors.0c01569There is no corresponding record for this reference.
- 28Hanley, M. B.; Lomas, W.; Mittar, D.; Maino, V.; Park, E. Detection of low abundance RNA molecules in individual cells by flow cytometry. PLoS One 2013, 8 (2), e57002 DOI: 10.1371/journal.pone.0057002There is no corresponding record for this reference.
From NLM.
- 29Warren, C. J.; Barbachano-Guerrero, A.; Huey, D.; Yang, Q.; Worden-Sapper, E. R.; Kuhn, J. H.; Sawyer, S. L. Quantification of virus-infected cells using RNA FISH-Flow. STAR Protoc 2023, 4 (2), 102291, DOI: 10.1016/j.xpro.2023.102291There is no corresponding record for this reference.
From NLM.
- 30Gaudin, R.; Barteneva, N. S. Sorting of small infectious virus particles by flow virometry reveals distinct infectivity profiles. Nat. Commun. 2015, 6 (1), 6022, DOI: 10.1038/ncomms7022There is no corresponding record for this reference.
- 31Actis, P.; Maalouf, M. M.; Kim, H. J.; Lohith, A.; Vilozny, B.; Seger, R. A.; Pourmand, N. Compartmental Genomics in Living Cells Revealed by Single-Cell Nanobiopsy. ACS Nano 2014, 8 (1), 546– 553, DOI: 10.1021/nn405097uThere is no corresponding record for this reference.
- 32Sahota, A.; Monteza Cabrejos, A.; Kwan, Z.; Paulose Nadappuram, B.; Ivanov, A. P.; Edel, J. B. Recent advances in single-cell subcellular sampling. Chem. Commun. (Camb) 2023, 59 (36), 5312– 5328, DOI: 10.1039/D3CC00573AThere is no corresponding record for this reference.
From NLM.
- 33Sahl, S. J.; Hell, S. W.; Jakobs, S. Fluorescence nanoscopy in cell biology. Nat. Rev. Mol. Cell Biol. 2017, 18 (11), 685– 701, DOI: 10.1038/nrm.2017.71There is no corresponding record for this reference.
- 34McIsaac, R. S.; Silverman, S. J.; Parsons, L.; Xu, P.; Briehof, R.; McClean, M. N.; Botstein, D. Visualization and analysis of mRNA molecules using fluorescence in situ hybridization in Saccharomyces cerevisiae. J. Vis Exp 2013, (76), e50382 DOI: 10.3791/50382There is no corresponding record for this reference.
From NLM.
- 35Elliott, A. D. Confocal Microscopy: Principles and Modern Practices. Curr. Protoc Cytom 2020, 92 (1), e68 DOI: 10.1002/cpcy.68There is no corresponding record for this reference.
From NLM.
- 36Singer, R. H.; Ward, D. C. Actin gene expression visualized in chicken muscle tissue culture by using in situ hybridization with a biotinated nucleotide analog. Proc. Natl. Acad. Sci. U. S. A. 1982, 79 (23), 7331– 7335, DOI: 10.1073/pnas.79.23.7331There is no corresponding record for this reference.
From NLM.
- 37Barakat, T. S.; Gribnau, J. Combined DNA-RNA fluorescent in situ hybridization (FISH) to study X chromosome inactivation in differentiated female mouse embryonic stem cells. J. Vis Exp 2014, 88, 51628, DOI: 10.3791/51628There is no corresponding record for this reference.
- 38Young, A. P.; Jackson, D. J.; Wyeth, R. C. A technical review and guide to RNA fluorescence in situ hybridization. PeerJ. 2020, 8, e8806 DOI: 10.7717/peerj.8806There is no corresponding record for this reference.
From NLM.
- 39Raj, A.; van den Bogaard, P.; Rifkin, S. A.; van Oudenaarden, A.; Tyagi, S. Imaging individual mRNA molecules using multiple singly labeled probes. Nat. Methods 2008, 5 (10), 877– 879, DOI: 10.1038/nmeth.1253There is no corresponding record for this reference.
From NLM.
- 40Haimovich, G.; Gerst, J. E. Single-molecule Fluorescence in situ Hybridization (smFISH) for RNA Detection in Adherent Animal Cells. Bio-protocol 2018, 8 (21), e3070 DOI: 10.21769/BioProtoc.3070There is no corresponding record for this reference.
- 41Femino, A. M.; Fay, F. S.; Fogarty, K.; Singer, R. H. Visualization of Single RNA Transcripts in Situ. Science 1998, 280 (5363), 585– 590, DOI: 10.1126/science.280.5363.585There is no corresponding record for this reference.
- 42Levesque, M. J.; Ginart, P.; Wei, Y.; Raj, A. Visualizing SNVs to quantify allele-specific expression in single cells. Nat. Methods 2013, 10 (9), 865– 867, DOI: 10.1038/nmeth.2589There is no corresponding record for this reference.
- 43Levesque, M. J.; Raj, A. Single-chromosome transcriptional profiling reveals chromosomal gene expression regulation. Nat. Methods 2013, 10 (3), 246– 248, DOI: 10.1038/nmeth.2372There is no corresponding record for this reference.
- 44Wang, F.; Flanagan, J.; Su, N.; Wang, L. C.; Bui, S.; Nielson, A.; Wu, X.; Vo, H. T.; Ma, X. J.; Luo, Y. RNAscope: a novel in situ RNA analysis platform for formalin-fixed, paraffin-embedded tissues. J. Mol. Diagn 2012, 14 (1), 22– 29, DOI: 10.1016/j.jmoldx.2011.08.002There is no corresponding record for this reference.
From NLM.
- 45Atout, S.; Shurrab, S.; Loveridge, C. Evaluation of the Suitability of RNAscope as a Technique to Measure Gene Expression in Clinical Diagnostics: A Systematic Review. Mol. Diagn Ther 2022, 26 (1), 19– 37, DOI: 10.1007/s40291-021-00570-2There is no corresponding record for this reference.
From NLM.
- 46Huang, T.; Guillotin, B.; Rahni, R.; Birnbaum, K. D.; Wagner, D. A rapid and sensitive, multiplex, whole mount RNA fluorescence in situ hybridization and immunohistochemistry protocol. Plant Methods 2023, 19 (1), 131, DOI: 10.1186/s13007-023-01108-9There is no corresponding record for this reference.
- 47Choi, H. M. T.; Beck, V. A.; Pierce, N. A. Next-Generation in Situ Hybridization Chain Reaction: Higher Gain, Lower Cost, Greater Durability. ACS Nano 2014, 8 (5), 4284– 4294, DOI: 10.1021/nn405717pThere is no corresponding record for this reference.
- 48Lovely, A. M.; Duerr, T. J.; Stein, D. F.; Mun, E. T.; Monaghan, J. R. Hybridization Chain Reaction Fluorescence In Situ Hybridization (HCR-FISH) in Ambystoma mexicanum Tissue. Methods Mol. Biol. 2023, 2562, 109– 122, DOI: 10.1007/978-1-0716-2659-7_6There is no corresponding record for this reference.
From NLM.
- 49Choi, H. M. T.; Schwarzkopf, M.; Fornace, M. E.; Acharya, A.; Artavanis, G.; Stegmaier, J.; Cunha, A.; Pierce, N. A. Third-generation in situ hybridization chain reaction: multiplexed, quantitative, sensitive, versatile, robust. Development 2018, 145 (12), dev165753, DOI: 10.1242/dev.165753There is no corresponding record for this reference.
- 50Deng, R.; Zhang, K.; Sun, Y.; Ren, X.; Li, J. Highly specific imaging of mRNA in single cells by target RNA-initiated rolling circle amplification. Chem. Sci. 2017, 8 (5), 3668– 3675, DOI: 10.1039/C7SC00292KThere is no corresponding record for this reference.
From NLM.
- 51Kurreck, J.; Wyszko, E.; Gillen, C.; Erdmann, V. A. Design of antisense oligonucleotides stabilized by locked nucleic acids. Nucleic Acids Res. 2002, 30 (9), 1911– 1918, DOI: 10.1093/nar/30.9.1911There is no corresponding record for this reference.
From NLM.
- 52Wu, C.; Simonetti, M.; Rossell, C.; Mignardi, M.; Mirzazadeh, R.; Annaratone, L.; Marchiò, C.; Sapino, A.; Bienko, M.; Crosetto, N.; Nilsson, M. RollFISH achieves robust quantification of single-molecule RNA biomarkers in paraffin-embedded tumor tissue samples. Communications Biology 2018, 1 (1), 209, DOI: 10.1038/s42003-018-0218-0There is no corresponding record for this reference.
- 53Lubeck, E.; Cai, L. Single-cell systems biology by super-resolution imaging and combinatorial labeling. Nat. Methods 2012, 9 (7), 743– 748, DOI: 10.1038/nmeth.2069There is no corresponding record for this reference.
- 54Lubeck, E.; Coskun, A. F.; Zhiyentayev, T.; Ahmad, M.; Cai, L. Single-cell in situ RNA profiling by sequential hybridization. Nat. Methods 2014, 11 (4), 360– 361, DOI: 10.1038/nmeth.2892There is no corresponding record for this reference.
- 55Shah, S.; Lubeck, E.; Zhou, W.; Cai, L. In Situ Transcription Profiling of Single Cells Reveals Spatial Organization of Cells in the Mouse Hippocampus. Neuron 2016, 92 (2), 342– 357, DOI: 10.1016/j.neuron.2016.10.001There is no corresponding record for this reference.
accessed 2026/02/05).
- 56Shah, S.; Takei, Y.; Zhou, W.; Lubeck, E.; Yun, J.; Eng, C.-H. L.; Koulena, N.; Cronin, C.; Karp, C.; Liaw, E. J.; Amin, M.; Cai, L. Dynamics and Spatial Genomics of the Nascent Transcriptome by Intron seqFISH. Cell 2018, 174 (2), 363– 376.e316, DOI: 10.1016/j.cell.2018.05.035There is no corresponding record for this reference.
accessed 2026/02/05).
- 57Eng, C.-H. L.; Shah, S.; Thomassie, J.; Cai, L. Profiling the transcriptome with RNA SPOTs. Nat. Methods 2017, 14 (12), 1153– 1155, DOI: 10.1038/nmeth.4500There is no corresponding record for this reference.
- 58Eng, C.-H. L.; Lawson, M.; Zhu, Q.; Dries, R.; Koulena, N.; Takei, Y.; Yun, J.; Cronin, C.; Karp, C.; Yuan, G.-C.; Cai, L. Transcriptome-scale super-resolved imaging in tissues by RNA seqFISH+. Nature 2019, 568 (7751), 235– 239, DOI: 10.1038/s41586-019-1049-yThere is no corresponding record for this reference.
- 59Chen, K. H.; Boettiger, A. N.; Moffitt, J. R.; Wang, S.; Zhuang, X. Spatially resolved, highly multiplexed RNA profiling in single cells. Science 2015, 348 (6233), aaa6090, DOI: 10.1126/science.aaa6090There is no corresponding record for this reference.
- 60Moffitt, J. R.; Hao, J.; Wang, G.; Chen, K. H.; Babcock, H. P.; Zhuang, X. High-throughput single-cell gene-expression profiling with multiplexed error-robust fluorescence in situ hybridization. Proc. Natl. Acad. Sci. U. S. A. 2016, 113 (39), 11046– 11051, DOI: 10.1073/pnas.1612826113There is no corresponding record for this reference.
- 61Moffitt, J. R.; Zhuang, X. Chapter One - RNA Imaging with Multiplexed Error-Robust Fluorescence In Situ Hybridization (MERFISH). In Methods in Enzymology, Filonov, G. S., Jaffrey, S. R., Eds.; Academic Press, 2016; Vol. 572, pp 1– 49.There is no corresponding record for this reference.
- 62Chen, F.; Tillberg, P. W.; Boyden, E. S. Expansion microscopy. Science 2015, 347 (6221), 543– 548, DOI: 10.1126/science.1260088There is no corresponding record for this reference.
- 63Liu, J.; Tran, V.; Vemuri, V. N. P.; Byrne, A.; Borja, M.; Kim, Y. J.; Agarwal, S.; Wang, R.; Awayan, K.; Murti, A.; Taychameekiatchai, A.; Wang, B.; Emanuel, G.; He, J.; Haliburton, J.; Oliveira Pisco, A.; Neff, N. F. Concordance of MERFISH spatial transcriptomics with bulk and single-cell RNA sequencing. Life Sci. Alliance 2023, 6 (1), e202201701, DOI: 10.26508/lsa.202201701There is no corresponding record for this reference.
- 64Fang, R.; Halpern, A.; Rahman, M. M.; Huang, Z.; Lei, Z.; Hell, S. J.; Dulac, C.; Zhuang, X. Three-dimensional single-cell transcriptome imaging of thick tissues. eLife 2024, 12, RP90029, DOI: 10.7554/eLife.90029There is no corresponding record for this reference.
- 65Braselmann, E.; Rathbun, C.; Richards, E. M.; Palmer, A. E. Illuminating RNA Biology: Tools for Imaging RNA in Live Mammalian Cells. Cell Chemical Biology 2020, 27 (8), 891– 903, DOI: 10.1016/j.chembiol.2020.06.010There is no corresponding record for this reference.
- 66Chiu, Y. F.; Huang, Y. W.; Chen, C. Y.; Chen, Y. C.; Gong, Y. N.; Kuo, R. L.; Huang, C. G.; Shih, S. R. Visualizing Influenza A Virus vRNA Replication. Front Microbiol 2022, 13, 812711, DOI: 10.3389/fmicb.2022.812711There is no corresponding record for this reference.
From NLM.
- 67Hu, Y.; Xu, J.; Gao, E.; Fan, X.; Wei, J.; Ye, B.; Xu, S.; Ma, W. Enhanced single RNA imaging reveals dynamic gene expression in live animals. eLife 2023, 12, e82178 DOI: 10.7554/eLife.82178There is no corresponding record for this reference.
- 68Peng, Y.; Shu, L.; Deng, X.; Huang, X.; Mo, X.; Du, F.; Tang, Z. Live-Cell Imaging of Endogenous RNA with a Genetically Encoded Fluorogenic Allosteric Aptamer. Anal. Chem. 2023, 95 (37), 13762– 13768, DOI: 10.1021/acs.analchem.2c05724There is no corresponding record for this reference.
- 69Constantin, T. P.; Silva, G. L.; Robertson, K. L.; Hamilton, T. P.; Fague, K.; Waggoner, A. S.; Armitage, B. A. Synthesis of New Fluorogenic Cyanine Dyes and Incorporation into RNA Fluoromodules. Org. Lett. 2008, 10 (8), 1561– 1564, DOI: 10.1021/ol702920eThere is no corresponding record for this reference.
- 70Huang, K.; Chen, X.; Li, C.; Song, Q.; Li, H.; Zhu, L.; Yang, Y.; Ren, A. Structure-based investigation of fluorogenic Pepper aptamer. Nat. Chem. Biol. 2021, 17 (12), 1289– 1295, DOI: 10.1038/s41589-021-00884-6There is no corresponding record for this reference.
- 71Zheng, H.; Liu, X.; Liu, L.; Hu, J.; Chen, X. Imaging of endogenous RNA in live cells using sequence-activated fluorescent RNA probes. Nucleic Acids Res. 2025, 53 (2), gkae1209, DOI: 10.1093/nar/gkae1209There is no corresponding record for this reference.
- 72Jansen, R.; Embden, J. D.; Gaastra, W.; Schouls, L. M. Identification of genes that are associated with DNA repeats in prokaryotes. Mol. Microbiol. 2002, 43 (6), 1565– 1575, DOI: 10.1046/j.1365-2958.2002.02839.xThere is no corresponding record for this reference.
From NLM.
- 73Wang, S. W.; Gao, C.; Zheng, Y. M.; Yi, L.; Lu, J. C.; Huang, X. Y.; Cai, J. B.; Zhang, P. F.; Cui, Y. H.; Ke, A. W. Current applications and future perspective of CRISPR/Cas9 gene editing in cancer. Mol. Cancer 2022, 21 (1), 57, DOI: 10.1186/s12943-022-01518-8There is no corresponding record for this reference.
From NLM.
- 74Chen, K.; Wang, Y. CRISPR/Cas systems for in situ imaging of intracellular nucleic acids: Concepts and applications. Biotechnol. Bioeng. 2023, 120 (12), 3446– 3464, DOI: 10.1002/bit.28543There is no corresponding record for this reference.
From NLM.
- 75Hillary, V. E.; Ceasar, S. A. A Review on the Mechanism and Applications of CRISPR/Cas9/Cas12/Cas13/Cas14 Proteins Utilized for Genome Engineering. Mol. Biotechnol 2023, 65 (3), 311– 325, DOI: 10.1007/s12033-022-00567-0There is no corresponding record for this reference.
From NLM.
- 76Cao, H.; Wang, Y.; Zhang, N.; Xia, S.; Tian, P.; Lu, L.; Du, J.; Du, Y. Progress of CRISPR-Cas13 Mediated Live-Cell RNA Imaging and Detection of RNA-Protein Interactions. Front Cell Dev Biol. 2022, 10, 866820, DOI: 10.3389/fcell.2022.866820There is no corresponding record for this reference.
From NLM.
- 77Jia, H.-Y.; Zhang, X.-Y.; Ye, B.-C.; Yin, B.-C. An Orthogonal CRISPR/dCas12a System for RNA Imaging in Live Cells. Anal. Chem. 2024, 96 (15), 5913– 5921, DOI: 10.1021/acs.analchem.3c05975There is no corresponding record for this reference.
- 78O’Connell, M. R.; Oakes, B. L.; Sternberg, S. H.; East-Seletsky, A.; Kaplan, M.; Doudna, J. A. Programmable RNA recognition and cleavage by CRISPR/Cas9. Nature 2014, 516 (7530), 263– 266, DOI: 10.1038/nature13769There is no corresponding record for this reference.
From NLM.
- 79Tang, H.; Peng, J.; Peng, S.; Wang, Q.; Jiang, X.; Xue, X.; Tao, Y.; Xiang, L.; Ji, Q.; Liu, S. M.; Weng, X.; Zhou, X. Live-cell RNA imaging using the CRISPR-dCas13 system with modified sgRNAs appended with fluorescent RNA aptamers. Chem. Sci. 2022, 13 (47), 14032– 14040, DOI: 10.1039/D2SC04656CThere is no corresponding record for this reference.
From NLM.
- 80Tyagi, S.; Kramer, F. R. Molecular Beacons: Probes that Fluoresce upon Hybridization. Nat. Biotechnol. 1996, 14 (3), 303– 308, DOI: 10.1038/nbt0396-303There is no corresponding record for this reference.
- 81Xia, C.; Colognori, D.; Jiang, X. S.; Xu, K.; Doudna, J. A. Single-molecule live-cell RNA imaging with CRISPR–Csm. Nat. Biotechnol. 2025, 43, 2023, DOI: 10.1038/s41587-024-02540-5There is no corresponding record for this reference.
- 82Bratu, D. P.; Cha, B.-J.; Mhlanga, M. M.; Kramer, F. R.; Tyagi, S. Visualizing the distribution and transport of mRNAs in living cells. Proc. Natl. Acad. Sci. U. S. A. 2003, 100 (23), 13308– 13313, DOI: 10.1073/pnas.2233244100There is no corresponding record for this reference.
- 83Turner-Bridger, B.; Jakobs, M.; Muresan, L.; Wong, H. H.-W.; Franze, K.; Harris, W. A.; Holt, C. E. Single-molecule analysis of endogenous β-actin mRNA trafficking reveals a mechanism for compartmentalized mRNA localization in axons. Proc. Natl. Acad. Sci. U. S. A. 2018, 115 (41), E9697– E9706, DOI: 10.1073/pnas.1806189115There is no corresponding record for this reference.
- 84Culley, S.; Caballero, A. C.; Burden, J. J.; Uhlmann, V. Made to measure: An introduction to quantifying microscopy data in the life sciences. J. Microsc 2024, 295 (1), 61– 82, DOI: 10.1111/jmi.13208There is no corresponding record for this reference.
From NLM.
- 85Schindelin, J.; Arganda-Carreras, I.; Frise, E.; Kaynig, V.; Longair, M.; Pietzsch, T.; Preibisch, S.; Rueden, C.; Saalfeld, S.; Schmid, B.; Tinevez, J. Y.; White, D. J.; Hartenstein, V.; Eliceiri, K.; Tomancak, P.; Cardona, A. Fiji: an open-source platform for biological-image analysis. Nat. Methods 2012, 9 (7), 676– 682, DOI: 10.1038/nmeth.2019There is no corresponding record for this reference.
From NLM.
- 86Carpenter, A. E.; Jones, T. R.; Lamprecht, M. R.; Clarke, C.; Kang, I. H.; Friman, O.; Guertin, D. A.; Chang, J. H.; Lindquist, R. A.; Moffat, J.; Golland, P.; Sabatini, D. M. CellProfiler: image analysis software for identifying and quantifying cell phenotypes. Genome Biology 2006, 7 (10), R100, DOI: 10.1186/gb-2006-7-10-r100There is no corresponding record for this reference.
- 87Waters, J. C. Accuracy and precision in quantitative fluorescence microscopy. J. Cell Biol. 2009, 185 (7), 1135– 1148, DOI: 10.1083/jcb.200903097There is no corresponding record for this reference.
From NLM.
- 88Sasaki, A. Recent advances in the standardization of fluorescence microscopy for quantitative image analysis. Biophys Rev. 2022, 14 (1), 33– 39, DOI: 10.1007/s12551-021-00871-0There is no corresponding record for this reference.
From NLM.
- 89Baskin, D. S.; Widmayer, M. A.; Sharpe, M. A. Quantification and calibration of images in fluorescence microscopy. Anal. Biochem. 2010, 404 (2), 118– 126, DOI: 10.1016/j.ab.2010.05.029There is no corresponding record for this reference.
- 90Barbini, P.; Cevenini, G.; Massai, M. R. Nearest-Neighbor Analysis of Spatial Point Patterns: Application to Biomedical Image Interpretation. Computers and Biomedical Research 1996, 29 (6), 482– 493, DOI: 10.1006/cbmr.1996.0035There is no corresponding record for this reference.
- 91Soltisz, A. M.; Craigmile, P. F.; Veeraraghavan, R. Spatial Pattern Analysis using Closest Events (SPACE)-A Nearest Neighbor Point Pattern Analysis Framework for Assessing Spatial Relationships from Digital Images. Microsc Microanal 2024, 30 (2), 306– 317, DOI: 10.1093/mam/ozae022There is no corresponding record for this reference.
From NLM.
- 92Kiskowski, M. A.; Hancock, J. F.; Kenworthy, A. K. On the use of Ripley’s K-function and its derivatives to analyze domain size. Biophys. J. 2009, 97 (4), 1095– 1103, DOI: 10.1016/j.bpj.2009.05.039There is no corresponding record for this reference.
From NLM.
- 93Ruan, Y.; Yin, P.; Li, F.; Li, D.; Lin, Q.; Li, K. The Accuracy of Determining Cluster Size by Analyzing Ripley’s K Function in Single Molecule Localization Microscopy. Applied Sciences 2019, 9 (16), 3271, DOI: 10.3390/app9163271There is no corresponding record for this reference.
- 94Peters, R.; Benthem Muñiz, M.; Griffié, J.; Williamson, D. J.; Ashdown, G. W.; Lorenz, C. D.; Owen, D. M. Quantification of fibrous spatial point patterns from single-molecule localization microscopy (SMLM) data. Bioinformatics 2017, 33 (11), 1703– 1711, DOI: 10.1093/bioinformatics/btx026There is no corresponding record for this reference.
(accessed 10/26/2025).
- 95Lee, C.; Roberts, S. E.; Gladfelter, A. S. Quantitative spatial analysis of transcripts in multinucleate cells using single-molecule FISH. Methods 2016, 98, 124– 133, DOI: 10.1016/j.ymeth.2015.12.007There is no corresponding record for this reference.
- 96Mancebo, A.; Mehra, D.; Banerjee, C.; Kim, D. H.; Puchner, E. M. Efficient Cross-Correlation Filtering of One- and Two-Color Single Molecule Localization Microscopy Data. Front Bioinform 2021, 1, 739769, DOI: 10.3389/fbinf.2021.739769There is no corresponding record for this reference.
From NLM.
- 97Schnitzbauer, J.; Wang, Y.; Zhao, S.; Bakalar, M.; Nuwal, T.; Chen, B.; Huang, B. Correlation analysis framework for localization-based superresolution microscopy. Proc. Natl. Acad. Sci. U. S. A. 2018, 115 (13), 3219– 3224, DOI: 10.1073/pnas.1711314115There is no corresponding record for this reference.
From NLM.
- 98Ester, M.; Kriegel, H.-P.; Sander, J.; Xu, X. A density-based algorithm for discovering clusters in large spatial databases with noise. kdd Proceedings 1996, 96, 226– 231There is no corresponding record for this reference.
- 99Huang, K.; Demirci, F.; Batish, M.; Treible, W.; Meyers, B. C.; Caplan, J. L. Quantitative, super-resolution localization of small RNAs with sRNA-PAINT. Nucleic Acids Res. 2020, 48 (16), e96– e96, DOI: 10.1093/nar/gkaa623There is no corresponding record for this reference.
(accessed 10/26/2025).
- 100Pageon, S. V.; Nicovich, P. R.; Mollazade, M.; Tabarin, T.; Gaus, K. Clus-DoC: a combined cluster detection and colocalization analysis for single-molecule localization microscopy data. Mol. Biol. Cell 2016, 27 (22), 3627– 3636, DOI: 10.1091/mbc.e16-07-0478There is no corresponding record for this reference.
From NLM.
- 101Bushra, A. A.; Kim, D.; Kan, Y.; Yi, G. AutoSCAN: automatic detection of DBSCAN parameters and efficient clustering of data in overlapping density regions. PeerJ. Comput. Sci. 2024, 10, e1921 DOI: 10.7717/peerj-cs.1921There is no corresponding record for this reference.
From NLM.
- 102Yin, L.; Hu, H.; Li, K.; Zheng, G.; Qu, Y.; Chen, H. Improvement of DBSCAN Algorithm Based on K-Dist Graph for Adaptive Determining Parameters. Electronics 2023, 12 (15), 3213, DOI: 10.3390/electronics12153213There is no corresponding record for this reference.
- 103Bushra, A. A.; Yi, G. Comparative Analysis Review of Pioneering DBSCAN and Successive Density-Based Clustering Algorithms. IEEE Access 2021, 9, 87918– 87935, DOI: 10.1109/ACCESS.2021.3089036There is no corresponding record for this reference.
- 104Marenda, M.; Lazarova, E.; van de Linde, S.; Gilbert, N.; Michieletto, D. Parameter-free molecular super-structures quantification in single-molecule localization microscopy. J. Cell Biol. 2021, 220 (5), e202010003, DOI: 10.1083/jcb.202010003There is no corresponding record for this reference.
- 105Mazouchi, A.; Milstein, J. N. Fast Optimized Cluster Algorithm for Localizations (FOCAL): a spatial cluster analysis for super-resolved microscopy. Bioinformatics 2016, 32 (5), 747– 754, DOI: 10.1093/bioinformatics/btv630There is no corresponding record for this reference.
(accessed 10/26/2025).
- 106Bertin, E.; Parazza, F.; Chassery, J. M. Segmentation and measurement based on 3D Voronoi diagram: Application to confocal microscopy. Computerized Medical Imaging and Graphics 1993, 17 (3), 175– 182, DOI: 10.1016/0895-6111(93)90041-KThere is no corresponding record for this reference.
- 107Eils, R.; Bertin, E.; Saracoglu, K.; Rinke, B.; Schröck, E.; Parazza, F.; Usson, Y.; Robert-Nicoud, M.; Stelzer, E. H.; Chassery, J. M. Application of confocal laser microscopy and three-dimensional Voronoi diagrams for volume and surface estimates of interphase chromosomes. J. Microsc 1995, 177 (Pt 2), 150– 161, DOI: 10.1111/j.1365-2818.1995.tb03545.xThere is no corresponding record for this reference.
From NLM.
- 108Andronov, L.; Orlov, I.; Lutz, Y.; Vonesch, J.-L.; Klaholz, B. P. ClusterViSu, a method for clustering of protein complexes by Voronoi tessellation in super-resolution microscopy. Sci. Rep. 2016, 6 (1), 24084, DOI: 10.1038/srep24084There is no corresponding record for this reference.
- 109Levet, F.; Hosy, E.; Kechkar, A.; Butler, C.; Beghin, A.; Choquet, D.; Sibarita, J.-B. SR-Tesseler: a method to segment and quantify localization-based super-resolution microscopy data. Nat. Methods 2015, 12 (11), 1065– 1071, DOI: 10.1038/nmeth.3579There is no corresponding record for this reference.
- 110Andronov, L.; Lutz, Y.; Vonesch, J. L.; Klaholz, B. P. SharpViSu: integrated analysis and segmentation of super-resolution microscopy data. Bioinformatics 2016, 32 (14), 2239– 2241, DOI: 10.1093/bioinformatics/btw123There is no corresponding record for this reference.
From NLM.
- 111Rubin-Delanchy, P.; Burn, G. L.; Griffié, J.; Williamson, D. J.; Heard, N. A.; Cope, A. P.; Owen, D. M. Bayesian cluster identification in single-molecule localization microscopy data. Nat. Methods 2015, 12 (11), 1072– 1076, DOI: 10.1038/nmeth.3612There is no corresponding record for this reference.
- 112Tang, Y.; Hendriks, J.; Gensch, T.; Dai, L.; Li, J. Automatic Bayesian single molecule identification for localization microscopy. Sci. Rep. 2016, 6 (1), 33521, DOI: 10.1038/srep33521There is no corresponding record for this reference.
- 113Griffié, J.; Shannon, M.; Bromley, C. L.; Boelen, L.; Burn, G. L.; Williamson, D. J.; Heard, N. A.; Cope, A. P.; Owen, D. M.; Rubin-Delanchy, P. A Bayesian cluster analysis method for single-molecule localization microscopy data. Nat. Protoc. 2016, 11 (12), 2499– 2514, DOI: 10.1038/nprot.2016.149There is no corresponding record for this reference.
- 114Hammer, J. L.; Devanny, A. J.; Kaufman, L. J. Bayesian optimized parameter selection for density-based clustering applied to single molecule localization microscopy. Communications Biology 2025, 8 (1), 902, DOI: 10.1038/s42003-025-08332-0There is no corresponding record for this reference.
- 115Andronov, L.; Han, M.; Zhu, Y.; Balaji, A.; Roy, A. R.; Barentine, A. E. S.; Patel, P.; Garhyan, J.; Qi, L. S.; Moerner, W. E. Nanoscale cellular organization of viral RNA and proteins in SARS-CoV-2 replication organelles. Nat. Commun. 2024, 15 (1), 4644, DOI: 10.1038/s41467-024-48991-xThere is no corresponding record for this reference.
- 116Mayerich, D.; Sun, R. Superresolution Image Processing. In Microscope Image Processing, Second ed.; Merchant, F. A., Castleman, K. R., Eds.; Academic Press, 2023; Chapter Twelve, pp 319– 334.There is no corresponding record for this reference.
- 117Jimenez, A.; Friedl, K.; Leterrier, C. About samples, giving examples: Optimized Single Molecule Localization Microscopy. Methods 2020, 174, 100– 114, DOI: 10.1016/j.ymeth.2019.05.008There is no corresponding record for this reference.
- 118Chen, F.; Wassie, A. T.; Cote, A. J.; Sinha, A.; Alon, S.; Asano, S.; Daugharthy, E. R.; Chang, J.-B.; Marblestone, A.; Church, G. M.; Raj, A.; Boyden, E. S. Nanoscale imaging of RNA with expansion microscopy. Nat. Methods 2016, 13 (8), 679– 684, DOI: 10.1038/nmeth.3899There is no corresponding record for this reference.
- 119Klar, T. A.; Jakobs, S.; Dyba, M.; Egner, A.; Hell, S. W. Fluorescence microscopy with diffraction resolution barrier broken by stimulated emission. Proc. Natl. Acad. Sci. U. S. A. 2000, 97 (15), 8206– 8210, DOI: 10.1073/pnas.97.15.8206There is no corresponding record for this reference.
- 120Balzarotti, F.; Eilers, Y.; Gwosch, K. C.; Gynnå, A. H.; Westphal, V.; Stefani, F. D.; Elf, J.; Hell, S. W. Nanometer resolution imaging and tracking of fluorescent molecules with minimal photon fluxes. Science 2017, 355 (6325), 606– 612, DOI: 10.1126/science.aak9913There is no corresponding record for this reference.
- 121Gustafsson, M. G. Surpassing the lateral resolution limit by a factor of two using structured illumination microscopy. J. Microsc. 2000, 198 (2), 82– 87, DOI: 10.1046/j.1365-2818.2000.00710.xThere is no corresponding record for this reference.
From NLM.
- 122Betzig, E.; Patterson, G. H.; Sougrat, R.; Lindwasser, O. W.; Olenych, S.; Bonifacino, J. S.; Davidson, M. W.; Lippincott-Schwartz, J.; Hess, H. F. Imaging intracellular fluorescent proteins at nanometer resolution. Science 2006, 313 (5793), 1642– 1645, DOI: 10.1126/science.1127344There is no corresponding record for this reference.
From NLM.
- 123Rust, M. J.; Bates, M.; Zhuang, X. Sub-diffraction-limit imaging by stochastic optical reconstruction microscopy (STORM). Nat. Methods 2006, 3 (10), 793– 796, DOI: 10.1038/nmeth929There is no corresponding record for this reference.
- 124Schnitzbauer, J.; Strauss, M. T.; Schlichthaerle, T.; Schueder, F.; Jungmann, R. Super-resolution microscopy with DNA-PAINT. Nat. Protoc. 2017, 12 (6), 1198– 1228, DOI: 10.1038/nprot.2017.024There is no corresponding record for this reference.
- 125Arnould, B.; Quillin, A. L.; Heemstra, J. M. Tracking the Message: Applying Single Molecule Localization Microscopy to Cellular RNA Imaging. Chembiochem 2023, 24 (10), e202300049 DOI: 10.1002/cbic.202300049There is no corresponding record for this reference.
From NLM.
- 126Lelek, M.; Gyparaki, M. T.; Beliu, G.; Schueder, F.; Griffié, J.; Manley, S.; Jungmann, R.; Sauer, M.; Lakadamyali, M.; Zimmer, C. Single-molecule localization microscopy. Nature Reviews Methods Primers 2021, 1 (1), 39, DOI: 10.1038/s43586-021-00038-xThere is no corresponding record for this reference.
- 127Hugelier, S.; Colosi, P. L.; Lakadamyali, M. Quantitative Single-Molecule Localization Microscopy. Annu. Rev. Biophys 2023, 52, 139– 160, DOI: 10.1146/annurev-biophys-111622-091212There is no corresponding record for this reference.
From NLM.
- 128Hyun, Y.; Kim, D. Recent development of computational cluster analysis methods for single-molecule localization microscopy images. Comput. Struct Biotechnol J. 2023, 21, 879– 888, DOI: 10.1016/j.csbj.2023.01.006There is no corresponding record for this reference.
From NLM.
- 129Khater, I. M.; Nabi, I. R.; Hamarneh, G. A Review of Super-Resolution Single-Molecule Localization Microscopy Cluster Analysis and Quantification Methods. Patterns (N Y) 2020, 1 (3), 100038, DOI: 10.1016/j.patter.2020.100038There is no corresponding record for this reference.
From NLM.
- 130Nieves, D. J.; Pike, J. A.; Levet, F.; Williamson, D. J.; Baragilly, M.; Oloketuyi, S.; de Marco, A.; Griffié, J.; Sage, D.; Cohen, E. A. K.; Sibarita, J.-B.; Heilemann, M.; Owen, D. M. A framework for evaluating the performance of SMLM cluster analysis algorithms. Nat. Methods 2023, 20 (2), 259– 267, DOI: 10.1038/s41592-022-01750-6There is no corresponding record for this reference.
- 131Ejdrup, A. L.; Lycas, M. D.; Lorenzen, N.; Konomi, A.; Herborg, F.; Madsen, K. L.; Gether, U. A density-based enrichment measure for assessing colocalization in single-molecule localization microscopy data. Nat. Commun. 2022, 13 (1), 4388, DOI: 10.1038/s41467-022-32064-yThere is no corresponding record for this reference.
- 132Ovesný, M.; Křížek, P.; Borkovec, J.; Svindrych, Z.; Hagen, G. M. ThunderSTORM: a comprehensive ImageJ plug-in for PALM and STORM data analysis and super-resolution imaging. Bioinformatics 2014, 30 (16), 2389– 2390, DOI: 10.1093/bioinformatics/btu202There is no corresponding record for this reference.
From NLM.
- 133Wolter, S.; Löschberger, A.; Holm, T.; Aufmkolk, S.; Dabauvalle, M.-C.; van de Linde, S.; Sauer, M. rapidSTORM: accurate, fast open-source software for localization microscopy. Nat. Methods 2012, 9 (11), 1040– 1041, DOI: 10.1038/nmeth.2224There is no corresponding record for this reference.
- 134Sage, D.; Kirshner, H.; Pengo, T.; Stuurman, N.; Min, J.; Manley, S.; Unser, M. Quantitative evaluation of software packages for single-molecule localization microscopy. Nat. Methods 2015, 12 (8), 717– 724, DOI: 10.1038/nmeth.3442There is no corresponding record for this reference.
- 135Liu, S.; Wang, C. Y.; Zheng, P.; Jia, B. B.; Zemke, N. R.; Ren, P.; Park, H. L.; Ren, B.; Zhuang, X. Cell type–specific 3D-genome organization and transcription regulation in the brain. Science Advances 2025, 11 (9), eadv2067 DOI: 10.1126/sciadv.adv2067There is no corresponding record for this reference.
- 136Shivanandan, A.; Deschout, H.; Scarselli, M.; Radenovic, A. Challenges in quantitative single molecule localization microscopy. FEBS letters 2014, 588 (19), 3595– 3602, DOI: 10.1016/j.febslet.2014.06.014There is no corresponding record for this reference.
- 137Olivier, N.; Keller, D.; Rajan, V. S.; Gönczy, P.; Manley, S. Simple buffers for 3D STORM microscopy. Biomed Opt Express 2013, 4 (6), 885– 899, DOI: 10.1364/BOE.4.000885There is no corresponding record for this reference.
From NLM.
- 138Shcherbakova, D. M.; Sengupta, P.; Lippincott-Schwartz, J.; Verkhusha, V. V. Photocontrollable fluorescent proteins for superresolution imaging. Annu. Rev. Biophys 2014, 43, 303– 329, DOI: 10.1146/annurev-biophys-051013-022836There is no corresponding record for this reference.
From NLM.
- 139Godin, A. G.; Lounis, B.; Cognet, L. Super-resolution microscopy approaches for live cell imaging. Biophys. J. 2014, 107 (8), 1777– 1784, DOI: 10.1016/j.bpj.2014.08.028There is no corresponding record for this reference.
From NLM.
- 140Xu, J.; Ma, H.; Liu, Y. Stochastic Optical Reconstruction Microscopy (STORM). Curr. Protoc Cytom 2017, 81, 12.46.11– 12.46.27, DOI: 10.1002/cpcy.23There is no corresponding record for this reference.
From NLM.
- 141Cho, W.-K.; Jayanth, N.; English, B. P.; Inoue, T.; Andrews, J. O.; Conway, W.; Grimm, J. B.; Spille, J.-H.; Lavis, L. D.; Lionnet, T.; Cisse, I. I. RNA Polymerase II cluster dynamics predict mRNA output in living cells. eLife 2016, 5, e13617 DOI: 10.7554/eLife.13617There is no corresponding record for this reference.
- 142Dankovich, T. M.; Rizzoli, S. O. Challenges facing quantitative large-scale optical super-resolution, and some simple solutions. iScience 2021, 24 (3), 102134, DOI: 10.1016/j.isci.2021.102134There is no corresponding record for this reference.
- 143Cho, W.-K.; Spille, J.-H.; Hecht, M.; Lee, C.; Li, C.; Grube, V.; Cisse, I. I. Mediator and RNA polymerase II clusters associate in transcription-dependent condensates. Science 2018, 361 (6400), 412– 415, DOI: 10.1126/science.aar4199There is no corresponding record for this reference.
- 144Cisse, I. I.; Izeddin, I.; Causse, S. Z.; Boudarene, L.; Senecal, A.; Muresan, L.; Dugast-Darzacq, C.; Hajj, B.; Dahan, M.; Darzacq, X. Real-Time Dynamics of RNA Polymerase II Clustering in Live Human Cells. Science 2013, 341 (6146), 664– 667, DOI: 10.1126/science.1239053There is no corresponding record for this reference.
- 145Yang, Y.; Qu, N.; Tan, J.; Rushdi, M. N.; Krueger, C. J.; Chen, A. K. Roles of Gag-RNA interactions in HIV-1 virus assembly deciphered by single-molecule localization microscopy. Proc. Natl. Acad. Sci. U. S. A. 2018, 115 (26), 6721– 6726, DOI: 10.1073/pnas.1805728115There is no corresponding record for this reference.
From NLM.
- 146Xie, L.; Dong, P.; Chen, X.; Hsieh, T. S.; Banala, S.; De Marzio, M.; English, B. P.; Qi, Y.; Jung, S. K.; Kieffer-Kwon, K. R.; Legant, W. R.; Hansen, A. S.; Schulmann, A.; Casellas, R.; Zhang, B.; Betzig, E.; Lavis, L. D.; Chang, H. Y.; Tjian, R.; Liu, Z. 3D ATAC-PALM: super-resolution imaging of the accessible genome. Nat. Methods 2020, 17 (4), 430– 436, DOI: 10.1038/s41592-020-0775-2There is no corresponding record for this reference.
From NLM.
- 147Sunwoo, H.; Wu, J. Y.; Lee, J. T. The Xist RNA-PRC2 complex at 20-nm resolution reveals a low Xist stoichiometry and suggests a hit-and-run mechanism in mouse cells. Proc. Natl. Acad. Sci. U. S. A. 2015, 112 (31), E4216– 4225, DOI: 10.1073/pnas.1503690112There is no corresponding record for this reference.
From NLM.
- 148Maclary, E.; Hinten, M.; Harris, C.; Sethuraman, S.; Gayen, S.; Kalantry, S. PRC2 represses transcribed genes on the imprinted inactive X chromosome in mice. Genome Biology 2017, 18 (1), 82, DOI: 10.1186/s13059-017-1211-5There is no corresponding record for this reference.
- 149Castells-Garcia, A.; Ed-daoui, I.; González-Almela, E.; Vicario, C.; Ottestrom, J.; Lakadamyali, M.; Neguembor, M. V.; Cosma, Maria P. Super resolution microscopy reveals how elongating RNA polymerase II and nascent RNA interact with nucleosome clutches. Nucleic Acids Res. 2022, 50 (1), 175– 190, DOI: 10.1093/nar/gkab1215There is no corresponding record for this reference.
(accessed 10/26/2025).
- 150Hilbert, L.; Sato, Y.; Kuznetsova, K.; Bianucci, T.; Kimura, H.; Jülicher, F.; Honigmann, A.; Zaburdaev, V.; Vastenhouw, N. L. Transcription organizes euchromatin via microphase separation. Nat. Commun. 2021, 12 (1), 1360, DOI: 10.1038/s41467-021-21589-3There is no corresponding record for this reference.
From NLM.
- 151Snijder, E. J.; Limpens, R.; de Wilde, A. H.; de Jong, A. W. M.; Zevenhoven-Dobbe, J. C.; Maier, H. J.; Faas, F.; Koster, A. J.; Bárcena, M. A unifying structural and functional model of the coronavirus replication organelle: Tracking down RNA synthesis. PLoS Biol. 2020, 18 (6), e3000715 DOI: 10.1371/journal.pbio.3000715There is no corresponding record for this reference.
From NLM.
- 152Jungmann, R.; Steinhauer, C.; Scheible, M.; Kuzyk, A.; Tinnefeld, P.; Simmel, F. C. Single-molecule kinetics and super-resolution microscopy by fluorescence imaging of transient binding on DNA origami. Nano Lett. 2010, 10 (11), 4756– 4761, DOI: 10.1021/nl103427wThere is no corresponding record for this reference.
From NLM.
- 153Jungmann, R.; Avendaño, M. S.; Woehrstein, J. B.; Dai, M.; Shih, W. M.; Yin, P. Multiplexed 3D cellular super-resolution imaging with DNA-PAINT and Exchange-PAINT. Nat. Methods 2014, 11 (3), 313– 318, DOI: 10.1038/nmeth.2835There is no corresponding record for this reference.
From NLM.
- 154Iinuma, R.; Ke, Y.; Jungmann, R.; Schlichthaerle, T.; Woehrstein, J. B.; Yin, P. Polyhedra self-assembled from DNA tripods and characterized with 3D DNA-PAINT. Science 2014, 344 (6179), 65– 69, DOI: 10.1126/science.1250944There is no corresponding record for this reference.
From NLM.
- 155Jungmann, R.; Avendaño, M. S.; Dai, M.; Woehrstein, J. B.; Agasti, S. S.; Feiger, Z.; Rodal, A.; Yin, P. Quantitative super-resolution imaging with qPAINT. Nat. Methods 2016, 13 (5), 439– 442, DOI: 10.1038/nmeth.3804There is no corresponding record for this reference.
From NLM.
- 156Schäfer, P.; Tenschert, W.; Schröter, M.; Gutensohn, K.; Laufs, R. False-Positive Results of Plasma PCR for Cytomegalovirus DNA due to Delayed Sample Preparation. Journal of Clinical Microbiology 2000, 38 (9), 3249– 3253, DOI: 10.1128/JCM.38.9.3249-3253.2000There is no corresponding record for this reference.
(accessed 2025/10/26).
- 157Faubert, A.; Bohsina, R.; You, E.; Hébert, M.; Griffith, M.; Aubin, M. J. Diagnostic Accuracy of Conjunctival Rt-Pcr in Sars-Cov-2: A Systematic Review and Diagnostic Accuracy Meta-Analysis. Ocul Immunol Inflamm 2024, 32 (8), 1621– 1632, DOI: 10.1080/09273948.2023.2272200There is no corresponding record for this reference.
From NLM.
- 158Ruijter, J. M.; Barnewall, R. J.; Marsh, I. B.; Szentirmay, A. N.; Quinn, J. C.; van Houdt, R.; Gunst, Q. D.; van den Hoff, M. J. B. Efficiency Correction Is Required for Accurate Quantitative PCR Analysis and Reporting. Clinical Chemistry 2021, 67 (6), 829– 842, DOI: 10.1093/clinchem/hvab052There is no corresponding record for this reference.
(accessed 10/26/2025).
- 159Louw, T. M.; Booth, C. S.; Pienaar, E.; TerMaat, J. R.; Whitney, S. E.; Viljoen, H. J. Experimental validation of a fundamental model for PCR efficiency. Chem. Eng. Sci. 2011, 66 (8), 1783– 1789, DOI: 10.1016/j.ces.2011.01.029There is no corresponding record for this reference.
- 160Zhao, L.; Qiu, M.; Li, X.; Yang, J.; Li, J. CRISPR-Cas13a system: A novel tool for molecular diagnostics. Front Microbiol 2022, 13, 1060947, DOI: 10.3389/fmicb.2022.1060947There is no corresponding record for this reference.
From NLM.
- 161Metcalf, G. A. D. MicroRNAs: circulating biomarkers for the early detection of imperceptible cancers via biosensor and machine-learning advances. Oncogene 2024, 43 (28), 2135– 2142, DOI: 10.1038/s41388-024-03076-3There is no corresponding record for this reference.
- 162Schofield, A. L.; Brown, J. P.; Brown, J.; Wilczynska, A.; Bell, C.; Glaab, W. E.; Hackl, M.; Howell, L.; Lee, S.; Dear, J. W.; Remes, M.; Reeves, P.; Zhang, E.; Allmer, J.; Norris, A.; Falciani, F.; Takeshita, L. Y.; Seyed Forootan, S.; Sutton, R.; Park, B. K.; Goldring, C. Systems analysis of miRNA biomarkers to inform drug safety. Arch. Toxicol. 2021, 95 (11), 3475– 3495, DOI: 10.1007/s00204-021-03150-9There is no corresponding record for this reference.
From NLM.
- 163Visone, R.; Croce, C. M. MiRNAs and cancer. Am. J. Pathol. 2009, 174 (4), 1131– 1138, DOI: 10.2353/ajpath.2009.080794There is no corresponding record for this reference.
From NLM.
- 164Kroh, E. M.; Parkin, R. K.; Mitchell, P. S.; Tewari, M. Analysis of circulating microRNA biomarkers in plasma and serum using quantitative reverse transcription-PCR (qRT-PCR). Methods 2010, 50 (4), 298– 301, DOI: 10.1016/j.ymeth.2010.01.032There is no corresponding record for this reference.
From NLM.
- 165Vester, B.; Wengel, J. LNA (locked nucleic acid): high-affinity targeting of complementary RNA and DNA. Biochemistry 2004, 43 (42), 13233– 13241, DOI: 10.1021/bi0485732There is no corresponding record for this reference.
From NLM.
- 166Woo, N.; Brown, C. M. Review of expansion microscopy combined with advanced imaging modalities. J. Microsc. 2026, 301, 335, DOI: 10.1111/jmi.70048There is no corresponding record for this reference.
- 167Xia, C.; Fan, J.; Emanuel, G.; Hao, J.; Zhuang, X. Spatial transcriptome profiling by MERFISH reveals subcellular RNA compartmentalization and cell cycle-dependent gene expression. Proc. Natl. Acad. Sci. U. S. A. 2019, 116 (39), 19490– 19499, DOI: 10.1073/pnas.1912459116There is no corresponding record for this reference.
- 168Stoldt, S.; Maass, F.; Weber, M.; Dennerlein, S.; Ilgen, P.; Gärtner, J.; Canfes, A.; Schweighofer, S. V.; Jans, D. C.; Rehling, P.; Jakobs, S. Super-resolution microscopy of mitochondrial mRNAs. Nat. Commun. 2025, 16 (1), 6391, DOI: 10.1038/s41467-025-61577-5There is no corresponding record for this reference.
- 169Rodermund, L.; Coker, H.; Oldenkamp, R.; Wei, G.; Bowness, J.; Rajkumar, B.; Nesterova, T.; Susano Pinto, D. M.; Schermelleh, L.; Brockdorff, N. Time-resolved structured illumination microscopy reveals key principles of Xist RNA spreading. Science 2021, 372 (6547), eabe7500 DOI: 10.1126/science.abe7500There is no corresponding record for this reference.
- 170Shao, W.; Zeng, S.-T.; Yu, Z.-Y.; Tang, G.-X.; Chen, S.-B.; Huang, Z.-S.; Chen, X.-C.; Tan, J.-H. Tracking Stress Granule Dynamics in Live Cells and In Vivo with a Small Molecule. Anal. Chem. 2021, 93 (49), 16297– 16301, DOI: 10.1021/acs.analchem.1c03577There is no corresponding record for this reference.
- 171Cawte, A. D.; Unrau, P. J.; Rueda, D. S. Live cell imaging of single RNA molecules with fluorogenic Mango II arrays. Nat. Commun. 2020, 11 (1), 1283, DOI: 10.1038/s41467-020-14932-7There is no corresponding record for this reference.
- 172Park, H. Y.; Trcek, T.; Wells, A. L.; Chao, J. A.; Singer, R. H. An Unbiased Analysis Method to Quantify mRNA Localization Reveals Its Correlation with Cell Motility. Cell Reports 2012, 1 (2), 179– 184, DOI: 10.1016/j.celrep.2011.12.009There is no corresponding record for this reference.
(accessed 2025/10/26).
- 173Tinevez, J. Y.; Perry, N.; Schindelin, J.; Hoopes, G. M.; Reynolds, G. D.; Laplantine, E.; Bednarek, S. Y.; Shorte, S. L.; Eliceiri, K. W. TrackMate: An open and extensible platform for single-particle tracking. Methods 2017, 115, 80– 90, DOI: 10.1016/j.ymeth.2016.09.016There is no corresponding record for this reference.
From NLM.
- 174Liu, J.; Cho, I.-H.; Cui, Y.; Irudayaraj, J. Second Harmonic Super-resolution Microscopy for Quantification of mRNA at Single Copy Sensitivity. ACS Nano 2014, 8 (12), 12418– 12427, DOI: 10.1021/nn505096tThere is no corresponding record for this reference.
- 175Campagnola, P. J.; Clark, H. A.; Mohler, W. A.; Lewis, A.; Loew, L. M. Second-harmonic imaging microscopy of living cells. J. Biomed Opt 2001, 6 (3), 277– 286, DOI: 10.1117/1.1383294There is no corresponding record for this reference.
From NLM.
- 176Aghigh, A.; Bancelin, S.; Rivard, M.; Pinsard, M.; Ibrahim, H.; Légaré, F. Second harmonic generation microscopy: a powerful tool for bio-imaging. Biophys Rev. 2023, 15 (1), 43– 70, DOI: 10.1007/s12551-022-01041-6There is no corresponding record for this reference.
From NLM.
- 177Chen, X.; Nadiarynkh, O.; Plotnikov, S.; Campagnola, P. J. Second harmonic generation microscopy for quantitative analysis of collagen fibrillar structure. Nat. Protoc 2012, 7 (4), 654– 669, DOI: 10.1038/nprot.2012.009There is no corresponding record for this reference.
From NLM.
- 178Stanciu, S. G.; Hristu, R.; Stanciu, G. A.; Tranca, D. E.; Eftimie, L.; Dumitru, A.; Costache, M.; Stenmark, H. A.; Manders, H.; Cherian, A.; Tark-Dame, M.; Manders, E. M. M. Super-resolution re-scan second harmonic generation microscopy. Proc. Natl. Acad. Sci. U. S. A. 2022, 119 (47), e2214662119 DOI: 10.1073/pnas.2214662119There is no corresponding record for this reference.
- 179Ding, J.; Lee, Y.-T.; Bhandari, Y.; Schwieters, C. D.; Fan, L.; Yu, P.; Tarosov, S. G.; Stagno, J. R.; Ma, B.; Nussinov, R.; Rein, A.; Zhang, J.; Wang, Y.-X. Visualizing RNA conformational and architectural heterogeneity in solution. Nat. Commun. 2023, 14 (1), 714, DOI: 10.1038/s41467-023-36184-xThere is no corresponding record for this reference.
- 180Degenhardt, M. F. S.; Degenhardt, H. F.; Bhandari, Y. R.; Lee, Y.-T.; Ding, J.; Yu, P.; Heinz, W. F.; Stagno, J. R.; Schwieters, C. D.; Watts, N. R.; Wingfield, P. T.; Rein, A.; Zhang, J.; Wang, Y.-X. Determining structures of RNA conformers using AFM and deep neural networks. Nature 2025, 637 (8048), 1234– 1243, DOI: 10.1038/s41586-024-07559-xThere is no corresponding record for this reference.
- 181Luo, B.; Zhang, C.; Ling, X.; Mukherjee, S.; Jia, G.; Xie, J.; Jia, X.; Liu, L.; Baulin, E. F.; Luo, Y.; Jiang, L.; Dong, H.; Wei, X.; Bujnicki, J. M.; Su, Z. Cryo-EM reveals dynamics of Tetrahymena group I intron self-splicing. Nature Catalysis 2023, 6 (4), 298– 309, DOI: 10.1038/s41929-023-00934-3There is no corresponding record for this reference.
- 182Liu, J.; McRae, E. K. S.; Zhang, M.; Geary, C.; Andersen, E. S.; Ren, G. Non-averaged single-molecule tertiary structures reveal RNA self-folding through individual-particle cryo-electron tomography. Nat. Commun. 2024, 15 (1), 9084, DOI: 10.1038/s41467-024-52914-1There is no corresponding record for this reference.
- 183Krieg, M.; Fläschner, G.; Alsteens, D.; Gaub, B. M.; Roos, W. H.; Wuite, G. J. L.; Gaub, H. E.; Gerber, C.; Dufrêne, Y. F.; Müller, D. J. Atomic force microscopy-based mechanobiology. Nature Reviews Physics 2019, 1 (1), 41– 57, DOI: 10.1038/s42254-018-0001-7There is no corresponding record for this reference.
- 184Berrevoets, E. S.; Kessler, L. F.; Balakrishnan, A.; Okuda, E. K.; Müller-McNicoll, M.; Rieger, B.; Stallinga, S.; Heilemann, M. Smart 3D super-resolution microscopy reveals the architecture of the RNA scaffold in a nuclear body. Nat. Commun. 2025, 16 (1), 10689, DOI: 10.1038/s41467-025-65723-xThere is no corresponding record for this reference.
- 185Autour, A.; C. Y. Jeng, S.; D. Cawte, A.; Abdolahzadeh, A.; Galli, A.; Panchapakesan, S. S. S.; Rueda, D.; Ryckelynck, M.; Unrau, P. J. Fluorogenic RNA Mango aptamers for imaging small non-coding RNAs in mammalian cells. Nat. Commun. 2018, 9 (1), 656, DOI: 10.1038/s41467-018-02993-8There is no corresponding record for this reference.
From NLM.



