
About the Cover:
Solved X-ray structure of the antibiotic cefditoren bound to a soluble construct of penicillin binding protein 1b (PBP1b). Spotlighted is a newly identified non-canonical pose of cefditoren observed from structure-based analysis of PBP inhibitors.
View the article.Introducing Our Authors
Introducing Our Authors
Jen Wagner
This publication is free to access through this site. Learn More
Letters
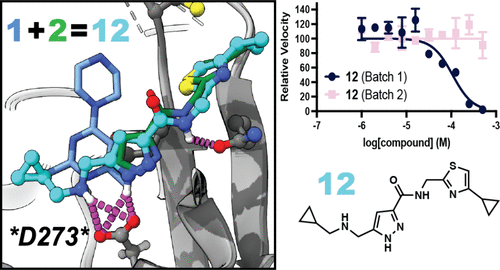
Fragment-Based Development of NSP14 Exonuclease Inhibitors Confounded by Batch-to-Batch Variability
Jesse A. Coker - ,
Rong Sun - ,
Paul M. Polzer - ,
Todd Romigh - ,
Christopher M. Goins - ,
Nancy S. Wang - ,
Jae U. Jung *- , and
Shaun R. Stauffer *
This publication is Open Access under the license indicated. Learn More
Point mutations in the exonuclease (ExoN) site of nonstructural protein 14 (NSP14) compromise the fitness of betacoronaviruses such as SARS-CoV-2, implicating NSP14 ExoN inhibition as an antiviral strategy. However, there are no advanced compounds that inhibit NSP14’s ExoN activity. Building upon the reported crystal structures of two fragments bound to NSP14’s ExoN site, we identified a series of 3,5-disubsituted pyrazoles that bound to and inhibited NSP14 ExoN. However, upon resynthesis, we discovered that these putative leads were false positives, perhaps due to contaminating divalent cations, which potently inhibit NSP14 ExoN. Our results provide a cautionary tale to the field about the sensitivity of NSP14 to divalent cations and illustrate the challenges associated with directly targeting the NSP14 ExoN site via fragment merging.
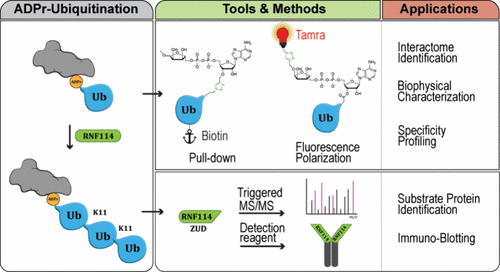
Development and Application of Chemical Biology Tools to Interrogate ADPr Ubiquitination
Max C. M. de Vries - and
Gerbrand J. van der Heden van Noort *
ADP-ribosyl ubiquitination is a unique form of crosstalk between post-translational modifications, where recently RNF114 has been identified as the first dedicated reader of this dual modification. New findings reveal that RNF114 extends the initial hybrid ADPr-Ub modification with K11-linked poly-Ub chains. This Letter highlights the recent advances made in methods and tools to study this hybrid signal and addresses some key questions that remain.
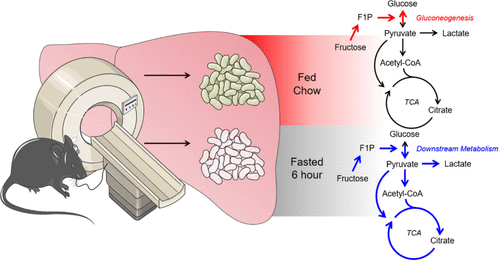
Hyperpolarized [U-2H, 2-13C]Fructose Distinguishes Direct Hepatic Gluconeogenesis Through Fructose-1-Phosphate Production in Fed and Fasted States
Celia Martínez de la Torre - ,
Grace Figlioli - ,
Mario C. Chang - ,
Quinlan Cullen - , and
Kayvan R. Keshari *
This publication is Open Access under the license indicated. Learn More
Hepatic fructose utilization depends on ketohexokinase mediated phosphorylation to generate fructose-1-phosphate and commit fructose carbons to additional metabolic steps. Since dysregulated fructose metabolism has been directly connected to the onset and progression of liver disease and cancer, there is considerable interest in identifying the contributions of fructose carbons in bioenergetic pathways. An essential technology for assessing fructose utilization has been the application of isotopically labeled fructose and magnetic resonance with the development of 13C hyperpolarized imaging with [2-13C]fructose allowing for in vivo assessments. While hyperpolarized imaging of [2-13C]fructose has achieved remarkable success in the detection of cancer metabolism, this approach has yet to be utilized to assess fed and fasted states in healthy livers. By challenging mice with a 6 h fast, we demonstrate that hyperpolarized [U-2H, 2-13C]fructose in vivo spectroscopy can clearly distinguish direct hepatic gluconeogenesis. Comprehensively, this work aims to establish a foundational methodology for the assessment of hepatic metabolism in vivo.
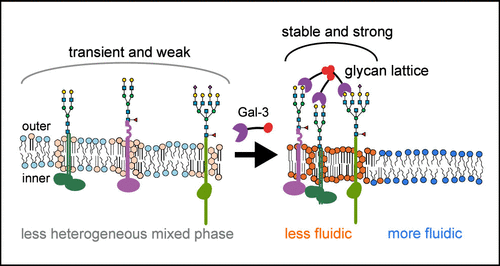
Galectin-3-Induced Glycan Lattices as Biophysical Modulators of Membrane Phase Behavior in Live Cells
Gil-Suk Yang - ,
Abigail E. Reeves - ,
Jia Meng Pang - ,
Shaheen A. Farhadi - ,
Arun Wanchoo - ,
Gregory Hudalla - , and
Mia L. Huang *
The plasma membrane is an important interface that integrates extracellular biochemical input with biophysical organization to regulate cell behavior. Galectin-3, a multivalent glycan-binding protein, can influence both events through the formation of extracellular glycan lattices on the surfaces of glycosylated cells. Although such lattices have been proposed to reshape membrane organization, their impact on nanoscale membrane phase behavior has remained difficult to quantify. Here, we establish a link between Galectin-3 lattice formation and the remodeling of plasma membranes by using imaging fluorescence correlation spectroscopy (ImFCS) to measure diffusion coefficients of a series of fluorescently labeled probes that partition into ordered or disordered regions of the cell membrane. Across several human cell types (BeWo, BxPC3, THP-1, and HEK293T), we observed that Galectin-3 induced significant changes in the lateral mobility of membranes in a manner dependent on the capacity of Galectin-3 to oligomerize and bind glycans, and that specific glycoproteins can play outsized contributing roles. Membrane regions enriched in Galectin-3 exhibited reduced diffusion, suggesting glycan lattices can serve as nucleation sites for ordered, raft-like microdomains. Finally, we also reveal that these Galectin-3-induced changes to membrane dynamics significantly amplifies Ca2+ triggered scrambling of phosphatidylserine exposure. Together, these findings identify Galectin-3 as an extracellular phase organizer that translates glycan recognition into nanoscale mechanical remodeling of the plasma membrane, potentially serving as a generalizable mechanism for fine-tuning cell behavior.
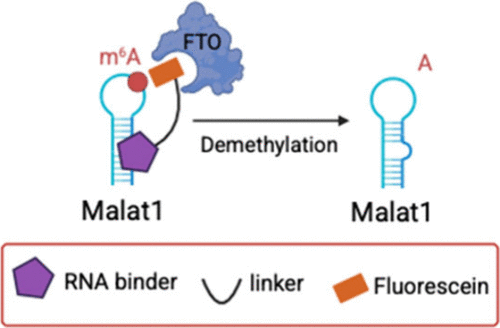
Cyclic γ-AApeptide-Based Molecular Glues for RNA m6A Editing
Chanjuan Dong - ,
Sihao Li - ,
Xinyu Xia - ,
Timothy H. Tran - ,
Satendra Kumar - ,
Huaxuan Yu - ,
Yu Yu Win - ,
Sining Li - ,
Yi He - ,
Jianfeng Cai *- , and
Fu-Sen Liang *
The m6A modification plays key roles in RNA metabolism and function and is implicated in various human diseases. In this study, we reported a novel molecular glue strategy for transcript-specific m6A editing using synthetic bifunctional molecules containing an RNA-targeting moiety and a ligand that recruit an endogenous m6A erasing enzyme. Through cyclic γ-AApeptide library screening, we identified a novel peptidomimetic binder to the long noncoding RNA MALAT1 A2577 region, which has a high m6A level. We developed a bifunctional molecular glue by coupling the identified MALAT1-binding cyclic γ-AApeptide to fluorescein, a reported binder to the m6A eraser FTO. We demonstrated that this bifunctional molecular glue successfully recruited FTO to the target RNA site, achieved the m6A erasing, disrupted HNRNPC-MALAT1 binding, and destabilized MALAT1. We anticipate that this novel molecular glue strategy will offer a new direction in developing molecules to regulate RNA modifications.
Articles
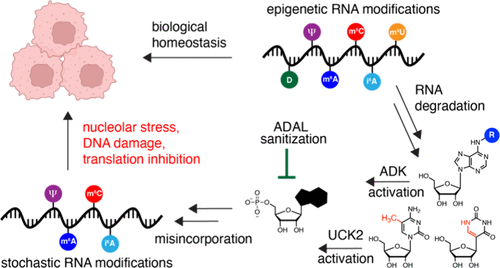
Metabolism of Epigenetic Ribonucleosides Leads to Nucleolar Stress and Cytotoxicity
Xuemeng Sun - ,
Anita Donlic - ,
Jacob A. Boyer - ,
Theodore E. Press - ,
Minjae Kim - ,
Neal K. Reddy - ,
Clifford P. Brangwynne - ,
Joshua D. Rabinowitz - , and
Ralph E. Kleiner *
This publication is Open Access under the license indicated. Learn More
Post-transcriptional RNA modifications are ubiquitous in biology, but the fate of epigenetic ribonucleotides after RNA turnover and the consequences of their metabolism and misincorporation into nucleic acids are largely unknown. Here, we explore epigenetic ribonucleoside metabolism in human cells by studying effects on cell growth, quantifying RNA misincorporation and identifying metabolic regulators, and exploring phenotypes associated with cytotoxicity. We find that bulky N6-modified adenosines (i.e., i6A) exhibit high levels of cytotoxicity and RNA misincorporation, whereas cells dramatically restrict the misincorporation of small N6-modified adenosines (i.e., m6A), partly through sanitization by enzymatic deamination, consistent with a recent report. Epigenetic ribopyrimidines also exhibit cytotoxicity, dependent on nucleoside kinase UCK2, but only at much higher concentrations than ribopurines. We further characterize the effects of cytotoxic ribonucleoside metabolism on nucleolar morphology and protein translation. Taken together, our work provides new insights into the metabolism of epigenetic ribonucleosides and mechanisms underlying their cytotoxicity to cells.
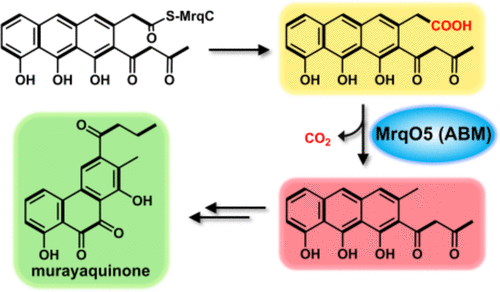
Decarboxylase Activity of an Antibiotic Biosynthesis Monooxygenase Family Protein in the Biosynthesis of the Type II Polyketide Murayaquinone
Jie Zhou - ,
Lin Lin - ,
Yaojie Gao - ,
Yan Gao - ,
Jinwei Zhu - ,
Xudong Qu - ,
Zixin Deng - , and
Ming Jiang *
Type II aromatic polyketides represent a structurally diverse class of natural products with medicinally relevant properties, and their biosynthesis usually involves biosynthetic intermediates with terminal carboxyl groups. In certain instances, terminal decarboxylation occurs, which can significantly impact the structural complexity. However, the enzymes and their involved mechanisms of terminal decarboxylation in type II aromatic polyketide biosynthesis have rarely been studied. This study has now shown that MrqO5, a member of the antibiotic biosynthesis monooxygenase (ABM) family, unexpectedly functions as a terminal decarboxylase involved in the biosynthesis of murayaquinone. Furthermore, an in vitro biochemical study demonstrated that two homologous proteins of MrqO5 exhibited similar decarboxylase activity. Therefore, the functional assignment and mechanistic investigation of this polyketide terminal decarboxylase elucidated an overlooked step in type II polyketide biosynthesis. Also, the discovery of this new family of decarboxylases expands the functions of the ABM superfamily proteins. Our structural characterizations, combined with site-directed mutagenesis studies, have unveiled the key residues involved in the decarboxylation and allowed an enzymatic decarboxylation mechanism to be proposed. Our studies advance the currently incomplete understanding of type II aromatic polyketide biosynthesis and gain the insight necessary for future engineering of these enzymes.
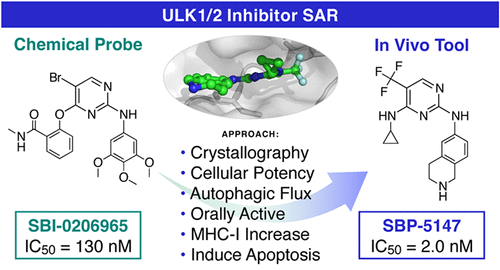
Synthesis and Characterization of ULK1/2 Kinase Inhibitors That Inhibit Autophagy and Upregulate Expression of Major Histocompatibility Complex I for the Treatment of Non-Small Cell Lung Cancer
Fabiana Izidro A. L. Layng - ,
Huiyu Ren - ,
Nicole A. Bakas - ,
Dhanya R. Panickar - ,
Lester J. Lambert - ,
Maria Celeridad - ,
Jiaqian Wu - ,
Laurent De Backer - ,
Preeti Chandrachud - ,
Allison S. Limpert - ,
Mitchell Vamos - ,
Apirat Chaikuad - ,
Betsaida B. Verdugo - ,
Patrick M. Hagan - ,
Sonja N. Brun - ,
Lutz Tautz - ,
Stefan Knapp - ,
Reuben J. Shaw - ,
Guy S. Salvesen - ,
Douglas J. Sheffler - , and
Nicholas D. P. Cosford *
Autophagy inhibition represents a promising therapeutic approach for the management of various cancers including nonsmall cell lung cancer (NSCLC). We previously reported SBP-7455, a dual inhibitor of unc-51-like kinase 1 (ULK1) and its homologue ULK2 and described its effects on triple-negative breast cancer (TNBC) cells. Herein we report the design, synthesis, and characterization of SBP-5147 and SBP-7501, two new dual ULK1/2 inhibitors that are cytotoxic against NSCLC cells, inhibit autophagic flux in A549 cells, and present greater oral exposure than SBP-7455 at a lower dose. In addition, SBP-5147 effectively modulates autophagy and increases the expression of major histocompatibility complex (MHC) class I in NSCLC cells, which may support the rationale for ULK1/2 inhibition as a strategy to overcome resistance to immunotherapy. Together these data support the use of ULK inhibitors as part of a cancer treatment strategy, either as a single agent or in combination with current therapies.
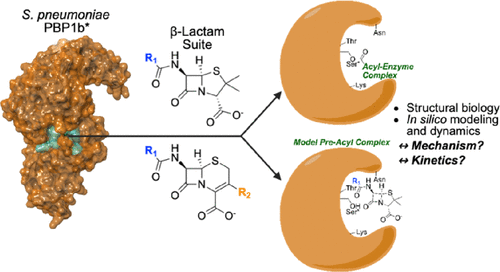
Structural and Dynamics Analyses of β-Lactam Inhibition of Streptococcus pneumoniae Penicillin-Binding Protein 1b (PBP1b) Guide Interrogation of Structure–Activity Relationships
Parker L. Flanders - ,
Jacob R. Gillingham - ,
Carlos Contreras-Martel - ,
Andréa Dessen - ,
Erin E. Carlson *- , and
Elizabeth A. Ambrose *
The Gram-positive pathogen Streptococcus pneumoniae, like the majority of bacteria, contains a peptidoglycan-based cell wall whose structure is highly dependent on the action of penicillin-binding proteins (PBPs). While the β-lactam antibiotics have been employed as an antimicrobial strategy for nearly a century, much remains unclear about how inhibitor structure informs potency and PBP isoform selectivity. Here, we obtained high-resolution structures (<2Å) of S. pneumoniae PBP1b cocrystallized with 6 β-lactams. Surprisingly, 2 structures feature a noncanonical conformation of the covalent “acyl-enzyme complex.” To clarify how protein–ligand interactions mediate inhibitor binding, we applied molecular modeling and molecular mechanics-based dynamics analyses. Our analyses illustrate how seemingly minimal changes to inhibitor structure modulate β-lactam binding mode and inhibitor potency, as described by the metric kinact/KI. Furthermore, we demonstrate that persistent interaction in the covalent acyl-enzyme complex between the inhibitor carboxylate and a highly conserved three-residue motif is not fully predictive of kinact/KI for PBP1b. In silico modeling suggests that the noncovalent preacyl complex may leverage this interaction, but a postacylation change in ligand conformation may accompany acylation in some inhibitors. The elucidation of key PBP1b ligand–receptor interactions pre- and postacylation will inform the rational design of novel PBP inhibitors and probes.
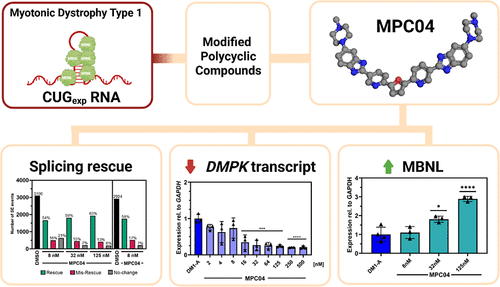
Modified Polycyclic Compounds Rescue Mis-splicing in Myotonic Dystrophy Type 1 Disease Models
Jesus A. Frias - ,
Sawyer M. Hicks - ,
Hormoz Mazdiyasni - ,
Subodh K. Mishra - ,
Kahini Sarkar - ,
Clara Yeboah - ,
Noah M. LeFever - ,
Marina M. Scotti - ,
Hana Zeghal - ,
Naomi Brandt - ,
Sweta Vangaveti - ,
Pramita Chakma - ,
Ting Wang - ,
Tammy S. Reid - ,
Omari McMichael - ,
Christopher Crumbaugh - ,
Marina Provenzano - ,
Melissa A. Hale - ,
John D. Cleary - ,
Nicholas E. Johnson - ,
Eric T. Wang - ,
Kaalak Reddy - , and
J. Andrew Berglund *
This publication is Open Access under the license indicated. Learn More
Myotonic dystrophy type 1 (DM1) is an autosomal dominant multisystemic disorder with no approved therapeutics targeting the disease mechanism. DM1 is caused by the expression of expanded CUG repeat RNA (CUGexp), which sequester the muscleblind-like (MBNL) family of RNA binding proteins leading to dysregulated alternative splicing and a host of downstream impacts. While previous studies showed that diamidines rescued DM1 dysregulated alternative splicing events, their potential was limited by toxicity and off-target effects. A new class of modified polycyclic compounds (MPCs), based on diamidines, were created and screened in DM1 patient-derived cell lines. This approach identified MPC03 and MPC04 as being capable of rescuing DM1 dysregulated splicing events at low nanomolar concentrations with no obvious toxicity and limited off-target effects. In a DM1 mouse model, treatment with MPC03 and MPC04 reduced CUGexp RNA levels and partially rescued DM1 mis-splicing. Binding data and modeling showed that lead MPCs bind to CUGexp RNA, and in cells lacking CUG repeats, MPC activity was absent, suggesting that these compounds displace sequestered MBNL proteins from CUGexp RNA. Taken together, MPCs show therapeutic promise across multiple DM1 models.
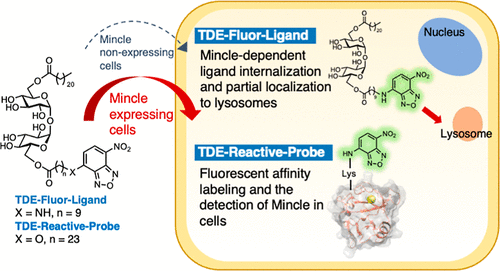
Live-Cell Imaging Analysis of the Intracellular Behavior of the C-Type Lectin Receptor Mincle and Its Ligand Using Newly Developed Fluorescence-Labeled Molecular Probes Based on Trehalose Diester
Takanori Matsumaru - ,
Kana Okubo - ,
Juri Takayama - ,
Yuki Hirano - ,
Yi-Ren Huo - ,
Satomi Toguchi - ,
Kenji Yoshida - ,
Ayumi Fujii - ,
Takehiro Suzuki - ,
Chun-Cheng Lin - ,
Naoshi Dohmae - ,
Sho Yamasaki - , and
Yukari Fujimoto *
Mincle, a member of the C-type lectin receptor (CLR) family, detects various glycolipids and glycerolipids such as trehalose dimycolate (TDM) from Mycobacterium tuberculosis, leading to the activation of the innate immune system. In this study, we developed new fluorescence-labeled molecular probes, TDE-Fluor-Ligand and TDE-Reactive-Probe, based on the structure of trehalose diester to elucidate the intracellular behavior of Mincle and its ligands. TDE-Fluor-Ligand was prepared for the ligand analysis, and TDE-Reactive-Probe was specifically designed to label Mincle by turn-on fluorescent affinity labeling. Live-cell imaging analysis using these probes revealed that TDE-Fluor-Ligand internalizes into the cell in a Mincle-dependent manner. Furthermore, imaging analysis using TDE-Reactive-Probe successfully detected Mincle in cells in a Mincle expression-dependent manner.
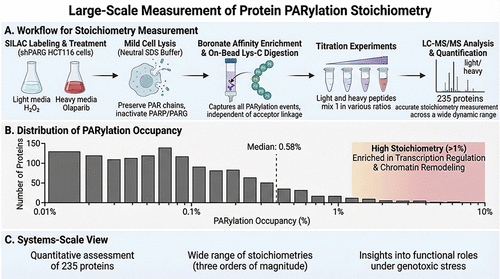
A Large-Scale Method to Measure the Stoichiometries of Protein Poly-ADP-Ribosylation
Peng Li - ,
Yajie Zhang - ,
Chiho Kim - , and
Yonghao Yu *
This publication is Open Access under the license indicated. Learn More
Poly-ADP-ribosylation (PARylation) is a reversible post-translational modification that occurs in higher eukaryotes. While thousands of PARylated substrates have been identified, the specific biological functions of most PARylated proteins remain elusive. PARylation stoichiometry is a critical parameter to assess the potential functions of a PARylated protein. Here, we developed a large-scale strategy to measure the stoichiometries of protein PARylation. By integrating chemically mild cell lysis conditions, boronate enrichment, and carefully designed titration experiments, we were able to determine the PARylation stoichiometries for a total of 235 proteins. Importantly, this approach enables the capture of all PARylation events, regardless of their amino acid acceptor linkages. We revealed that PARylation occupancy spans over 3 orders of magnitude. However, most PARylation events occur at low stoichiometric values (median 0.58%). Notably, we observed that high-stoichiometry PARylation (>1%) predominantly targets proteins involved in transcription regulation and chromatin remodeling. Thus, our study provides a system-scale, quantitative view of PARylation stoichiometries under genotoxic conditions, which serves as an invaluable resource for future functional studies of this important protein post-translational modification.
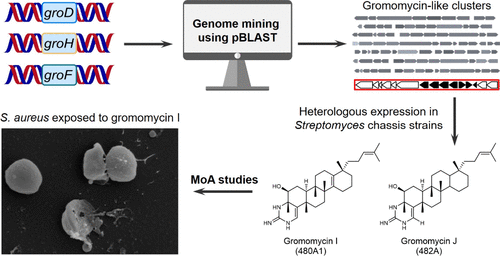
Genome Mining-Driven Isolation of New Gromomycins and Insights into Their Mode of Action
Dmytro Bratiichuk - ,
Franziska Fries - ,
Marc Stierhof - ,
Leon Morguet - ,
Josef Zapp - ,
Mathias Müsken - ,
Yuriy Rebets - ,
Maksym Myronovskyi - ,
Rolf Müller - ,
Jennifer Herrmann *- , and
Andriy Luzhetskyy *
This publication is Open Access under the license indicated. Learn More
The growing threat of multidrug-resistant bacterial infections highlights the urgent need for antibiotics with novel mechanisms of action. Gromomycins, a newly identified class of triterpene antibiotics, exhibit potent activity against Gram-positive bacteria, including drug-resistant species, through a previously uncharacterized mode of action. Here, we report the discovery of a gromomycin-like biosynthetic gene cluster in the Actinoplanes genus through a genome mining approach, leading to the isolation and characterization of new bioactive derivatives that overcome resistance to clinically used drugs in vancomycin-resistant enterococci. Mechanistic studies revealed that gromomycins induce rapid potassium ion leakage and depolarization of the bacterial membrane, resulting in bactericidal activity against Staphylococcus aureus. Gromomycins disrupt the integrity of the cytoplasmic membrane, as evidenced by large pore formation, leakage of intracellular contents, and subsequent cell lysis. Supplementation with membrane lipids and fatty acids neutralized their antibacterial activity, suggesting a direct membrane-targeting mechanism, further supported by the inability to raise gromomycin resistance and their toxic effects on eukaryotic cells. Collectively, these findings deepen our understanding of gromomycin activity and demonstrate the utility of genome mining to uncover structurally novel and biologically active natural products.
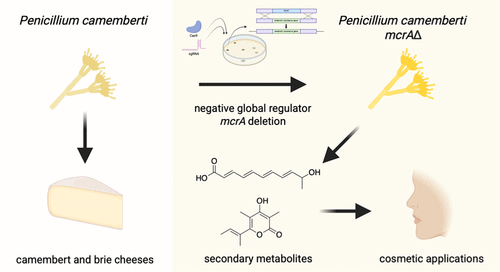
Manipulation of Global Regulator mcrA Activates the Fumigermin Pathway in Penicillium camemberti IMV00769
Jennifer Shyong - ,
Quoc-Dung Tran Huynh - ,
Victoria Feng - ,
Stella Dziedzic - ,
Pei Fang Chiu - ,
Youhan Liu - ,
Jason E. Stajich - ,
Ching-Kuo Lee - , and
Clay C. C. Wang *
Fungal secondary metabolites have historically provided important applications in a variety of industries. Penicillium camemberti, a fungus with a role in cheese production, was domesticated to food use partly due to its metabolically depleted characteristic, minimizing the risk of toxic compound formation. However, antiSMASH analysis of the genome reveals that strains of the species do contain various cryptic biosynthetic gene clusters and, thus, have the potential capability of producing multiple secondary metabolites despite its limited compound production under normal laboratory conditions. Here, we genetically engineered Penicillium camemberti strain IMV00769, which is genetically similar to cheese-making isolates, by deleting negative global regulator, mcrA. This deletion resulted in the production of secondary metabolites not previously produced by this strain, including fumigermin, a compound patented for cosmetic applications for the reduction of skin wrinkles, enhancement of skin elasticity, and skin whitening. Our findings highlight the power of global regulator manipulation to activate cryptic biosynthetic pathways and expand the range of natural products accessible from domesticated fungal strains.
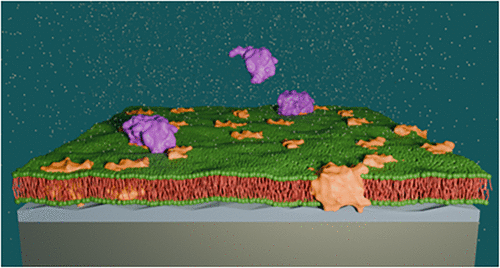
Avoiding Mitochondrial Apoptosis by the Bcl-2-Driven Bax Oligomerization on Membrane Surfaces
Sophie E. Ayscough - ,
Luke A. Clifton *- ,
Jörgen Ådén - ,
Sebastian Köhler - ,
Nicolò Paracini - ,
James Doutch - ,
Éilís C. Bragginton - ,
Anna E. Leung - ,
Oliver Bogojevic - ,
Jia-Fei Poon - ,
Tamás Milán Nagy - ,
Hanna P. Wacklin-Knecht - , and
Gerhard Gröbner *
This publication is Open Access under the license indicated. Learn More
The Bcl-2 family of proteins governs mitochondrial outer membrane (MOM) permeabilization, a critical step in apoptosis that is dysfunctional in many cancers. Although cellular studies have long implicated direct interactions between the pore-forming apoptotic Bax protein and its opponent, the antiapoptotic Bcl-2 protein in apoptosis regulation, the underlying basic principles behind this control remained unresolved. To provide in-depth insight, we carried out a systematic biophysical study in which we utilized neutron reflectometry (NR) and ATR-FTIR to elucidate the molecular communication between those proteins in and around the mitochondrial membrane environment. The spatial and temporal changes across model MOM surfaces were resolved during the interaction of Bax with Bcl-2. The NR-derived membrane surface Bax distributions suggested that Bcl-2 mediated Bax sequestration through both Bcl-2/Bax heterodimerization and Bax/Bax oligomerization. Kinetic analysis revealed a two-step process: rapid formation of Bcl-2/Bax heterodimers, followed by slower Bax oligomerization on these complexes. Importantly, this sequestration mechanism was also observed in the presence of cardiolipin, a lipid known to promote the formation of an apoptotic pore by Bax in the absence of Bcl-2. These findings suggest a fundamental mechanism by which cancer cells may evade apoptosis by exploiting Bcl-2’s ability to neutralize Bax through structural entrapment, even if excess Bax is present, either in response to treatment or natural death signals.
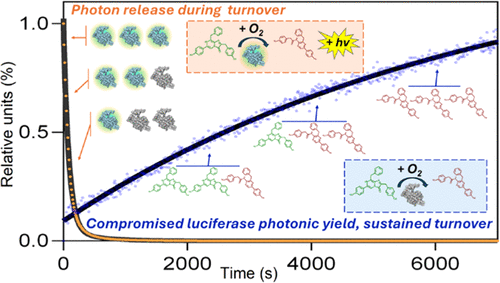
Elucidating Catalytic Bioluminescence of Mammalian Gaussia Luciferase through Mutant and Ancestral Analysis
Raina M. Borum - ,
Michael Lanzillotti - ,
Aniruddha Sahasrabuddhe - ,
John O. Hui - ,
Victoria Cochran Xie - , and
John Ferbas *
A mechanistic basis for luciferase bioluminescence provides a glimpse into its evolutionary role for organism survival, as it provides a blueprint to engineer luciferase enzymes for advanced technological applications. Gaussia Luciferase is among the brightest natural luciferases, but (1) the evolutionary development of its luminescence behavior remains unclear, (2) recent fundamental studies utilized Escherichia coli expression systems instead of eukaryotic expression systems, and (3) notable mutants have been discovered but not integrated into a comprehensive mechanistic analysis. We describe new mechanistic observations from GLuc by addressing these gaps. We monitored the fluorescent coelenterazine-to-coelenteramide conversion to study turnover kinetics of mammalian-derived GLuc; this assay characterized the positive cooperativity kinetics of GLuc. The nonluminescent mutants, R76A and R147A, still turn over the substrate with high efficiency, each demonstrating sustained positive cooperativity. Through mass spectrometry, mutational analysis, and analytical liquid chromatography, we demonstrate that GLuc undergoes methionine oxidation during substrate turnover and that this impacts the luciferase’s flash-type luminescence; we did not observe indications of covalent attachment with the substrate, product, or their intermediates. Chromatography of luciferases derived from ancestral sequence reconstruction highlighted that the extent of methionine-induced surface changes was greater for earlier ancestral luciferases. Ancestral sequence reconstruction also revealed that earlier ancestral copepod luciferases produced less light when compared to GLuc.
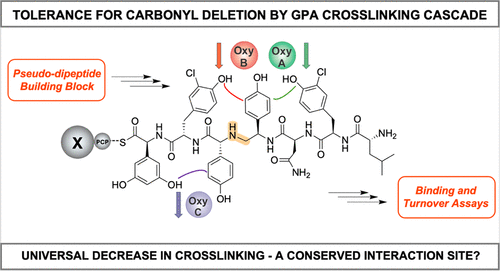
Methylene Peptide Backbone Substitution Significantly Affects the Glycopeptide Antibiotic Cross-Linking Cascade
Jemma Gullick - ,
Edward Marschall - ,
Sam Tucker - ,
Ralf B. Schittenhelm - ,
Lara R. Malins - ,
Julien Tailhades *- , and
Max J. Cryle *
Glycopeptide antibiotics (GPAs) are clinically important antibiotics characterized by a rigid, highly cross-linked structure. The cross-links in GPAs are installed by the activity of several cytochrome P450 (Oxy) enzymes, which are recruited to their peptide substrates by a unique domain, the X-domain. Given that this cross-linking cascade is the source of both the antibiotic activity and the synthetic complexity of GPAs, it remains a central point for exploring the tolerance of the Oxy enzymes for altered peptide substrates. In this study, we have investigated the ability of the Oxy enzymes to cross-link peptides with changes to their amide backbone, specifically a [Ψ[CH2NH]Tpg] methylene linkage that was inspired by synthetic efforts showing that such analogues can recover antibiotic activity toward resistant bacteria. Our results show that the Oxy enzymes are extremely sensitive to the presence of a methylene linkage in their peptide substrates, which suggests that these backbone carbonyl groups play a crucial role in maintaining the correct binding of peptide substrates to the P450 enzymes within the GPA cross-linking cascade.
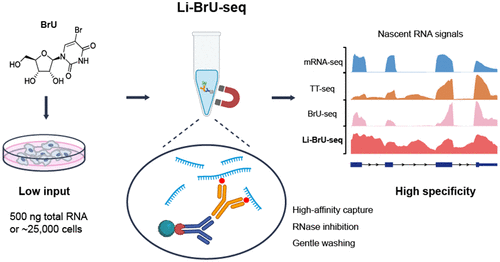
Li-BrU-seq: A Low-Input and Simplified Metabolic Labeling Method for Nascent RNA Sequencing
Yi-Feng Huang - ,
Jun-Tong He - ,
Ye-Lin Lan - ,
Zhi-Hao Zhang - ,
Fu Wu - ,
Chen-Long Wang - ,
Qiu-Yu Wang - ,
Guan-Zheng Luo *- , and
Zhang Zhang *
Precise regulation of transcriptional dynamics underlies gene expression programs, governing critical biological processes such as cell fate determination, tissue development, and stress responses. While nascent RNA sequencing technologies offer powerful tools for dissecting these mechanisms, existing methods remain constrained by complex workflows, high cellular input requirements, and cytotoxicity. Here, we present Li-BrU-seq, a systematically optimized 5-bromouridine (BrU)-based profiling strategy designed for low-input samples. Rigorous benchmarking demonstrates that Li-BrU-seq outperforms previous protocols in both enrichment specificity and sensitivity. By streamlining the enrichment workflow, the method enables high-quality transcriptomic profiling from low-input material (500 ng total RNA or ∼25,000 cells). Furthermore, Li-BrU-seq supports flexible temporal resolution ranging from ultrashort pulses to long-term tracking, free from the stress-induced artifacts inherent to 4sU. Additionally, it offers tailored workflows compatible with diverse downstream applications. Li-BrU-seq provides an accessible and versatile platform that expands nascent RNA analysis to low-input, rare, and physiologically sensitive biological systems.
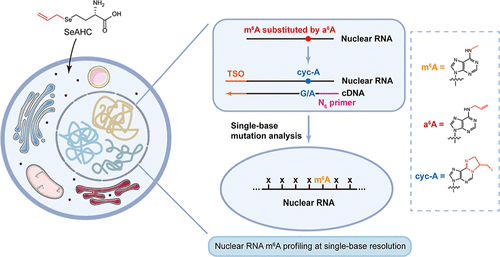
Nuclear-m6A-Label-Seq Enables Transcriptome-Wide Nuclear m6A Profiling at Single-Base Resolution
Chenyang Huang - ,
Xiner Ying - ,
Xiao Shu *- , and
Jianzhao Liu *
N6-Methyladenosine (m6A) on mature mRNA has been extensively characterized, yet its precise mapping and functions in nuclear noncoding RNAs remain elusive. To address this issue, we recently developed Nuclear-m6A-label-seq, a metabolic labeling-based method for transcriptome-wide nuclear m6A profiling at single-base resolution. This approach builds on the prototypical m6A-label-seq principle, in which an allyl group, instead of methyl group, is metabolically installed at N6-position at supposed RNA m6A-generating adenosines and the resultant N6-allyl adenosine is subsequently converted into 1, N6-cyclized adenosine (cyc-A) by mild iodination reaction. During RNA reverse transcription, HIV reverse transcriptase is employed to introduce a base misincorporation at cyc-A sites while enabling a template switch to incorporate adapter sequences to the complementary DNA end in a single step. Through this strategy, library construction is shortened to about 6 h, and the required cell-labeling total RNA input is reduced to 5 μg of total nuclear RNA, representing a 100-fold reduction compared to the prototypical protocol. Both polyadenylated and nonpolyadenylated nuclear transcripts are captured through the sequential nuclear RNA isolation and rRNA depletion. Following high-throughput sequencing, reads from human cells are aligned with the complete T2T-CHM13 genome, enabling accurate mapping of repetitive regions. Aligned reads are then analyzed using the user-friendly rMATS-DVR pipeline to identify high-confidence m6A sites based on cyc-A-induced misincorporation patterns. Here, we provide a detailed step-by-step protocol for Nuclear-m6A-label-seq, which stands for a direct and high-resolution approach for profiling the nuclear m6A epitranscriptome.
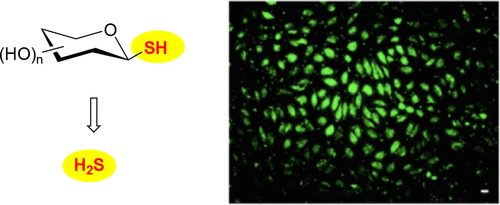
Evaluation of Thiosaccharides as Carbohydrate-Based Slow-Releasing Hydrogen Sulfide Donors
Stephen Lindahl - ,
Meg Shieh - ,
Iris Yang - ,
Masana Yazaki - ,
Taiming Liu - ,
Charlotte Stang - ,
Austin Sarker-Young - ,
Benjamin Kuran - ,
Jiajun Huang - ,
Yinsheng Wan - ,
Xiang Ni - ,
Mieko Arisawa - , and
Ming Xian *
Hydrogen sulfide (H2S) is an important gasotransmitter that has shown many physiological effects, ranging from anti-inflammation to antioxidation. To advance research on H2S, donor compounds that can slowly release H2S in biological conditions while producing minimal bioactive byproducts are essential. Herein, we report the evaluation of thiosaccharides as hydrolysis-based H2S donors. These compounds were found to slowly produce H2S over days, in aqueous buffers and in cells. Their H2S release rates could be affected by the hydroxyl protection groups of thiosaccharides, with faster release by electron-donating groups and slower release by electron-withdrawing groups. We also demonstrated the vasodilatory effect of 1-thioglucose using arterial rings isolated from adult ewes, which is likely due to H2S release. Altogether, thiosaccharides might be suitable slow-releasing H2S donors for biological applications.
Additions and Corrections
Correction to “Introducing Our Authors”
Jen Wagner
This publication is free to access through this site. Learn More
Mastheads
Issue Editorial Masthead
This publication is free to access through this site. Learn More
Issue Publication Information
This publication is free to access through this site. Learn More
