Applications of Raman Spectroscopy in Pandemic Virology: A Comprehensive ReviewClick to copy article linkArticle link copied!
- Hulya YilmazHulya YilmazMember of Leibniz Health Technologies, Member of the Leibniz Centre for Photonics in Infection Research (LPI), Leibniz Institute of Photonic Technology, Albert-Einstein-Straße 9, 07745 Jena, GermanyMember of the Leibniz Centre for Photonics in Infection Research (LPI), Institute of Physical Chemistry (IPC) and Abbe Center of Photonics (ACP), Friedrich Schiller University Jena, Helmholtzweg 4, 07743 Jena, GermanyMore by Hulya Yilmaz
- Anuradha RamojiAnuradha RamojiMember of Leibniz Health Technologies, Member of the Leibniz Centre for Photonics in Infection Research (LPI), Leibniz Institute of Photonic Technology, Albert-Einstein-Straße 9, 07745 Jena, GermanyMember of the Leibniz Centre for Photonics in Infection Research (LPI), Institute of Physical Chemistry (IPC) and Abbe Center of Photonics (ACP), Friedrich Schiller University Jena, Helmholtzweg 4, 07743 Jena, GermanyMore by Anuradha Ramoji
- Andreea WinterfeldAndreea WinterfeldMember of Leibniz Health Technologies, Member of the Leibniz Centre for Photonics in Infection Research (LPI), Leibniz Institute of Photonic Technology, Albert-Einstein-Straße 9, 07745 Jena, GermanyMore by Andreea Winterfeld
- Hamideh SalehiHamideh SalehiUniversity of Strasbourg, ICube UMR 7357, CNRS, Inserm, Strasbourg 67000, FranceMore by Hamideh Salehi
- Aykut OzkulAykut OzkulDepartment of Virology, Faculty of Veterinary Medicine, Ankara University, Ankara 06070, TurkeyMore by Aykut Ozkul
- Jürgen Popp*Jürgen Popp*Email: [email protected]Member of Leibniz Health Technologies, Member of the Leibniz Centre for Photonics in Infection Research (LPI), Leibniz Institute of Photonic Technology, Albert-Einstein-Straße 9, 07745 Jena, GermanyMember of the Leibniz Centre for Photonics in Infection Research (LPI), Institute of Physical Chemistry (IPC) and Abbe Center of Photonics (ACP), Friedrich Schiller University Jena, Helmholtzweg 4, 07743 Jena, GermanyInfectoGnostics Research Campus Jena, Center of Applied Research, Philosophenweg 7, Jena 07743, GermanyMore by Jürgen Popp
Abstract
Raman spectroscopy, with its ability to provide molecular fingerprinting of biological samples, has recently emerged as a powerful technique in virology. Unfortunately, emerging and re-emerging viral pathogens with a rising threat highlight the urgent requirement for rapid, sensitive, and label-free diagnostic tools to monitor both viral presence and host responses. Recently, Raman spectroscopy use has increased not only in nonpandemic viruses but also in pandemic-prone viruses. This review presents a comprehensive analysis of Raman spectroscopy and its enhanced variants, such as surface-enhanced Raman spectroscopy (SERS), tip-enhanced Raman spectroscopy (TERS), and coherent Raman spectroscopies (CARS/SRS), in the context of pandemic virus research. By improving sensitivity, spatial resolution, and acquisition speed, these variants allow for quick and, in some cases, real-time analysis of viral particles and virus-host interactions in clinical samples and infected cells. A systematic evaluation of Raman-based methodologies was conducted across several virus families with pandemic potential, including Orthomyxoviridae, Coronaviridae, Filoviridae, Flaviviridae, and others, selected for their roles in past pandemics. Furthermore, prominent technical and virological obstacles hindering the expanded application of these methods were identified, alongside a discussion of prospective future directions for research and development. This effort aims to consolidate the current status of Raman spectroscopy in pandemic virology to enhance its integration into worldwide research on pandemic virus surveillance, pathogenesis, and diagnostics.
This publication is licensed under
License Summary*
You are free to share(copy and redistribute) this article in any medium or format within the parameters below:
Creative Commons (CC): This is a Creative Commons license.
Attribution (BY): Credit must be given to the creator.
Non-Commercial (NC): Only non-commercial uses of the work are permitted.
No Derivatives (ND): Derivative works may be created for non-commercial purposes, but sharing is prohibited.
*Disclaimer
This summary highlights only some of the key features and terms of the actual license. It is not a license and has no legal value. Carefully review the actual license before using these materials.
License Summary*
You are free to share(copy and redistribute) this article in any medium or format within the parameters below:
Creative Commons (CC): This is a Creative Commons license.
Attribution (BY): Credit must be given to the creator.
Non-Commercial (NC): Only non-commercial uses of the work are permitted.
No Derivatives (ND): Derivative works may be created for non-commercial purposes, but sharing is prohibited.
*Disclaimer
This summary highlights only some of the key features and terms of the actual license. It is not a license and has no legal value. Carefully review the actual license before using these materials.
License Summary*
You are free to share(copy and redistribute) this article in any medium or format within the parameters below:
Creative Commons (CC): This is a Creative Commons license.
Attribution (BY): Credit must be given to the creator.
Non-Commercial (NC): Only non-commercial uses of the work are permitted.
No Derivatives (ND): Derivative works may be created for non-commercial purposes, but sharing is prohibited.
*Disclaimer
This summary highlights only some of the key features and terms of the actual license. It is not a license and has no legal value. Carefully review the actual license before using these materials.
License Summary*
You are free to share(copy and redistribute) this article in any medium or format within the parameters below:
Creative Commons (CC): This is a Creative Commons license.
Attribution (BY): Credit must be given to the creator.
Non-Commercial (NC): Only non-commercial uses of the work are permitted.
No Derivatives (ND): Derivative works may be created for non-commercial purposes, but sharing is prohibited.
*Disclaimer
This summary highlights only some of the key features and terms of the actual license. It is not a license and has no legal value. Carefully review the actual license before using these materials.
License Summary*
You are free to share(copy and redistribute) this article in any medium or format within the parameters below:
Creative Commons (CC): This is a Creative Commons license.
Attribution (BY): Credit must be given to the creator.
Non-Commercial (NC): Only non-commercial uses of the work are permitted.
No Derivatives (ND): Derivative works may be created for non-commercial purposes, but sharing is prohibited.
*Disclaimer
This summary highlights only some of the key features and terms of the actual license. It is not a license and has no legal value. Carefully review the actual license before using these materials.
Introduction
Pandemic Viruses
Figure 1
Figure 1. Overview of virion classification and their properties (Ramoji, A., Pahlow, S., Pistiki, A., Rueger, J., Shaik, T. A., Shen, H., Wichmann, C., Krafft, C.,& Popp, J. (2022). Understanding viruses and viral infections by biophotonic methods. Translational Biophotonics, 4(1–2), e202100008. 10.1002/tbio.202100008). Reprinted from ref (2).
| viral family | type of nucleic acid in the genome | virus | reservoir | human infection from reservoir and human |
|---|---|---|---|---|
| Orthomyxoviridae | –ssRNA | Influenza virus | Wild waterfowl | – Airborne or close contact with infected animals |
| Filoviridae | –ssRNA | Marburg virus | Bats | – Contact/consumption of infected bats or intermediary hosts |
| – Direct contact with body fluids | ||||
| Ebola virus | Bats | – Contact/consumption of infected bats or intermediary hosts | ||
| – Direct contact with body fluids | ||||
| Flaviviridae | –ssRNA | Dengue Virus | Arthropods | – Aedes species mosquitoes (Ae. aegypti or Ae. albopictus). |
| Zika virus | Arthropods | – Arthropod-borne (mosquito bite) Perinatal, sexual contact | ||
| Picornaviridae | +ssRNA | Echovirus | Anthroponotic | – Fecal-oral transmission, respiratory droplets |
| Polio virus | Anthroponotic | – N/A, -Fecal-oral transmission | ||
| Poxviridae | dsDNA | Smallpox virus | Anthroponotic | – N/A, -Droplets (respiratory secretions) |
| Monkeypox virus | Most likely small mammals | – Direct contact with infected animals | ||
| – Close, personal contact; fomites; respiratory secretions | ||||
| Nairoviridae | –ssRNA | Crimean-Congo hemorrhagic fever | Arthropods (ticks) | – Arthropod-borne |
| – Direct contact with body fluids | ||||
| Arenaviridae | –ssRNA | Lassa virus | Rodents (Mastomys species) | – Contact with rodent urine or feces |
| – Direct contact with body fluids | ||||
| Coronaviridae | +nsRNA | SARS-CoV-2 | Bats | – Airborne or close contact with infected animals |
| MERS | Bats, camels | – Direct contact with intermediary hosts such as camels | ||
| SARS | Bats | – Unknown (possibly close contact with intermediary hosts), airborne | ||
| Paramyxoviridae | –ssRNA | Nipah virus | Bats | – Consumption of infected bats or pigs, virus-contaminated fruits |
| – Consumption of infected bats or pigs, virus-contaminated fruits-Respiratory secretions | ||||
| Phenuiviridae | –ssRNA | Rift Valley fever virus | Arthropods (mosquitoes, ticks) | – Close contact with infected livestock |
| – No human-to-human transmission |
MERS: Middle East respiratory syndrome, SARS: Severe acute respiratory syndrome, EVD: Ebola virus disease, DCDRS: Drop-coating deposition Raman spectroscopy, – ssRNA: Negative-stranded RNA, + ssRNA: Positive-stranded RNA, + nsRNA: nonsegmented positive-strand RNA, dsDNA: double-stranded DNA.
Conventional Techniques for Virus Structure, Function, and Host Interaction
| techniques | types | advantages | limitations | references |
|---|---|---|---|---|
| electron microscopy | microscopy | – High-resolution imaging of viral particles - No need for specific reagents tailored to identify the pathogenic organism. | – Requires extensive sample preparation and expensive equipment | (3,13) |
| confocal microscopy | – Visualization of virus localization in cells and tissue | – Fluorescent labels requirement -Limited depth penetration - Potential for photobleaching - Requires extensive sample preparation and expensive equipment | ||
| – Axial resolution in the range of 100–150 nm. | ||||
| viral genome sequencing | molecular biology | – Precise identification of viral strains and mutations | – Requires high-quality nucleic acids and bioinformatics | (15,14,21) |
| – Single-molecule long-read sequencing | ||||
| reverse genetics | – Enables manipulation of viral genomes to study gene function | – Technically complex; requires a specialized system | ||
| reporter assays | – Real-time monitoring of viral infection using reporter genes | – May require transgenic systems or cell line engineering | ||
| gene knockdown/knockout | – Targeted study of gene function in host-virus interactions | – Potential off-target effects; requires validation | ||
| mass spectrometry | proteomics and metabolomics | – Sensitive detection of viral proteins and metabolites | – High cost - Complex sample preparation and data analysis | (6,16) |
| – Identifies direct and indirect molecular interactions | ||||
| enzyme-linked immunosorbent assay | immunological techniques | – Sensitive, high-throughput quantification of antigens or antibodies | – May produce false positives; requires careful controls | (17,18) |
| flow cytometry | – Rapid, multiparametric analysis of infected cells | – Requires labeled antibodies; limited to known targets | ||
| immunohistochemistry | – Localizes viral antigens in tissue samples | – Relatively low throughput; dependent on antibody specificity | ||
| viral entry assays | functional assays | – Quantifies viral entry efficiency into host cells | – Limited to in vitro systems; may not mimic in vivo conditions | (7,19) |
| viral replication assays | – Measures efficiency of viral genome replication | – May not distinguish between infectious and noninfectious particles | ||
| cell viability assays | – Assesses cell health and cytopathic effects of viral infection | – May not directly measure viral activity | ||
| computational modeling | systems biology | – Simulates infection dynamics; integrates large data sets | – Model assumptions require experimental validation | (20) |
Spontaneous Raman Spectroscopy and Its Variants
Figure 2
Figure 2. (A) Illustration of Stokes and anti-Stokes Raman scattering, showing energy transitions during spontaneous inelastic light scattering. (Anti-Stokes scattering is shown for completeness of the Raman process.) (B) SERS mechanism, where localized surface plasmons on metallic nanostructures enhance the EM and Raman signal. (C) Schematic of CARS, highlighting the nonlinear four-wave mixing process generating anti-Stokes emission (D) TERS, utilizing a plasmonic metallic tip to confine and enhance the EM field at the nanoscale for high-resolution Raman imaging. (Jones, R.R., Hooper, DC. Zhang, L. et al. Raman Techniques: Fundamentals and Frontiers. Nanoscale Res Lett 14, 231 (2019). 10.1186/s11671-019-3039-2). Adapted from ref (22).
| technique | description | advantages | limitations | refs |
|---|---|---|---|---|
| Spontaneous Raman | – Inelastic light scattering to capture molecular vibrations | – Nondestructive | – Weak signal | (32) |
| – Water insensitive | – Fluorescence interference | |||
| RRS | – Laser tuned to molecular transitions | – Selective enhancement for chromophores | – Not suitable for all molecules | (30) |
| SERS | – Nanostructure | – High sensitivity (single-molecule level) | – Requires metal substrates | (24,33) |
| – Enhanced Raman scattering | – Label-free | – Complex preparation | ||
| TERS | – Raman with scanning probe tip | – Nanoscale spatial resolution | – Requires flat samples | (24,26,27,34,35) |
| – Challenging tip control and its cost | ||||
| CARS | – Nonlinear Raman imaging | – Real-time | – Complex laser setup | (24,25,28) |
| – 3D imaging of live cells | ||||
| SRS | – Stimulated molecular vibrations with two lasers | – Fast, quantitative chemical imaging | – Expensive, complex instrumentation | (24,30) |
PAndemic Viruses’ Studies by Raman Spectroscopy and Its Variants
Orthomyxoviridae Family
Influenza Virus
Filoviridae Family
Marburg Virus (MARV)
Ebola Virus
Flaviviridae Family
Dengue Virus (DENV)
Figure 3
Figure 3. (A) Mean SERS spectra of NS1-positive dengue samples, NS1-negative febrile controls, and healthy individuals, highlighting distinct biochemical signatures. (B) PCA score plot showing clear separation of the three groups based on SERS spectral features. (C) PC1 loading plot indicating Raman bands contributing most strongly to classification. (Gahlaut, S. K.; Savargaonkar, D.; Sharan, C.; Yadav, Sarjana, Mishra, P.; Singh, J. P. SERS Platform for Dengue Diagnosis from Clinical Samples Employing a Hand Held Raman Spectrometer. Anal. Chem.2020, 92 (3), 2527–2534. 10.1021/acs.analchem.9b04129) Adapted from ref (145).
| Raman shift (cm–1) | assignment | interpretation |
|---|---|---|
| ∼488 | Disulfide bond (S–S stretching) | Protein tertiary structure |
| ∼636 | N-acetyl glucosamine/phenyl ring | Glycoprotein-associated signatures |
| ∼720–735 | Adenine ring breathing | Viral genome presence; increased host transcription |
| ∼780–785 | Cytosine/uracil ring breathing | Viral nucleic acid fingerprint |
| ∼830–850 | Tyrosine residues | Immune signaling, phosphorylation |
| ∼860 | Nucleic acids/phosphodiester | Genome/backbone-associated signatures in infection |
| ∼892 | Deoxyribose (in-plane mode) | Nucleic-acid–associated changes |
| ∼940–960 | Protein backbone (α-helix) | Conformational protein changes |
| ∼1000–1006 | Phenylalanine | Structural proteins; total protein content |
| ∼1080–1100 | PO2– stretching | Membrane remodeling during replication |
| ∼1120–1130 | C–C stretching of lipids/proteins | Enveloped virus identification |
| ∼1156 | C–C stretch mode of β-carotene | Carotenoid depletion associated with viral infection |
| ∼1200–1210 | Tryptophan/Amide III | Protein/aromatic amino-acid changes; infection signature |
| ∼1240–1270 | Amide III β | Protein secondary structure changes |
| ∼1330–1360 | CH deformation/nucleic acids | Metabolic activation; RNA/protein overlap |
| ∼1425–1450 | CH2/CH3 deformation | Lipid metabolism alteration |
| ∼1516 | β-carotene (C = C stretching) | Carotenoid depletion associated with viral infection |
| ∼1520–1560 | Amide II/aromatic residues | Viral protein composition; variant effects |
| ∼ 1570–1610 | Tryptophan | Protein-associated changes |
| ∼1650–1665 | Amide I | Protein folding, inflammation |
| ∼1730–1750 | C = O ester stretches | Lipid oxidation, membrane damage |
| ∼2850–3000 | C–H stretching (lipids/proteins) | Membrane remodeling/lipid–protein content |
Band positions may shift depending on excitation wavelength, enhancement strategy, and local chemical environment.
Zika Virus
Figure 4
Figure 4. Multiplexed SERS assay where Zika and Dengue test lines are separately painted. SERS spectra are shown for Zika (A) and Dengue (B) test lines and a combined Zika-Dengue test line (C). (Sánchez-Purrà M, Carré-Camps M, de Puig H, Bosch I, Gehrke L, Hamad-Schifferli K. (2017) Surface-Enhanced Raman Spectroscopy-Based Sandwich Immunoassays for Multiplexed Detection of Zika and Dengue Viral Biomarkers. ACS Infect Dis.;3(10):767–776. doi: 10.1021/acsinfecdis.7b00110). Adapted from ref (63).
Picornaviridae Family
Echovirus
Poliovirus
Poxviridae Family
Smallpox Virus
Monkeypox Virus (MPXV)
Nairoviridae Family
Crimean-Congo Hemorrhagic Fever Virus (CCHF)
Arenaviridae Family
Lassa Virus
Coronaviridae Family
SARS-CoV-2
Figure 5
Figure 5. Development of a Raman spectral database for the SARS-CoV-2 spike protein. (a) Schematic overview illustrating the structural organization of the spectral database. (b) Standard Raman spectra obtained for the full-length spike (S) protein, its subunits S1 and S2, and RBD. (c) Comparative analysis between computed and experimentally recorded Raman spectra for the S1 subunit. (d) Optical image of the S protein dried on a silver-coated silicon substrate; scale bar: 300 μm. (e) Scanning electron microscopy image depicting the Au nanoparticle array used for SERS measurements; scale bar: 200 nm. (f) Comparative Raman profiles of the spike proteins from SARS-CoV-2, SARS-CoV, and MERS-CoV, expressed in insect cells using the baculovirus system. (g) Raman spectral differences in SARS-CoV-2 S protein before and after thermal treatment. (h) SERS spectra of the SARS-CoV-2 spike protein. (Huang, J., Wen, J., Zhou, M., Ni, S., Le, W., Chen, G., Wei, L., Zeng, Y., Qi, D., Pan, M., Xu, J., Wu, Y., Li, Z., Feng, Y., Zhao, Z., He, Z., Li, B., Zhao, S., Zhang, B., Xue, P., He, S., Fang, K., Zhao, Y.,& Du, K. (2021). On-site detection of SARS-CoV-2 antigen by deep learning-based surface-enhanced Raman spectroscopy and its biochemical foundations. Analytical Chemistry, 93(26), 9174–9182.) (Adapted from Analytical Chemistry, 2021, 93, 9174–9182). Reprinted from ref (87).
Figure 6
Figure 6. (A) PCA of confocal Raman spectra extracted from intracellular compartments of Control, measles virus (MeV)-infected, and SARS-CoV-2-infected Vero E6 cells. (A, B) Cytoplasm region: (A) 3D PCA score plot and (B) corresponding PC loadings. (C, D) Golgi-mitochondria region: (C) PCA score plot and (D) PC loadings. (E, F) Nucleus region: (E) PCA score plot and (F) PC loadings. (Salehi, H.; Ramoji, A.; Mougari, S.; Merida, P.; Neyret, A.; Popp, J.; Horvat, B.; Muriaux, D.; Cuisinier, F. Specific intracellular signature of SARS-CoV-2 infection using confocal Raman microscopy. Commun Chem2022, 5 (1), 85. DOI: 10.1038/s42004–022–00702–7.) Reprinted from ref (97).
Figure 7
Figure 7. (A) Representative SERS spectra of SARS-CoV-2, human coronavirus OC43 (HCoV-OC43), SARS-CoV-2 Omicron variant BA.5.1, and CCoV, acquired on AuNP films under 785 nm excitation. Major diagnostic bands are labeled, and the background trace corresponds to virus-free control samples. (B) Three-dimensional PCA score plot showing clear clustering and separation of the four coronavirus types based on their SERS spectral signatures. (C) Corresponding PCA loading vectors for PC1, PC2, and PC3, highlighting the Raman bands contributing most strongly to virus discrimination. (Zhou, L.; Vestri, A.; Marchesano, V.; Rippa, M.; Sagnelli, D.; Picazio, G.; Fusco, G.; Han, J.; Zhou, J.; Petti, L. The Label-Free Detection and Identification of SARS-CoV-2 Using Surface-Enhanced Raman Spectroscopy and Principal Component Analysis. Biosensors (Basel) 2023, 13 (12). DOI: 10.3390/bios13121014) Adapted from ref (104).
MERS-CoV
SARS-CoV
Paramyxoviridae Family
Nipah Virus (NiV)
Phenuiviridae Family
Rift Valley Fever Virus
Challenges and Future Directions
Technical Challenges in Raman Spectroscopy for Virology
| challenge | strategy | mechanism |
|---|---|---|
| Autofluorescence/background | Near-IR excitation | Reduces fluorophore excitation |
| Weak Raman signal | SERS assays (LFA/microdevices) | Plasmonic field enhancement |
| Low capture/poor repeatability | Affinity capture (“virus-trap” etc.) | Target enrichment at hotspots |
| Spectral congestion/variability | Preprocessing, chemometrics | Baseline removal, variance separation |
| Subtle class separation | ML/deep learning | Nonlinear spectral pattern learning |
| Fluorescence-dominated imaging | Coherent Raman (CARS/SRS) | Fast nonlinear contrast; low fluorescence |
| Biosafety limits | Surrogates (inactivated/VLP/pseudotype) | Lower-BSL method development |
Virological and Biosafety Challenges in Pandemic Virus Research
Discussion
Conclusions
Acknowledgments
The authors acknowledge Dr. Cavit Agca for assistance with reference formatting.
References
This article references 150 other publications.
- 1Neumann, G.; Kawaoka, Y. Which Virus Will Cause the Next Pandemic?. Viruses 2023, 15 (1), 199, DOI: 10.3390/v15010199Google ScholarThere is no corresponding record for this reference.
- 2Ramoji, A.; Pahlow, S.; Pistiki, A.; Rueger, J.; Shaik, T. A.; Shen, H.; Wichmann, C.; Krafft, C.; Popp, J. Understanding viruses and viral infections by biophotonic methods. Translational Biophotonics 2022, 4 (1–2), e202100008, DOI: 10.1002/tbio.202100008Google ScholarThere is no corresponding record for this reference.
- 3Liu, D.; Pan, L.; Zhai, H.; Qiu, H. J.; Sun, Y. Virus tracking technologies and their applications in viral life cycle: research advances and future perspectives. Front. Immunol. 2023, 14, 1204730 DOI: 10.3389/fimmu.2023.1204730Google ScholarThere is no corresponding record for this reference.
- 4Maginnis, M. S. Virus-Receptor Interactions: The Key to Cellular Invasion. J. Mol. Biol. 2018, 430 (17), 2590– 2611, DOI: 10.1016/j.jmb.2018.06.024Google ScholarThere is no corresponding record for this reference.
- 5Jones, J. E.; Le Sage, V.; Lakdawala, S. S. Viral and host heterogeneity and their effects on the viral life cycle. Nat. Rev. Microbiol. 2021, 19 (4), 272– 282, DOI: 10.1038/s41579-020-00449-9Google ScholarThere is no corresponding record for this reference.
- 6Idrees, S.; Chen, H.; Panth, N.; Paudel, K. R.; Hansbro, P. M. Exploring Viral-Host Protein Interactions as Antiviral Therapies: A Computational Perspective. Microorganisms 2024, 12 (3). 630 DOI: 10.3390/microorganisms12030630 .Google ScholarThere is no corresponding record for this reference.
- 7Datta, S.; Hett, E. C.; Vora, K. A.; Hazuda, D. J.; Oslund, R. C.; Fadeyi, O. O.; Emili, A. The chemical biology of coronavirus host-cell interactions. RSC Chem. Biol. 2021, 2 (1), 30– 46, DOI: 10.1039/D0CB00197JGoogle ScholarThere is no corresponding record for this reference.
- 8Merenich, D. K. E. V. M.-B. C. J. K. A. M. Advanced microscopy techniques for the visualization and analysis of cell behaviors. In Cell Movement in Health and Disease; Michael Schnoor, L.-M. Y.; Sean, X. Sun., Eds.; Academic Press, 2022.Google ScholarThere is no corresponding record for this reference.
- 9Kianzad, S.; SeyedAlinaghi, S.; Asadollahi-Amin, A.; Dadras, O.; Karimi, A.; Afsahi, A. M.; MohsseniPour, M.; Barzegary, A.; Mirzapour, P.; Mirghaderi, S. P. Comparison of SARS-CoV-2 (Coronavirus) with other Similar Viruses Based on Current Evidence J. Iranian Med. Council 2022 DOI: 10.18502/jimc.v5i1.9565 .Google ScholarThere is no corresponding record for this reference.
- 10Islam, M. T.; Quispe, C.; Herrera-Bravo, J.; Sarkar, C.; Sharma, R.; Garg, N.; Fredes, L. I.; Martorell, M.; Alshehri, M. M.; Sharifi-Rad, J. Production, Transmission, Pathogenesis, and Control of Dengue Virus: A Literature-Based Undivided Perspective. Biomed. Res. Int. 2021, 2021, 4224816 DOI: 10.1155/2021/4224816Google ScholarThere is no corresponding record for this reference.
- 11Ruokola, P.; Dadu, E.; Kazmertsuk, A.; Hakkanen, H.; Marjomaki, V.; Ihalainen, J. A. Raman spectroscopic signatures of echovirus 1 uncoating. J. Virol. 2014, 88 (15), 8504– 8513, DOI: 10.1128/JVI.03398-13Google ScholarThere is no corresponding record for this reference.
- 12Organization, W. H. WHO reports CDC , 2014https://www.bbc.com/news/world-africa-29769782.Google ScholarThere is no corresponding record for this reference.
- 13Goldsmith, C. S.; Miller, S. E. Modern uses of electron microscopy for detection of viruses. Clin. Microbiol Rev. 2009, 22 (4), 552– 563, DOI: 10.1128/CMR.00027-09Google ScholarThere is no corresponding record for this reference.
- 14Schaack, G. A.; Mehle, A. Experimental Approaches to Identify Host Factors Important for Influenza Virus. Cold Spring Harb Perspect Med. 2020, 10 (12), a038521, DOI: 10.1101/cshperspect.a038521Google ScholarThere is no corresponding record for this reference.
- 15Jansz, N.; Faulkner, G. J. Viral genome sequencing methods: benefits and pitfalls of current approaches. Biochem. Soc. Trans. 2024, 52 (3), 1431– 1447, DOI: 10.1042/BST20231322Google ScholarThere is no corresponding record for this reference.
- 16Mahmud, I.; Garrett, T. J. Mass Spectrometry Techniques in Emerging Pathogens Studies: COVID-19 Perspectives. J. Am. Soc. Mass Spectrom. 2020, 31 (10), 2013– 2024, DOI: 10.1021/jasms.0c00238Google ScholarThere is no corresponding record for this reference.
- 17Garg, T.; Weiss, C. R.; Sheth, R. A. Techniques for Profiling the Cellular Immune Response and Their Implications for Interventional Oncology. Cancers 2022, 14 (15). 3628 DOI: 10.3390/cancers14153628 .Google ScholarThere is no corresponding record for this reference.
- 18Oumarou Hama, H.; Aboudharam, G.; Barbieri, R.; Lepidi, H.; Drancourt, M. Immunohistochemical diagnosis of human infectious diseases: a review. Diagn. Pathol. 2022, 17 (1), 17, DOI: 10.1186/s13000-022-01197-5Google ScholarThere is no corresponding record for this reference.
- 19Iwanami, S.; Kitagawa, K.; Ohashi, H.; Asai, Y.; Shionoya, K.; Saso, W.; Nishioka, K.; Inaba, H.; Nakaoka, S.; Wakita, T. Should a viral genome stay in the host cell or leave? A quantitative dynamics study of how hepatitis C virus deals with this dilemma. PLoS Biol. 2020, 18 (7), e3000562 DOI: 10.1371/journal.pbio.3000562Google ScholarThere is no corresponding record for this reference.
- 20Ferrarini, M. G.; Lal, A.; Rebollo, R.; Gruber, A. J.; Guarracino, A.; Gonzalez, I. M.; Floyd, T.; de Oliveira, D. S.; Shanklin, J.; Beausoleil, E. Genome-wide bioinformatic analyses predict key host and viral factors in SARS-CoV-2 pathogenesis. Commun. Biol. 2021, 4 (1), 590, DOI: 10.1038/s42003-021-02095-0Google ScholarThere is no corresponding record for this reference.
- 21Ashmore-Harris, C.; Iafrate, M.; Saleem, A.; Fruhwirth, G. O. Non-invasive Reporter Gene Imaging of Cell Therapies, including T Cells and Stem Cells. Mol. Ther. 2020, 28 (6), 1392– 1416, DOI: 10.1016/j.ymthe.2020.03.016Google ScholarThere is no corresponding record for this reference.
- 22Jones, R. R.; Hooper, D. C.; Zhang, L.; Wolverson, D.; Valev, V. K. Raman Techniques: Fundamentals and Frontiers. Nanoscale Res. Lett. 2019, 14 (1), 231, DOI: 10.1186/s11671-019-3039-2Google ScholarThere is no corresponding record for this reference.
- 23Fernández-Galiana, Á.; Bibikova, O.; Vilms Pedersen, S.; Stevens, M. M. Fundamentals and Applications of Raman-Based Techniques for the Design and Development of Active Biomedical Materials. Adv. Mater. 2024, 36 (43), e2210807 DOI: 10.1002/adma.202210807Google ScholarThere is no corresponding record for this reference.
- 24Cialla-May, D.; Krafft, C.; Rosch, P.; Deckert-Gaudig, T.; Frosch, T.; Jahn, I. J.; Pahlow, S.; Stiebing, C.; Meyer-Zedler, T.; Bocklitz, T. Raman Spectroscopy and Imaging in Bioanalytics. Anal. Chem. 2022, 94 (1), 86– 119, DOI: 10.1021/acs.analchem.1c03235Google ScholarThere is no corresponding record for this reference.
- 25Deckert, V.; Deckert-Gaudig, T.; Cialla-May, D.; Popp, J.; Zell, R.; Deinhard-Emmer, S.; Sokolov, A. V.; Yi, Z.; Scully, M. O. Laser spectroscopic technique for direct identification of a single virus I: FASTER CARS. Proc. Natl. Acad. Sci. U. S. A. 2020, 117 (45), 27820– 27824, DOI: 10.1073/pnas.2013169117Google ScholarThere is no corresponding record for this reference.
- 26Xu, D.; Liang, B.; Xu, Y.; Liu, M. Recent advances in tip-enhanced Raman spectroscopy probe designs. Nano Res. 2023, 16 (4), 5555– 5571, DOI: 10.1007/s12274-022-5220-7Google ScholarThere is no corresponding record for this reference.
- 27Höppener, C.; Aizpurua, J.; Chen, H.; Gräfe, S.; Jorio, A.; Kupfer, S.; Zhang, Z.; Deckert, V. Tip-enhanced Raman scattering. Nature Reviews Methods Primers 2024, 4 (1), 47, DOI: 10.1038/s43586-024-00323-5Google ScholarThere is no corresponding record for this reference.
- 28Tian, Z. Y. M.; VB; ; Zhang, X.; Khalil, M.; Ayejoto, D. A. Coherent Anti-Stokes Raman Scattering. In Advanced Diagnostics in Combustion Science; Springer, 2023.Google ScholarThere is no corresponding record for this reference.
- 29Zhang, C.; Zhang, D.; Cheng, J. X. Coherent Raman Scattering Microscopy in Biology and Medicine. Annu. Rev. Biomed Eng. 2015, 17, 415– 445, DOI: 10.1146/annurev-bioeng-071114-040554Google ScholarThere is no corresponding record for this reference.
- 30Cheng, Q.; Miao, Y.; Wild, J.; Min, W.; Yang, Y. Emerging applications of stimulated Raman scattering microscopy in materials science. Matter 2021, 4 (5), 1460– 1483, DOI: 10.1016/j.matt.2021.02.013Google ScholarThere is no corresponding record for this reference.
- 31Orlando, A.; Franceschini, F.; Muscas, C.; Pidkova, S.; Bartoli, M.; Rovere, M.; Tagliaferro, A. A Comprehensive Review on Raman Spectroscopy Applications. Chemosensors 2021, 9 (9). 262 DOI: 10.3390/chemosensors9090262 .Google ScholarThere is no corresponding record for this reference.
- 32Lukose, J.; Barik, A. K.; Mithun, N.; Sanoop Pavithran, M.; George, S. D.; Murukeshan, V. M.; Chidangil, S. Raman spectroscopy for viral diagnostics. Biophys Rev. 2023, 15 (2), 199– 221, DOI: 10.1007/s12551-023-01059-4Google ScholarThere is no corresponding record for this reference.
- 33Lisyansky, A. A. A.; E S; ; Vinogradov, A. P.; Shishkov, V. Y. Surface-Enhanced Raman Scattering. In: Quantum Optics of Light Scattering. In Quantum Optics of Light Scattering, Springer Series in Optical Sciences; Springer, 2024.Google ScholarThere is no corresponding record for this reference.
- 34Kumar, N.; Mignuzzi, S.; Su, W.; Roy, D. Tip-enhanced Raman spectroscopy: principles and applications. EPJ Techniques Instrumentation 2015, 2 (1), 9, DOI: 10.1140/epjti/s40485-015-0019-5Google ScholarThere is no corresponding record for this reference.
- 35Wang, X.; Li, S.; Qu, H.; Hao, L.; Shao, T.; Wang, K.; Xia, Z.; Li, Z.; Li, Q. SERS-based immunomagnetic bead for rapid detection of H5N1 influenza virus. Influenza Other Respir Viruses 2023, 17 (3), e13114 DOI: 10.1111/irv.13114Google ScholarThere is no corresponding record for this reference.
- 36Park, H. J.; Yang, S. C.; Choo, J. Early Diagnosis of Influenza Virus A Using Surface-enhanced Raman Scattering-based Lateral Flow Assay. Bulletin Korean Chem. Society 2016, 37 (12), 2019– 2024, DOI: 10.1002/bkcs.11021Google ScholarThere is no corresponding record for this reference.
- 37Xiao, M.; Xie, K.; Dong, X.; Wang, L.; Huang, C.; Xu, F.; Xiao, W.; Jin, M.; Huang, B.; Tang, Y. Ultrasensitive detection of avian influenza A (H7N9) virus using surface-enhanced Raman scattering-based lateral flow immunoassay strips. Anal. Chim. Acta 2019, 1053, 139– 147, DOI: 10.1016/j.aca.2018.11.056Google ScholarThere is no corresponding record for this reference.
- 38Wang, C.; Wang, C.; Wang, X.; Wang, K.; Zhu, Y.; Rong, Z.; Wang, W.; Xiao, R.; Wang, S. Magnetic SERS Strip for Sensitive and Simultaneous Detection of Respiratory Viruses. ACS Appl. Mater. Interfaces 2019, 11 (21), 19495– 19505, DOI: 10.1021/acsami.9b03920Google ScholarThere is no corresponding record for this reference.
- 39Zhang, D.; Huang, L.; Liu, B.; Ge, Q.; Dong, J.; Zhao, X. Rapid and Ultrasensitive Quantification of Multiplex Respiratory Tract Infection Pathogen via Lateral Flow Microarray based on SERS Nanotags. Theranostics 2019, 9 (17), 4849– 4859, DOI: 10.7150/thno.35824Google ScholarThere is no corresponding record for this reference.
- 40Lim, J. Y.; Nam, J. S.; Yang, S. E.; Shin, H.; Jang, Y. H.; Bae, G. U.; Kang, T.; Lim, K. I.; Choi, Y. Identification of Newly Emerging Influenza Viruses by Surface-Enhanced Raman Spectroscopy. Anal. Chem. 2015, 87 (23), 11652– 11659, DOI: 10.1021/acs.analchem.5b02661Google ScholarThere is no corresponding record for this reference.
- 41Dardir, K.; Wang, H.; Martin, B. E.; Atzampou, M.; Brooke, C. B.; Fabris, L. SERS Nanoprobe for Intracellular Monitoring of Viral Mutations. J. Phys. Chem. C 2020, 124 (5), 3211– 3217, DOI: 10.1021/acs.jpcc.9b09253Google ScholarThere is no corresponding record for this reference.
- 42Sun, Y.; Xu, L.; Zhang, F.; Song, Z.; Hu, Y.; Ji, Y.; Shen, J.; Li, B.; Lu, H.; Yang, H. A promising magnetic SERS immunosensor for sensitive detection of avian influenza virus. Biosens. Bioelectron. 2017, 89 (Pt 2), 906– 912, DOI: 10.1016/j.bios.2016.09.100Google ScholarThere is no corresponding record for this reference.
- 43Kukushkin, V. I.; Ivanov, N. M.; Novoseltseva, A. A.; Gambaryan, A. S.; Yaminsky, I. V.; Kopylov, A. M.; Zavyalova, E. G. Highly sensitive detection of influenza virus with SERS aptasensor. PLoS One 2019, 14 (4), e0216247 DOI: 10.1371/journal.pone.0216247Google ScholarThere is no corresponding record for this reference.
- 44Negri, P.; Chen, G.; Kage, A.; Nitsche, A.; Naumann, D.; Xu, B.; Dluhy, R. A. Direct optical detection of viral nucleoprotein binding to an anti-influenza aptamer. Anal. Chem. 2012, 84 (13), 5501– 5508, DOI: 10.1021/ac202427eGoogle ScholarThere is no corresponding record for this reference.
- 45Zhang, Z.; Jiang, S.; Wang, X.; Dong, T.; Wang, Y.; Li, D.; Gao, X.; Qu, Z.; Li, Y. A novel enhanced substrate for label-free detection of SARS-CoV-2 based on surface-enhanced Raman scattering. Sens Actuators B Chem. 2022, 359, 131568 DOI: 10.1016/j.snb.2022.131568Google ScholarThere is no corresponding record for this reference.
- 46Tabarov, A.; Vitkin, V.; Andreeva, O.; Shemanaeva, A.; Popov, E.; Dobroslavin, A.; Kurikova, V.; Kuznetsova, O.; Grigorenko, K.; Tzibizov, I.; Detection of A and B Influenza Viruses by Surface-Enhanced Raman Scattering Spectroscopy and Machine Learning. Biosensors 2022, 12 (12). 1065 DOI: 10.3390/bios12121065 .Google ScholarThere is no corresponding record for this reference.
- 47Eom, G.; Hwang, A.; Kim, H.; Yang, S.; Lee, D. K.; Song, S.; Ha, K.; Jeong, J.; Jung, J.; Lim, E. K.; Kang, T. Diagnosis of Tamiflu-Resistant Influenza Virus in Human Nasal Fluid and Saliva Using Surface-Enhanced Raman Scattering. ACS Sens 2019, 4 (9), 2282– 2287, DOI: 10.1021/acssensors.9b00697Google ScholarThere is no corresponding record for this reference.
- 48Chen, H.; Das, A.; Bi, L.; Choi, N.; Moon, J.-I.; Wu, Y.; Park, S.; Choo, J. Recent advances in surface-enhanced Raman scattering-based microdevices for point-of-care diagnosis of viruses and bacteria. Nanoscale 2020, 12 (42), 21560– 21570, DOI: 10.1039/D0NR06340AGoogle ScholarThere is no corresponding record for this reference.
- 49Paria, D.; Kwok, K. S.; Raj, P.; Zheng, P.; Gracias, D. H.; Barman, I. Label-Free Spectroscopic SARS-CoV-2 Detection on Versatile Nanoimprinted Substrates. Nano Lett. 2022, 22 (9), 3620– 3627, DOI: 10.1021/acs.nanolett.1c04722Google ScholarThere is no corresponding record for this reference.
- 50Sebba, D.; Lastovich, A. G.; Kuroda, M.; Fallows, E.; Johnson, J.; Ahouidi, A. N.; Honko, A. N.; Fu, H.; Carruthers, E.; Diédhiou, C.; Ahmadou, B.; Soropogui, B.; Ruedas, J.; Peters, K.; Bartkowiak, M.; Mboup, S.; Ben Amor, Y.; Connor, J. H.; Weidemaier, K. A point-of-care diagnostic for differentiating Ebola from endemic febrile diseases. Science Translational Medicine. Sci. Transl. Med. 2018, 10, eaat0944 DOI: 10.1126/scitranslmed.aat0944Google ScholarThere is no corresponding record for this reference.
- 51Song, C.; Zhang, J.; Liu, Y.; Guo, X.; Guo, Y.; Jiang, X.; Wang, L. Highly sensitive SERS assay of DENV gene via a cascade signal amplification strategy of localized catalytic hairpin assembly and hybridization chain reaction. Sens Actuators B Chem. 2020, 325, 128970 DOI: 10.1016/j.snb.2020.128970Google ScholarThere is no corresponding record for this reference.
- 52Ngo, H. T.; Wang, H. N.; Fales, A. M.; Nicholson, B. P.; Woods, C. W.; Vo-Dinh, T. DNA bioassay-on-chip using SERS detection for dengue diagnosis. Analyst 2014, 139 (22), 5655– 5659, DOI: 10.1039/C4AN01077AGoogle ScholarThere is no corresponding record for this reference.
- 53Saleem, M.; Bilal, M.; Anwar, S.; Rehman, A.; Ahmed, M. Optical diagnosis of dengue virus infection in human blood serum using Raman spectroscopy. Laser Physics Letters 2013, 10 (3). 035602 DOI: 10.1088/1612-2011/10/3/035602 .Google ScholarThere is no corresponding record for this reference.
- 54Khan, S.; Ullah, R.; Saleem, M.; Bilal, M.; Rashid, R.; Khan, I.; Mahmood, A.; Nawaz, M. Raman spectroscopic analysis of dengue virus infection in human blood sera. Optik 2016, 127 (4), 2086– 2088, DOI: 10.1016/j.ijleo.2015.11.060Google ScholarThere is no corresponding record for this reference.
- 55Khan, S.; Ullah, R.; Khan, A.; Wahab, N.; Bilal, M.; Ahmed, M. Analysis of dengue infection based on Raman spectroscopy and support vector machine (SVM). Biomed Opt Express 2016, 7 (6), 2249– 2256, DOI: 10.1364/BOE.7.002249Google ScholarThere is no corresponding record for this reference.
- 56Khan, S.; Ullah, R.; Khan, A.; Sohail, A.; Wahab, N.; Bilal, M.; Ahmed, M. Random Forest-Based Evaluation of Raman Spectroscopy for Dengue Fever Analysis. Appl. Spectrosc. 2017, 71 (9), 2111– 2117, DOI: 10.1177/0003702817695571Google ScholarThere is no corresponding record for this reference.
- 57Mahmood, T.; Nawaz, H.; Ditta, A.; Majeed, M. I.; Hanif, M. A.; Rashid, N.; Bhatti, H. N.; Nargis, H. F.; Saleem, M.; Bonnier, F.; Byrne, H. Raman spectral analysis for rapid screening of dengue infection. Spectrochim Acta A Mol. Biomol. Spectrosc. 2018, 200, 136– 142, DOI: 10.1016/j.saa.2018.04.018Google ScholarThere is no corresponding record for this reference.
- 58Naseer, K.; Amin, A.; Saleem, M.; Qazi, J. Raman spectroscopy based differentiation of typhoid and dengue fever in infected human sera. Spectrochim Acta A Mol. Biomol. Spectrosc. 2019, 206, 197– 201, DOI: 10.1016/j.saa.2018.08.008Google ScholarThere is no corresponding record for this reference.
- 59Bilal, M.; Bilal, M.; Saleem, M.; Khurram, M.; Khan, S.; Ullah, R.; Ali, H.; Ahmed, M.; Shahzada, S.; Khan, E. U. Raman spectroscopy based investigation of molecular changes associated with an early stage of dengue virus infection. Laser Physics 2017, 27 (4). 045601 DOI: 10.1088/1555-6611/aa5a94 .Google ScholarThere is no corresponding record for this reference.
- 60Othman, N. H. L.; Khuan, Y.; Radzol, A. R. M.; Mansor, W.; Hisham, N. I. A. PCA-QDA Model Selection for Detecting NS1 Related Diseases from SERS Spectra of Salivary Mixtures. In World Congress on Medical Physics and Biomedical Engineering; Lhotska, L.; Sukupova, L.; Lacković, I.; Ibbott, G. S., Eds.; Springer, 2019.Google ScholarThere is no corresponding record for this reference.
- 61Radzol, A. R. M. L.; K Y; Mansor, W.; Ariffin, N. Biostatistical analysis of principal component of salivary Raman spectra for NS1 infection. IEEE EMBS Conference on Biomedical Engineering and Sciences 2016, 13– 18, DOI: 10.1109/IECBES.2016.7843406Google ScholarThere is no corresponding record for this reference.
- 62Desai, S.; Mishra, S. V.; Joshi, A.; Sarkar, D.; Hole, A.; Mishra, R.; Dutt, S.; Chilakapati, M. K.; Gupta, S.; Dutt, A. Raman spectroscopy-based detection of RNA viruses in saliva: A preliminary report. J. Biophotonics 2020, 13 (10), e202000189 DOI: 10.1002/jbio.202000189Google ScholarThere is no corresponding record for this reference.
- 63Sánchez-Purrà, M.; Carre-Camps, M.; de Puig, H.; Bosch, I.; Gehrke, L.; Hamad-Schifferli, K. Surface-Enhanced Raman Spectroscopy-Based Sandwich Immunoassays for Multiplexed Detection of Zika and Dengue Viral Biomarkers. ACS Infect. Dis. 2017, 3 (10), 767– 776, DOI: 10.1021/acsinfecdis.7b00110Google ScholarThere is no corresponding record for this reference.
- 64Camacho, S. A.; Sobral-Filho, R. G.; Aoki, P. H. B.; Constantino, C. J. L.; Brolo, A. G. Zika Immunoassay Based on Surface-Enhanced Raman Scattering Nanoprobes. ACS Sens 2018, 3 (3), 587– 594, DOI: 10.1021/acssensors.7b00639Google ScholarThere is no corresponding record for this reference.
- 65Tripathi, M. N.; Jangir, P.; Aakriti; Rai, S.; Gangwar, M.; Nath, G.; Saxena, P. S.; Srivastava, A. A novel approach for rapid and sensitive detection of Zika virus utilizing silver nanoislands as SERS platform. Spectrochim Acta A Mol. Biomol Spectrosc. 2023, 302, 123045 DOI: 10.1016/j.saa.2023.123045Google ScholarThere is no corresponding record for this reference.
- 66Silge, A.; Bocklitz, T.; Becker, B.; Matheis, W.; Popp, J.; Bekeredjian-Ding, I. Raman spectroscopy-based identification of toxoid vaccine products. NPJ Vaccines 2018, 3, 50, DOI: 10.1038/s41541-018-0088-yGoogle ScholarThere is no corresponding record for this reference.
- 67Global Poliovirus Containment Action Plan 2022–2024; World Health Organization, 2022.Google ScholarThere is no corresponding record for this reference.
- 68Alexander, T. A. Development of methodology based on commercialized SERS-active substrates for rapid discrimination of Poxviridae virions. Anal. Chem. 2008, 80 (8), 2817– 2825, DOI: 10.1021/ac702464wGoogle ScholarThere is no corresponding record for this reference.
- 69Alexander, T. A. Surface-enhanced Raman spectroscopy: A new approach to rapid identification of intact viruses. Spectroscopy 2008, 23 (7), 36, DOI: 10.1021/ac702455nGoogle ScholarThere is no corresponding record for this reference.
- 70Zhang, Z.; Jiang, H.; Jiang, S.; Dong, T.; Wang, X.; Wang, Y.; Li, Y. Rapid Detection of the Monkeypox Virus Genome and Antigen Proteins Based on Surface-Enhanced Raman Spectroscopy. ACS Appl. Mater. Interfaces 2023, 15 (29), 34419– 34426, DOI: 10.1021/acsami.3c04285Google ScholarThere is no corresponding record for this reference.
- 71Yu, Q.; Li, J.; Zheng, S.; Xia, X.; Xu, C.; Wang, C.; Wang, C.; Gu, B. Molybdenum disulfide-loaded multilayer AuNPs with colorimetric-SERS dual-signal enhancement activities for flexible immunochromatographic diagnosis of monkeypox virus. J. Hazard Mater. 2023, 459, 132136 DOI: 10.1016/j.jhazmat.2023.132136Google ScholarThere is no corresponding record for this reference.
- 72Lv, X.; Zhang, Z.; Zhao, Y.; Sun, X.; Jiang, H.; Zhang, S.; Sun, X.; Qiu, X.; Li, Y. Label-free detection of virus based on surface-enhanced Raman scattering. Spectrochim Acta A Mol. Biomol Spectrosc 2023, 302, 123087 DOI: 10.1016/j.saa.2023.123087Google ScholarThere is no corresponding record for this reference.
- 73Cornish, N. E.; Anderson, N. L.; Arambula, D. G.; Arduino, M. J.; Bryan, A.; Burton, N. C.; Chen, B.; Dickson, B. A.; Giri, J. G.; Griffith, N. K. Clinical Laboratory Biosafety Gaps: Lessons Learned from Past Outbreaks Reveal a Path to a Safer Future. Clin Microbiol Rev. 2021, 34 (3), e0012618 DOI: 10.1128/CMR.00126-18Google ScholarThere is no corresponding record for this reference.
- 74Mazzola, L. T.; Kelly-Cirino, C. Diagnostics for Lassa fever virus: a genetically diverse pathogen found in low-resource settings. BMJ Glob Health 2019, 4 (Suppl 2), e001116 DOI: 10.1136/bmjgh-2018-001116Google ScholarThere is no corresponding record for this reference.
- 75Filip, R.; Gheorghita Puscaselu, R.; Anchidin-Norocel, L.; Dimian, M.; Savage, W. K. Global Challenges to Public Health Care Systems during the COVID-19 Pandemic: A Review of Pandemic Measures and Problems. J. Pers Med. 2022, 12 (8). 1295 DOI: 10.3390/jpm12081295 .Google ScholarThere is no corresponding record for this reference.
- 76Goulart, A. C. C.; Silveira, L., Jr.; Carvalho, H. C.; Dorta, C. B.; Pacheco, M. T. T.; Zangaro, R. A. Diagnosing COVID-19 in human serum using Raman spectroscopy. Lasers Med. Sci. 2022, 37 (4), 2217– 2226, DOI: 10.1007/s10103-021-03488-7Google ScholarThere is no corresponding record for this reference.
- 77Awada, C.; Abdullah, M. M. B.; Traboulsi, H.; Dab, C.; Alshoaibi, A. SARS-CoV-2 Receptor Binding Domain as a Stable-Potential Target for SARS-CoV-2 Detection by Surface-Enhanced Raman Spectroscopy. Sensors 2021, 21 (13). 4617 DOI: 10.3390/s21134617 .Google ScholarThere is no corresponding record for this reference.
- 78Yin, G.; Li, L.; Lu, S.; Yin, Y.; Su, Y.; Zeng, Y.; Luo, M.; Ma, M.; Zhou, H.; Orlandini, L. An efficient primary screening of COVID-19 by serum Raman spectroscopy. J. Raman Spectrosc. 2021, 52 (5), 949– 958, DOI: 10.1002/jrs.6080Google ScholarThere is no corresponding record for this reference.
- 79Carlomagno, C.; Bertazioli, D.; Gualerzi, A.; Picciolini, S.; Banfi, P. I.; Lax, A.; Messina, E.; Navarro, J.; Bianchi, L.; Caronni, A. COVID-19 salivary Raman fingerprint: innovative approach for the detection of current and past SARS-CoV-2 infections. Sci. Rep. 2021, 11 (1), 4943 DOI: 10.1038/s41598-021-84565-3Google ScholarThere is no corresponding record for this reference.
- 80Ember, K.; Daoust, F.; Mahfoud, M.; Dallaire, F.; Ahmad, E. Z.; Tran, T.; Plante, A.; Diop, M. K.; Nguyen, T.; St-Georges-Robillard, A. Saliva-based detection of COVID-19 infection in a real-world setting using reagent-free Raman spectroscopy and machine learning. J. Biomed. Opt. 2022, 27 (2), 025002, DOI: 10.1117/1.JBO.27.2.025002Google ScholarThere is no corresponding record for this reference.
- 81Farhane, Z.; Bonnier, F.; Casey, A.; Maguire, A.; O’Neill, L.; Byrne, H. J. Cellular discrimination using in vitro Raman micro spectroscopy: the role of the nucleolus. Analyst 2015, 140 (17), 5908– 5919, DOI: 10.1039/C5AN01157DGoogle ScholarThere is no corresponding record for this reference.
- 82Goulart, A. C. C.; Zangaro, R. A.; Carvalho, H. C.; Lednev, I. K.; Silveira, L., Jr. Diagnosing COVID-19 in nasopharyngeal secretion through Raman spectroscopy: a feasibility study. Lasers Med. Sci. 2023, 38 (1), 210, DOI: 10.1007/s10103-023-03871-6Google ScholarThere is no corresponding record for this reference.
- 83Goulart, A. C. C.; Zangaro, R. A.; Carvalho, H. C.; Silveira, L., Jr. Diagnosing COVID-19 in human sera with detected immunoglobulins IgM and IgG by means of Raman spectroscopy. J. Raman Spectrosc. 2021, 52 (12), 2671– 2682, DOI: 10.1002/jrs.6235Google ScholarThere is no corresponding record for this reference.
- 84Guan, P. C.; Zhang, H.; Li, Z. Y.; Xu, S. S.; Sun, M.; Tian, X. M.; Ma, Z.; Lin, J. S.; Gu, M. M.; Wen, H. Rapid Point-of-Care Assay by SERS Detection of SARS-CoV-2 Virus and Its Variants. Anal. Chem. 2022, 94 (51), 17795– 17802, DOI: 10.1021/acs.analchem.2c03437Google ScholarThere is no corresponding record for this reference.
- 85Pezzotti, G.; Boschetto, F.; Ohgitani, E.; Fujita, Y.; Shin-Ya, M.; Adachi, T.; Yamamoto, T.; Kanamura, N.; Marin, E.; Zhu, W. Raman Molecular Fingerprints of SARS-CoV-2 British Variant and the Concept of Raman Barcode. Adv. Sci. 2022, 9 (3), e2103287 DOI: 10.1002/advs.202103287Google ScholarThere is no corresponding record for this reference.
- 86Rumaling, M. I.; Chee, F. P.; Bade, A.; Goh, L. P. W.; Juhim, F. Biofingerprint detection of corona virus using Raman spectroscopy: a novel approach. SN Appl. Sci. 2023, 5 (7), 197, DOI: 10.1007/s42452-023-05419-3Google ScholarThere is no corresponding record for this reference.
- 87Huang, J.; Wen, J.; Zhou, M.; Ni, S.; Le, W.; Chen, G.; Wei, L.; Zeng, Y.; Qi, D.; Pan, M. On-Site Detection of SARS-CoV-2 Antigen by Deep Learning-Based Surface-Enhanced Raman Spectroscopy and Its Biochemical Foundations. Anal. Chem. 2021, 93 (26), 9174– 9182, DOI: 10.1021/acs.analchem.1c01061Google ScholarThere is no corresponding record for this reference.
- 88Yang, Y.; Peng, Y.; Lin, C.; Long, L.; Hu, J.; He, J.; Zeng, H.; Huang, Z.; Li, Z. Y.; Tanemura, M. Human ACE2-Functionalized Gold ̀̀Virus-Trap″ Nanostructures for Accurate Capture of SARS-CoV-2 and Single-Virus SERS Detection. Nanomicro Lett. 2021, 13, 109, DOI: 10.1007/s40820-021-00620-8Google ScholarThere is no corresponding record for this reference.
- 89Yeh, Y. J.; Le, T. N.; Hsiao, W. W.; Tung, K. L.; Ostrikov, K. K.; Chiang, W. H. Plasmonic nanostructure-enhanced Raman scattering for detection of SARS-CoV-2 nucleocapsid protein and spike protein variants. Anal. Chim. Acta 2023, 1239, 340651 DOI: 10.1016/j.aca.2022.340651Google ScholarThere is no corresponding record for this reference.
- 90Garsuault, D.; El Messaoudi, S.; Prabakaran, M.; Cheong, I.; Boulanger, A.; Schmitt-Boulanger, M. Detection of several respiratory viruses with Surface-Enhanced Raman Spectroscopy coupled with Artificial Intelligence. Clinical Spectroscopy 2023, 5. 100025 DOI: 10.1016/j.clispe.2023.100025 .Google ScholarThere is no corresponding record for this reference.
- 91Khlebtsov, B.; Khlebtsov, N. Surface-Enhanced Raman Scattering-Based Lateral-Flow Immunoassay. Nanomaterials 2020, 10 (11). 2228 DOI: 10.3390/nano10112228 .Google ScholarThere is no corresponding record for this reference.
- 92Chen, H.; Park, S. G.; Choi, N.; Kwon, H. J.; Kang, T.; Lee, M. K.; Choo, J. Sensitive Detection of SARS-CoV-2 Using a SERS-Based Aptasensor. ACS Sens 2021, 6 (6), 2378– 2385, DOI: 10.1021/acssensors.1c00596Google ScholarThere is no corresponding record for this reference.
- 93Yang, Y.; Zhao, R.; Wang, Y.; Song, D.; Jiang, B.; Guo, X.; Liu, W.; Long, F.; Song, H.; Hao, R. Rapid and universal detection of SARS-CoV-2 and influenza A virus using a reusable dual-channel optic fiber immunosensor. J. Med. Virol. 2022, 94 (11), 5325– 5335, DOI: 10.1002/jmv.28015Google ScholarThere is no corresponding record for this reference.
- 94Chen, J.; Yu, Q.; Lu, M.; Jeon, C. S.; Pyun, S. H.; Choo, J. A strategy to enhance SERS detection sensitivity through the use of SiO(2) beads in a 1536-well plate. Anal Bioanal Chem. 2023, 415 (24), 5939– 5948, DOI: 10.1007/s00216-023-04896-0Google ScholarThere is no corresponding record for this reference.
- 95Mousavi, S. M.; Hashemi, S. A.; Rahmanian, V.; Kalashgrani, M. Y.; Gholami, A.; Omidifar, N.; Chiang, W. H. Highly Sensitive Flexible SERS-Based Sensing Platform for Detection of COVID-19. Biosensors 2022, 12 (7). 466 DOI: 10.3390/bios12070466 .Google ScholarThere is no corresponding record for this reference.
- 96Ebbah, E.; Amissah, A.; Kim, J. H.; Driskell, J. D. SERS-based immunoassay on a plasmonic syringe filter for improved sampling and labeling efficiency of biomarkers. Analyst 2023, 149 (1), 221– 230, DOI: 10.1039/D3AN01899GGoogle ScholarThere is no corresponding record for this reference.
- 97Salehi, H.; Ramoji, A.; Mougari, S.; Merida, P.; Neyret, A.; Popp, J.; Horvat, B.; Muriaux, D.; Cuisinier, F. Specific intracellular signature of SARS-CoV-2 infection using confocal Raman microscopy. Commun. Chem. 2022, 5 (1), 85, DOI: 10.1038/s42004-022-00702-7Google ScholarThere is no corresponding record for this reference.
- 98Peng, Y.; Lin, C.; Long, L.; Masaki, T.; Tang, M.; Yang, L.; Liu, J.; Huang, Z.; Li, Z.; Luo, X. Charge-Transfer Resonance and Electromagnetic Enhancement Synergistically Enabling MXenes with Excellent SERS Sensitivity for SARS-CoV-2 S Protein Detection. Nanomicro Lett. 2021, 13, 52, DOI: 10.1007/s40820-020-00565-4Google ScholarThere is no corresponding record for this reference.
- 99Sanchez, J. E.; Jaramillo, S. A.; Settles, E.; Velazquez Salazar, J. J.; Lehr, A.; Gonzalez, J.; Rodriguez Aranda, C.; Navarro-Contreras, H. R.; Raniere, M. O.; Harvey, M. Detection of SARS-CoV-2 and its S and N proteins using surface enhanced Raman spectroscopy. RSC Adv. 2021, 11 (41), 25788– 25794, DOI: 10.1039/D1RA03481BGoogle ScholarThere is no corresponding record for this reference.
- 100Song, K.; Xue, W.; Li, X.; Chang, Y.; Liu, M. Self-Assembly of Single-Virus SERS Hotspots for Highly Sensitive In Situ Detection of SARS-CoV-2 on Solid Surfaces. Anal. Chem. 2024, 96 (21), 8830– 8836, DOI: 10.1021/acs.analchem.4c01607Google ScholarThere is no corresponding record for this reference.
- 101Jadhav, S. A.; Biji, P.; Panthalingal, M. K.; Murali Krishna, C.; Rajkumar, S.; Joshi, D. S.; Sundaram, N. Development of integrated microfluidic platform coupled with Surface-enhanced Raman Spectroscopy for diagnosis of COVID-19. Med. Hypotheses 2021, 146, 110356 DOI: 10.1016/j.mehy.2020.110356Google ScholarThere is no corresponding record for this reference.
- 102Sitjar, J.; Xu, H. Z.; Liu, C. Y.; Wang, J. R.; Liao, J. D.; Tsai, H. P.; Lee, H.; Liu, B. H.; Chang, C. W. Synergistic surface-enhanced Raman scattering effect to distinguish live SARS-CoV-2 S pseudovirus. Anal. Chim. Acta 2022, 1193, 339406 DOI: 10.1016/j.aca.2021.339406Google ScholarThere is no corresponding record for this reference.
- 103Sitjar, J.; Liao, J. D.; Lee, H.; Tsai, H. P.; Wang, J. R.; Chen, C. H.; Wang, H.; Liu, B. H. Detection of live SARS-CoV-2 virus and its variants by specially designed SERS-active substrates and spectroscopic analyses. Anal. Chim. Acta 2023, 1256, 341151 DOI: 10.1016/j.aca.2023.341151Google ScholarThere is no corresponding record for this reference.
- 104Zhou, L.; Vestri, A.; Marchesano, V.; Rippa, M.; Sagnelli, D.; Picazio, G.; Fusco, G.; Han, J.; Zhou, J.; Petti, L. The Label-Free Detection and Identification of SARS-CoV-2 Using Surface-Enhanced Raman Spectroscopy and Principal Component Analysis. Biosensors 2023, 13 (12). 1014 DOI: 10.3390/bios13121014 .Google ScholarThere is no corresponding record for this reference.
- 105Akdeniz, M.; Al-Shaebi, Z.; Altunbek, M.; Bayraktar, C.; Kayabolen, A.; Bagci-Onder, T.; Aydin, O. Characterization and discrimination of spike protein in SARS-CoV-2 virus-like particles via surface-enhanced Raman spectroscopy. Biotechnol. J. 2024, 19 (1), e2300191 DOI: 10.1002/biot.202300191Google ScholarThere is no corresponding record for this reference.
- 106Robertson, J. L.; Senger, R. S.; Talty, J.; Du, P.; Sayed-Issa, A.; Avellar, M. L.; Ngo, L. T.; Gomez De La Espriella, M.; Fazili, T. N.; Jackson-Akers, J. Y. Alterations in the molecular composition of COVID-19 patient urine, detected using Raman spectroscopic/computational analysis. PLoS One 2022, 17 (7), e0270914 DOI: 10.1371/journal.pone.0270914Google ScholarThere is no corresponding record for this reference.
- 107Pahlow, S.; Richard-Lacroix, M.; Hornung, F.; Kose-Vogel, N.; Mayerhofer, T. G.; Hniopek, J.; Ryabchykov, O.; Bocklitz, T.; Weber, K.; Ehricht, R.; Simple, Fast and Convenient Magnetic Bead-Based Sample Preparation for Detecting Viruses via Raman-Spectroscopy. Biosensors 2023, 13 (6). 594 DOI: 10.3390/bios13060594 .Google ScholarThere is no corresponding record for this reference.
- 108Zhang, D.; Zhang, X.; Ma, R.; Deng, S.; Wang, X.; Wang, X.; Zhang, X.; Huang, X.; Liu, Y.; Li, G. Ultra-fast and onsite interrogation of Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2) in waters via surface enhanced Raman scattering (SERS). Water Res. 2021, 200, 117243 DOI: 10.1016/j.watres.2021.117243Google ScholarThere is no corresponding record for this reference.
- 109Kim, K.; Kashefi-Kheyrabadi, L.; Joung, Y.; Kim, K.; Dang, H.; Chavan, S. G.; Lee, M. H.; Choo, J. Recent advances in sensitive surface-enhanced Raman scattering-based lateral flow assay platforms for point-of-care diagnostics of infectious diseases. Sens Actuators B Chem. 2021, 329, 129214 DOI: 10.1016/j.snb.2020.129214Google ScholarThere is no corresponding record for this reference.
- 110Mo, W.; Wen, J.; Huang, J.; Yang, Y.; Zhou, M.; Ni, S.; Le, W.; Wei, L.; Qi, D.; Wang, S. Classification of Coronavirus Spike Proteins by Deep-Learning-Based Raman Spectroscopy and its Interpretative Analysis. J. Appl. Spectrosc. 2023, 89 (6), 1203– 1211, DOI: 10.1007/s10812-023-01487-wGoogle ScholarThere is no corresponding record for this reference.
- 111Lu, X.; Liu, Q.; Benavides-Montano, J. A.; Nicola, A. V.; Aston, D. E.; Rasco, B. A.; Aguilar, H. C. Detection of receptor-induced glycoprotein conformational changes on enveloped virions by using confocal micro-Raman spectroscopy. J. Virol 2013, 87 (6), 3130– 3142, DOI: 10.1128/JVI.03220-12Google ScholarThere is no corresponding record for this reference.
- 112Neng, J.; Li, Y.; Driscoll, A. J.; Wilson, W. C.; Johnson, P. A. Detection of Multiple Pathogens in Serum Using Silica-Encapsulated Nanotags in a Surface-Enhanced Raman Scattering-Based Immunoassay. J. Agric. Food Chem. 2018, 66 (22), 5707– 5712, DOI: 10.1021/acs.jafc.8b00026Google ScholarThere is no corresponding record for this reference.
- 113Zhang, H.; Harpster, M. H.; Wilson, W. C.; Johnson, P. A. Surface-enhanced Raman scattering detection of DNAs derived from virus genomes using Au-coated paramagnetic nanoparticles. Langmuir 2012, 28 (8), 4030– 4037, DOI: 10.1021/la204890tGoogle ScholarThere is no corresponding record for this reference.
- 114Neng, J.; Harpster, M. H.; Wilson, W. C.; Johnson, P. A. Surface-enhanced Raman scattering (SERS) detection of multiple viral antigens using magnetic capture of SERS-active nanoparticles. Biosens. Bioelectron. 2013, 41, 316– 321, DOI: 10.1016/j.bios.2012.08.048Google ScholarThere is no corresponding record for this reference.
- 115Zong, C.; Xu, M.; Xu, L. J.; Wei, T.; Ma, X.; Zheng, X. S.; Hu, R.; Ren, B. Surface-Enhanced Raman Spectroscopy for Bioanalysis: Reliability and Challenges. Chem. Rev. 2018, 118 (10), 4946– 4980, DOI: 10.1021/acs.chemrev.7b00668Google ScholarThere is no corresponding record for this reference.
- 116Kharbach, M.; Alaoui Mansouri, M.; Taabouz, M.; Yu, H. Current Application of Advancing Spectroscopy Techniques in Food Analysis: Data Handling with Chemometric Approaches. Foods 2023, 12 (14). 2753 DOI: 10.3390/foods12142753 .Google ScholarThere is no corresponding record for this reference.
- 117Marzi, J.; Fuhrmann, E.; Brauchle, E.; Singer, V.; Pfannstiel, J.; Schmidt, I.; Hartmann, H. Non-Invasive Three-Dimensional Cell Analysis in Bioinks by Raman Imaging. ACS Appl. Mater. Interfaces 2022, 14 (27), 30455– 30465, DOI: 10.1021/acsami.1c24463Google ScholarThere is no corresponding record for this reference.
- 118Eberhardt, K. S.; Matthäus, C.; Schmitt, C.; Popp, M. J. Advantages and limitations of Raman spectroscopy for molecular diagnostics: An update. Expert Rev. Mol. Diagn. 2015, 15 (6), 773– 787, DOI: 10.1586/14737159.2015.1036744Google ScholarThere is no corresponding record for this reference.
- 119Storozhuk, D.; Ryabchykov, O.; Popp, J.; Bocklitz, T. RAMANMETRIX: A Delightful Way to Analyze Raman Spectra , 2022, arXiv:2201.07586. arXiv.org e-Print archive https://arxiv.org/abs/2201.07586.Google ScholarThere is no corresponding record for this reference.
- 120Ramanmetrix. RAMANMETRIX Software Documentation , 2021.Google ScholarThere is no corresponding record for this reference.
- 121Zu, Y.; Chang, H.; Cui, Z. Molecular point-of-care testing technologies: Current status and challenges. Nexus 2025, 2 (2), 100059 DOI: 10.1016/j.ynexs.2025.100059Google ScholarThere is no corresponding record for this reference.
- 122Food, U. S.; Drug, A. Overview of Device Regulation Device Advice: Comprehensive Regulatory Assistance , 2025.Google ScholarThere is no corresponding record for this reference.
- 123Regulation (EU) 2017/746 of the European Parliament and of the Council of 5 April 2017 on in vitro diagnostic medical devices and repealing Directive 98/79/EC and Commission Decision 2010/227/EU . 2010, Vol. L 117 176 332.Google ScholarThere is no corresponding record for this reference.
- 124Food, U. S.; Drug, A. Artificial Intelligence in Software as a Medical Device. Software as a Medical Device (SaMD)─Device Advice: Comprehensive Regulatory Assistance . 2025, (acccessed 2025/08/21).Google ScholarThere is no corresponding record for this reference.
- 125Bottini, M.; Ryu, S. J.; Terander, A. E.; Voglis, S.; Maldaner, N.; Bellut, D.; Regli, L.; Serra, C.; Staartjes, V. E. The Ever-Evolving Regulatory Landscape Concerning Development and Clinical Application of Machine Intelligence: Practical Consequences for Spine Artificial Intelligence Research. Neurospine 2025, 22 (1), 134– 143, DOI: 10.14245/ns.2449186.593Google ScholarThere is no corresponding record for this reference.
- 126Artika, I. M.; Ma’roef, C. N. Laboratory biosafety for handling emerging viruses. Asian Pac. J. Trop. Biomed. 2017, 7 (5), 483– 491, DOI: 10.1016/j.apjtb.2017.01.020Google ScholarThere is no corresponding record for this reference.
- 127Dolskiy, A. A.; Grishchenko, I. V.; Yudkin, D. V. Cell Cultures for Virology: Usability, Advantages, and Prospects. Int. J. Mol. Sci. 2020, 21 (21), 7978, DOI: 10.3390/ijms21217978Google ScholarThere is no corresponding record for this reference.
- 128Selo, M. A.; Sake, J. A.; Kim, K. J.; Ehrhardt, C. In vitro and ex vivo models in inhalation biopharmaceutical research - advances, challenges and future perspectives. Adv. Drug Delivery Rev. 2021, 177, 113862 DOI: 10.1016/j.addr.2021.113862Google ScholarThere is no corresponding record for this reference.
- 129Colby, L. A.; Quenee, L. E.; Zitzow, L. A. Considerations for infectious disease research studies using animals. In Comparative Medicine 2017; Vol. 67, pp 222– 231.Google ScholarThere is no corresponding record for this reference.
- 130Klingstrom, T.; Bongcam-Rudloff, E.; Reichel, J. Legal & ethical compliance when sharing biospecimen. Brief Funct. Genomics 2018, 17 (1), 1– 7, DOI: 10.1093/bfgp/elx008Google ScholarThere is no corresponding record for this reference.
- 131Srivastava, S.; Sharma, D.; Kumar, S.; Sharma, A.; Rijal, R.; Asija, A.; Adhikari, S.; Rustagi, S.; Sah, S.; Al-Qaim, Z. H. Emergence of Marburg virus: a global perspective on fatal outbreaks and clinical challenges. Front. Microbiol. 2023, 14, 1239079 DOI: 10.3389/fmicb.2023.1239079Google ScholarThere is no corresponding record for this reference.
- 132Vitti, J. N.; Vitti, R.; Chu, K.; Mellis, S. The ethics of clinical research in the era of COVID-19. Front Public Health 2024, 12, 1359654 DOI: 10.3389/fpubh.2024.1359654Google ScholarThere is no corresponding record for this reference.
- 133Selgelid, M. J. Gain-of-Function Research: Ethical Analysis. Sci. Eng. Ethics 2016, 22 (4), 923– 964, DOI: 10.1007/s11948-016-9810-1Google ScholarThere is no corresponding record for this reference.
- 134Belser, J. A.; Kieran, T. J.; Mitchell, Z. A.; Sun, X.; Mayfield, K.; Tumpey, T. M.; Spengler, J. R.; Maines, T. R. Key considerations to improve the normalization, interpretation and reproducibility of morbidity data in mammalian models of viral disease. Dis. Model Mech. 2024, 17 (3), dmm050511, DOI: 10.1242/dmm.050511Google ScholarThere is no corresponding record for this reference.
- 135Masopust, D.; Sivula, C. P.; Jameson, S. C. Of Mice, Dirty Mice, and Men: Using Mice To Understand Human Immunology. J. Immunol. 2017, 199 (2), 383– 388, DOI: 10.4049/jimmunol.1700453Google ScholarThere is no corresponding record for this reference.
- 136Zabidi, N. Z.; Liew, H. L.; Farouk, I. A.; Puniyamurti, A.; Yip, A. J. W.; Wijesinghe, V. N.; Low, Z. Y.; Tang, J. W.; Chow, V. T. K.; Lal, S. K. Evolution of SARS-CoV-2 Variants: Implications on Immune Escape, Vaccination, Therapeutic and Diagnostic Strategies. Viruses 2023, 15 (4). 944 DOI: 10.3390/v15040944 .Google ScholarThere is no corresponding record for this reference.
- 137Hartman, K. A.; Clayton, N.; Thomas, G. J., Jr. Studies of Virus Structure by Raman Spectroscopy. I. R17 Virus and R17 RNA. Biochem. Biophys. Res. Commun. 1973, 50 (3), 942– 949, DOI: 10.1016/0006-291X(73)91336-3Google ScholarThere is no corresponding record for this reference.
- 138Thomas, G. J., Jr.; Prescott, B.; McDonald-Ordzie, P. E.; Hartman, K. A. Studies of Virus Structure by Laser Raman Spectroscopy. II. MS2 Phage, MS2 Capsids, and MS2 RNA in Aqueous Solutions. J. Mol. Biol. 1976, 102 (1), 103– 124, DOI: 10.1016/0022-2836(76)90076-0Google ScholarThere is no corresponding record for this reference.
- 139Thomas, G. J.; Prescott, B.; Day, L. A. Structure Similarity, Difference, and Variability in the Filamentous Viruses fd, If1, IKe, Pf1, and Xf Investigated by Laser Raman Spectroscopy. J. Mol. Biol. 1983, 165 (2), 321– 356, DOI: 10.1016/S0022-2836(83)80260-5Google ScholarThere is no corresponding record for this reference.
- 140Wen, Z.-Q.; Thomas, G. J., Jr. UV Resonance Raman Spectroscopy of DNA and Protein Constituents of Viruses: Assignments and Cross Sections for Excitations at 257, 244, 238, and 229 nm. Biopolymers 1998, 45 (3), 247– 256, DOI: 10.1002/(SICI)1097-0282(199803)45:3Google ScholarThere is no corresponding record for this reference.
- 141Moor, K.; Terada, Y.; Taketani, A.; Matsuyoshi, H.; Ohtani, K.; Sato, H. Early Detection of Virus Infection in Live Human Cells Using Raman Spectroscopy. J. Biomed. Opt. 2018, 23 (9), 097001 DOI: 10.1117/1.JBO.23.9.097001Google ScholarThere is no corresponding record for this reference.
- 142El-Mashtoly, S. F.; Gerwert, K. Diagnostics and Therapy Assessment Using Label-Free Raman Imaging. Anal. Chem. 2022, 94 (1), 120– 142, DOI: 10.1021/acs.analchem.1c04483Google ScholarThere is no corresponding record for this reference.
- 143Marotta, N. E. Patterned Nanoarray SERS Substrates for Pathogen Detection. Ph.D. Thesis. Georgia Institute of Technology: Atlanta, GA, 2010.Google ScholarThere is no corresponding record for this reference.
- 144Sigle, M.; Rohlfing, K.; Kenny, M.; Scheuermann, S.; Sun, N.; Graeßner, U.; Haug, V.; Sudmann, J.; Seitz, C. M.; Heinzmann, D.; Schenke-Layland, K.; Maguire, P. B.; Walch, A.; Marzi, J.; Gawaz, M. P. Translating Genomic Tools to Raman Spectroscopy Analysis Enables High-Dimensional Tissue Characterization at Molecular Resolution. Nat. Commun. 2023, 14, 5799 DOI: 10.1038/s41467-023-41417-0Google ScholarThere is no corresponding record for this reference.
- 145Gahlaut, S. K.; Savargaonkar, D.; Sharan, C.; Yadav, S.; Mishra, P.; Singh, J. P. SERS Platform for Dengue Diagnosis from Clinical Samples Employing a Hand Held Raman Spectrometer. Anal. Chem. 2020, 92 (3), 2527– 2534, DOI: 10.1021/acs.analchem.9b04129Google ScholarThere is no corresponding record for this reference.
- 146Gordon, J.; Mousavi, S. D.; Li, J.; Filippone, N.; Walter, L.; Cheng, H.-W.; Skeete, Z.; Feldman, H.; Hakimi, S.; Cappuccio, K.; Wang, S.; Hader, M.; Shang, G.; Turner, J.; Cameron, A.; Bane, S.; Poliks, M.; Lu, S.; Yuan, T. T.; Zhong, C.-J. Plasmonic Nanoprobe-Enabled SERS Detection of SARS-CoV-2 Proteins and Virus Samples on Wax-Printed Paper Substrates. Anal. Chem. 2025, 97 (46), 21303– 21313, DOI: 10.1021/acs.analchem.5c02355Google ScholarThere is no corresponding record for this reference.
- 147Plavec, Z.; Domanska, A.; Liu, X.; Laine, P.; Paulin, L.; Varjosalo, M.; Auvinen, P.; Wolf, S. G.; Anastasina, M.; Butcher, S. J. SARS-CoV-2 Production, Purification Methods and UV Inactivation for Proteomics and Structural Studies. Viruses 2022, 14 (9), 1989, DOI: 10.3390/v14091989Google ScholarThere is no corresponding record for this reference.
- 148Pestov, D.; Murawski, R. K.; Ariunbold, G. O.; Wang, X.; Zhi, M.; Sokolov, A. V.; Sautenkov, V. A.; Rostovtsev, Y. V.; Dogariu, A.; Huang, Y.; Scully, M. O. Optimizing the laser-pulse configuration for coherent Raman spectroscopy. Science. 2007, 316 (5822), 265– 268, DOI: 10.1126/science.1139055Google ScholarThere is no corresponding record for this reference.
- 149Wang, Y.; Ruan, Q.; Lei, Z. C.; Lin, S. C.; Zhu, Z.; Zhou, L.; Yang, C. Highly Sensitive and Automated Surface Enhanced Raman Scattering-based Immunoassay for H5N1 Detection with Digital Microfluidics. Anal. Chem. 2018, 90, 5224– 5231, DOI: 10.1021/acs.analchem.8b00002Google ScholarThere is no corresponding record for this reference.
- 150Chang, Y.; Wang, Y.; Li, W.; Wei, Z.; Tang, S.; Chen, R. Mechanisms, Techniques and Devices of Airborne Virus Detection: A Review. Int. J. Environ. Res. Public Health 2023, 20, 5471, DOI: 10.3390/ijerph20085471Google ScholarThere is no corresponding record for this reference.
Cited By
This article has not yet been cited by other publications.
Article Views
Altmetric
Citations
Article Views are the COUNTER-compliant sum of full text article downloads since November 2008 (both PDF and HTML) across all institutions and individuals. These metrics are regularly updated to reflect usage leading up to the last few days.
Citations are the number of other articles citing this article, calculated by Crossref and updated daily. Find more information about Crossref citation counts.
The Altmetric Attention Score is a quantitative measure of the attention that a research article has received online. Clicking on the donut icon will load a page at altmetric.com with additional details about the score and the social media presence for the given article. Find more information on the Altmetric Attention Score and how the score is calculated.
Recommended Articles
Abstract

Figure 1
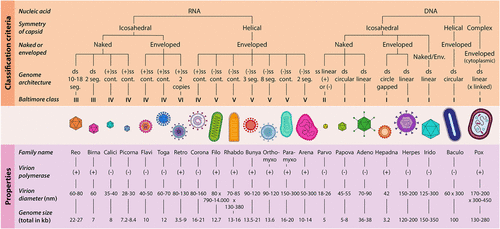
Figure 1. Overview of virion classification and their properties (Ramoji, A., Pahlow, S., Pistiki, A., Rueger, J., Shaik, T. A., Shen, H., Wichmann, C., Krafft, C.,& Popp, J. (2022). Understanding viruses and viral infections by biophotonic methods. Translational Biophotonics, 4(1–2), e202100008. 10.1002/tbio.202100008). Reprinted from ref (2).
Figure 2
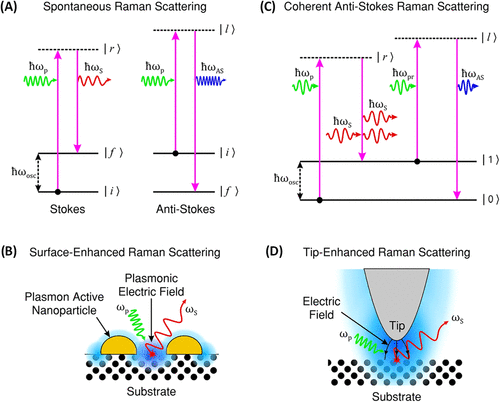
Figure 2. (A) Illustration of Stokes and anti-Stokes Raman scattering, showing energy transitions during spontaneous inelastic light scattering. (Anti-Stokes scattering is shown for completeness of the Raman process.) (B) SERS mechanism, where localized surface plasmons on metallic nanostructures enhance the EM and Raman signal. (C) Schematic of CARS, highlighting the nonlinear four-wave mixing process generating anti-Stokes emission (D) TERS, utilizing a plasmonic metallic tip to confine and enhance the EM field at the nanoscale for high-resolution Raman imaging. (Jones, R.R., Hooper, DC. Zhang, L. et al. Raman Techniques: Fundamentals and Frontiers. Nanoscale Res Lett 14, 231 (2019). 10.1186/s11671-019-3039-2). Adapted from ref (22).
Figure 3

Figure 3. (A) Mean SERS spectra of NS1-positive dengue samples, NS1-negative febrile controls, and healthy individuals, highlighting distinct biochemical signatures. (B) PCA score plot showing clear separation of the three groups based on SERS spectral features. (C) PC1 loading plot indicating Raman bands contributing most strongly to classification. (Gahlaut, S. K.; Savargaonkar, D.; Sharan, C.; Yadav, Sarjana, Mishra, P.; Singh, J. P. SERS Platform for Dengue Diagnosis from Clinical Samples Employing a Hand Held Raman Spectrometer. Anal. Chem.2020, 92 (3), 2527–2534. 10.1021/acs.analchem.9b04129) Adapted from ref (145).
Figure 4

Figure 4. Multiplexed SERS assay where Zika and Dengue test lines are separately painted. SERS spectra are shown for Zika (A) and Dengue (B) test lines and a combined Zika-Dengue test line (C). (Sánchez-Purrà M, Carré-Camps M, de Puig H, Bosch I, Gehrke L, Hamad-Schifferli K. (2017) Surface-Enhanced Raman Spectroscopy-Based Sandwich Immunoassays for Multiplexed Detection of Zika and Dengue Viral Biomarkers. ACS Infect Dis.;3(10):767–776. doi: 10.1021/acsinfecdis.7b00110). Adapted from ref (63).
Figure 5

Figure 5. Development of a Raman spectral database for the SARS-CoV-2 spike protein. (a) Schematic overview illustrating the structural organization of the spectral database. (b) Standard Raman spectra obtained for the full-length spike (S) protein, its subunits S1 and S2, and RBD. (c) Comparative analysis between computed and experimentally recorded Raman spectra for the S1 subunit. (d) Optical image of the S protein dried on a silver-coated silicon substrate; scale bar: 300 μm. (e) Scanning electron microscopy image depicting the Au nanoparticle array used for SERS measurements; scale bar: 200 nm. (f) Comparative Raman profiles of the spike proteins from SARS-CoV-2, SARS-CoV, and MERS-CoV, expressed in insect cells using the baculovirus system. (g) Raman spectral differences in SARS-CoV-2 S protein before and after thermal treatment. (h) SERS spectra of the SARS-CoV-2 spike protein. (Huang, J., Wen, J., Zhou, M., Ni, S., Le, W., Chen, G., Wei, L., Zeng, Y., Qi, D., Pan, M., Xu, J., Wu, Y., Li, Z., Feng, Y., Zhao, Z., He, Z., Li, B., Zhao, S., Zhang, B., Xue, P., He, S., Fang, K., Zhao, Y.,& Du, K. (2021). On-site detection of SARS-CoV-2 antigen by deep learning-based surface-enhanced Raman spectroscopy and its biochemical foundations. Analytical Chemistry, 93(26), 9174–9182.) (Adapted from Analytical Chemistry, 2021, 93, 9174–9182). Reprinted from ref (87).
Figure 6

Figure 6. (A) PCA of confocal Raman spectra extracted from intracellular compartments of Control, measles virus (MeV)-infected, and SARS-CoV-2-infected Vero E6 cells. (A, B) Cytoplasm region: (A) 3D PCA score plot and (B) corresponding PC loadings. (C, D) Golgi-mitochondria region: (C) PCA score plot and (D) PC loadings. (E, F) Nucleus region: (E) PCA score plot and (F) PC loadings. (Salehi, H.; Ramoji, A.; Mougari, S.; Merida, P.; Neyret, A.; Popp, J.; Horvat, B.; Muriaux, D.; Cuisinier, F. Specific intracellular signature of SARS-CoV-2 infection using confocal Raman microscopy. Commun Chem2022, 5 (1), 85. DOI: 10.1038/s42004–022–00702–7.) Reprinted from ref (97).
Figure 7

Figure 7. (A) Representative SERS spectra of SARS-CoV-2, human coronavirus OC43 (HCoV-OC43), SARS-CoV-2 Omicron variant BA.5.1, and CCoV, acquired on AuNP films under 785 nm excitation. Major diagnostic bands are labeled, and the background trace corresponds to virus-free control samples. (B) Three-dimensional PCA score plot showing clear clustering and separation of the four coronavirus types based on their SERS spectral signatures. (C) Corresponding PCA loading vectors for PC1, PC2, and PC3, highlighting the Raman bands contributing most strongly to virus discrimination. (Zhou, L.; Vestri, A.; Marchesano, V.; Rippa, M.; Sagnelli, D.; Picazio, G.; Fusco, G.; Han, J.; Zhou, J.; Petti, L. The Label-Free Detection and Identification of SARS-CoV-2 Using Surface-Enhanced Raman Spectroscopy and Principal Component Analysis. Biosensors (Basel) 2023, 13 (12). DOI: 10.3390/bios13121014) Adapted from ref (104).
References
This article references 150 other publications.
- 1Neumann, G.; Kawaoka, Y. Which Virus Will Cause the Next Pandemic?. Viruses 2023, 15 (1), 199, DOI: 10.3390/v15010199There is no corresponding record for this reference.
- 2Ramoji, A.; Pahlow, S.; Pistiki, A.; Rueger, J.; Shaik, T. A.; Shen, H.; Wichmann, C.; Krafft, C.; Popp, J. Understanding viruses and viral infections by biophotonic methods. Translational Biophotonics 2022, 4 (1–2), e202100008, DOI: 10.1002/tbio.202100008There is no corresponding record for this reference.
- 3Liu, D.; Pan, L.; Zhai, H.; Qiu, H. J.; Sun, Y. Virus tracking technologies and their applications in viral life cycle: research advances and future perspectives. Front. Immunol. 2023, 14, 1204730 DOI: 10.3389/fimmu.2023.1204730There is no corresponding record for this reference.
- 4Maginnis, M. S. Virus-Receptor Interactions: The Key to Cellular Invasion. J. Mol. Biol. 2018, 430 (17), 2590– 2611, DOI: 10.1016/j.jmb.2018.06.024There is no corresponding record for this reference.
- 5Jones, J. E.; Le Sage, V.; Lakdawala, S. S. Viral and host heterogeneity and their effects on the viral life cycle. Nat. Rev. Microbiol. 2021, 19 (4), 272– 282, DOI: 10.1038/s41579-020-00449-9There is no corresponding record for this reference.
- 6Idrees, S.; Chen, H.; Panth, N.; Paudel, K. R.; Hansbro, P. M. Exploring Viral-Host Protein Interactions as Antiviral Therapies: A Computational Perspective. Microorganisms 2024, 12 (3). 630 DOI: 10.3390/microorganisms12030630 .There is no corresponding record for this reference.
- 7Datta, S.; Hett, E. C.; Vora, K. A.; Hazuda, D. J.; Oslund, R. C.; Fadeyi, O. O.; Emili, A. The chemical biology of coronavirus host-cell interactions. RSC Chem. Biol. 2021, 2 (1), 30– 46, DOI: 10.1039/D0CB00197JThere is no corresponding record for this reference.
- 8Merenich, D. K. E. V. M.-B. C. J. K. A. M. Advanced microscopy techniques for the visualization and analysis of cell behaviors. In Cell Movement in Health and Disease; Michael Schnoor, L.-M. Y.; Sean, X. Sun., Eds.; Academic Press, 2022.There is no corresponding record for this reference.
- 9Kianzad, S.; SeyedAlinaghi, S.; Asadollahi-Amin, A.; Dadras, O.; Karimi, A.; Afsahi, A. M.; MohsseniPour, M.; Barzegary, A.; Mirzapour, P.; Mirghaderi, S. P. Comparison of SARS-CoV-2 (Coronavirus) with other Similar Viruses Based on Current Evidence J. Iranian Med. Council 2022 DOI: 10.18502/jimc.v5i1.9565 .There is no corresponding record for this reference.
- 10Islam, M. T.; Quispe, C.; Herrera-Bravo, J.; Sarkar, C.; Sharma, R.; Garg, N.; Fredes, L. I.; Martorell, M.; Alshehri, M. M.; Sharifi-Rad, J. Production, Transmission, Pathogenesis, and Control of Dengue Virus: A Literature-Based Undivided Perspective. Biomed. Res. Int. 2021, 2021, 4224816 DOI: 10.1155/2021/4224816There is no corresponding record for this reference.
- 11Ruokola, P.; Dadu, E.; Kazmertsuk, A.; Hakkanen, H.; Marjomaki, V.; Ihalainen, J. A. Raman spectroscopic signatures of echovirus 1 uncoating. J. Virol. 2014, 88 (15), 8504– 8513, DOI: 10.1128/JVI.03398-13There is no corresponding record for this reference.
- 12Organization, W. H. WHO reports CDC , 2014https://www.bbc.com/news/world-africa-29769782.There is no corresponding record for this reference.
- 13Goldsmith, C. S.; Miller, S. E. Modern uses of electron microscopy for detection of viruses. Clin. Microbiol Rev. 2009, 22 (4), 552– 563, DOI: 10.1128/CMR.00027-09There is no corresponding record for this reference.
- 14Schaack, G. A.; Mehle, A. Experimental Approaches to Identify Host Factors Important for Influenza Virus. Cold Spring Harb Perspect Med. 2020, 10 (12), a038521, DOI: 10.1101/cshperspect.a038521There is no corresponding record for this reference.
- 15Jansz, N.; Faulkner, G. J. Viral genome sequencing methods: benefits and pitfalls of current approaches. Biochem. Soc. Trans. 2024, 52 (3), 1431– 1447, DOI: 10.1042/BST20231322There is no corresponding record for this reference.
- 16Mahmud, I.; Garrett, T. J. Mass Spectrometry Techniques in Emerging Pathogens Studies: COVID-19 Perspectives. J. Am. Soc. Mass Spectrom. 2020, 31 (10), 2013– 2024, DOI: 10.1021/jasms.0c00238There is no corresponding record for this reference.
- 17Garg, T.; Weiss, C. R.; Sheth, R. A. Techniques for Profiling the Cellular Immune Response and Their Implications for Interventional Oncology. Cancers 2022, 14 (15). 3628 DOI: 10.3390/cancers14153628 .There is no corresponding record for this reference.
- 18Oumarou Hama, H.; Aboudharam, G.; Barbieri, R.; Lepidi, H.; Drancourt, M. Immunohistochemical diagnosis of human infectious diseases: a review. Diagn. Pathol. 2022, 17 (1), 17, DOI: 10.1186/s13000-022-01197-5There is no corresponding record for this reference.
- 19Iwanami, S.; Kitagawa, K.; Ohashi, H.; Asai, Y.; Shionoya, K.; Saso, W.; Nishioka, K.; Inaba, H.; Nakaoka, S.; Wakita, T. Should a viral genome stay in the host cell or leave? A quantitative dynamics study of how hepatitis C virus deals with this dilemma. PLoS Biol. 2020, 18 (7), e3000562 DOI: 10.1371/journal.pbio.3000562There is no corresponding record for this reference.
- 20Ferrarini, M. G.; Lal, A.; Rebollo, R.; Gruber, A. J.; Guarracino, A.; Gonzalez, I. M.; Floyd, T.; de Oliveira, D. S.; Shanklin, J.; Beausoleil, E. Genome-wide bioinformatic analyses predict key host and viral factors in SARS-CoV-2 pathogenesis. Commun. Biol. 2021, 4 (1), 590, DOI: 10.1038/s42003-021-02095-0There is no corresponding record for this reference.
- 21Ashmore-Harris, C.; Iafrate, M.; Saleem, A.; Fruhwirth, G. O. Non-invasive Reporter Gene Imaging of Cell Therapies, including T Cells and Stem Cells. Mol. Ther. 2020, 28 (6), 1392– 1416, DOI: 10.1016/j.ymthe.2020.03.016There is no corresponding record for this reference.
- 22Jones, R. R.; Hooper, D. C.; Zhang, L.; Wolverson, D.; Valev, V. K. Raman Techniques: Fundamentals and Frontiers. Nanoscale Res. Lett. 2019, 14 (1), 231, DOI: 10.1186/s11671-019-3039-2There is no corresponding record for this reference.
- 23Fernández-Galiana, Á.; Bibikova, O.; Vilms Pedersen, S.; Stevens, M. M. Fundamentals and Applications of Raman-Based Techniques for the Design and Development of Active Biomedical Materials. Adv. Mater. 2024, 36 (43), e2210807 DOI: 10.1002/adma.202210807There is no corresponding record for this reference.
- 24Cialla-May, D.; Krafft, C.; Rosch, P.; Deckert-Gaudig, T.; Frosch, T.; Jahn, I. J.; Pahlow, S.; Stiebing, C.; Meyer-Zedler, T.; Bocklitz, T. Raman Spectroscopy and Imaging in Bioanalytics. Anal. Chem. 2022, 94 (1), 86– 119, DOI: 10.1021/acs.analchem.1c03235There is no corresponding record for this reference.
- 25Deckert, V.; Deckert-Gaudig, T.; Cialla-May, D.; Popp, J.; Zell, R.; Deinhard-Emmer, S.; Sokolov, A. V.; Yi, Z.; Scully, M. O. Laser spectroscopic technique for direct identification of a single virus I: FASTER CARS. Proc. Natl. Acad. Sci. U. S. A. 2020, 117 (45), 27820– 27824, DOI: 10.1073/pnas.2013169117There is no corresponding record for this reference.
- 26Xu, D.; Liang, B.; Xu, Y.; Liu, M. Recent advances in tip-enhanced Raman spectroscopy probe designs. Nano Res. 2023, 16 (4), 5555– 5571, DOI: 10.1007/s12274-022-5220-7There is no corresponding record for this reference.
- 27Höppener, C.; Aizpurua, J.; Chen, H.; Gräfe, S.; Jorio, A.; Kupfer, S.; Zhang, Z.; Deckert, V. Tip-enhanced Raman scattering. Nature Reviews Methods Primers 2024, 4 (1), 47, DOI: 10.1038/s43586-024-00323-5There is no corresponding record for this reference.
- 28Tian, Z. Y. M.; VB; ; Zhang, X.; Khalil, M.; Ayejoto, D. A. Coherent Anti-Stokes Raman Scattering. In Advanced Diagnostics in Combustion Science; Springer, 2023.There is no corresponding record for this reference.
- 29Zhang, C.; Zhang, D.; Cheng, J. X. Coherent Raman Scattering Microscopy in Biology and Medicine. Annu. Rev. Biomed Eng. 2015, 17, 415– 445, DOI: 10.1146/annurev-bioeng-071114-040554There is no corresponding record for this reference.
- 30Cheng, Q.; Miao, Y.; Wild, J.; Min, W.; Yang, Y. Emerging applications of stimulated Raman scattering microscopy in materials science. Matter 2021, 4 (5), 1460– 1483, DOI: 10.1016/j.matt.2021.02.013There is no corresponding record for this reference.
- 31Orlando, A.; Franceschini, F.; Muscas, C.; Pidkova, S.; Bartoli, M.; Rovere, M.; Tagliaferro, A. A Comprehensive Review on Raman Spectroscopy Applications. Chemosensors 2021, 9 (9). 262 DOI: 10.3390/chemosensors9090262 .There is no corresponding record for this reference.
- 32Lukose, J.; Barik, A. K.; Mithun, N.; Sanoop Pavithran, M.; George, S. D.; Murukeshan, V. M.; Chidangil, S. Raman spectroscopy for viral diagnostics. Biophys Rev. 2023, 15 (2), 199– 221, DOI: 10.1007/s12551-023-01059-4There is no corresponding record for this reference.
- 33Lisyansky, A. A. A.; E S; ; Vinogradov, A. P.; Shishkov, V. Y. Surface-Enhanced Raman Scattering. In: Quantum Optics of Light Scattering. In Quantum Optics of Light Scattering, Springer Series in Optical Sciences; Springer, 2024.There is no corresponding record for this reference.
- 34Kumar, N.; Mignuzzi, S.; Su, W.; Roy, D. Tip-enhanced Raman spectroscopy: principles and applications. EPJ Techniques Instrumentation 2015, 2 (1), 9, DOI: 10.1140/epjti/s40485-015-0019-5There is no corresponding record for this reference.
- 35Wang, X.; Li, S.; Qu, H.; Hao, L.; Shao, T.; Wang, K.; Xia, Z.; Li, Z.; Li, Q. SERS-based immunomagnetic bead for rapid detection of H5N1 influenza virus. Influenza Other Respir Viruses 2023, 17 (3), e13114 DOI: 10.1111/irv.13114There is no corresponding record for this reference.
- 36Park, H. J.; Yang, S. C.; Choo, J. Early Diagnosis of Influenza Virus A Using Surface-enhanced Raman Scattering-based Lateral Flow Assay. Bulletin Korean Chem. Society 2016, 37 (12), 2019– 2024, DOI: 10.1002/bkcs.11021There is no corresponding record for this reference.
- 37Xiao, M.; Xie, K.; Dong, X.; Wang, L.; Huang, C.; Xu, F.; Xiao, W.; Jin, M.; Huang, B.; Tang, Y. Ultrasensitive detection of avian influenza A (H7N9) virus using surface-enhanced Raman scattering-based lateral flow immunoassay strips. Anal. Chim. Acta 2019, 1053, 139– 147, DOI: 10.1016/j.aca.2018.11.056There is no corresponding record for this reference.
- 38Wang, C.; Wang, C.; Wang, X.; Wang, K.; Zhu, Y.; Rong, Z.; Wang, W.; Xiao, R.; Wang, S. Magnetic SERS Strip for Sensitive and Simultaneous Detection of Respiratory Viruses. ACS Appl. Mater. Interfaces 2019, 11 (21), 19495– 19505, DOI: 10.1021/acsami.9b03920There is no corresponding record for this reference.
- 39Zhang, D.; Huang, L.; Liu, B.; Ge, Q.; Dong, J.; Zhao, X. Rapid and Ultrasensitive Quantification of Multiplex Respiratory Tract Infection Pathogen via Lateral Flow Microarray based on SERS Nanotags. Theranostics 2019, 9 (17), 4849– 4859, DOI: 10.7150/thno.35824There is no corresponding record for this reference.
- 40Lim, J. Y.; Nam, J. S.; Yang, S. E.; Shin, H.; Jang, Y. H.; Bae, G. U.; Kang, T.; Lim, K. I.; Choi, Y. Identification of Newly Emerging Influenza Viruses by Surface-Enhanced Raman Spectroscopy. Anal. Chem. 2015, 87 (23), 11652– 11659, DOI: 10.1021/acs.analchem.5b02661There is no corresponding record for this reference.
- 41Dardir, K.; Wang, H.; Martin, B. E.; Atzampou, M.; Brooke, C. B.; Fabris, L. SERS Nanoprobe for Intracellular Monitoring of Viral Mutations. J. Phys. Chem. C 2020, 124 (5), 3211– 3217, DOI: 10.1021/acs.jpcc.9b09253There is no corresponding record for this reference.
- 42Sun, Y.; Xu, L.; Zhang, F.; Song, Z.; Hu, Y.; Ji, Y.; Shen, J.; Li, B.; Lu, H.; Yang, H. A promising magnetic SERS immunosensor for sensitive detection of avian influenza virus. Biosens. Bioelectron. 2017, 89 (Pt 2), 906– 912, DOI: 10.1016/j.bios.2016.09.100There is no corresponding record for this reference.
- 43Kukushkin, V. I.; Ivanov, N. M.; Novoseltseva, A. A.; Gambaryan, A. S.; Yaminsky, I. V.; Kopylov, A. M.; Zavyalova, E. G. Highly sensitive detection of influenza virus with SERS aptasensor. PLoS One 2019, 14 (4), e0216247 DOI: 10.1371/journal.pone.0216247There is no corresponding record for this reference.
- 44Negri, P.; Chen, G.; Kage, A.; Nitsche, A.; Naumann, D.; Xu, B.; Dluhy, R. A. Direct optical detection of viral nucleoprotein binding to an anti-influenza aptamer. Anal. Chem. 2012, 84 (13), 5501– 5508, DOI: 10.1021/ac202427eThere is no corresponding record for this reference.
- 45Zhang, Z.; Jiang, S.; Wang, X.; Dong, T.; Wang, Y.; Li, D.; Gao, X.; Qu, Z.; Li, Y. A novel enhanced substrate for label-free detection of SARS-CoV-2 based on surface-enhanced Raman scattering. Sens Actuators B Chem. 2022, 359, 131568 DOI: 10.1016/j.snb.2022.131568There is no corresponding record for this reference.
- 46Tabarov, A.; Vitkin, V.; Andreeva, O.; Shemanaeva, A.; Popov, E.; Dobroslavin, A.; Kurikova, V.; Kuznetsova, O.; Grigorenko, K.; Tzibizov, I.; Detection of A and B Influenza Viruses by Surface-Enhanced Raman Scattering Spectroscopy and Machine Learning. Biosensors 2022, 12 (12). 1065 DOI: 10.3390/bios12121065 .There is no corresponding record for this reference.
- 47Eom, G.; Hwang, A.; Kim, H.; Yang, S.; Lee, D. K.; Song, S.; Ha, K.; Jeong, J.; Jung, J.; Lim, E. K.; Kang, T. Diagnosis of Tamiflu-Resistant Influenza Virus in Human Nasal Fluid and Saliva Using Surface-Enhanced Raman Scattering. ACS Sens 2019, 4 (9), 2282– 2287, DOI: 10.1021/acssensors.9b00697There is no corresponding record for this reference.
- 48Chen, H.; Das, A.; Bi, L.; Choi, N.; Moon, J.-I.; Wu, Y.; Park, S.; Choo, J. Recent advances in surface-enhanced Raman scattering-based microdevices for point-of-care diagnosis of viruses and bacteria. Nanoscale 2020, 12 (42), 21560– 21570, DOI: 10.1039/D0NR06340AThere is no corresponding record for this reference.
- 49Paria, D.; Kwok, K. S.; Raj, P.; Zheng, P.; Gracias, D. H.; Barman, I. Label-Free Spectroscopic SARS-CoV-2 Detection on Versatile Nanoimprinted Substrates. Nano Lett. 2022, 22 (9), 3620– 3627, DOI: 10.1021/acs.nanolett.1c04722There is no corresponding record for this reference.
- 50Sebba, D.; Lastovich, A. G.; Kuroda, M.; Fallows, E.; Johnson, J.; Ahouidi, A. N.; Honko, A. N.; Fu, H.; Carruthers, E.; Diédhiou, C.; Ahmadou, B.; Soropogui, B.; Ruedas, J.; Peters, K.; Bartkowiak, M.; Mboup, S.; Ben Amor, Y.; Connor, J. H.; Weidemaier, K. A point-of-care diagnostic for differentiating Ebola from endemic febrile diseases. Science Translational Medicine. Sci. Transl. Med. 2018, 10, eaat0944 DOI: 10.1126/scitranslmed.aat0944There is no corresponding record for this reference.
- 51Song, C.; Zhang, J.; Liu, Y.; Guo, X.; Guo, Y.; Jiang, X.; Wang, L. Highly sensitive SERS assay of DENV gene via a cascade signal amplification strategy of localized catalytic hairpin assembly and hybridization chain reaction. Sens Actuators B Chem. 2020, 325, 128970 DOI: 10.1016/j.snb.2020.128970There is no corresponding record for this reference.
- 52Ngo, H. T.; Wang, H. N.; Fales, A. M.; Nicholson, B. P.; Woods, C. W.; Vo-Dinh, T. DNA bioassay-on-chip using SERS detection for dengue diagnosis. Analyst 2014, 139 (22), 5655– 5659, DOI: 10.1039/C4AN01077AThere is no corresponding record for this reference.
- 53Saleem, M.; Bilal, M.; Anwar, S.; Rehman, A.; Ahmed, M. Optical diagnosis of dengue virus infection in human blood serum using Raman spectroscopy. Laser Physics Letters 2013, 10 (3). 035602 DOI: 10.1088/1612-2011/10/3/035602 .There is no corresponding record for this reference.
- 54Khan, S.; Ullah, R.; Saleem, M.; Bilal, M.; Rashid, R.; Khan, I.; Mahmood, A.; Nawaz, M. Raman spectroscopic analysis of dengue virus infection in human blood sera. Optik 2016, 127 (4), 2086– 2088, DOI: 10.1016/j.ijleo.2015.11.060There is no corresponding record for this reference.
- 55Khan, S.; Ullah, R.; Khan, A.; Wahab, N.; Bilal, M.; Ahmed, M. Analysis of dengue infection based on Raman spectroscopy and support vector machine (SVM). Biomed Opt Express 2016, 7 (6), 2249– 2256, DOI: 10.1364/BOE.7.002249There is no corresponding record for this reference.
- 56Khan, S.; Ullah, R.; Khan, A.; Sohail, A.; Wahab, N.; Bilal, M.; Ahmed, M. Random Forest-Based Evaluation of Raman Spectroscopy for Dengue Fever Analysis. Appl. Spectrosc. 2017, 71 (9), 2111– 2117, DOI: 10.1177/0003702817695571There is no corresponding record for this reference.
- 57Mahmood, T.; Nawaz, H.; Ditta, A.; Majeed, M. I.; Hanif, M. A.; Rashid, N.; Bhatti, H. N.; Nargis, H. F.; Saleem, M.; Bonnier, F.; Byrne, H. Raman spectral analysis for rapid screening of dengue infection. Spectrochim Acta A Mol. Biomol. Spectrosc. 2018, 200, 136– 142, DOI: 10.1016/j.saa.2018.04.018There is no corresponding record for this reference.
- 58Naseer, K.; Amin, A.; Saleem, M.; Qazi, J. Raman spectroscopy based differentiation of typhoid and dengue fever in infected human sera. Spectrochim Acta A Mol. Biomol. Spectrosc. 2019, 206, 197– 201, DOI: 10.1016/j.saa.2018.08.008There is no corresponding record for this reference.
- 59Bilal, M.; Bilal, M.; Saleem, M.; Khurram, M.; Khan, S.; Ullah, R.; Ali, H.; Ahmed, M.; Shahzada, S.; Khan, E. U. Raman spectroscopy based investigation of molecular changes associated with an early stage of dengue virus infection. Laser Physics 2017, 27 (4). 045601 DOI: 10.1088/1555-6611/aa5a94 .There is no corresponding record for this reference.
- 60Othman, N. H. L.; Khuan, Y.; Radzol, A. R. M.; Mansor, W.; Hisham, N. I. A. PCA-QDA Model Selection for Detecting NS1 Related Diseases from SERS Spectra of Salivary Mixtures. In World Congress on Medical Physics and Biomedical Engineering; Lhotska, L.; Sukupova, L.; Lacković, I.; Ibbott, G. S., Eds.; Springer, 2019.There is no corresponding record for this reference.
- 61Radzol, A. R. M. L.; K Y; Mansor, W.; Ariffin, N. Biostatistical analysis of principal component of salivary Raman spectra for NS1 infection. IEEE EMBS Conference on Biomedical Engineering and Sciences 2016, 13– 18, DOI: 10.1109/IECBES.2016.7843406There is no corresponding record for this reference.
- 62Desai, S.; Mishra, S. V.; Joshi, A.; Sarkar, D.; Hole, A.; Mishra, R.; Dutt, S.; Chilakapati, M. K.; Gupta, S.; Dutt, A. Raman spectroscopy-based detection of RNA viruses in saliva: A preliminary report. J. Biophotonics 2020, 13 (10), e202000189 DOI: 10.1002/jbio.202000189There is no corresponding record for this reference.
- 63Sánchez-Purrà, M.; Carre-Camps, M.; de Puig, H.; Bosch, I.; Gehrke, L.; Hamad-Schifferli, K. Surface-Enhanced Raman Spectroscopy-Based Sandwich Immunoassays for Multiplexed Detection of Zika and Dengue Viral Biomarkers. ACS Infect. Dis. 2017, 3 (10), 767– 776, DOI: 10.1021/acsinfecdis.7b00110There is no corresponding record for this reference.
- 64Camacho, S. A.; Sobral-Filho, R. G.; Aoki, P. H. B.; Constantino, C. J. L.; Brolo, A. G. Zika Immunoassay Based on Surface-Enhanced Raman Scattering Nanoprobes. ACS Sens 2018, 3 (3), 587– 594, DOI: 10.1021/acssensors.7b00639There is no corresponding record for this reference.
- 65Tripathi, M. N.; Jangir, P.; Aakriti; Rai, S.; Gangwar, M.; Nath, G.; Saxena, P. S.; Srivastava, A. A novel approach for rapid and sensitive detection of Zika virus utilizing silver nanoislands as SERS platform. Spectrochim Acta A Mol. Biomol Spectrosc. 2023, 302, 123045 DOI: 10.1016/j.saa.2023.123045There is no corresponding record for this reference.
- 66Silge, A.; Bocklitz, T.; Becker, B.; Matheis, W.; Popp, J.; Bekeredjian-Ding, I. Raman spectroscopy-based identification of toxoid vaccine products. NPJ Vaccines 2018, 3, 50, DOI: 10.1038/s41541-018-0088-yThere is no corresponding record for this reference.
- 67Global Poliovirus Containment Action Plan 2022–2024; World Health Organization, 2022.There is no corresponding record for this reference.
- 68Alexander, T. A. Development of methodology based on commercialized SERS-active substrates for rapid discrimination of Poxviridae virions. Anal. Chem. 2008, 80 (8), 2817– 2825, DOI: 10.1021/ac702464wThere is no corresponding record for this reference.
- 69Alexander, T. A. Surface-enhanced Raman spectroscopy: A new approach to rapid identification of intact viruses. Spectroscopy 2008, 23 (7), 36, DOI: 10.1021/ac702455nThere is no corresponding record for this reference.
- 70Zhang, Z.; Jiang, H.; Jiang, S.; Dong, T.; Wang, X.; Wang, Y.; Li, Y. Rapid Detection of the Monkeypox Virus Genome and Antigen Proteins Based on Surface-Enhanced Raman Spectroscopy. ACS Appl. Mater. Interfaces 2023, 15 (29), 34419– 34426, DOI: 10.1021/acsami.3c04285There is no corresponding record for this reference.
- 71Yu, Q.; Li, J.; Zheng, S.; Xia, X.; Xu, C.; Wang, C.; Wang, C.; Gu, B. Molybdenum disulfide-loaded multilayer AuNPs with colorimetric-SERS dual-signal enhancement activities for flexible immunochromatographic diagnosis of monkeypox virus. J. Hazard Mater. 2023, 459, 132136 DOI: 10.1016/j.jhazmat.2023.132136There is no corresponding record for this reference.
- 72Lv, X.; Zhang, Z.; Zhao, Y.; Sun, X.; Jiang, H.; Zhang, S.; Sun, X.; Qiu, X.; Li, Y. Label-free detection of virus based on surface-enhanced Raman scattering. Spectrochim Acta A Mol. Biomol Spectrosc 2023, 302, 123087 DOI: 10.1016/j.saa.2023.123087There is no corresponding record for this reference.
- 73Cornish, N. E.; Anderson, N. L.; Arambula, D. G.; Arduino, M. J.; Bryan, A.; Burton, N. C.; Chen, B.; Dickson, B. A.; Giri, J. G.; Griffith, N. K. Clinical Laboratory Biosafety Gaps: Lessons Learned from Past Outbreaks Reveal a Path to a Safer Future. Clin Microbiol Rev. 2021, 34 (3), e0012618 DOI: 10.1128/CMR.00126-18There is no corresponding record for this reference.
- 74Mazzola, L. T.; Kelly-Cirino, C. Diagnostics for Lassa fever virus: a genetically diverse pathogen found in low-resource settings. BMJ Glob Health 2019, 4 (Suppl 2), e001116 DOI: 10.1136/bmjgh-2018-001116There is no corresponding record for this reference.
- 75Filip, R.; Gheorghita Puscaselu, R.; Anchidin-Norocel, L.; Dimian, M.; Savage, W. K. Global Challenges to Public Health Care Systems during the COVID-19 Pandemic: A Review of Pandemic Measures and Problems. J. Pers Med. 2022, 12 (8). 1295 DOI: 10.3390/jpm12081295 .There is no corresponding record for this reference.
- 76Goulart, A. C. C.; Silveira, L., Jr.; Carvalho, H. C.; Dorta, C. B.; Pacheco, M. T. T.; Zangaro, R. A. Diagnosing COVID-19 in human serum using Raman spectroscopy. Lasers Med. Sci. 2022, 37 (4), 2217– 2226, DOI: 10.1007/s10103-021-03488-7There is no corresponding record for this reference.
- 77Awada, C.; Abdullah, M. M. B.; Traboulsi, H.; Dab, C.; Alshoaibi, A. SARS-CoV-2 Receptor Binding Domain as a Stable-Potential Target for SARS-CoV-2 Detection by Surface-Enhanced Raman Spectroscopy. Sensors 2021, 21 (13). 4617 DOI: 10.3390/s21134617 .There is no corresponding record for this reference.
- 78Yin, G.; Li, L.; Lu, S.; Yin, Y.; Su, Y.; Zeng, Y.; Luo, M.; Ma, M.; Zhou, H.; Orlandini, L. An efficient primary screening of COVID-19 by serum Raman spectroscopy. J. Raman Spectrosc. 2021, 52 (5), 949– 958, DOI: 10.1002/jrs.6080There is no corresponding record for this reference.
- 79Carlomagno, C.; Bertazioli, D.; Gualerzi, A.; Picciolini, S.; Banfi, P. I.; Lax, A.; Messina, E.; Navarro, J.; Bianchi, L.; Caronni, A. COVID-19 salivary Raman fingerprint: innovative approach for the detection of current and past SARS-CoV-2 infections. Sci. Rep. 2021, 11 (1), 4943 DOI: 10.1038/s41598-021-84565-3There is no corresponding record for this reference.
- 80Ember, K.; Daoust, F.; Mahfoud, M.; Dallaire, F.; Ahmad, E. Z.; Tran, T.; Plante, A.; Diop, M. K.; Nguyen, T.; St-Georges-Robillard, A. Saliva-based detection of COVID-19 infection in a real-world setting using reagent-free Raman spectroscopy and machine learning. J. Biomed. Opt. 2022, 27 (2), 025002, DOI: 10.1117/1.JBO.27.2.025002There is no corresponding record for this reference.
- 81Farhane, Z.; Bonnier, F.; Casey, A.; Maguire, A.; O’Neill, L.; Byrne, H. J. Cellular discrimination using in vitro Raman micro spectroscopy: the role of the nucleolus. Analyst 2015, 140 (17), 5908– 5919, DOI: 10.1039/C5AN01157DThere is no corresponding record for this reference.
- 82Goulart, A. C. C.; Zangaro, R. A.; Carvalho, H. C.; Lednev, I. K.; Silveira, L., Jr. Diagnosing COVID-19 in nasopharyngeal secretion through Raman spectroscopy: a feasibility study. Lasers Med. Sci. 2023, 38 (1), 210, DOI: 10.1007/s10103-023-03871-6There is no corresponding record for this reference.
- 83Goulart, A. C. C.; Zangaro, R. A.; Carvalho, H. C.; Silveira, L., Jr. Diagnosing COVID-19 in human sera with detected immunoglobulins IgM and IgG by means of Raman spectroscopy. J. Raman Spectrosc. 2021, 52 (12), 2671– 2682, DOI: 10.1002/jrs.6235There is no corresponding record for this reference.
- 84Guan, P. C.; Zhang, H.; Li, Z. Y.; Xu, S. S.; Sun, M.; Tian, X. M.; Ma, Z.; Lin, J. S.; Gu, M. M.; Wen, H. Rapid Point-of-Care Assay by SERS Detection of SARS-CoV-2 Virus and Its Variants. Anal. Chem. 2022, 94 (51), 17795– 17802, DOI: 10.1021/acs.analchem.2c03437There is no corresponding record for this reference.
- 85Pezzotti, G.; Boschetto, F.; Ohgitani, E.; Fujita, Y.; Shin-Ya, M.; Adachi, T.; Yamamoto, T.; Kanamura, N.; Marin, E.; Zhu, W. Raman Molecular Fingerprints of SARS-CoV-2 British Variant and the Concept of Raman Barcode. Adv. Sci. 2022, 9 (3), e2103287 DOI: 10.1002/advs.202103287There is no corresponding record for this reference.
- 86Rumaling, M. I.; Chee, F. P.; Bade, A.; Goh, L. P. W.; Juhim, F. Biofingerprint detection of corona virus using Raman spectroscopy: a novel approach. SN Appl. Sci. 2023, 5 (7), 197, DOI: 10.1007/s42452-023-05419-3There is no corresponding record for this reference.
- 87Huang, J.; Wen, J.; Zhou, M.; Ni, S.; Le, W.; Chen, G.; Wei, L.; Zeng, Y.; Qi, D.; Pan, M. On-Site Detection of SARS-CoV-2 Antigen by Deep Learning-Based Surface-Enhanced Raman Spectroscopy and Its Biochemical Foundations. Anal. Chem. 2021, 93 (26), 9174– 9182, DOI: 10.1021/acs.analchem.1c01061There is no corresponding record for this reference.
- 88Yang, Y.; Peng, Y.; Lin, C.; Long, L.; Hu, J.; He, J.; Zeng, H.; Huang, Z.; Li, Z. Y.; Tanemura, M. Human ACE2-Functionalized Gold ̀̀Virus-Trap″ Nanostructures for Accurate Capture of SARS-CoV-2 and Single-Virus SERS Detection. Nanomicro Lett. 2021, 13, 109, DOI: 10.1007/s40820-021-00620-8There is no corresponding record for this reference.
- 89Yeh, Y. J.; Le, T. N.; Hsiao, W. W.; Tung, K. L.; Ostrikov, K. K.; Chiang, W. H. Plasmonic nanostructure-enhanced Raman scattering for detection of SARS-CoV-2 nucleocapsid protein and spike protein variants. Anal. Chim. Acta 2023, 1239, 340651 DOI: 10.1016/j.aca.2022.340651There is no corresponding record for this reference.
- 90Garsuault, D.; El Messaoudi, S.; Prabakaran, M.; Cheong, I.; Boulanger, A.; Schmitt-Boulanger, M. Detection of several respiratory viruses with Surface-Enhanced Raman Spectroscopy coupled with Artificial Intelligence. Clinical Spectroscopy 2023, 5. 100025 DOI: 10.1016/j.clispe.2023.100025 .There is no corresponding record for this reference.
- 91Khlebtsov, B.; Khlebtsov, N. Surface-Enhanced Raman Scattering-Based Lateral-Flow Immunoassay. Nanomaterials 2020, 10 (11). 2228 DOI: 10.3390/nano10112228 .There is no corresponding record for this reference.
- 92Chen, H.; Park, S. G.; Choi, N.; Kwon, H. J.; Kang, T.; Lee, M. K.; Choo, J. Sensitive Detection of SARS-CoV-2 Using a SERS-Based Aptasensor. ACS Sens 2021, 6 (6), 2378– 2385, DOI: 10.1021/acssensors.1c00596There is no corresponding record for this reference.
- 93Yang, Y.; Zhao, R.; Wang, Y.; Song, D.; Jiang, B.; Guo, X.; Liu, W.; Long, F.; Song, H.; Hao, R. Rapid and universal detection of SARS-CoV-2 and influenza A virus using a reusable dual-channel optic fiber immunosensor. J. Med. Virol. 2022, 94 (11), 5325– 5335, DOI: 10.1002/jmv.28015There is no corresponding record for this reference.
- 94Chen, J.; Yu, Q.; Lu, M.; Jeon, C. S.; Pyun, S. H.; Choo, J. A strategy to enhance SERS detection sensitivity through the use of SiO(2) beads in a 1536-well plate. Anal Bioanal Chem. 2023, 415 (24), 5939– 5948, DOI: 10.1007/s00216-023-04896-0There is no corresponding record for this reference.
- 95Mousavi, S. M.; Hashemi, S. A.; Rahmanian, V.; Kalashgrani, M. Y.; Gholami, A.; Omidifar, N.; Chiang, W. H. Highly Sensitive Flexible SERS-Based Sensing Platform for Detection of COVID-19. Biosensors 2022, 12 (7). 466 DOI: 10.3390/bios12070466 .There is no corresponding record for this reference.
- 96Ebbah, E.; Amissah, A.; Kim, J. H.; Driskell, J. D. SERS-based immunoassay on a plasmonic syringe filter for improved sampling and labeling efficiency of biomarkers. Analyst 2023, 149 (1), 221– 230, DOI: 10.1039/D3AN01899GThere is no corresponding record for this reference.
- 97Salehi, H.; Ramoji, A.; Mougari, S.; Merida, P.; Neyret, A.; Popp, J.; Horvat, B.; Muriaux, D.; Cuisinier, F. Specific intracellular signature of SARS-CoV-2 infection using confocal Raman microscopy. Commun. Chem. 2022, 5 (1), 85, DOI: 10.1038/s42004-022-00702-7There is no corresponding record for this reference.
- 98Peng, Y.; Lin, C.; Long, L.; Masaki, T.; Tang, M.; Yang, L.; Liu, J.; Huang, Z.; Li, Z.; Luo, X. Charge-Transfer Resonance and Electromagnetic Enhancement Synergistically Enabling MXenes with Excellent SERS Sensitivity for SARS-CoV-2 S Protein Detection. Nanomicro Lett. 2021, 13, 52, DOI: 10.1007/s40820-020-00565-4There is no corresponding record for this reference.
- 99Sanchez, J. E.; Jaramillo, S. A.; Settles, E.; Velazquez Salazar, J. J.; Lehr, A.; Gonzalez, J.; Rodriguez Aranda, C.; Navarro-Contreras, H. R.; Raniere, M. O.; Harvey, M. Detection of SARS-CoV-2 and its S and N proteins using surface enhanced Raman spectroscopy. RSC Adv. 2021, 11 (41), 25788– 25794, DOI: 10.1039/D1RA03481BThere is no corresponding record for this reference.
- 100Song, K.; Xue, W.; Li, X.; Chang, Y.; Liu, M. Self-Assembly of Single-Virus SERS Hotspots for Highly Sensitive In Situ Detection of SARS-CoV-2 on Solid Surfaces. Anal. Chem. 2024, 96 (21), 8830– 8836, DOI: 10.1021/acs.analchem.4c01607There is no corresponding record for this reference.
- 101Jadhav, S. A.; Biji, P.; Panthalingal, M. K.; Murali Krishna, C.; Rajkumar, S.; Joshi, D. S.; Sundaram, N. Development of integrated microfluidic platform coupled with Surface-enhanced Raman Spectroscopy for diagnosis of COVID-19. Med. Hypotheses 2021, 146, 110356 DOI: 10.1016/j.mehy.2020.110356There is no corresponding record for this reference.
- 102Sitjar, J.; Xu, H. Z.; Liu, C. Y.; Wang, J. R.; Liao, J. D.; Tsai, H. P.; Lee, H.; Liu, B. H.; Chang, C. W. Synergistic surface-enhanced Raman scattering effect to distinguish live SARS-CoV-2 S pseudovirus. Anal. Chim. Acta 2022, 1193, 339406 DOI: 10.1016/j.aca.2021.339406There is no corresponding record for this reference.
- 103Sitjar, J.; Liao, J. D.; Lee, H.; Tsai, H. P.; Wang, J. R.; Chen, C. H.; Wang, H.; Liu, B. H. Detection of live SARS-CoV-2 virus and its variants by specially designed SERS-active substrates and spectroscopic analyses. Anal. Chim. Acta 2023, 1256, 341151 DOI: 10.1016/j.aca.2023.341151There is no corresponding record for this reference.
- 104Zhou, L.; Vestri, A.; Marchesano, V.; Rippa, M.; Sagnelli, D.; Picazio, G.; Fusco, G.; Han, J.; Zhou, J.; Petti, L. The Label-Free Detection and Identification of SARS-CoV-2 Using Surface-Enhanced Raman Spectroscopy and Principal Component Analysis. Biosensors 2023, 13 (12). 1014 DOI: 10.3390/bios13121014 .There is no corresponding record for this reference.
- 105Akdeniz, M.; Al-Shaebi, Z.; Altunbek, M.; Bayraktar, C.; Kayabolen, A.; Bagci-Onder, T.; Aydin, O. Characterization and discrimination of spike protein in SARS-CoV-2 virus-like particles via surface-enhanced Raman spectroscopy. Biotechnol. J. 2024, 19 (1), e2300191 DOI: 10.1002/biot.202300191There is no corresponding record for this reference.
- 106Robertson, J. L.; Senger, R. S.; Talty, J.; Du, P.; Sayed-Issa, A.; Avellar, M. L.; Ngo, L. T.; Gomez De La Espriella, M.; Fazili, T. N.; Jackson-Akers, J. Y. Alterations in the molecular composition of COVID-19 patient urine, detected using Raman spectroscopic/computational analysis. PLoS One 2022, 17 (7), e0270914 DOI: 10.1371/journal.pone.0270914There is no corresponding record for this reference.
- 107Pahlow, S.; Richard-Lacroix, M.; Hornung, F.; Kose-Vogel, N.; Mayerhofer, T. G.; Hniopek, J.; Ryabchykov, O.; Bocklitz, T.; Weber, K.; Ehricht, R.; Simple, Fast and Convenient Magnetic Bead-Based Sample Preparation for Detecting Viruses via Raman-Spectroscopy. Biosensors 2023, 13 (6). 594 DOI: 10.3390/bios13060594 .There is no corresponding record for this reference.
- 108Zhang, D.; Zhang, X.; Ma, R.; Deng, S.; Wang, X.; Wang, X.; Zhang, X.; Huang, X.; Liu, Y.; Li, G. Ultra-fast and onsite interrogation of Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2) in waters via surface enhanced Raman scattering (SERS). Water Res. 2021, 200, 117243 DOI: 10.1016/j.watres.2021.117243There is no corresponding record for this reference.
- 109Kim, K.; Kashefi-Kheyrabadi, L.; Joung, Y.; Kim, K.; Dang, H.; Chavan, S. G.; Lee, M. H.; Choo, J. Recent advances in sensitive surface-enhanced Raman scattering-based lateral flow assay platforms for point-of-care diagnostics of infectious diseases. Sens Actuators B Chem. 2021, 329, 129214 DOI: 10.1016/j.snb.2020.129214There is no corresponding record for this reference.
- 110Mo, W.; Wen, J.; Huang, J.; Yang, Y.; Zhou, M.; Ni, S.; Le, W.; Wei, L.; Qi, D.; Wang, S. Classification of Coronavirus Spike Proteins by Deep-Learning-Based Raman Spectroscopy and its Interpretative Analysis. J. Appl. Spectrosc. 2023, 89 (6), 1203– 1211, DOI: 10.1007/s10812-023-01487-wThere is no corresponding record for this reference.
- 111Lu, X.; Liu, Q.; Benavides-Montano, J. A.; Nicola, A. V.; Aston, D. E.; Rasco, B. A.; Aguilar, H. C. Detection of receptor-induced glycoprotein conformational changes on enveloped virions by using confocal micro-Raman spectroscopy. J. Virol 2013, 87 (6), 3130– 3142, DOI: 10.1128/JVI.03220-12There is no corresponding record for this reference.
- 112Neng, J.; Li, Y.; Driscoll, A. J.; Wilson, W. C.; Johnson, P. A. Detection of Multiple Pathogens in Serum Using Silica-Encapsulated Nanotags in a Surface-Enhanced Raman Scattering-Based Immunoassay. J. Agric. Food Chem. 2018, 66 (22), 5707– 5712, DOI: 10.1021/acs.jafc.8b00026There is no corresponding record for this reference.
- 113Zhang, H.; Harpster, M. H.; Wilson, W. C.; Johnson, P. A. Surface-enhanced Raman scattering detection of DNAs derived from virus genomes using Au-coated paramagnetic nanoparticles. Langmuir 2012, 28 (8), 4030– 4037, DOI: 10.1021/la204890tThere is no corresponding record for this reference.
- 114Neng, J.; Harpster, M. H.; Wilson, W. C.; Johnson, P. A. Surface-enhanced Raman scattering (SERS) detection of multiple viral antigens using magnetic capture of SERS-active nanoparticles. Biosens. Bioelectron. 2013, 41, 316– 321, DOI: 10.1016/j.bios.2012.08.048There is no corresponding record for this reference.
- 115Zong, C.; Xu, M.; Xu, L. J.; Wei, T.; Ma, X.; Zheng, X. S.; Hu, R.; Ren, B. Surface-Enhanced Raman Spectroscopy for Bioanalysis: Reliability and Challenges. Chem. Rev. 2018, 118 (10), 4946– 4980, DOI: 10.1021/acs.chemrev.7b00668There is no corresponding record for this reference.
- 116Kharbach, M.; Alaoui Mansouri, M.; Taabouz, M.; Yu, H. Current Application of Advancing Spectroscopy Techniques in Food Analysis: Data Handling with Chemometric Approaches. Foods 2023, 12 (14). 2753 DOI: 10.3390/foods12142753 .There is no corresponding record for this reference.
- 117Marzi, J.; Fuhrmann, E.; Brauchle, E.; Singer, V.; Pfannstiel, J.; Schmidt, I.; Hartmann, H. Non-Invasive Three-Dimensional Cell Analysis in Bioinks by Raman Imaging. ACS Appl. Mater. Interfaces 2022, 14 (27), 30455– 30465, DOI: 10.1021/acsami.1c24463There is no corresponding record for this reference.
- 118Eberhardt, K. S.; Matthäus, C.; Schmitt, C.; Popp, M. J. Advantages and limitations of Raman spectroscopy for molecular diagnostics: An update. Expert Rev. Mol. Diagn. 2015, 15 (6), 773– 787, DOI: 10.1586/14737159.2015.1036744There is no corresponding record for this reference.
- 119Storozhuk, D.; Ryabchykov, O.; Popp, J.; Bocklitz, T. RAMANMETRIX: A Delightful Way to Analyze Raman Spectra , 2022, arXiv:2201.07586. arXiv.org e-Print archive https://arxiv.org/abs/2201.07586.There is no corresponding record for this reference.
- 120Ramanmetrix. RAMANMETRIX Software Documentation , 2021.There is no corresponding record for this reference.
- 121Zu, Y.; Chang, H.; Cui, Z. Molecular point-of-care testing technologies: Current status and challenges. Nexus 2025, 2 (2), 100059 DOI: 10.1016/j.ynexs.2025.100059There is no corresponding record for this reference.
- 122Food, U. S.; Drug, A. Overview of Device Regulation Device Advice: Comprehensive Regulatory Assistance , 2025.There is no corresponding record for this reference.
- 123Regulation (EU) 2017/746 of the European Parliament and of the Council of 5 April 2017 on in vitro diagnostic medical devices and repealing Directive 98/79/EC and Commission Decision 2010/227/EU . 2010, Vol. L 117 176 332.There is no corresponding record for this reference.
- 124Food, U. S.; Drug, A. Artificial Intelligence in Software as a Medical Device. Software as a Medical Device (SaMD)─Device Advice: Comprehensive Regulatory Assistance . 2025, (acccessed 2025/08/21).There is no corresponding record for this reference.
- 125Bottini, M.; Ryu, S. J.; Terander, A. E.; Voglis, S.; Maldaner, N.; Bellut, D.; Regli, L.; Serra, C.; Staartjes, V. E. The Ever-Evolving Regulatory Landscape Concerning Development and Clinical Application of Machine Intelligence: Practical Consequences for Spine Artificial Intelligence Research. Neurospine 2025, 22 (1), 134– 143, DOI: 10.14245/ns.2449186.593There is no corresponding record for this reference.
- 126Artika, I. M.; Ma’roef, C. N. Laboratory biosafety for handling emerging viruses. Asian Pac. J. Trop. Biomed. 2017, 7 (5), 483– 491, DOI: 10.1016/j.apjtb.2017.01.020There is no corresponding record for this reference.
- 127Dolskiy, A. A.; Grishchenko, I. V.; Yudkin, D. V. Cell Cultures for Virology: Usability, Advantages, and Prospects. Int. J. Mol. Sci. 2020, 21 (21), 7978, DOI: 10.3390/ijms21217978There is no corresponding record for this reference.
- 128Selo, M. A.; Sake, J. A.; Kim, K. J.; Ehrhardt, C. In vitro and ex vivo models in inhalation biopharmaceutical research - advances, challenges and future perspectives. Adv. Drug Delivery Rev. 2021, 177, 113862 DOI: 10.1016/j.addr.2021.113862There is no corresponding record for this reference.
- 129Colby, L. A.; Quenee, L. E.; Zitzow, L. A. Considerations for infectious disease research studies using animals. In Comparative Medicine 2017; Vol. 67, pp 222– 231.There is no corresponding record for this reference.
- 130Klingstrom, T.; Bongcam-Rudloff, E.; Reichel, J. Legal & ethical compliance when sharing biospecimen. Brief Funct. Genomics 2018, 17 (1), 1– 7, DOI: 10.1093/bfgp/elx008There is no corresponding record for this reference.
- 131Srivastava, S.; Sharma, D.; Kumar, S.; Sharma, A.; Rijal, R.; Asija, A.; Adhikari, S.; Rustagi, S.; Sah, S.; Al-Qaim, Z. H. Emergence of Marburg virus: a global perspective on fatal outbreaks and clinical challenges. Front. Microbiol. 2023, 14, 1239079 DOI: 10.3389/fmicb.2023.1239079There is no corresponding record for this reference.
- 132Vitti, J. N.; Vitti, R.; Chu, K.; Mellis, S. The ethics of clinical research in the era of COVID-19. Front Public Health 2024, 12, 1359654 DOI: 10.3389/fpubh.2024.1359654There is no corresponding record for this reference.
- 133Selgelid, M. J. Gain-of-Function Research: Ethical Analysis. Sci. Eng. Ethics 2016, 22 (4), 923– 964, DOI: 10.1007/s11948-016-9810-1There is no corresponding record for this reference.
- 134Belser, J. A.; Kieran, T. J.; Mitchell, Z. A.; Sun, X.; Mayfield, K.; Tumpey, T. M.; Spengler, J. R.; Maines, T. R. Key considerations to improve the normalization, interpretation and reproducibility of morbidity data in mammalian models of viral disease. Dis. Model Mech. 2024, 17 (3), dmm050511, DOI: 10.1242/dmm.050511There is no corresponding record for this reference.
- 135Masopust, D.; Sivula, C. P.; Jameson, S. C. Of Mice, Dirty Mice, and Men: Using Mice To Understand Human Immunology. J. Immunol. 2017, 199 (2), 383– 388, DOI: 10.4049/jimmunol.1700453There is no corresponding record for this reference.
- 136Zabidi, N. Z.; Liew, H. L.; Farouk, I. A.; Puniyamurti, A.; Yip, A. J. W.; Wijesinghe, V. N.; Low, Z. Y.; Tang, J. W.; Chow, V. T. K.; Lal, S. K. Evolution of SARS-CoV-2 Variants: Implications on Immune Escape, Vaccination, Therapeutic and Diagnostic Strategies. Viruses 2023, 15 (4). 944 DOI: 10.3390/v15040944 .There is no corresponding record for this reference.
- 137Hartman, K. A.; Clayton, N.; Thomas, G. J., Jr. Studies of Virus Structure by Raman Spectroscopy. I. R17 Virus and R17 RNA. Biochem. Biophys. Res. Commun. 1973, 50 (3), 942– 949, DOI: 10.1016/0006-291X(73)91336-3There is no corresponding record for this reference.
- 138Thomas, G. J., Jr.; Prescott, B.; McDonald-Ordzie, P. E.; Hartman, K. A. Studies of Virus Structure by Laser Raman Spectroscopy. II. MS2 Phage, MS2 Capsids, and MS2 RNA in Aqueous Solutions. J. Mol. Biol. 1976, 102 (1), 103– 124, DOI: 10.1016/0022-2836(76)90076-0There is no corresponding record for this reference.
- 139Thomas, G. J.; Prescott, B.; Day, L. A. Structure Similarity, Difference, and Variability in the Filamentous Viruses fd, If1, IKe, Pf1, and Xf Investigated by Laser Raman Spectroscopy. J. Mol. Biol. 1983, 165 (2), 321– 356, DOI: 10.1016/S0022-2836(83)80260-5There is no corresponding record for this reference.
- 140Wen, Z.-Q.; Thomas, G. J., Jr. UV Resonance Raman Spectroscopy of DNA and Protein Constituents of Viruses: Assignments and Cross Sections for Excitations at 257, 244, 238, and 229 nm. Biopolymers 1998, 45 (3), 247– 256, DOI: 10.1002/(SICI)1097-0282(199803)45:3There is no corresponding record for this reference.
- 141Moor, K.; Terada, Y.; Taketani, A.; Matsuyoshi, H.; Ohtani, K.; Sato, H. Early Detection of Virus Infection in Live Human Cells Using Raman Spectroscopy. J. Biomed. Opt. 2018, 23 (9), 097001 DOI: 10.1117/1.JBO.23.9.097001There is no corresponding record for this reference.
- 142El-Mashtoly, S. F.; Gerwert, K. Diagnostics and Therapy Assessment Using Label-Free Raman Imaging. Anal. Chem. 2022, 94 (1), 120– 142, DOI: 10.1021/acs.analchem.1c04483There is no corresponding record for this reference.
- 143Marotta, N. E. Patterned Nanoarray SERS Substrates for Pathogen Detection. Ph.D. Thesis. Georgia Institute of Technology: Atlanta, GA, 2010.There is no corresponding record for this reference.
- 144Sigle, M.; Rohlfing, K.; Kenny, M.; Scheuermann, S.; Sun, N.; Graeßner, U.; Haug, V.; Sudmann, J.; Seitz, C. M.; Heinzmann, D.; Schenke-Layland, K.; Maguire, P. B.; Walch, A.; Marzi, J.; Gawaz, M. P. Translating Genomic Tools to Raman Spectroscopy Analysis Enables High-Dimensional Tissue Characterization at Molecular Resolution. Nat. Commun. 2023, 14, 5799 DOI: 10.1038/s41467-023-41417-0There is no corresponding record for this reference.
- 145Gahlaut, S. K.; Savargaonkar, D.; Sharan, C.; Yadav, S.; Mishra, P.; Singh, J. P. SERS Platform for Dengue Diagnosis from Clinical Samples Employing a Hand Held Raman Spectrometer. Anal. Chem. 2020, 92 (3), 2527– 2534, DOI: 10.1021/acs.analchem.9b04129There is no corresponding record for this reference.
- 146Gordon, J.; Mousavi, S. D.; Li, J.; Filippone, N.; Walter, L.; Cheng, H.-W.; Skeete, Z.; Feldman, H.; Hakimi, S.; Cappuccio, K.; Wang, S.; Hader, M.; Shang, G.; Turner, J.; Cameron, A.; Bane, S.; Poliks, M.; Lu, S.; Yuan, T. T.; Zhong, C.-J. Plasmonic Nanoprobe-Enabled SERS Detection of SARS-CoV-2 Proteins and Virus Samples on Wax-Printed Paper Substrates. Anal. Chem. 2025, 97 (46), 21303– 21313, DOI: 10.1021/acs.analchem.5c02355There is no corresponding record for this reference.
- 147Plavec, Z.; Domanska, A.; Liu, X.; Laine, P.; Paulin, L.; Varjosalo, M.; Auvinen, P.; Wolf, S. G.; Anastasina, M.; Butcher, S. J. SARS-CoV-2 Production, Purification Methods and UV Inactivation for Proteomics and Structural Studies. Viruses 2022, 14 (9), 1989, DOI: 10.3390/v14091989There is no corresponding record for this reference.
- 148Pestov, D.; Murawski, R. K.; Ariunbold, G. O.; Wang, X.; Zhi, M.; Sokolov, A. V.; Sautenkov, V. A.; Rostovtsev, Y. V.; Dogariu, A.; Huang, Y.; Scully, M. O. Optimizing the laser-pulse configuration for coherent Raman spectroscopy. Science. 2007, 316 (5822), 265– 268, DOI: 10.1126/science.1139055There is no corresponding record for this reference.
- 149Wang, Y.; Ruan, Q.; Lei, Z. C.; Lin, S. C.; Zhu, Z.; Zhou, L.; Yang, C. Highly Sensitive and Automated Surface Enhanced Raman Scattering-based Immunoassay for H5N1 Detection with Digital Microfluidics. Anal. Chem. 2018, 90, 5224– 5231, DOI: 10.1021/acs.analchem.8b00002There is no corresponding record for this reference.
- 150Chang, Y.; Wang, Y.; Li, W.; Wei, Z.; Tang, S.; Chen, R. Mechanisms, Techniques and Devices of Airborne Virus Detection: A Review. Int. J. Environ. Res. Public Health 2023, 20, 5471, DOI: 10.3390/ijerph20085471There is no corresponding record for this reference.



