Spherical Nucleic Acids: Turning Synthetic Advances and Fundamental Discovery into Translational Breakthroughs in Chemistry, Materials Development, Biology, and MedicineClick to copy article linkArticle link copied!
- Connor M. ForsythConnor M. ForsythInterdisciplinary Biological Sciences Graduate Program, Northwestern University, Evanston, Illinois 60208, United StatesInternational Institute for Nanotechnology, Northwestern University, Evanston, Illinois 60208, United StatesMore by Connor M. Forsyth
- Rachel R. ChanRachel R. ChanDepartment of Chemistry, Northwestern University, Evanston, Illinois 60208, United StatesInternational Institute for Nanotechnology, Northwestern University, Evanston, Illinois 60208, United StatesMore by Rachel R. Chan
- Tanner D. FinkTanner D. FinkDepartment of Chemistry, Northwestern University, Evanston, Illinois 60208, United StatesInternational Institute for Nanotechnology, Northwestern University, Evanston, Illinois 60208, United StatesMore by Tanner D. Fink
- Janice KangJanice KangDepartment of Chemistry, Northwestern University, Evanston, Illinois 60208, United StatesInternational Institute for Nanotechnology, Northwestern University, Evanston, Illinois 60208, United StatesMore by Janice Kang
- Jacob D. CohenJacob D. CohenInterdisciplinary Biological Sciences Graduate Program, Northwestern University, Evanston, Illinois 60208, United StatesInternational Institute for Nanotechnology, Northwestern University, Evanston, Illinois 60208, United StatesMore by Jacob D. Cohen
- Sarah Hurst PetroskoSarah Hurst PetroskoDepartment of Chemistry, Northwestern University, Evanston, Illinois 60208, United StatesInternational Institute for Nanotechnology, Northwestern University, Evanston, Illinois 60208, United StatesMore by Sarah Hurst Petrosko
- Chad A. Mirkin*Chad A. Mirkin*Email: [email protected]Interdisciplinary Biological Sciences Graduate Program, Northwestern University, Evanston, Illinois 60208, United StatesDepartment of Chemistry, Northwestern University, Evanston, Illinois 60208, United StatesInternational Institute for Nanotechnology, Northwestern University, Evanston, Illinois 60208, United StatesMore by Chad A. Mirkin
Abstract
Conspectus
Early research in nanoscience and nanotechnology focused on gaining synthetic control over the size, shape, and composition of nanostructures, as well as exploring their fundamental properties. Over the past few decades, these capabilities have become increasingly sophisticated. Today, we have well-established synthetic toolkits and methodologies that enable the design of nanostructures with tailored properties and functions, guided by sets of design rules, for use in many areas spanning biology and medicine to energy, the environment, and catalysis.
To illustrate this paradigm, where synthesis and fundamental discovery drive engineering and technological innovation, we examine spherical nucleic acids (SNAs) as a case study. SNAs are nanoconstructs consisting of a nanoparticle core densely functionalized with a radially oriented oligonucleotide shell. Over the past 30 years, the evolution of SNAs has spanned their invention, the development of increasingly advanced syntheses enabling the creation of dozens of SNA classes (and related DNA-functionalized anisotropic materials, often termed programmable atom equivalents [PAEs]), the discovery of novel phenomena that have reshaped core chemical principles, and their translation into nanomedicines, biological labels, and synthons in materials science.
SNAs were first developed in 1996 as gold nanoparticle–DNA conjugates. Since then, extensive study has revealed common structural features that are tied to their unique properties, defining SNAs as a distinct materials class. Most SNAs feature a core (typically a nanoparticle, though recent advances involve molecular scaffolds) that concentrate nucleic acid strands into close proximity. This architecture confers several distinctive properties: enhanced binding affinity to complementary DNA (both free and surface-bound), resistance to enzymatic degradation, reduced immune activation (unless specifically designed for immunostimulation), and efficient cellular uptake without requiring transfection agents.
These synthetic and fundamental advances offer significant advantages in biomedical probe and therapeutic design. Due to their modularity, stability, biocompatibility, and ability to access intracellular compartments, SNAs have been applied as intracellular and extracellular probes, tools for gene regulation, vaccines, and gene editing platforms (especially when coupled with CRISPR/Cas9 technology). In parallel, SNAs serve as foundational elements in a new class of programmable matter: DNA-mediated colloidal crystals. Here, sequence-specific DNA interactions are used to organize SNAs into three-dimensional, periodic structures. This line of inquiry has enabled the design and synthesis of thousands of crystal variations, with different lattice symmetries, parameters, and nanoparticle compositions, unlocking the potential for novel optical and mechanical metamaterials and catalysts with exceptional properties, such as negative refractive indices, shape memory, and second harmonic generation. In sum, SNAs exemplify how synthetic mastery and fundamental discovery can catalyze innovation across disciplines, providing a framework that chemists can use in developing transformative new materials.
This publication is licensed for personal use by The American Chemical Society.
Key References
Mirkin, C. A.; Letsinger, R. L.; Mucic, R. C.; Storhoff, J. J. A DNA-based method for rationally assembling nanoparticles into macroscopic materials. Nature 1996, 382, 607–609. (1) This paper introduces spherical nucleic acids (SNAs) and the concept of the nanoparticle “atom” and the DNA “bond”. It discusses SNA synthesis and characterization as well as their use in colloidal crystal engineering with DNA to generate programmable matter.Rosi, N. L. ; Oligonucleotide-Modified Gold Nanoparticles for Intracellular Gene Regulation. Science 2006, 312, 1027–1030. (7) This paper highlights the first demonstration of the ability of SNAs to enter cells in high quantities as single entity agents, an unexpected result at the time. This discovery laid the foundation for SNAs as intracellular probes and therapeutic agents.Macfarlane, R. J. ; Nanoparticle Superlattice Engineering with DNA. Science 2011, 334, 204–208. (10) This paper describes the first design rules for SNA-based colloidal crystals, enabling control over crystallographic parameters (composition, symmetry, and lattice constant). It introduces the complementary contact model (CCM) for designing structures based upon particle arrangements that maximize DNA bonding interactions.Wang, S. ; Rational vaccinology with spherical nucleic acids. Proc. Natl. Acad. Sci. U. S. A. 2019, 116, 10473–10481. (6) This paper introduces rational vaccinology, where the structures of SNA immunotherapeutics, not solely the composition (i.e., medicinal components), dictate their potency through control of target engagement and signaling kinetics. It lays the foundation for the broader field of structural nanomedicine.
I. Introduction
Figure 1
Figure 1. Synthetic advances with SNAs. (A) Schematic shows a gold-core SNA radially functionalized with strands of oligonucleotides. (B) SNAs exhibit a high degree of oligonucleotide and core shape modularity. Functional oligonucleotides typically contain three segments: a functional oligonucleotide sequence, a spacer segment, and an attachment group for nanoparticle surface binding. SNA-like properties can be achieved using anisotropic nanoparticle cores including truncated octahedron, truncated octahedral nanoframes, cubes, and triangular prisms. Adapted with permission from ref (206). Copyright 2022 Springer Nature Ltd. (C) SNA diameter is dependent on the core size and oligonucleotide shell length. Both parameters can be tuned independently. SNAs of the same diameter can be prepared using different core sizes and DNA shell lengths. Adapted with permission from ref (10). Copyright 2011 American Association for the Advancement of Science. (D) To date, SNAs have been synthesized using a variety of core materials that allow access to vastly different physicochemical properties. Core materials have included both inorganic and organic cores, and they endow the SNA structures with specific properties (e.g., plasmonic and enzymatic) based on composition. Pictured are a selection of different SNAs with a diversity of core materials, including molecular, protein, gold, liposomal, and lipid nanoparticle cores ranging in size from ∼1 to >150 nm. The representation of a CRISPR SNA is adapted from PDB ID 8UZA. ePMV software was used in the generation of panels A, B, and D, ref (250).
II. Synthetic Advances Underpinning Scientific Discovery with SNAs
II.i. Synthetic Advance: Ability to Control SNA Size
II.ii. Synthetic Advance: Ability to Independently Control Composition and Structure of SNA Core and Shell
II.iii. Synthetic Advance: Ability to Control Shape (i.e., Prepare Anisotropic Analogues to SNAs)
III. From Scientific Discovery to Engineering Outcomes with SNAs
III.i. Scientific Discovery: Spherical Nucleic Acids Exhibiting Distance-Dependent Optical Properties. Engineering Outcome: Colorimetric Detection
Figure 2
Figure 2. Fundamental research and engineering development with SNA-based probes. (A) Solutions of gold nanoparticle-core SNAs before (left; dispersed particles are red) and after the addition of a target DNA sequence links the SNAs (right; aggregated particles are purple). (B) Melting curves show free DNA and the same DNA attached to a gold-core SNA and a core-free polyvalent nucleic acid nanostructure (PNAN) (left); enthalpies and entropies of DNA hybridization to linear DNA and SNAs are derived from a concentration-dependent fluorescence hybridization assay (right). Panel B, left: Adapted with permission from ref (59). Copyright 2011 American Chemical Society. Panel B, right: Reprinted with permission from ref (131). Copyright 2018 American Chemical Society. (C) Example shows polynucleotide target detection using the colorimetric Northwestern Spot Test (left). Solutions of SNA probes were mixed with a fully complementary target strand (Target), without the target (No Target) and with a target contaning a single base pair mismatch (Mismatch Target) and heated to the indicated temperature. Note the high degree of discrimination between the fully complementary and mismatched targets, allowing for the detection of single nucleotide mismatches. Example shows the first SNA-based scanometric detection system wherein oligonucleotide arrays were treated with target strands (either fully complementary [A] or with a single base pair mismatch, substituting A for G, T, or C) and SNA probes were developed at the indicated temperatures (right). Panel C, left: Adapted with permission from ref (112). Copyright 1997 American Association for the Advancement of Science. Panel C, right: Reprinted with permission from ref (126). Copyright 2000 American Association for the Advancement of Science. (D) Chip-based multiplexed scanometric detection of barcode-DNA strands are released from SNA probes after incubation with DNA targets derived from the human immunodeficiency virus (HIV), Ebola virus (EV), variola virus (VV), and hepatitis B virus (HBV), or a mixture of all four (Mix). Adapted with permission from ref (139). Copyright 2006 WILEY-VCH Verlag GmbH & Co. KGaA, Weinheim. ePMV software was used in the generation of panel A, ref (250).
III.ii. Scientific Discovery: Nucleic Acids on Particle Surfaces Exhibiting Cooperative, Higher Affinity Target Binding Properties. Engineering Outcome: High-Sensitivity and High-Selectivity Detection
III.iii. Scientific Discovery: Gold Core of SNA Gold Nanoparticle Conjugates Catalyzes Cross-Linking of Alkyne-Based Adsorbates. Engineering Outcome: Preparation of Hollow SNAs and All-Nucleic Acid Nanomaterials
III.iv. Scientific Discovery: SNA Resistance to Nuclease Degradation and no Significant Immune Response (Unless Designed to do so Based on Sequence). Engineering Outcome: Stability in in Vivo and in Vitro Biological Environments and High Compatibility with Cells and Tissues
Figure 3
Figure 3. Scientific discoveries and engineering outcomes of SNAs in nanomedicine research. (A) The dense 3D DNA shell of SNAs confers enhanced resistance to nuclease degradation relative to linear oligonucleotides. This protection arises from slower enzymatic hydrolysis, as the negatively charged and sterically bulky shell inhibits nuclease activity. The protein corona that forms on SNAs in biological media is not depicted. FAN1 Nuclease PDB ID 8S5A. (B) Compared to linear or transfected oligonucleotides, SNAs exhibit high cellular uptake due to their multivalent interactions with class A scavenger receptors, which facilitate endocytosis via a lipid-raft-dependent, caveolae-mediated pathway. The representation of scavenger receptors is adapted from PDB IDs 2QIH and 7DPX. (C) Their high uptake efficiency, stability, and limited activation of host immune responses enable SNAs to function broadly as tools in both in vitro and in vivo systems. This versatility supports their use in intracellular detection, imaging, gene regulation, and immunotherapy. (D) Detection probes, including nanoflares, sticky-flares, and FIT-flares, have been engineered to sense intracellular RNA, ions, small molecules, and diseased tissue in vivo, as well as to perform live-cell genetic and metabolic analysis. These probes incorporate fluorophores that are either released or activated upon target binding, generating sensitive and responsive readouts. (E) As gene regulation agents, SNAs can be constructed from DNA or RNA to modulate expression through antisense or RNA interference pathways. As immunotherapies, they stimulate both innate and adaptive immune responses and have shown promise as nanomedicines for treating diverse cancers and infectious disease. The representation of a CRISPR SNA is adapted from PDB ID 8UZA. ePMV software was used in the generation of panels A, B, D, and E, ref (250).
III.v. Scientific Discovery: SNAs Exhibiting Cellular Entry in High Quantities Without the Need for Transfection Agents. Engineering Outcomes: Intracellular Detection/Imaging and Therapeutics
III.vi. Scientific Discovery: SNAs Binding with TLRs More Strongly Than Linear Strands and Stimulating the Immune System. Engineering Outcome: Immunotherapeutic Development, Based on Structure in Addition to Composition (Structural Immunotherapy, Specifically “Rational Vaccinology”, a Subfield of Structural Nanomedicine)
Figure 4
Figure 4. Fundamental research and engineering development with SNA-based immunotherapeutics. Key scientific discoveries such as (A) the multivalent display of CpG DNA on SNAs potently stimulating TLR9 have pioneered the new field of structural immunotherapy. The representation of TLR9 is adapted from PDB IDs 5Y3M and 6US8. (B) Compositionally identical yet structurally distinct SNAs suppress tumor growth differentially, demonstrating the importance of nanoarchitecture in next-generation therapeutic development. Adapted with permission from ref (6). Copyright 2019 National Academy of Sciences. (C) These structure–function relationships extend to more complex therapeutics. The differential placements of MHC-I- and MHC-II-restricted antigens in DA-SNA 1 and 2 substantially alter tumor suppression. Adapted with permission from ref (22). Copyright 2023 Springer Nature Ltd. (D) The oligonucleotide shell of SNAs dictates their circulation and biodistribution, leading to the discovery of new agents that cross the blood–brain barrier (BBB). (E) Gold SNAs bearing Bcl2L12 siRNA labeled with Gd(III) localize in glioma tumor tissue following intracranial injection as shown in magnetic resonance (MR) images (top), hematoxylin and eosin-stained brain sections (middle), and the 3D reconstruction of MR data (bottom). Reprinted with permission from ref (195). Copyright 2013 American Association for the Advancement of Science (AAAS). (F) β-galactosidase-based ProSNA functionalized with transferrin aptamer (upper left) successfully accumulates within the brain (lower left), quantified by radiant efficiency (right). Reprinted with permission from ref (41). Copyright 2022 American Chemical Society. ePMV software was used in the generation of panels A, B, and C, ref (250).
III.vii. Scientific Discovery: SNA Biodistribution and Circulation Time Controlled via SNA Structure and SNAs Crossing the Blood–Brain and Blood–Tumor Barriers. Engineering Outcome: Organ Targeting with SNA Therapies
III.viii. Scientific Discovery: DNA Programmability and Annealing Processes Used to Drive the Formation of Colloidal Crystals and Tune Lattice Parameters (with Nanometer-Scale Precision), Symmetries, and Crystal Habits in Solution and on Surfaces. Engineering Outcomes: Variety of Interesting Optical, Mechanical, and Catalytic Colloidal Crystals
Figure 5
Figure 5. Scientific discoveries, materials design, and engineering outcomes through the crystallization of SNAs and other types of programmable atom equivalents (PAEs). (A) The colloidal crystal design space arises from the synthetic advances in SNA preparation, where enthalpy-dominated DNA interactions direct crystallization. Metal nanoparticle core (shape and size) and DNA design determine the lattice symmetry and crystal habit. Scale bars = 125, 75, and 1000 nm from left to right. (B) Enthalpic DNA bonding interactions are maximized with facet registry between anisotropic nanoparticle cores, such as cubes. Scale bar = 500 nm. (C) DNA bonding is maximized along the edges of nanoframes, forming a crystalline symmetry from a non-space-filling polyhedral. Scale bar = 1 μm. Reprinted with permission from ref (206). Copyright 2022 Springer Nature Ltd. (D) Thin films and Winterbottom constructions can be formed from heterogeneous crystal growth on substrates. Reprinted with permission from ref (226). Copyright 2017 American Chemical Society. (E) The refractive index of colloidal crystals can be designed as a function of NP shape, NP size, volume fraction, and symmetry. Reprinted with permission from ref (206). Copyright 2022 Springer Nature Ltd. (F) Colloidal crystals components can be stabilized and used as heterogeneous catalysts. (G) Deterministic placement is shown for single PAEs to couple photonic response (determined by spacing, P) and plasmonic response (distance between Au cube and Au substrate). Reprinted with permission from ref (243). Copyright 2015 American Chemical Society. (H) Colloidal crystals exhibit shape memory upon dehydration and rehydration. The mechanical response of the colloidal crystal due to a reversible change in refractive index results in tunable optical response. ePMV software was used in the generation of panel A, ref (250).
III.ix. Scientific Discovery: Face-to-Face Bonding Driving Crystallization through the Complementary Contact Model. Engineering Outcomes: Expanding Structural Possibilities for Optically Active Colloidal Crystals Engineered with DNA
III.x. Scientific Discovery: Edge-Bonding Driving Crystallization of Hollow, Nanoframe Materials via Complementary Contact Model. Engineering Outcomes: Variety of Interesting Optical and Mechanical Colloidal Crystals
IV. Conclusions
Biographies
Connor M. Forsyth
Connor M. Forsyth is an M.D./Ph.D. candidate in the Medical Scientist Training Program at Northwestern University, advised by Profs. Chad A. Mirkin and Alexander H. Stegh. He holds a B.S. in Molecular and Cellular Biology from the University of Illinois (2018) and an M.S. in Biotechnology from Northwestern University (2020). His research focuses on developing nanomedicine-based immunotherapies and investigating their fundamental interactions with biological systems.
Rachel R. Chan
Rachel R. Chan received her B.S. in Chemistry from the University of California, Berkeley (2021). She is currently pursuing her Ph.D. in Chemistry under the guidance of Prof. Chad Mirkin at Northwestern University. Her research focuses on designing defects into colloidal crystals engineered with DNA and integrating nanoparticles into larger, nanophotonic structures.
Tanner D. Fink
Tanner D. Fink received his B.S. degree in Chemical Engineering from Iowa State University (2016) and Ph.D. in Chemical Engineering from Rensselaer Polytechnic Institute (2022). Now, he is an International Institute for Nanotechnology Willens Nano-oncology Postdoctoral Fellow under Prof. Chad Mirkin at Northwestern University. His research focuses on utilizing the spherical nucleic acid platform for the development of cancer vaccine technologies.
Janice Kang
Janice Kang received her B.A. degree in Chemistry from Hamilton College (2020) and is currently pursuing her Ph.D. in Chemistry with Prof. Chad A. Mirkin at Northwestern University. Her work integrates chemistry with structural nanomedicine for next-generation cancer immunotherapy.
Jacob D. Cohen
Jacob D. Cohen received his B.Scs. in Biochemistry and Neuroscience from Indiana University Bloomington (2018). He is currently a Ph.D. candidate under the supervision of Prof. Chad A. Mirkin at Northwestern University. His work primarily focuses on studying nanoscale vaccine processing kinetics using spherical nucleic acids and the development of breast cancer therapeutics.
Sarah Hurst Petrosko
Sarah Hurst Petrosko received B.S. degrees in Chemistry and in Physics from the University of Florida in 2003 and a Ph.D. in Chemistry from Northwestern University (NU) in 2009. She is currently a Research Professor in the Department of Chemistry at NU and the Associate Director of NU’s International Institute for Nanotechnology.
Chad A. Mirkin
Chad A. Mirkin obtained his B.S. degree in 1986 from Dickinson College and his Ph.D. in 1989 from The Pennsylvania State University. He joined Northwestern University as an Assistant Professor in 1991. He currently serves as the Director of the International Institute for Nanotechnology and a Professor of Chemistry, Materials Science and Engineering, Biomedical Engineering, Chemical and Biological Engineering, and Medicine at Northwestern University. He is the inventor and developer of spherical nucleic acids as well as several additive manufacturing, nanopatterning, and materials discovery methodologies (including Dip Pen Nanolithography, Polymer Pen Lithography, Beam Pen Lithography, and High Area Rapid Printing), and he has cofounded 11 companies. Professor Mirkin has been elected to all three U.S. National Academies, as well as the American Academy of Arts and Sciences. He cofounded the journal Small, and has served on the editorial advisory boards of more than 35 scholarly journals.
Acknowledgments
Research reported in this publication was supported by the National Cancer Institute of the National Institutes of Health under Awards R01CA257926 and R01CA275430 and the Research Development Program under P50CA221747. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health. This manuscript is the result of funding in whole or in part by the National Institutes of Health (NIH) and is subject to the NIH Public Access Policy. Through acceptance of this federal funding, NIH has been given a right to make this manuscript publicly available in PubMed Central upon the Official Date of Publication, as defined by NIH. This material is also based upon work supported by the Air Force Office of Scientific Research Award FA9550-22-1-0300, the Lefkofsky Family Foundation, and Edgar H. Bachrach through the Bachrach Family Foundation. R.R.C. was partially supported by the National Science Foundation Graduate Research Fellowship Program Grant DGE-2234667. J.D.C. was partially supported by the Department of Defense through the National Defense Science & Engineering Graduate (NDSEG) Fellowship Program. C.M.F. was partially supported by a research grant from the Melanoma Research Foundation. Molecular graphics were performed with UCSF ChimeraX, developed by the Resource for Biocomputing, Visualization, and Informatics at the University of California, San Francisco, with support from National Institutes of Health R01-GM129325 and the Office of Cyber Infrastructure and Computational Biology, National Institute of Allergy and Infectious Diseases. Some figures were created with BioRender.
References
This article references 250 other publications.
- 1Mirkin, C. A.; Letsinger, R. L.; Mucic, R. C.; Storhoff, J. J. A DNA-Based Method for Rationally Assembling Nanoparticles into Macroscopic Materials. Nature 1996, 382 (6592), 607– 609, DOI: 10.1038/382607a0Google ScholarThere is no corresponding record for this reference.
- 2Cutler, J. I.; Auyeung, E.; Mirkin, C. A. Spherical Nucleic Acids. J. Am. Chem. Soc. 2012, 134 (3), 1376– 1391, DOI: 10.1021/ja209351uGoogle ScholarThere is no corresponding record for this reference.
- 3Mirkin, C. A.; Petrosko, S. H. Inspired Beyond Nature: Three Decades of Spherical Nucleic Acids and Colloidal Crystal Engineering with DNA. ACS Nano 2023, 17 (17), 16291– 16307, DOI: 10.1021/acsnano.3c06564Google ScholarThere is no corresponding record for this reference.
- 4Macfarlane, R. J.; O’Brien, M. N.; Petrosko, S. H.; Mirkin, C. A. Nucleic Acid-Modified Nanostructures as Programmable Atom Equivalents: Forging a New ″Table of Elements″. Angew. Chem., Int. Ed. 2013, 52 (22), 5688– 5698, DOI: 10.1002/anie.201209336Google ScholarThere is no corresponding record for this reference.
- 5Samanta, D.; Zhou, W.; Ebrahimi, S. B.; Petrosko, S. H.; Mirkin, C. A. Programmable Matter: The Nanoparticle Atom and DNA Bond. Adv. Mater. 2022, 34 (12), 2107875, DOI: 10.1002/adma.202107875Google ScholarThere is no corresponding record for this reference.
- 6Wang, S.; Qin, L.; Yamankurt, G.; Skakuj, K.; Huang, Z.; Chen, P.-C.; Dominguez, D.; Lee, A.; Zhang, B.; Mirkin, C. A. Rational Vaccinology with Spherical Nucleic Acids. Proc. Natl. Acad. Soc. U. S. A. 2019, 116 (21), 10473– 10481, DOI: 10.1073/pnas.1902805116Google ScholarThere is no corresponding record for this reference.
- 7Rosi, N. L.; Giljohann, D. A.; Thaxton, C. S.; Lytton-Jean, A. K. R.; Han, M. S.; Mirkin, C. A. Oligonucleotide-Modified Gold Nanoparticles for Intracellular Gene Regulation. Science 2006, 312 (5776), 1027– 1030, DOI: 10.1126/science.1125559Google ScholarThere is no corresponding record for this reference.
- 8Mirkin, C. A.; Langer, R.; Mrksich, M.; Margolin, A. A.; Petrosko, S. H.; Artzi, N. Blueprints for Better Drugs: The Structural Revolution in Nanomedicine. ACS Nano 2025, 19 (20), 18889– 18901, DOI: 10.1021/acsnano.5c06380Google ScholarThere is no corresponding record for this reference.
- 9Mirkin, C. A.; Mrksich, M.; Artzi, N. The Emerging Era of Structural Nanomedicine. Nat. Rev. Bioeng. 2025, 3 (7), 526– 528, DOI: 10.1038/s44222-025-00306-5Google ScholarThere is no corresponding record for this reference.
- 10Macfarlane, R. J.; Lee, B.; Jones, M. R.; Harris, N.; Schatz, G. C.; Mirkin, C. A. Nanoparticle Superlattice Engineering with DNA. Science 2011, 334 (6053), 204– 208, DOI: 10.1126/science.1210493Google ScholarThere is no corresponding record for this reference.
- 11Park, J.; Evangelopoulos, M.; Vasher, M. K.; Kudruk, S.; Ramani, N.; Mayer, V.; Solivan, A. C.; Lee, A.; Mirkin, C. A. Enhancing Endosomal Escape and Gene Regulation Activity for Spherical Nucleic Acids. Small 2024, 20 (11), 2306902, DOI: 10.1002/smll.202306902Google ScholarThere is no corresponding record for this reference.
- 12Samanta, D.; Ebrahimi, S. B.; Kusmierz, C. D.; Cheng, H. F.; Mirkin, C. A. Protein Spherical Nucleic Acids for Live-Cell Chemical Analysis. J. Am. Chem. Soc. 2020, 142 (31), 13350– 13355, DOI: 10.1021/jacs.0c06866Google ScholarThere is no corresponding record for this reference.
- 13Seferos, D. S.; Giljohann, D. A.; Hill, H. D.; Prigodich, A. E.; Mirkin, C. A. Nano-Flares: Probes for Transfection and mRNA Detection in Living Cells. J. Am. Chem. Soc. 2007, 129 (50), 15477– 15479, DOI: 10.1021/ja0776529Google ScholarThere is no corresponding record for this reference.
- 14Briley, W. E.; Bondy, M. H.; Randeria, P. S.; Dupper, T. J.; Mirkin, C. A. Quantification and Real-Time Tracking of RNA in Live Cells Using Sticky-Flares. Proc. Natl. Acad. Soc. U. S. A. 2015, 112 (31), 9591– 9595, DOI: 10.1073/pnas.1510581112Google ScholarThere is no corresponding record for this reference.
- 15Yamankurt, G.; Stawicki, R. J.; Posadas, D. M.; Nguyen, J. Q.; Carthew, R. W.; Mirkin, C. A. The Effector Mechanism of siRNA Spherical Nucleic Acids. Proc. Natl. Acad. Soc. U. S. A. 2020, 117 (3), 1312– 1320, DOI: 10.1073/pnas.1915907117Google ScholarThere is no corresponding record for this reference.
- 16Li, J.; Mao, X.; Zhao, T.; Fang, W.; Jin, Y.; Liu, M.; Fan, C.; Tian, Y. Tetrahedral DNA Framework-Based Spherical Nucleic Acids for Efficient siRNA Delivery. Angew. Chem., Int. Ed. 2025, 64 (5), e202416988 DOI: 10.1002/anie.202416988Google ScholarThere is no corresponding record for this reference.
- 17Randeria, P. S.; Seeger, M. A.; Wang, X.-Q.; Wilson, H.; Shipp, D.; Mirkin, C. A.; Paller, A. S. siRNA-Based Spherical Nucleic Acids Reverse Impaired Wound Healing in Diabetic Mice by Ganglioside GM3 Synthase Knockdown. Proc. Natl. Acad. Soc. U. S. A. 2015, 112 (18), 5573– 5578, DOI: 10.1073/pnas.1505951112Google ScholarThere is no corresponding record for this reference.
- 18Kumthekar, P.; Ko, C. H.; Paunesku, T.; Dixit, K.; Sonabend, A. M.; Bloch, O.; Tate, M.; Schwartz, M.; Zuckerman, L.; Lezon, R. A First-in-Human Phase 0 Clinical Study of RNA Interference-Based Spherical Nucleic Acids in Patients with Recurrent Glioblastoma. Sci. Transl. Med. 2021, 13 (584), eabb3945 DOI: 10.1126/scitranslmed.abb3945Google ScholarThere is no corresponding record for this reference.
- 19Huang, C.; Han, Z.; Evangelopoulos, M.; Mirkin, C. A. CRISPR Spherical Nucleic Acids. J. Am. Chem. Soc. 2022, 144 (41), 18756– 18760, DOI: 10.1021/jacs.2c07913Google ScholarThere is no corresponding record for this reference.
- 20Han, Z.; Huang, C.; Luo, T.; Mirkin, C. A. A General Genome Editing Strategy Using CRISPR Lipid Nanoparticle Spherical Nucleic Acids. Proc. Natl. Acad. Soc. U. S. A. 2025, 122 (36), e2426094122 DOI: 10.1073/pnas.2426094122Google ScholarThere is no corresponding record for this reference.
- 21Qin, L.; Wang, S.; Dominguez, D.; Long, A.; Chen, S.; Fan, J.; Ahn, J.; Skakuj, K.; Huang, Z.; Lee, A. Development of Spherical Nucleic Acids for Prostate Cancer Immunotherapy. Front. Immunol. 2020, 11, 1333, DOI: 10.3389/fimmu.2020.01333Google ScholarThere is no corresponding record for this reference.
- 22Teplensky, M. H.; Evangelopoulos, M.; Dittmar, J. W.; Forsyth, C. M.; Sinegra, A. J.; Wang, S.; Mirkin, C. A. Multi-Antigen Spherical Nucleic Acid Cancer Vaccines. Nat. Biomed. Eng. 2023, 7 (7), 911– 927, DOI: 10.1038/s41551-022-01000-2Google ScholarThere is no corresponding record for this reference.
- 23Hwang, J.; Dittmar, J. W.; Kang, J.; Ocampo, T.; Evangelopoulos, M.; Han, Z.; Kudruk, S.; Lorch, J.; Mirkin, C. A. DNA Anchoring Strength Directly Correlates with Spherical Nucleic Acid-Based HPV E7 Cancer Vaccine Potency. Nano Lett. 2024, 24 (25), 7629– 7636, DOI: 10.1021/acs.nanolett.4c01392Google ScholarThere is no corresponding record for this reference.
- 24Auyeung, E.; Morris, W.; Mondloch, J. E.; Hupp, J. T.; Farha, O. K.; Mirkin, C. A. Controlling Structure and Porosity in Catalytic Nanoparticle Superlattices with DNA. J. Am. Chem. Soc. 2015, 137 (4), 1658– 1662, DOI: 10.1021/ja512116pGoogle ScholarThere is no corresponding record for this reference.
- 25Sun, L.; Lin, H.; Kohlstedt, K. L.; Schatz, G. C.; Mirkin, C. A. Design Principles for Photonic Crystals Based on Plasmonic Nanoparticle Superlattices. Proc. Natl. Acad. Soc. U. S. A. 2018, 115 (28), 7242– 7247, DOI: 10.1073/pnas.1800106115Google ScholarThere is no corresponding record for this reference.
- 26Lee, S.; Calcaterra, H. A.; Lee, S.; Hadibrata, W.; Lee, B.; Oh, E.; Aydin, K.; Glotzer, S. C.; Mirkin, C. A. Shape Memory in Self-Adapting Colloidal Crystals. Nature 2022, 610 (7933), 674– 679, DOI: 10.1038/s41586-022-05232-9Google ScholarThere is no corresponding record for this reference.
- 27Michelson, A.; Shani, L.; Kahn, J. S.; Redeker, D. C.; Lee, W.-I.; DeOlivares, K. R.; Kisslinger, K.; Tiwale, N.; Yan, H.; Pattammattel, A. Scalable Fabrication of Chip-Integrated 3D-Nanostructured Electronic Devices via DNA-Programmable Assembly. Sci. Adv. 2025, 11 (13), eadt5620 DOI: 10.1126/sciadv.adt5620Google ScholarThere is no corresponding record for this reference.
- 28Daniel, W. L.; Lorch, U.; Mix, S.; Bexon, A. S. A First-in-Human Phase 1 Study of Cavrotolimod, a TLR9 Agonist Spherical Nucleic Acid, in Healthy Participants: Evidence of Immune Activation. Front. Immunol. 2022, 13, 1073777, DOI: 10.3389/fimmu.2022.1073777Google ScholarThere is no corresponding record for this reference.
- 29Milhem, M. M.; Wise-Draper, T. M.; Chandra, S.; Hanna, G. J.; Laux, D. E.; Medina, T. M.; Ansstas, G.; Daud, A.; Kelly, C. M.; O’Day, S. J. Phase 1b/2 Study Evaluating Safety, Efficacy and Immune Effects of TLR9 Agonist Cavrotolimod with Anti-PD-1 Antibodies Among Patients with Advanced Solid Tumors. J. Immunother. Cancer 2025, 13 (7), e011651 DOI: 10.1136/jitc-2025-011651Google ScholarThere is no corresponding record for this reference.
- 30Zhang, C.; Macfarlane, R. J.; Young, K. L.; Choi, C. H. J.; Hao, L.; Auyeung, E.; Liu, G.; Zhou, X.; Mirkin, C. A. A General Approach to DNA-Programmable Atom Equivalents. Nat. Mater. 2013, 12 (8), 741– 746, DOI: 10.1038/nmat3647Google ScholarThere is no corresponding record for this reference.
- 31Mucic, R. C.; Storhoff, J. J.; Mirkin, C. A.; Letsinger, R. L. DNA-Directed Synthesis of Binary Nanoparticle Network Materials. J. Am. Chem. Soc. 1998, 120 (48), 12674– 12675, DOI: 10.1021/ja982721sGoogle ScholarThere is no corresponding record for this reference.
- 32Callmann, C. E.; Vasher, M. K.; Das, A.; Kusmierz, C. D.; Mirkin, C. A. In Vivo Behavior of Ultrasmall Spherical Nucleic Acids. Small 2023, 19 (24), 2300097, DOI: 10.1002/smll.202300097Google ScholarThere is no corresponding record for this reference.
- 33Hill, H. D.; Millstone, J. E.; Banholzer, M. J.; Mirkin, C. A. The Role Radius of Curvature Plays in Thiolated Oligonucleotide Loading on Gold Nanoparticles. ACS Nano 2009, 3 (2), 418– 424, DOI: 10.1021/nn800726eGoogle ScholarThere is no corresponding record for this reference.
- 34PELCO Gold Nanoparticles─Gold Colloids; Ted Pella. https://www.tedpella.com/gold_html/pelco-gold-nanoparticles.aspx#Passive (accessed 2025-03-22).Google ScholarThere is no corresponding record for this reference.
- 35Frens, G. Controlled Nucleation for the Regulation of the Particle Size in Monodisperse Gold Suspensions. Nat. Phys. Sci. 1973, 241 (105), 20– 22, DOI: 10.1038/physci241020a0Google ScholarThere is no corresponding record for this reference.
- 36Hayat, M. A. Colloidal Gold: Principles, Methods, and Applications; Academic Press: New York, 1989.Google ScholarThere is no corresponding record for this reference.
- 37Hill, H. D.; Macfarlane, R. J.; Senesi, A. J.; Lee, B.; Park, S. Y.; Mirkin, C. A. Controlling the Lattice Parameters of Gold Nanoparticle FCC Crystals with Duplex DNA Linkers. Nano Lett. 2008, 8 (8), 2341– 2344, DOI: 10.1021/nl8011787Google ScholarThere is no corresponding record for this reference.
- 38Wang, M. D. Ruler of Life. Nat. Phys. 2021, 17 (8), 976– 976, DOI: 10.1038/s41567-021-01300-5Google ScholarThere is no corresponding record for this reference.
- 39Banga, R. J.; Chernyak, N.; Narayan, S. P.; Nguyen, S. T.; Mirkin, C. A. Liposomal Spherical Nucleic Acids. J. Am. Chem. Soc. 2014, 136 (28), 9866– 9869, DOI: 10.1021/ja504845fGoogle ScholarThere is no corresponding record for this reference.
- 40Yamankurt, G.; Berns, E. J.; Xue, A.; Lee, A.; Bagheri, N.; Mrksich, M.; Mirkin, C. A. Exploration of the Nanomedicine-Design Space with High-Throughput Screening and Machine Learning. Nat. Biomed. Eng. 2019, 3 (4), 318– 327, DOI: 10.1038/s41551-019-0351-1Google ScholarThere is no corresponding record for this reference.
- 41Kusmierz, C. D.; Callmann, C. E.; Kudruk, S.; Distler, M. E.; Mirkin, C. A. Transferrin Aptamers Increase the In Vivo Blood-Brain Barrier Targeting of Protein Spherical Nucleic Acids. Bioconjugate Chem. 2022, 33 (10), 1803– 1810, DOI: 10.1021/acs.bioconjchem.2c00389Google ScholarThere is no corresponding record for this reference.
- 42Ebrahimi, S. B.; Samanta, D.; Kusmierz, C. D.; Mirkin, C. A. Protein Transfection via Spherical Nucleic Acids. Nat. Protoc. 2022, 17 (2), 327– 357, DOI: 10.1038/s41596-021-00642-xGoogle ScholarThere is no corresponding record for this reference.
- 43Girard, M.; Wang, S.; Du, J. S.; Das, A.; Huang, Z.; Dravid, V. P.; Lee, B.; Mirkin, C. A.; Olvera de la Cruz, M. Particle Analogs of Electrons in Colloidal Crystals. Science 2019, 364 (6446), 1174– 1178, DOI: 10.1126/science.aaw8237Google ScholarThere is no corresponding record for this reference.
- 44Daniel, M.-C.; Astruc, D. Gold Nanoparticles: Assembly, Supramolecular Chemistry, Quantum-Size-Related Properties, and Applications toward Biology, Catalysis, and Nanotechnology. Chem. Rev. 2004, 104 (1), 293– 346, DOI: 10.1021/cr030698+Google ScholarThere is no corresponding record for this reference.
- 45Lee, J.-S.; Lytton-Jean, A. K. R.; Hurst, S. J.; Mirkin, C. A. Silver Nanoparticle-Oligonucleotide Conjugates Based on DNA with Triple Cyclic Disulfide Moieties. Nano Lett. 2007, 7 (7), 2112– 2115, DOI: 10.1021/nl071108gGoogle ScholarThere is no corresponding record for this reference.
- 46Xue, C.; Chen, X.; Hurst, S. J.; Mirkin, C. A. Self-Assembled Monolayer Mediated Silica Coating of Silver Triangular Nanoprisms. Adv. Mater. 2007, 19 (22), 4071– 4074, DOI: 10.1002/adma.200701506Google ScholarThere is no corresponding record for this reference.
- 47Young, K. L.; Scott, A. W.; Hao, L.; Mirkin, S. E.; Liu, G.; Mirkin, C. A. Hollow Spherical Nucleic Acids for Intracellular Gene Regulation Based upon Biocompatible Silica Shells. Nano Lett. 2012, 12 (7), 3867– 3871, DOI: 10.1021/nl3020846Google ScholarThere is no corresponding record for this reference.
- 48Cutler, J. I.; Zheng, D.; Xu, X.; Giljohann, D. A.; Mirkin, C. A. Polyvalent Oligonucleotide Iron Oxide Nanoparticle “Click” Conjugates. Nano Lett. 2010, 10 (4), 1477– 1480, DOI: 10.1021/nl100477mGoogle ScholarThere is no corresponding record for this reference.
- 49Mitchell, G. P.; Mirkin, C. A.; Letsinger, R. L. Programmed Assembly of DNA Functionalized Quantum Dots. J. Am. Chem. Soc. 1999, 121 (35), 8122– 8123, DOI: 10.1021/ja991662vGoogle ScholarThere is no corresponding record for this reference.
- 50Sun, D.; Gang, O. DNA-Functionalized Quantum Dots: Fabrication, Structural, and Physicochemical Properties. Langmuir 2013, 29 (23), 7038– 7046, DOI: 10.1021/la4000186Google ScholarThere is no corresponding record for this reference.
- 51Mulvaney, P. Surface Plasmon Spectroscopy of Nanosized Metal Particles. Langmuir 1996, 12 (3), 788– 800, DOI: 10.1021/la9502711Google ScholarThere is no corresponding record for this reference.
- 52Gao, C.; Hu, Y.; Wang, M.; Chi, M.; Yin, Y. Fully Alloyed Ag/Au Nanospheres: Combining the Plasmonic Property of Ag with the Stability of Au. J. Am. Chem. Soc. 2014, 136 (20), 7474– 7479, DOI: 10.1021/ja502890cGoogle ScholarThere is no corresponding record for this reference.
- 53Banga, R. J.; Meckes, B.; Narayan, S. P.; Sprangers, A. J.; Nguyen, S. T.; Mirkin, C. A. Cross-Linked Micellar Spherical Nucleic Acids from Thermoresponsive Templates. J. Am. Chem. Soc. 2017, 139 (12), 4278– 4281, DOI: 10.1021/jacs.6b13359Google ScholarThere is no corresponding record for this reference.
- 54Zhang, C.; Hao, L.; Calabrese, C. M.; Zhou, Y.; Choi, C. H. J.; Xing, H.; Mirkin, C. A. Biodegradable DNA-Brush Block Copolymer Spherical Nucleic Acids Enable Transfection Agent-Free Intracellular Gene Regulation. Small 2015, 11 (40), 5360– 5368, DOI: 10.1002/smll.201501573Google ScholarThere is no corresponding record for this reference.
- 55Chen, H.; Ding, Q.; Li, L.; Wei, P.; Niu, Z.; Kong, T.; Fu, P.; Wang, Y.; Li, J.; Wang, K. Extracellular Vesicle Spherical Nucleic Acids. JACS Au 2024, 4 (6), 2381– 2392, DOI: 10.1021/jacsau.4c00338Google ScholarThere is no corresponding record for this reference.
- 56Zhu, S.; Xing, H.; Gordiichuk, P.; Park, J.; Mirkin, C. A. PLGA Spherical Nucleic Acids. Adv. Mater. 2018, 30 (22), 1707113, DOI: 10.1002/adma.201707113Google ScholarThere is no corresponding record for this reference.
- 57Brodin, J. D.; Sprangers, A. J.; McMillan, J. R.; Mirkin, C. A. DNA-Mediated Cellular Delivery of Functional Enzymes. J. Am. Chem. Soc. 2015, 137 (47), 14838– 14841, DOI: 10.1021/jacs.5b09711Google ScholarThere is no corresponding record for this reference.
- 58Sinegra, A. J.; Evangelopoulos, M.; Park, J.; Huang, Z.; Mirkin, C. A. Lipid Nanoparticle Spherical Nucleic Acids for Intracellular DNA and RNA Delivery. Nano Lett. 2021, 21 (15), 6584– 6591, DOI: 10.1021/acs.nanolett.1c01973Google ScholarThere is no corresponding record for this reference.
- 59Cutler, J. I.; Zhang, K.; Zheng, D.; Auyeung, E.; Prigodich, A. E.; Mirkin, C. A. Polyvalent Nucleic Acid Nanostructures. J. Am. Chem. Soc. 2011, 133 (24), 9254– 9257, DOI: 10.1021/ja203375nGoogle ScholarThere is no corresponding record for this reference.
- 60Auyeung, E.; Cutler, J. I.; Macfarlane, R. J.; Jones, M. R.; Wu, J.; Liu, G.; Zhang, K.; Osberg, K. D.; Mirkin, C. A. Synthetically Programmable Nanoparticle Superlattices Using a Hollow Three-Dimensional Spacer Approach. Nat. Nanotechnol. 2012, 7 (1), 24– 28, DOI: 10.1038/nnano.2011.222Google ScholarThere is no corresponding record for this reference.
- 61Teplensky, M. H.; Distler, M. E.; Kusmierz, C. D.; Evangelopoulos, M.; Gula, H.; Elli, D.; Tomatsidou, A.; Nicolaescu, V.; Gelarden, I.; Yeldandi, A. Spherical Nucleic Acids as an Infectious Disease Vaccine Platform. Proc. Natl. Acad. Sci. U. S. A. 2022, 119 (14), e2119093119 DOI: 10.1073/pnas.2119093119Google ScholarThere is no corresponding record for this reference.
- 62Giljohann, D. A.; Seferos, D. S.; Prigodich, A. E.; Patel, P. C.; Mirkin, C. A. Gene Regulation with Polyvalent siRNA-Nanoparticle Conjugates. J. Am. Chem. Soc. 2009, 131 (6), 2072– 2073, DOI: 10.1021/ja808719pGoogle ScholarThere is no corresponding record for this reference.
- 63Liu, L.; Lu, H.; Shi, R.; Peng, X.-X.; Xiang, Q.; Wang, B.; Wan, Q.-Q.; Sun, Y.; Yang, F.; Zhang, G.-J. Synergy of Peptide-Nucleic Acid and Spherical Nucleic Acid Enabled Quantitative and Specific Detection of Tumor Exosomal MicroRNA. Anal. Chem. 2019, 91 (20), 13198– 13205, DOI: 10.1021/acs.analchem.9b03622Google ScholarThere is no corresponding record for this reference.
- 64Lytton-Jean, A. K. R.; Gibbs-Davis, J. M.; Long, H.; Schatz, G. C.; Mirkin, C. A.; Nguyen, S. T. Highly Cooperative Behavior of Peptide Nucleic Acid-Linked DNA-Modified Gold-Nanoparticle and Comb-Polymer Aggregates. Adv. Mater. 2009, 21 (6), 706– 709, DOI: 10.1002/adma.200801724Google ScholarThere is no corresponding record for this reference.
- 65McKenzie, F.; Faulds, K.; Graham, D. Sequence-Specific DNA Detection Using High-Affinity LNA-Functionalized Gold Nanoparticles. Small 2007, 3 (11), 1866– 1868, DOI: 10.1002/smll.200700225Google ScholarThere is no corresponding record for this reference.
- 66Seferos, D. S.; Giljohann, D. A.; Rosi, N. L.; Mirkin, C. A. Locked Nucleic Acid-Nanoparticle Conjugates. ChemBioChem. 2007, 8 (11), 1230– 1232, DOI: 10.1002/cbic.200700262Google ScholarThere is no corresponding record for this reference.
- 67Dittmar, J. W.; Teplensky, M. H.; Evangelopoulos, M.; Qin, L.; Zhang, B.; Mirkin, C. A. Tuning DNA Dissociation from Spherical Nucleic Acids for Enhanced Immunostimulation. ACS Nano 2023, 17 (18), 17996– 18007, DOI: 10.1021/acsnano.3c04333Google ScholarThere is no corresponding record for this reference.
- 68Zhang, K.; Hao, L.; Hurst, S. J.; Mirkin, C. A. Antibody-Linked Spherical Nucleic Acids for Cellular Targeting. J. Am. Chem. Soc. 2012, 134 (40), 16488– 16491, DOI: 10.1021/ja306854dGoogle ScholarThere is no corresponding record for this reference.
- 69Zhang, W.; Callmann, C. E.; Meckes, B.; Mirkin, C. A. Tumor-Associated Enzyme-Activatable Spherical Nucleic Acids. ACS Nano 2022, 16 (7), 10931– 10942, DOI: 10.1021/acsnano.2c03323Google ScholarThere is no corresponding record for this reference.
- 70Choi, C. H. J.; Hao, L.; Narayan, S. P.; Auyeung, E.; Mirkin, C. A. Mechanism for the Endocytosis of Spherical Nucleic Acid Nanoparticle Conjugates. Proc. Natl. Acad. Sci. U. S. A. 2013, 110 (19), 7625– 7630, DOI: 10.1073/pnas.1305804110Google ScholarThere is no corresponding record for this reference.
- 71Wu, X. A.; Choi, C. H. J.; Zhang, C.; Hao, L.; Mirkin, C. A. Intracellular Fate of Spherical Nucleic Acid Nanoparticle Conjugates. J. Am. Chem. Soc. 2014, 136 (21), 7726– 7733, DOI: 10.1021/ja503010aGoogle ScholarThere is no corresponding record for this reference.
- 72Han, Z.; Guo, A. X.; Luo, T.; Cai, T.; Mirkin, C. A. Biomineralization of Semiconductor Quantum Dots Using DNA-Functionalized Protein Nanoreactors. Sci. Adv. 2025, 11 (19), eadv6906 DOI: 10.1126/sciadv.adv6906Google ScholarThere is no corresponding record for this reference.
- 73Hsiao, J. C.; Buryska, T.; Kim, E.; Howes, P. D.; deMello, A. J. Tuning DNA-Nanoparticle Conjugate Properties Allows Modulation of Nuclease Activity. Nanoscale 2021, 13 (9), 4956– 4970, DOI: 10.1039/D0NR08668AGoogle ScholarThere is no corresponding record for this reference.
- 74Seferos, D. S.; Prigodich, A. E.; Giljohann, D. A.; Patel, P. C.; Mirkin, C. A. Polyvalent DNA Nanoparticle Conjugates Stabilize Nucleic Acids. Nano Lett. 2009, 9 (1), 308– 311, DOI: 10.1021/nl802958fGoogle ScholarThere is no corresponding record for this reference.
- 75Kyriazi, M.-E.; El-Sagheer, A. H.; Medintz, I. L.; Brown, T.; Kanaras, A. G. An Investigation into the Resistance of Spherical Nucleic Acids against DNA Enzymatic Degradation. Bioconjugate Chem. 2022, 33 (1), 219– 225, DOI: 10.1021/acs.bioconjchem.1c00540Google ScholarThere is no corresponding record for this reference.
- 76Giljohann, D. A.; Seferos, D. S.; Patel, P. C.; Millstone, J. E.; Rosi, N. L.; Mirkin, C. A. Oligonucleotide Loading Determines Cellular Uptake of DNA-Modified Gold Nanoparticles. Nano Lett. 2007, 7 (12), 3818– 3821, DOI: 10.1021/nl072471qGoogle ScholarThere is no corresponding record for this reference.
- 77Hurst, S. J.; Lytton-Jean, A. K. R.; Mirkin, C. A. Maximizing DNA Loading on a Range of Gold Nanoparticle Sizes. Anal. Chem. 2006, 78 (24), 8313– 8318, DOI: 10.1021/ac0613582Google ScholarThere is no corresponding record for this reference.
- 78Liu, B.; Liu, J. Freezing-Driven DNA Adsorption on Gold Nanoparticles: Tolerating Extremely Low Salt Concentration but Requiring High DNA Concentration. Langmuir 2019, 35 (19), 6476– 6482, DOI: 10.1021/acs.langmuir.9b00746Google ScholarThere is no corresponding record for this reference.
- 79Hao, Y.; Li, Y.; Song, L.; Deng, Z. Flash Synthesis of Spherical Nucleic Acids with Record DNA Density. J. Am. Chem. Soc. 2021, 143 (8), 3065– 3069, DOI: 10.1021/jacs.1c00568Google ScholarThere is no corresponding record for this reference.
- 80Ye, Y.; Hao, Y.; Ye, M.; Song, X.; Deng, Z. Evaporative Drying: A General and Readily Scalable Route to Spherical Nucleic Acids with Quantitative, Fully Tunable, and Record-High DNA Loading. Small 2022, 18 (24), 2202458, DOI: 10.1002/smll.202202458Google ScholarThere is no corresponding record for this reference.
- 81Liu, B.; Liu, J. Freezing Directed Construction of Bio/Nano Interfaces: Reagentless Conjugation, Denser Spherical Nucleic Acids, and Better Nanoflares. J. Am. Chem. Soc. 2017, 139 (28), 9471– 9474, DOI: 10.1021/jacs.7b04885Google ScholarThere is no corresponding record for this reference.
- 82Eller, M. J.; Chandra, K.; Coughlin, E. E.; Odom, T. W.; Schweikert, E. A. Label Free Particle-by-Particle Quantification of DNA Loading on Sorted Gold Nanostars. Anal. Chem. 2019, 91 (9), 5566– 5572, DOI: 10.1021/acs.analchem.8b03715Google ScholarThere is no corresponding record for this reference.
- 83Lee, J.; Lee, S. Non-Invasive, Reliable, and Fast Quantification of DNA Loading on Gold Nanoparticles by a One-Step Optical Measurement. Anal. Chem. 2023, 95 (3), 1856– 1866, DOI: 10.1021/acs.analchem.2c03378Google ScholarThere is no corresponding record for this reference.
- 84Lee, S.; Pandit, S.; Gilman, G.; Bhattacharya, A.; Samanta, D. Label-Free Quantification of DNA Loading on Centrifugation-Resistant Spherical Nucleic Acids. Anal. Chem. 2025, 97 (23), 12267– 12275, DOI: 10.1021/acs.analchem.5c01112Google ScholarThere is no corresponding record for this reference.
- 85Li, H.; Zhang, B.; Lu, X.; Tan, X.; Jia, F.; Xiao, Y.; Cheng, Z.; Li, Y.; Silva, D. O.; Schrekker, H. S. Molecular Spherical Nucleic Acids. Proc. Natl. Acad. Sci. U. S. A. 2018, 115 (17), 4340– 4344, DOI: 10.1073/pnas.1801836115Google ScholarThere is no corresponding record for this reference.
- 86Cavaliere, J. P.; Forsyth, C. M.; Guo, A. X.; Kang, J.; Ngo, K. H.; Mirkin, C. A. Amphiphilic Polymeric Micelle Spherical Nucleic Acids (SNAs) as Drug Delivery Vehicles. Bioconjugate Chem. 2026, 37, 83, DOI: 10.1021/acs.bioconjchem.5c00507Google ScholarThere is no corresponding record for this reference.
- 87Distler, M. E.; Teplensky, M. H.; Bujold, K. E.; Kusmierz, C. D.; Evangelopoulos, M.; Mirkin, C. A. DNA Dendrons as Agents for Intracellular Delivery. J. Am. Chem. Soc. 2021, 143 (34), 13513– 13518, DOI: 10.1021/jacs.1c07240Google ScholarThere is no corresponding record for this reference.
- 88Chinen, A. B.; Guan, C. M.; Ko, C. H.; Mirkin, C. A. The Impact of Protein Corona Formation on the Macrophage Cellular Uptake and Biodistribution of Spherical Nucleic Acids. Small 2017, 13 (16), 1603847, DOI: 10.1002/smll.201603847Google ScholarThere is no corresponding record for this reference.
- 89Mahajan, A. S.; Stegh, A. H. Spherical Nucleic Acids as Precision Therapeutics for the Treatment of Cancer-From Bench to Bedside. Cancers 2022, 14 (7), 1615, DOI: 10.3390/cancers14071615Google ScholarThere is no corresponding record for this reference.
- 90Yan, R.; Chen, J.; Wang, J.; Rao, J.; Du, X.; Liu, Y.; Zhang, L.; Qiu, L.; Liu, B.; Zhao, Y.-D. A NanoFlare-Based Strategy for In Situ Tumor Margin Demarcation and Neoadjuvant Gene/Photothermal Therapy. Small 2018, 14 (50), 1802745, DOI: 10.1002/smll.201802745Google ScholarThere is no corresponding record for this reference.
- 91Radovic-Moreno, A. F.; Chernyak, N.; Mader, C. C.; Nallagatla, S.; Kang, R. S.; Hao, L.; Walker, D. A.; Halo, T. L.; Merkel, T. J.; Rische, C. H. Immunomodulatory Spherical Nucleic Acids. Proc. Natl. Acad. Sci. U. S. A. 2015, 112 (13), 3892– 3897, DOI: 10.1073/pnas.1502850112Google ScholarThere is no corresponding record for this reference.
- 92O’Brien, M. N.; Girard, M.; Lin, H.-X.; Millan, J. A.; Olvera de la Cruz, M.; Lee, B.; Mirkin, C. A. Exploring the Zone of Anisotropy and Broken Symmetries in DNA-Mediated Nanoparticle Crystallization. Proc. Natl. Acad. Sci. U. S. A. 2016, 113 (38), 10485– 10490, DOI: 10.1073/pnas.1611808113Google ScholarThere is no corresponding record for this reference.
- 93Jones, M. R.; Macfarlane, R. J.; Lee, B.; Zhang, J.; Young, K. L.; Senesi, A. J.; Mirkin, C. A. DNA-Nanoparticle Superlattices Formed from Anisotropic Building Blocks. Nat. Mater. 2010, 9 (11), 913– 917, DOI: 10.1038/nmat2870Google ScholarThere is no corresponding record for this reference.
- 94O’Brien, M. N.; Lin, H.-X.; Girard, M.; Olvera de la Cruz, M.; Mirkin, C. A. Programming Colloidal Crystal Habit with Anisotropic Nanoparticle Building Blocks and DNA Bonds. J. Am. Chem. Soc. 2016, 138 (44), 14562– 14565, DOI: 10.1021/jacs.6b09704Google ScholarThere is no corresponding record for this reference.
- 95O’Brien, M. N.; Jones, M. R.; Lee, B.; Mirkin, C. A. Anisotropic Nanoparticle Complementarity in DNA-Mediated Co-Crystallization. Nat. Mater. 2015, 14 (8), 833– 839, DOI: 10.1038/nmat4293Google ScholarThere is no corresponding record for this reference.
- 96Chang, C.-C.; Wu, H.-L.; Kuo, C.-H.; Huang, M. H. Hydrothermal Synthesis of Monodispersed Octahedral Gold Nanocrystals with Five Different Size Ranges and Their Self-Assembled Structures. Chem. Mater. 2008, 20 (24), 7570– 7574, DOI: 10.1021/cm8021984Google ScholarThere is no corresponding record for this reference.
- 97Jin, R.; Cao, Y.; Mirkin, C. A.; Kelly, K. L.; Schatz, G. C.; Zheng, J. G. Photoinduced Conversion of Silver Nanospheres to Nanoprisms. Science 2001, 294 (5548), 1901– 1903, DOI: 10.1126/science.1066541Google ScholarThere is no corresponding record for this reference.
- 98Jana, N. R.; Gearheart, L.; Murphy, C. J. Wet Chemical Synthesis of High Aspect Ratio Cylindrical Gold Nanorods. J. Phys. Chem. B 2001, 105 (19), 4065– 4067, DOI: 10.1021/jp0107964Google ScholarThere is no corresponding record for this reference.
- 99O’Brien, M. N.; Jones, M. R.; Brown, K. A.; Mirkin, C. A. Universal Noble Metal Nanoparticle Seeds Realized Through Iterative Reductive Growth and Oxidative Dissolution Reactions. J. Am. Chem. Soc. 2014, 136 (21), 7603– 7606, DOI: 10.1021/ja503509kGoogle ScholarThere is no corresponding record for this reference.
- 100Akbulut, O.; Mace, C. R.; Martinez, R. V.; Kumar, A. A.; Nie, Z.; Patton, M. R.; Whitesides, G. M. Separation of Nanoparticles in Aqueous Multiphase Systems through Centrifugation. Nano Lett. 2012, 12 (8), 4060– 4064, DOI: 10.1021/nl301452xGoogle ScholarThere is no corresponding record for this reference.
- 101Jana, N. R. Nanorod Shape Separation Using Surfactant Assisted Self-Assembly. Chem. Commun. 2003, (15), 1950– 1951, DOI: 10.1039/b303103aGoogle ScholarThere is no corresponding record for this reference.
- 102Scarabelli, L.; Coronado-Puchau, M.; Giner-Casares, J. J.; Langer, J.; Liz-Marzán, L. M. Monodisperse Gold Nanotriangles: Size Control, Large-Scale Self-Assembly, and Performance in Surface-Enhanced Raman Scattering. ACS Nano 2014, 8 (6), 5833– 5842, DOI: 10.1021/nn500727wGoogle ScholarThere is no corresponding record for this reference.
- 103Cheng, Z.; Jones, M. R. Assembly of Planar Chiral Superlattices from Achiral Building Blocks. Nat. Commun. 2022, 13 (1), 4207, DOI: 10.1038/s41467-022-31868-2Google ScholarThere is no corresponding record for this reference.
- 104Millstone, J. E.; Georganopoulou, D. G.; Xu, X.; Wei, W.; Li, S.; Mirkin, C. A. DNA-Gold Triangular Nanoprism Conjugates. Small 2008, 4 (12), 2176– 2180, DOI: 10.1002/smll.200800931Google ScholarThere is no corresponding record for this reference.
- 105Zhou, W.; Li, Y.; Je, K.; Vo, T.; Lin, H.; Partridge, B. E.; Huang, Z.; Glotzer, S. C.; Mirkin, C. A. Space-Tiled Colloidal Crystals from DNA-Forced Shape-Complementary Polyhedra Pairing. Science 2024, 383 (6680), 312– 319, DOI: 10.1126/science.adj1021Google ScholarThere is no corresponding record for this reference.
- 106Li, Y.; Lin, H.; Zhou, W.; Sun, L.; Samanta, D.; Mirkin, C. A. Corner-, Edge-, and Facet-Controlled Growth of Nanocrystals. Sci. Adv. 2021, 7 (3), eabf1410 DOI: 10.1126/sciadv.abf1410Google ScholarThere is no corresponding record for this reference.
- 107Brodin, J. D.; Auyeung, E.; Mirkin, C. A. DNA-Mediated Engineering of Multicomponent Enzyme Crystals. Proc. Nat. Acad. Sci. U. S. A. 2015, 112 (15), 4564– 4569, DOI: 10.1073/pnas.1503533112Google ScholarThere is no corresponding record for this reference.
- 108Wang, Y.; Wang, Y.; Breed, D. R.; Manoharan, V. N.; Feng, L.; Hollingsworth, A. D.; Weck, M.; Pine, D. J. Colloids with Valence and Specific Directional Bonding. Nature 2012, 491 (7422), 51– 55, DOI: 10.1038/nature11564Google ScholarThere is no corresponding record for this reference.
- 109Chen, G.; Gibson, K. J.; Liu, D.; Rees, H. C.; Lee, J.-H.; Xia, W.; Lin, R.; Xin, H. L.; Gang, O.; Weizmann, Y. Regioselective Surface Encoding of Nanoparticles for Programmable Self-Assembly. Nat. Mater. 2019, 18 (2), 169– 174, DOI: 10.1038/s41563-018-0231-1Google ScholarThere is no corresponding record for this reference.
- 110Tian, Y.; Zhang, Y.; Wang, T.; Xin, H. L.; Li, H.; Gang, O. Lattice Engineering through Nanoparticle-DNA Frameworks. Nat. Mater. 2016, 15 (6), 654– 661, DOI: 10.1038/nmat4571Google ScholarThere is no corresponding record for this reference.
- 111Papavassiliou, G. C. Optical Properties of Small Inorganic and Organic Metal Particles. Prog. Solid State Chem. 1979, 12 (3), 185– 271, DOI: 10.1016/0079-6786(79)90001-3Google ScholarThere is no corresponding record for this reference.
- 112Elghanian, R.; Storhoff, J. J.; Mucic, R. C.; Letsinger, R. L.; Mirkin, C. A. Selective Colorimetric Detection of Polynucleotides Based on the Distance-Dependent Optical Properties of Gold Nanoparticles. Science 1997, 277 (5329), 1078– 1081, DOI: 10.1126/science.277.5329.1078Google ScholarThere is no corresponding record for this reference.
- 113Storhoff, J. J.; Lazarides, A. A.; Mucic, R. C.; Mirkin, C. A.; Letsinger, R. L.; Schatz, G. C. What Controls the Optical Properties of DNA-Linked Gold Nanoparticle Assemblies?. J. Am. Chem. Soc. 2000, 122 (19), 4640– 4650, DOI: 10.1021/ja993825lGoogle ScholarThere is no corresponding record for this reference.
- 114Xia, C.; Cheng, H.; Hou, X.; Zhang, Y.; Zhou, X.; Yan, Q.; Cao, S. Spherical Nucleic Acids for Biomedical Applications. Adv. Sens. Energy Mater. 2024, 3 (4), 100117, DOI: 10.1016/j.asems.2024.100117Google ScholarThere is no corresponding record for this reference.
- 115Lee, J.-S.; Han, M. S.; Mirkin, C. A. Colorimetric Detection of Mercuric Ion (Hg2+) in Aqueous Media Using DNA-Functionalized Gold Nanoparticles. Angew. Chem., Int. Ed. 2007, 46 (22), 4093– 4096, DOI: 10.1002/anie.200700269Google ScholarThere is no corresponding record for this reference.
- 116Lee, J.-S.; Mirkin, C. A. Chip-Based Scanometric Detection of Mercuric Ion Using DNA-Functionalized Gold Nanoparticles. Anal. Chem. 2008, 80 (17), 6805– 6808, DOI: 10.1021/ac801046aGoogle ScholarThere is no corresponding record for this reference.
- 117Xu, X.; Daniel, W. L.; Wei, W.; Mirkin, C. A. Colorimetric Cu2+ Detection Using DNA-Modified Gold-Nanoparticle Aggregates as Probes and Click Chemistry. Small 2010, 6 (5), 623– 626, DOI: 10.1002/smll.200901691Google ScholarThere is no corresponding record for this reference.
- 118Han, M. S.; Lytton-Jean, A. K. R.; Oh, B.-K.; Heo, J.; Mirkin, C. A. Colorimetric Screening of DNA-Binding Molecules with Gold Nanoparticle Probes. Angew. Chem., Int. Ed. 2006, 45 (11), 1807– 1810, DOI: 10.1002/anie.200504277Google ScholarThere is no corresponding record for this reference.
- 119Nam, J.-M.; Thaxton, C. S.; Mirkin, C. A. Nanoparticle-Based Bio-Bar Codes for the Ultrasensitive Detection of Proteins. Science 2003, 301 (5641), 1884– 1886, DOI: 10.1126/science.1088755Google ScholarThere is no corresponding record for this reference.
- 120Pavlov, V.; Xiao, Y.; Shlyahovsky, B.; Willner, I. Aptamer-Functionalized Au Nanoparticles for the Amplified Optical Detection of Thrombin. J. Am. Chem. Soc. 2004, 126 (38), 11768– 11769, DOI: 10.1021/ja046970uGoogle ScholarThere is no corresponding record for this reference.
- 121Kim, D.; Daniel, W. L.; Mirkin, C. A. Microarray-Based Multiplexed Scanometric Immunoassay for Protein Cancer Markers Using Gold Nanoparticle Probes. Anal. Chem. 2009, 81 (21), 9183– 9187, DOI: 10.1021/ac9018389Google ScholarThere is no corresponding record for this reference.
- 122Xu, X.; Han, M. S.; Mirkin, C. A. A Gold-Nanoparticle-Based Real-Time Colorimetric Screening Method for Endonuclease Activity and Inhibition. Angew. Chem., Int. Ed. 2007, 46 (19), 3468– 3470, DOI: 10.1002/anie.200605249Google ScholarThere is no corresponding record for this reference.
- 123Zheng, G.; Daniel, W. L.; Mirkin, C. A. A New Approach to Amplified Telomerase Detection with Polyvalent Oligonucleotide Nanoparticle Conjugates. J. Am. Chem. Soc. 2008, 130 (30), 9644– 9645, DOI: 10.1021/ja803035pGoogle ScholarThere is no corresponding record for this reference.
- 124Chen, L.; Yuan, A.; Zhang, D.; Xie, W.; Peng, H. Fluorescence and Colorimetric Analysis of β-Estradiol Based on Aptamer Assembled Spherical Nucleic Acids. Anal. Methods 2024, 16 (37), 6356– 6363, DOI: 10.1039/D4AY01283FGoogle ScholarThere is no corresponding record for this reference.
- 125Storhoff, J. J.; Elghanian, R.; Mucic, R. C.; Mirkin, C. A.; Letsinger, R. L. One-Pot Colorimetric Differentiation of Polynucleotides with Single Base Imperfections Using Gold Nanoparticle Probes. J. Am. Chem. Soc. 1998, 120 (9), 1959– 1964, DOI: 10.1021/ja972332iGoogle ScholarThere is no corresponding record for this reference.
- 126Taton, T. A.; Mirkin, C. A.; Letsinger, R. L. Scanometric DNA Array Detection with Nanoparticle Probes. Science 2000, 289 (5485), 1757– 1760, DOI: 10.1126/science.289.5485.1757Google ScholarThere is no corresponding record for this reference.
- 127Alhasan, A. H.; Kim, D. Y.; Daniel, W. L.; Watson, E.; Meeks, J. J.; Thaxton, C. S.; Mirkin, C. A. Scanometric MicroRNA Array Profiling of Prostate Cancer Markers Using Spherical Nucleic Acid-Gold Nanoparticle Conjugates. Anal. Chem. 2012, 84 (9), 4153– 4160, DOI: 10.1021/ac3004055Google ScholarThere is no corresponding record for this reference.
- 128The Verigene System; Diasorin. https://us.diasorin.com/en/molecular-diagnostics/tools/verigene-system?f%5B0%5D=mdx_disease%3A249 (accessed 2025-03-24).Google ScholarThere is no corresponding record for this reference.
- 129Hurst, S. J.; Hill, H. D.; Mirkin, C. A. “Three-Dimensional Hybridization” with Polyvalent DNA-Gold Nanoparticle Conjugates. J. Am. Chem. Soc. 2008, 130 (36), 12192– 12200, DOI: 10.1021/ja804266jGoogle ScholarThere is no corresponding record for this reference.
- 130Lytton-Jean, A. K. R.; Mirkin, C. A. A Thermodynamic Investigation into the Binding Properties of DNA Functionalized Gold Nanoparticle Probes and Molecular Fluorophore Probes. J. Am. Chem. Soc. 2005, 127 (37), 12754– 12755, DOI: 10.1021/ja052255oGoogle ScholarThere is no corresponding record for this reference.
- 131Fong, L.-K.; Wang, Z.; Schatz, G. C.; Luijten, E.; Mirkin, C. A. The Role of Structural Enthalpy in Spherical Nucleic Acid Hybridization. J. Am. Chem. Soc. 2018, 140 (20), 6226– 6230, DOI: 10.1021/jacs.8b03459Google ScholarThere is no corresponding record for this reference.
- 132Randeria, P. S.; Jones, M. R.; Kohlstedt, K. L.; Banga, R. J.; Olvera de la Cruz, M.; Schatz, G. C.; Mirkin, C. A. What Controls the Hybridization Thermodynamics of Spherical Nucleic Acids?. J. Am. Chem. Soc. 2015, 137 (10), 3486– 3489, DOI: 10.1021/jacs.5b00670Google ScholarThere is no corresponding record for this reference.
- 133Wang, X.; Huang, K.; Cheng, N.; Liu, J. Thermal Drying Synthesized Density-Tunable and High Stability Nonthiolated Spherical Nucleic Acids for Split Aptamer and Lateral Flow Biosensors. Biosens. Bioelectron. 2025, 289, 117863, DOI: 10.1016/j.bios.2025.117863Google ScholarThere is no corresponding record for this reference.
- 134Nam, J.-M.; Park, S.-J.; Mirkin, C. A. Bio-Barcodes Based on Oligonucleotide-Modified Nanoparticles. J. Am. Chem. Soc. 2002, 124 (15), 3820– 3821, DOI: 10.1021/ja0178766Google ScholarThere is no corresponding record for this reference.
- 135Nam, J.-M.; Stoeva, S. I.; Mirkin, C. A. Bio-Bar-Code-Based DNA Detection with PCR-like Sensitivity. J. Am. Chem. Soc. 2004, 126 (19), 5932– 5933, DOI: 10.1021/ja049384+Google ScholarThere is no corresponding record for this reference.
- 136Thaxton, C. S.; Hill, H. D.; Georganopoulou, D. G.; Stoeva, S. I.; Mirkin, C. A. A Bio-Bar-Code Assay Based upon Dithiothreitol-Induced Oligonucleotide Release. Anal. Chem. 2005, 77 (24), 8174– 8178, DOI: 10.1021/ac0514265Google ScholarThere is no corresponding record for this reference.
- 137Hill, H. D.; Mirkin, C. A. The Bio-Barcode Assay for the Detection of Protein and Nucleic Acid Targets Using DTT-Induced Ligand Exchange. Nat. Protoc. 2006, 1 (1), 324– 336, DOI: 10.1038/nprot.2006.51Google ScholarThere is no corresponding record for this reference.
- 138Goluch, E. D.; Savka, S. I.; Georganopoulou, D. G.; Nam, J. M.; Shaikh, K. A.; Ryu, K. S.; Chiesl, T. N.; Barron, A. E.; Mirkin, C. A.; Liu, C. Chip-Based High-Sensitivity Detection of Multiple Disease Biomarkers. 19th IEEE International Conference on Micro Electro Mechanical Systems, Istanbul, Turkey; IEEE, 2006; pp 478– 481. DOI: 10.1109/memsys.2006.1627840 .Google ScholarThere is no corresponding record for this reference.
- 139Stoeva, S. I.; Lee, J.-S.; Thaxton, C. S.; Mirkin, C. A. Multiplexed DNA Detection with Biobarcoded Nanoparticle Probes. Angew. Chem., Int. Ed. 2006, 45 (20), 3303– 3306, DOI: 10.1002/anie.200600124Google ScholarThere is no corresponding record for this reference.
- 140Stoeva, S. I.; Lee, J.-S.; Smith, J. E.; Rosen, S. T.; Mirkin, C. A. Multiplexed Detection of Protein Cancer Markers with Biobarcoded Nanoparticle Probes. J. Am. Chem. Soc. 2006, 128 (26), 8378– 8379, DOI: 10.1021/ja0613106Google ScholarThere is no corresponding record for this reference.
- 141Thaxton, C. S.; Elghanian, R.; Thomas, A. D.; Stoeva, S. I.; Lee, J.-S.; Smith, N. D.; Schaeffer, A. J.; Klocker, H.; Horninger, W.; Bartsch, G. Nanoparticle-Based Bio-Barcode Assay Redefines “Undetectable” PSA and Biochemical Recurrence after Radical Prostatectomy. Proc. Natl. Acad. Sci. U. S. A. 2009, 106 (44), 18437– 18442, DOI: 10.1073/pnas.0904719106Google ScholarThere is no corresponding record for this reference.
- 142Lee, J. H.; Wang, Z.; Liu, J.; Lu, Y. Highly Sensitive and Selective Colorimetric Sensors for Uranyl (UO22+): Development and Comparison of Labeled and Label-Free DNAzyme-Gold Nanoparticle Systems. J. Am. Chem. Soc. 2008, 130 (43), 14217– 14226, DOI: 10.1021/ja803607zGoogle ScholarThere is no corresponding record for this reference.
- 143Wu, P.; Hwang, K.; Lan, T.; Lu, Y. A DNAzyme-Gold Nanoparticle Probe for Uranyl Ion in Living Cells. J. Am. Chem. Soc. 2013, 135 (14), 5254– 5257, DOI: 10.1021/ja400150vGoogle ScholarThere is no corresponding record for this reference.
- 144Li, Y.; Liu, M.-L.; Liang, W.-B.; Zhuo, Y.; He, X.-J. Spherical Nucleic Acid Enzyme Programmed Network to Accelerate CRISPR Assays for Electrochemiluminescence Biosensing Applications. Biosens. Bioelectron. 2023, 238, 115589, DOI: 10.1016/j.bios.2023.115589Google ScholarThere is no corresponding record for this reference.
- 145Yang, X.; Yuan, L.; Xu, Y.; He, B. Target-Catalyzed Self-Assembled Spherical G-Quadruplex/Hemin DNAzymes for Highly Sensitive Colorimetric Detection of MicroRNA in Serum. Anal. Chim. Acta 2023, 1247, 340879, DOI: 10.1016/j.aca.2023.340879Google ScholarThere is no corresponding record for this reference.
- 146Sun, Y.; Shi, L.; Wang, Q.; Mi, L.; Li, T. Spherical Nucleic Acid Enzyme (SNAzyme) Boosted Chemiluminescence miRNA Imaging Using a Smartphone. Anal. Chem. 2019, 91 (5), 3652– 3658, DOI: 10.1021/acs.analchem.8b05696Google ScholarThere is no corresponding record for this reference.
- 147Shi, L.; Sun, Y.; Mi, L.; Li, T. Target-Catalyzed Self-Growing Spherical Nucleic Acid Enzyme (SNAzyme) as a Double Amplifier for Ultrasensitive Chemiluminescence MicroRNA Detection. ACS Sens. 2019, 4 (12), 3219– 3226, DOI: 10.1021/acssensors.9b01655Google ScholarThere is no corresponding record for this reference.
- 148Shi, L.; Liu, C.; Wang, H.; Zheng, J.; Wang, Q.; Shi, L.; Li, T. Framework and Spherical Nucleic Acids Synergistically Enhanced Electrochemiluminescence Nanosensors for Rapidly Diagnosing Acute Myocardial Infarction Based on Circulating MicroRNA Levels. Anal. Chem. 2022, 94 (41), 14394– 14401, DOI: 10.1021/acs.analchem.2c03144Google ScholarThere is no corresponding record for this reference.
- 149Wang, H.; Shi, L.; Wang, Q.; Shi, L.; Li, T. Robust Noncovalent Spherical Nucleic Acid Enzymes (SNAzymes) for Ultrasensitive Unamplified Electrochemiluminescence Detection of Endogenous Myocardial MicroRNAs. Biosens. Bioelectron. 2023, 241, 115687, DOI: 10.1016/j.bios.2023.115687Google ScholarThere is no corresponding record for this reference.
- 150Qu, S.; Shi, L.; Li, H.; Li, T. A Label-Free Silicon-Based Spherical Nucleic Acid Enzyme (SNAzyme) for Ultrasensitive Chemiluminescence Detection of Acute Myocardial Infarction-Related Nucleic Acids. Sens. Diagn. 2023, 2 (1), 194– 202, DOI: 10.1039/D2SD00170EGoogle ScholarThere is no corresponding record for this reference.
- 151Wang, K.; Zhu, W.; Li, X.; Ling, G.; Zhang, P. Programmable DNAzyme Walkers on Gold Nanoplatforms: From Rational Design to Bioimaging/Biosensing Breakthroughs. Acta Biomater. 2025, 206, 74– 87, DOI: 10.1016/j.actbio.2025.09.020Google ScholarThere is no corresponding record for this reference.
- 152Zhang, K.; Cutler, J. I.; Zhang, J.; Zheng, D.; Auyeung, E.; Mirkin, C. A. Nanopod Formation through Gold Nanoparticle Templated and Catalyzed Cross-linking of Polymers Bearing Pendant Propargyl Ethers. J. Am. Chem. Soc. 2010, 132 (43), 15151– 15153, DOI: 10.1021/ja107224sGoogle ScholarThere is no corresponding record for this reference.
- 153Zhang, K.; Zheng, D.; Hao, L.; Cutler, J. I.; Auyeung, E.; Mirkin, C. A. ImmunoPods: Polymer Shells with Native Antibody Cross-Links. Angew. Chem., Int. Ed. 2012, 51 (5), 1169– 1172, DOI: 10.1002/anie.201106313Google ScholarThere is no corresponding record for this reference.
- 154Massich, M. D.; Giljohann, D. A.; Seferos, D. S.; Ludlow, L. E.; Horvath, C. M.; Mirkin, C. A. Regulating Immune Response Using Polyvalent Nucleic Acid-Gold Nanoparticle Conjugates. Mol. Pharmaceutics 2009, 6 (6), 1934– 1940, DOI: 10.1021/mp900172mGoogle ScholarThere is no corresponding record for this reference.
- 155Massich, M. D.; Giljohann, D. A.; Schmucker, A. L.; Patel, P. C.; Mirkin, C. A. Cellular Response of Polyvalent Oligonucleotide-Gold Nanoparticle Conjugates. ACS Nano 2010, 4 (10), 5641– 5646, DOI: 10.1021/nn102228sGoogle ScholarThere is no corresponding record for this reference.
- 156Zhao, J.; He, J.; Ding, X.; Zhou, Y.; Liu, M.; Chen, X.; Quan, W.; Hua, D.; Tong, J.; Li, J. DENV Peptides Delivered as Spherical Nucleic Acid Constructs Enhance Antigen Presentation and Immunogenicity in Vitro and in Vivo. Int. J. Nanomedicine 2024, 19, 9757– 9770, DOI: 10.2147/IJN.S467427Google ScholarThere is no corresponding record for this reference.
- 157Sparwasser, T.; Hültner, L.; Koch, E. S.; Luz, A.; Lipford, G. B.; Wagner, H. Immunostimulatory CpG-Oligodeoxynucleotides Cause Extramedullary Murine Hemopoiesis. J. Immunol. 1999, 162 (4), 2368– 2374, DOI: 10.4049/jimmunol.162.4.2368Google ScholarThere is no corresponding record for this reference.
- 158Sparwasser, T.; Miethke, T.; Lipford, G.; Borschert, K.; Häcker, H.; Heeg, K.; Wagner, H. Bacterial DNA Causes Septic Shock. Nature 1997, 386 (6623), 336– 337, DOI: 10.1038/386336a0Google ScholarThere is no corresponding record for this reference.
- 159Ferrer, J. R.; Sinegra, A. J.; Ivancic, D.; Yeap, X. Y.; Qiu, L.; Wang, J.-J.; Zhang, Z. J.; Wertheim, J. A.; Mirkin, C. A. Structure-Dependent Biodistribution of Liposomal Spherical Nucleic Acids. ACS Nano 2020, 14 (2), 1682– 1693, DOI: 10.1021/acsnano.9b07254Google ScholarThere is no corresponding record for this reference.
- 160Asohan, J.; Fakih, H. H.; Das, T.; Sleiman, H. F. Control of the Assembly and Disassembly of Spherical Nucleic Acids Is Critical for Enhanced Gene Silencing. ACS Nano 2024, 18 (5), 3996– 4007, DOI: 10.1021/acsnano.3c05940Google ScholarThere is no corresponding record for this reference.
- 161Meckes, B.; Banga, R. J.; Nguyen, S. T.; Mirkin, C. A. Enhancing the Stability and Immunomodulatory Activity of Liposomal Spherical Nucleic Acids through Lipid-Tail DNA Modifications. Small 2018, 14 (5), 1702909, DOI: 10.1002/smll.201702909Google ScholarThere is no corresponding record for this reference.
- 162Callmann, C. E.; Kusmierz, C. D.; Dittmar, J. W.; Broger, L.; Mirkin, C. A. Impact of Liposomal Spherical Nucleic Acid Structure on Immunotherapeutic Function. ACS Cent. Sci. 2021, 7 (5), 892– 899, DOI: 10.1021/acscentsci.1c00181Google ScholarThere is no corresponding record for this reference.
- 163Huang, Z. N.; Cole, L. E.; Callmann, C. E.; Wang, S.; Mirkin, C. A. Sequence Multiplicity within Spherical Nucleic Acids. ACS Nano 2020, 14 (1), 1084– 1092, DOI: 10.1021/acsnano.9b08750Google ScholarThere is no corresponding record for this reference.
- 164Liu, H.; Moynihan, K. D.; Zheng, Y.; Szeto, G. L.; Li, A. V.; Huang, B.; Van Egeren, D. S.; Park, C.; Irvine, D. J. Structure-Based Programming of Lymph-Node Targeting in Molecular Vaccines. Nature 2014, 507 (7493), 519– 522, DOI: 10.1038/nature12978Google ScholarThere is no corresponding record for this reference.
- 165Smith, A. A. A.; Gale, E. C.; Roth, G. A.; Maikawa, C. L.; Correa, S.; Yu, A. C.; Appel, E. A. Nanoparticles Presenting Potent TLR7/8 Agonists Enhance Anti-PD-L1 Immunotherapy in Cancer Treatment. Biomacromolecules 2020, 21 (9), 3704– 3712, DOI: 10.1021/acs.biomac.0c00812Google ScholarThere is no corresponding record for this reference.
- 166Oyler-Yaniv, A.; Oyler-Yaniv, J.; Whitlock, B. M.; Liu, Z.; Germain, R. N.; Huse, M.; Altan-Bonnet, G.; Krichevsky, O. A Tunable Diffusion-Consumption Mechanism of Cytokine Propagation Enables Plasticity in Cell-to-Cell Communication in the Immune System. Immunity 2017, 46 (4), 609– 620, DOI: 10.1016/j.immuni.2017.03.011Google ScholarThere is no corresponding record for this reference.
- 167Kusmierz, C. D.; Bujold, K. E.; Callmann, C. E.; Mirkin, C. A. Defining the Design Parameters for in Vivo Enzyme Delivery Through Protein Spherical Nucleic Acids. ACS Cent. Sci. 2020, 6 (5), 815– 822, DOI: 10.1021/acscentsci.0c00313Google ScholarThere is no corresponding record for this reference.
- 168Narayan, S. P.; Choi, C. H. J.; Hao, L.; Calabrese, C. M.; Auyeung, E.; Zhang, C.; Goor, O. J. G. M.; Mirkin, C. A. The Sequence-Specific Cellular Uptake of Spherical Nucleic Acid Nanoparticle Conjugates. Small 2015, 11 (33), 4173– 4182, DOI: 10.1002/smll.201500027Google ScholarThere is no corresponding record for this reference.
- 169Pearson, A. M.; Rich, A.; Krieger, M. Polynucleotide Binding to Macrophage Scavenger Receptors Depends on the Formation of Base-Quartet-Stabilized Four-Stranded Helices. J. Biol. Chem. 1993, 268 (5), 3546– 3554, DOI: 10.1016/S0021-9258(18)53729-7Google ScholarThere is no corresponding record for this reference.
- 170Paranandi, K. S.; Lee, A.; Stanic, N.; Mirkin, C. A. Cellular Export Fate of Liposomal Spherical Nucleic Acids. ACS Nano 2023, 17 (19), 19000– 19010, DOI: 10.1021/acsnano.3c04608Google ScholarThere is no corresponding record for this reference.
- 171Wiraja, C.; Mori, Y.; Ichimura, T.; Hwang, J.; Xu, C.; Bonventre, J. V. Nephrotoxicity Assessment with Human Kidney Tubuloids Using Spherical Nucleic Acid-Based mRNA Nanoflares. Nano Lett. 2021, 21 (13), 5850– 5858, DOI: 10.1021/acs.nanolett.1c01840Google ScholarThere is no corresponding record for this reference.
- 172Jayagopal, A.; Halfpenny, K. C.; Perez, J. W.; Wright, D. W. Hairpin DNA-Functionalized Gold Colloids for the Imaging of mRNA in Live Cells. J. Am. Chem. Soc. 2010, 132 (28), 9789– 9796, DOI: 10.1021/ja102585vGoogle ScholarThere is no corresponding record for this reference.
- 173Liu, L.; Li, N.; Huang, Z.-M.; Tang, L.-J.; Ying, Z.-M.; Jiang, J.-H. Gold Nanoflares with Computing Function as Smart Diagnostic Automata for Multi-miRNA Patterns in Living Cells. Anal. Chem. 2020, 92 (16), 10925– 10929, DOI: 10.1021/acs.analchem.0c02325Google ScholarThere is no corresponding record for this reference.
- 174Li, L.; Feng, J.; Fan, Y.; Tang, B. Simultaneous Imaging of Zn2+ and Cu2+ in Living Cells Based on DNAzyme Modified Gold Nanoparticle. Anal. Chem. 2015, 87 (9), 4829– 4835, DOI: 10.1021/acs.analchem.5b00204Google ScholarThere is no corresponding record for this reference.
- 175Zheng, D.; Seferos, D. S.; Giljohann, D. A.; Patel, P. C.; Mirkin, C. A. Aptamer Nano-flares for Molecular Detection in Living Cells. Nano Lett. 2009, 9 (9), 3258– 3261, DOI: 10.1021/nl901517bGoogle ScholarThere is no corresponding record for this reference.
- 176Chen, L.; Chao, J.; Qu, X.; Zhang, H.; Zhu, D.; Su, S.; Aldalbahi, A.; Wang, L.; Pei, H. Probing Cellular Molecules with PolyA-Based Engineered Aptamer Nanobeacon. ACS Appl. Mater. Interfaces 2017, 9 (9), 8014– 8020, DOI: 10.1021/acsami.6b16764Google ScholarThere is no corresponding record for this reference.
- 177Yeo, D. C.; Wiraja, C.; Paller, A. S.; Mirkin, C. A.; Xu, C. Abnormal Scar Identification with Spherical-Nucleic-Acid Technology. Nat. Biomed. Eng. 2018, 2 (4), 227– 238, DOI: 10.1038/s41551-018-0218-xGoogle ScholarThere is no corresponding record for this reference.
- 178Liu, Z.; Zhao, J.; Zhang, R.; Han, G.; Zhang, C.; Liu, B.; Zhang, Z.; Han, M.-Y.; Gao, X. Cross-Platform Cancer Cell Identification Using Telomerase-Specific Spherical Nucleic Acids. ACS Nano 2018, 12 (4), 3629– 3637, DOI: 10.1021/acsnano.8b00743Google ScholarThere is no corresponding record for this reference.
- 179Conde, J.; Oliva, N.; Artzi, N. Implantable Hydrogel Embedded Dark-Gold Nanoswitch as a Theranostic Probe to Sense and Overcome Cancer Multidrug Resistance. Proc. Natl. Acad. Sci. U. S. A. 2015, 112 (11), E1278-E1287 DOI: 10.1073/pnas.1421229112Google ScholarThere is no corresponding record for this reference.
- 180Halo, T. L.; McMahon, K. M.; Angeloni, N. L.; Xu, Y.; Wang, W.; Chinen, A. B.; Malin, D.; Strekalova, E.; Cryns, V. L.; Cheng, C. NanoFlares for the Detection, Isolation, and Culture of Live Tumor Cells from Human Blood. Proc. Natl. Acad. Sci. U. S. A. 2014, 111 (48), 17104– 17109, DOI: 10.1073/pnas.1418637111Google ScholarThere is no corresponding record for this reference.
- 181Zhu, D.; Li, X.; Zhu, Y.; Wei, Q.; Hu, Y.; Su, S.; Chao, J.; Wang, L.; Weng, L. Spatiotemporal Monitoring of Subcellular mRNAs In Situ via Polyadenine-Mediated Dual-Color Sticky Flares. ACS Appl. Mater. Interfaces 2023, 15 (12), 15250– 15259, DOI: 10.1021/acsami.3c01242Google ScholarThere is no corresponding record for this reference.
- 182Lin, M.; Yi, X.; Huang, F.; Ma, X.; Zuo, X.; Xia, F. Photoactivated Nanoflares for mRNA Detection in Single Living Cells. Anal. Chem. 2019, 91 (3), 2021– 2027, DOI: 10.1021/acs.analchem.8b04434Google ScholarThere is no corresponding record for this reference.
- 183Song, S.; Liang, Z.; Zhang, J.; Wang, L.; Li, G.; Fan, C. Gold-Nanoparticle-Based Multicolor Nanobeacons for Sequence-Specific DNA Analysis. Angew. Chem., Int. Ed. 2009, 48 (46), 8670– 8674, DOI: 10.1002/anie.200901887Google ScholarThere is no corresponding record for this reference.
- 184Pan, W.; Zhang, T.; Yang, H.; Diao, W.; Li, N.; Tang, B. Multiplexed Detection and Imaging of Intracellular mRNAs Using a Four-Color Nanoprobe. Anal. Chem. 2013, 85 (21), 10581– 10588, DOI: 10.1021/ac402700sGoogle ScholarThere is no corresponding record for this reference.
- 185Melnychuk, N.; Klymchenko, A. S. DNA-Functionalized Dye-Loaded Polymeric Nanoparticles: Ultrabright FRET Platform for Amplified Detection of Nucleic Acids. J. Am. Chem. Soc. 2018, 140 (34), 10856– 10865, DOI: 10.1021/jacs.8b05840Google ScholarThere is no corresponding record for this reference.
- 186Zhao, Y.; Zhao, J.; Zhang, J.; Sun, Y.; Li, L.; Li, Z.; Li, M. Enzymatically Controlled Nanoflares for Specific Molecular Recognition and Biosensing. Anal. Chem. 2022, 94 (25), 8883– 8889, DOI: 10.1021/acs.analchem.2c00166Google ScholarThere is no corresponding record for this reference.
- 187Chen, M.; Duan, R.; Xu, S.; Duan, Z.; Yuan, Q.; Xia, F.; Huang, F. Photoactivated DNA Walker Based on DNA Nanoflares for Signal-Amplified MicroRNA Imaging in Single Living Cells. Anal. Chem. 2021, 93 (48), 16264– 16272, DOI: 10.1021/acs.analchem.1c04505Google ScholarThere is no corresponding record for this reference.
- 188Li, J.; Liu, S.; Wang, J.; Liu, R.; Yang, X.; Wang, K.; Huang, J. Photocaged Amplified FRET Nanoflares: Spatiotemporal Controllable of mRNA-Powered Nanomachines for Precise and Sensitive MicroRNA Imaging in Live Cells. Nucleic Acids Res. 2022, 50 (7), e40 DOI: 10.1093/nar/gkab1258Google ScholarThere is no corresponding record for this reference.
- 189Yang, Y.; Huang, J.; Yang, X.; Quan, K.; Wang, H.; Ying, L.; Xie, N.; Ou, M.; Wang, K. FRET Nanoflares for Intracellular mRNA Detection: Avoiding False Positive Signals and Minimizing Effects of System Fluctuations. J. Am. Chem. Soc. 2015, 137 (26), 8340– 8343, DOI: 10.1021/jacs.5b04007Google ScholarThere is no corresponding record for this reference.
- 190Wang, N.; Song, L.; Xing, H.; Zhang, K.; Yang, R.; Li, J. A Spherical Nucleic Acid-Based Two-Photon Nanoprobe for RNase H Activity Assay in Living Cells and Tissues. Nanoscale 2019, 11 (17), 8133– 8137, DOI: 10.1039/C9NR00880BGoogle ScholarThere is no corresponding record for this reference.
- 191Zeng, T.; Fang, J.; Jiang, Y.; Xing, C.; Lu, C.; Yang, H. Spherical Nucleic Acid Probe Based on 2′-Fluorinated DNA Functionalization for High-Fidelity Intracellular Sensing. Anal. Chem. 2022, 94 (51), 18009– 18016, DOI: 10.1021/acs.analchem.2c04294Google ScholarThere is no corresponding record for this reference.
- 192Xiao, F.; Lin, L.; Chao, Z.; Shao, C.; Chen, Z.; Wei, Z.; Lu, J.; Huang, Y.; Li, L.; Liu, Q. Organic Spherical Nucleic Acids for the Transport of a NIR-II-Emitting Dye Across the Blood-Brain Barrier. Angew. Chem., Int. Ed. 2020, 59 (24), 9702– 9710, DOI: 10.1002/anie.202002312Google ScholarThere is no corresponding record for this reference.
- 193Fakih, H. H.; Katolik, A.; Malek-Adamian, E.; Fakhoury, J. J.; Kaviani, S.; Damha, M. J.; Sleiman, H. F. Design and Enhanced Gene Silencing Activity of Spherical 2′-Fluoroarabinose Nucleic Acids (FANA-SNAs). Chem. Sci. 2021, 12 (8), 2993– 3003, DOI: 10.1039/D0SC06645AGoogle ScholarThere is no corresponding record for this reference.
- 194Chen, P.; Wang, D.; Wang, Y.; Zhang, L.; Wang, Q.; Liu, L.; Li, J.; Sun, X.; Ren, M.; Wang, R. Maximizing TLR9 Activation in Cancer Immunotherapy with Dual-Adjuvanted Spherical Nucleic Acids. Nano Lett. 2022, 22 (10), 4058– 4066, DOI: 10.1021/acs.nanolett.2c00723Google ScholarThere is no corresponding record for this reference.
- 195Jensen, S. A.; Day, E. S.; Ko, C. H.; Hurley, L. A.; Luciano, J. P.; Kouri, F. M.; Merkel, T. J.; Luthi, A. J.; Patel, P. C.; Cutler, J. I. Spherical Nucleic Acid Nanoparticle Conjugates as an RNAi-Based Therapy for Glioblastoma. Sci. Transl. Med. 2013, 5 (209), 209ra152, DOI: 10.1126/scitranslmed.3006839Google ScholarThere is no corresponding record for this reference.
- 196Zheng, D.; Giljohann, D. A.; Chen, D. L.; Massich, M. D.; Wang, X.-Q.; Iordanov, H.; Mirkin, C. A.; Paller, A. S. Topical Delivery of siRNA-Based Spherical Nucleic Acid Nanoparticle Conjugates for Gene Regulation. Proc. Natl. Acad. Sci. U. S. A. 2012, 109 (30), 11975– 11980, DOI: 10.1073/pnas.1118425109Google ScholarThere is no corresponding record for this reference.
- 197Fang, Y.; Lu, X.; Wang, D.; Cai, J.; Wang, Y.; Chen, P.; Ren, M.; Lu, H.; Union, J.; Zhang, L. Spherical Nucleic Acids for Topical Treatment of Hyperpigmentation. J. Am. Chem. Soc. 2021, 143 (3), 1296– 1300, DOI: 10.1021/jacs.0c12044Google ScholarThere is no corresponding record for this reference.
- 198Huang, Z.; Callmann, C. E.; Wang, S.; Vasher, M. K.; Evangelopoulos, M.; Petrosko, S. H.; Mirkin, C. A. Rational Vaccinology: Harnessing Nanoscale Chemical Design for Cancer Immunotherapy. ACS Cent. Sci. 2022, 8 (6), 692– 704, DOI: 10.1021/acscentsci.2c00227Google ScholarThere is no corresponding record for this reference.
- 199Skakuj, K.; Wang, S.; Qin, L.; Lee, A.; Zhang, B.; Mirkin, C. A. Conjugation Chemistry-Dependent T-Cell Activation with Spherical Nucleic Acids. J. Am. Chem. Soc. 2018, 140 (4), 1227– 1230, DOI: 10.1021/jacs.7b12579Google ScholarThere is no corresponding record for this reference.
- 200Skakuj, K.; Teplensky, M. H.; Wang, S.; Dittmar, J. W.; Mirkin, C. A. Chemically Tuning the Antigen Release Kinetics from Spherical Nucleic Acids Maximizes Immune Stimulation. ACS Cent. Sci. 2021, 7 (11), 1838– 1846, DOI: 10.1021/acscentsci.1c00779Google ScholarThere is no corresponding record for this reference.
- 201Teplensky, M. H.; Dittmar, J. W.; Qin, L.; Wang, S.; Evangelopoulos, M.; Zhang, B.; Mirkin, C. A. Spherical Nucleic Acid Vaccine Structure Markedly Influences Adaptive Immune Responses of Clinically Utilized Prostate Cancer Targets. Adv. Healthcare Mater. 2021, 10 (22), 2101262, DOI: 10.1002/adhm.202101262Google ScholarThere is no corresponding record for this reference.
- 202Chinen, A. B.; Ferrer, J. R.; Merkel, T. J.; Mirkin, C. A. Relationships between Poly(ethylene glycol) Modifications on RNA-Spherical Nucleic Acid Conjugates and Cellular Uptake and Circulation Time. Bioconjugate Chem. 2016, 27 (11), 2715– 2721, DOI: 10.1021/acs.bioconjchem.6b00483Google ScholarThere is no corresponding record for this reference.
- 203Park, S. Y.; Lytton-Jean, A. K. R.; Lee, B.; Weigand, S.; Schatz, G. C.; Mirkin, C. A. DNA-Programmable Nanoparticle Crystallization. Nature 2008, 451 (7178), 553– 556, DOI: 10.1038/nature06508Google ScholarThere is no corresponding record for this reference.
- 204Laramy, C. R.; O’Brien, M. N.; Mirkin, C. A. Crystal Engineering with DNA. Nat. Rev. Mater. 2019, 4 (3), 201– 224, DOI: 10.1038/s41578-019-0087-2Google ScholarThere is no corresponding record for this reference.
- 205Wang, S.; Lee, S.; Du, J. S.; Partridge, B. E.; Cheng, H. F.; Zhou, W.; Dravid, V. P.; Lee, B.; Glotzer, S. C.; Mirkin, C. A. The Emergence of Valency in Colloidal Crystals through Electron Equivalents. Nat. Mater. 2022, 21 (5), 580– 587, DOI: 10.1038/s41563-021-01170-5Google ScholarThere is no corresponding record for this reference.
- 206Li, Y.; Zhou, W.; Tanriover, I.; Hadibrata, W.; Partridge, B. E.; Lin, H.; Hu, X.; Lee, B.; Liu, J.; Dravid, V. P. Open-Channel Metal Particle Superlattices. Nature 2022, 611 (7937), 695– 701, DOI: 10.1038/s41586-022-05291-yGoogle ScholarThere is no corresponding record for this reference.
- 207Cheng, H. F.; Distler, M. E.; Lee, B.; Zhou, W.; Weigand, S.; Mirkin, C. A. Nanoparticle Superlattices through Template-Encoded DNA Dendrimers. J. Am. Chem. Soc. 2021, 143 (41), 17170– 17179, DOI: 10.1021/jacs.1c07858Google ScholarThere is no corresponding record for this reference.
- 208Distler, M. E.; Landy, K. M.; Gibson, K. J.; Lee, B.; Weigand, S.; Mirkin, C. A. Symmetry-Breaking Dendrimer Synthons in Colloidal Crystal Engineering with DNA. J. Am. Chem. Soc. 2023, 145 (2), 841– 850, DOI: 10.1021/jacs.2c08599Google ScholarThere is no corresponding record for this reference.
- 209Li, Y.; Zhou, W.; Zhou, Y.; Cheng, H. F.; Lee, B.; Hu, X.; Roth, E. W.; Dravid, V. P.; Glotzer, S. C.; Mirkin, C. A. Cocrystals Combining Order and Correlated Disorder via Colloidal Crystal Engineering with DNA. Sci. Adv. 2025, 11 (16), eadu4919 DOI: 10.1126/sciadv.adu4919Google ScholarThere is no corresponding record for this reference.
- 210Liu, H.; Matthies, M.; Russo, J.; Rovigatti, L.; Narayanan, R. P.; Diep, T.; McKeen, D.; Gang, O.; Stephanopoulos, N.; Sciortino, F. Inverse Design of a Pyrochlore Lattice of DNA Origami through Model-Driven Experiments. Science 2024, 384 (6697), 776– 781, DOI: 10.1126/science.adl5549Google ScholarThere is no corresponding record for this reference.
- 211Kahn, J. S.; Minevich, B.; Michelson, A.; Emamy, H.; Wu, J.; Ji, H.; Yun, A.; Kisslinger, K.; Xiang, S.; Yu, N. Encoding Hierarchical 3D Architecture through Inverse Design of Programmable Bonds. Nat. Mater. 2025, 24 (8), 1273– 1282, DOI: 10.1038/s41563-025-02263-1Google ScholarThere is no corresponding record for this reference.
- 212He, M.; Gales, J. P.; Ducrot, É.; Gong, Z.; Yi, G.-R.; Sacanna, S.; Pine, D. J. Colloidal Diamond. Nature 2020, 585 (7826), 524– 529, DOI: 10.1038/s41586-020-2718-6Google ScholarThere is no corresponding record for this reference.
- 213Lin, H.; Lee, S.; Sun, L.; Spellings, M.; Engel, M.; Glotzer, S. C.; Mirkin, C. A. Clathrate Colloidal Crystals. Science 2017, 355 (6328), 931– 935, DOI: 10.1126/science.aal3919Google ScholarThere is no corresponding record for this reference.
- 214Zhou, W.; Lim, Y.; Lin, H.; Lee, S.; Li, Y.; Huang, Z.; Du, J. S.; Lee, B.; Wang, S.; Sánchez-Iglesias, A. Colloidal Quasicrystals Engineered with DNA. Nat. Mater. 2024, 23 (3), 424– 428, DOI: 10.1038/s41563-023-01706-xGoogle ScholarThere is no corresponding record for this reference.
- 215Liu, W.; Tagawa, M.; Xin, H. L.; Wang, T.; Emamy, H.; Li, H.; Yager, K. G.; Starr, F. W.; Tkachenko, A. V.; Gang, O. Diamond Family of Nanoparticle Superlattices. Science 2016, 351 (6273), 582– 586, DOI: 10.1126/science.aad2080Google ScholarThere is no corresponding record for this reference.
- 216Kim, Y.; Macfarlane, R. J.; Jones, M. R.; Mirkin, C. A. Transmutable Nanoparticles with Reconfigurable Surface Ligands. Science 2016, 351 (6273), 579– 582, DOI: 10.1126/science.aad2212Google ScholarThere is no corresponding record for this reference.
- 217Rogers, W. B.; Manoharan, V. N. Programming Colloidal Phase Transitions with DNA Strand Displacement. Science 2015, 347 (6222), 639– 642, DOI: 10.1126/science.1259762Google ScholarThere is no corresponding record for this reference.
- 218Oh, J. S.; Yi, G.-R.; Pine, D. J. Reconfigurable Self-Assembly and Kinetic Control of Multiprogrammed DNA-Coated Particles. ACS Nano 2020, 14 (4), 4595– 4600, DOI: 10.1021/acsnano.0c00164Google ScholarThere is no corresponding record for this reference.
- 219Zhu, J.; Lin, H.; Kim, Y.; Yang, M.; Skakuj, K.; Du, J. S.; Lee, B.; Schatz, G. C.; Van Duyne, R. P.; Mirkin, C. A. Light-Responsive Colloidal Crystals Engineered with DNA. Adv. Mater. 2020, 32 (8), 1906600, DOI: 10.1002/adma.201906600Google ScholarThere is no corresponding record for this reference.
- 220Zhu, J.; Kim, Y.; Lin, H.; Wang, S.; Mirkin, C. A. pH-Responsive Nanoparticle Superlattices with Tunable DNA Bonds. J. Am. Chem. Soc. 2018, 140 (15), 5061– 5064, DOI: 10.1021/jacs.8b02793Google ScholarThere is no corresponding record for this reference.
- 221Mason, J. A.; Laramy, C. R.; Lai, C.-T.; O’Brien, M. N.; Lin, Q.-Y.; Dravid, V. P.; Schatz, G. C.; Mirkin, C. A. Contraction and Expansion of Stimuli-Responsive DNA Bonds in Flexible Colloidal Crystals. J. Am. Chem. Soc. 2016, 138 (28), 8722– 8725, DOI: 10.1021/jacs.6b05430Google ScholarThere is no corresponding record for this reference.
- 222Hensley, A.; Jacobs, W. M.; Rogers, W. B. Self-Assembly of Photonic Crystals by Controlling the Nucleation and Growth of DNA-Coated Colloids. Proc. Natl. Acad. Sci. U. S. A. 2022, 119 (1), e2114050118 DOI: 10.1073/pnas.2114050118Google ScholarThere is no corresponding record for this reference.
- 223Hensley, A.; Videbæk, T. E.; Seyforth, H.; Jacobs, W. M.; Rogers, W. B. Macroscopic Photonic Single Crystals via Seeded Growth of DNA-Coated Colloids. Nat. Commun. 2023, 14 (1), 4237, DOI: 10.1038/s41467-023-39992-3Google ScholarThere is no corresponding record for this reference.
- 224Auyeung, E.; Li, T. I. N. G.; Senesi, A. J.; Schmucker, A. L.; Pals, B. C.; de la Cruz, M. O.; Mirkin, C. A. DNA-Mediated Nanoparticle Crystallization into Wulff Polyhedra. Nature 2014, 505 (7481), 73– 77, DOI: 10.1038/nature12739Google ScholarThere is no corresponding record for this reference.
- 225Hellstrom, S. L.; Kim, Y.; Fakonas, J. S.; Senesi, A. J.; Macfarlane, R. J.; Mirkin, C. A.; Atwater, H. A. Epitaxial Growth of DNA-Assembled Nanoparticle Superlattices on Patterned Substrates. Nano Lett. 2013, 13 (12), 6084– 6090, DOI: 10.1021/nl4033654Google ScholarThere is no corresponding record for this reference.
- 226Wang, M. X.; Seo, S. E.; Gabrys, P. A.; Fleischman, D.; Lee, B.; Kim, Y.; Atwater, H. A.; Macfarlane, R. J.; Mirkin, C. A. Epitaxy: Programmable Atom Equivalents Versus Atoms. ACS Nano 2017, 11 (1), 180– 185, DOI: 10.1021/acsnano.6b06584Google ScholarThere is no corresponding record for this reference.
- 227Lewis, D. J.; Zornberg, L. Z.; Carter, D. J. D.; Macfarlane, R. J. Single-Crystal Winterbottom Constructions of Nanoparticle Superlattices. Nat. Mater. 2020, 19 (7), 719– 724, DOI: 10.1038/s41563-020-0643-6Google ScholarThere is no corresponding record for this reference.
- 228Senesi, A. J.; Eichelsdoerfer, D. J.; Macfarlane, R. J.; Jones, M. R.; Auyeung, E.; Lee, B.; Mirkin, C. A. Stepwise Evolution of DNA-Programmable Nanoparticle Superlattices. Angew. Chem., Int. Ed. 2013, 52 (26), 6624– 6628, DOI: 10.1002/anie.201301936Google ScholarThere is no corresponding record for this reference.
- 229Sun, L.; Lin, H.; Li, Y.; Zhou, W.; Du, J. S.; Mirkin, C. A. Position- and Orientation-Controlled Growth of Wulff-Shaped Colloidal Crystals Engineered with DNA. Adv. Mater. 2020, 32 (47), 2005316, DOI: 10.1002/adma.202005316Google ScholarThere is no corresponding record for this reference.
- 230Gabrys, P. A.; Seo, S. E.; Wang, M. X.; Oh, E.; Macfarlane, R. J.; Mirkin, C. A. Lattice Mismatch in Crystalline Nanoparticle Thin Films. Nano Lett. 2018, 18 (1), 579– 585, DOI: 10.1021/acs.nanolett.7b04737Google ScholarThere is no corresponding record for this reference.
- 231Young, K. L.; Ross, M. B.; Blaber, M. G.; Rycenga, M.; Jones, M. R.; Zhang, C.; Senesi, A. J.; Lee, B.; Schatz, G. C.; Mirkin, C. A. Using DNA to Design Plasmonic Metamaterials with Tunable Optical Properties. Adv. Mater. 2014, 26 (4), 653– 659, DOI: 10.1002/adma.201302938Google ScholarThere is no corresponding record for this reference.
- 232Zornberg, L. Z.; Lewis, D. J.; Mertiri, A.; Hueckel, T.; Carter, D. J. D.; Macfarlane, R. J. Self-Assembling Systems for Optical Out-of-Plane Coupling Devices. ACS Nano 2023, 17 (4), 3394– 3400, DOI: 10.1021/acsnano.2c08344Google ScholarThere is no corresponding record for this reference.
- 233Lewis, D. J.; Carter, D. J. D.; Macfarlane, R. J. Using DNA to Control the Mechanical Response of Nanoparticle Superlattices. J. Am. Chem. Soc. 2020, 142 (45), 19181– 19188, DOI: 10.1021/jacs.0c08790Google ScholarThere is no corresponding record for this reference.
- 234Kim, S.; Svetlizky, I.; Weitz, D. A.; Spaepen, F. Work Hardening in Colloidal Crystals. Nature 2024, 630 (8017), 648– 653, DOI: 10.1038/s41586-024-07453-6Google ScholarThere is no corresponding record for this reference.
- 235Seo, S. E.; Wang, M. X.; Shade, C. M.; Rouge, J. L.; Brown, K. A.; Mirkin, C. A. Modulating the Bond Strength of DNA-Nanoparticle Superlattices. ACS Nano 2016, 10 (2), 1771– 1779, DOI: 10.1021/acsnano.5b07103Google ScholarThere is no corresponding record for this reference.
- 236Samanta, D.; Iscen, A.; Laramy, C. R.; Ebrahimi, S. B.; Bujold, K. E.; Schatz, G. C.; Mirkin, C. A. Multivalent Cation-Induced Actuation of DNA-Mediated Colloidal Superlattices. J. Am. Chem. Soc. 2019, 141 (51), 19973– 19977, DOI: 10.1021/jacs.9b09900Google ScholarThere is no corresponding record for this reference.
- 237Michelson, A.; Subramanian, A.; Kisslinger, K.; Tiwale, N.; Xiang, S.; Shen, E.; Kahn, J. S.; Nykypanchuk, D.; Yan, H.; Nam, C.-Y. Three-Dimensional Nanoscale Metal, Metal Oxide, and Semiconductor Frameworks through DNA-Programmable Assembly and Templating. Sci. Adv. 2024, 10 (2), eadl0604 DOI: 10.1126/sciadv.adl0604Google ScholarThere is no corresponding record for this reference.
- 238Auyeung, E.; Macfarlane, R. J.; Choi, C. H. J.; Cutler, J. I.; Mirkin, C. A. Transitioning DNA-Engineered Nanoparticle Superlattices from Solution to the Solid State. Adv. Mater. 2012, 24 (38), 5181– 5186, DOI: 10.1002/adma.201202069Google ScholarThere is no corresponding record for this reference.
- 239Wang, S.; Park, S. S.; Buru, C. T.; Lin, H.; Chen, P.-C.; Roth, E. W.; Farha, O. K.; Mirkin, C. A. Colloidal Crystal Engineering with Metal-Organic Framework Nanoparticles and DNA. Nat. Commun. 2020, 11 (1), 2495, DOI: 10.1038/s41467-020-16339-wGoogle ScholarThere is no corresponding record for this reference.
- 240Kim, S.; Zheng, C. Y.; Schatz, G. C.; Aydin, K.; Kim, K.-H.; Mirkin, C. A. Mie-Resonant Three-Dimensional Metacrystals. Nano Lett. 2020, 20 (11), 8096– 8101, DOI: 10.1021/acs.nanolett.0c03089Google ScholarThere is no corresponding record for this reference.
- 241Zhang, Y.; Xu, D. D.; Tanriover, I.; Zhou, W.; Li, Y.; López-Arteaga, R.; Aydin, K.; Mirkin, C. A. Nonlinear Optical Colloidal Metacrystals. Nat. Photonics 2025, 19 (1), 20– 27, DOI: 10.1038/s41566-024-01558-0Google ScholarThere is no corresponding record for this reference.
- 242Li, Z.; Lim, Y.; Tanriover, I.; Zhou, W.; Li, Y.; Zhang, Y.; Aydin, K.; Glotzer, S. C.; Mirkin, C. A. DNA-Mediated Assembly of Au Bipyramids into Anisotropic Light Emitting Kagome Superlattices. Sci. Adv. 2024, 10 (29), eadp3756 DOI: 10.1126/sciadv.adp3756Google ScholarThere is no corresponding record for this reference.
- 243Lin, Q.-Y.; Li, Z.; Brown, K. A.; O’Brien, M. N.; Ross, M. B.; Zhou, Y.; Butun, S.; Chen, P.-C.; Schatz, G. C.; Dravid, V. P. Strong Coupling between Plasmonic Gap Modes and Photonic Lattice Modes in DNA-Assembled Gold Nanocube Arrays. Nano Lett. 2015, 15 (7), 4699– 4703, DOI: 10.1021/acs.nanolett.5b01548Google ScholarThere is no corresponding record for this reference.
- 244Zheng, C. Y.; Palacios, E.; Zhou, W.; Hadibrata, W.; Sun, L.; Huang, Z.; Schatz, G. C.; Aydin, K.; Mirkin, C. A. Tunable Fluorescence from Dye-Modified DNA-Assembled Plasmonic Nanocube Arrays. Adv. Mater. 2019, 31 (41), 1904448, DOI: 10.1002/adma.201904448Google ScholarThere is no corresponding record for this reference.
- 245Zheng, C. Y.; Hadibrata, W.; Kim, S.; Schatz, G. C.; Aydin, K.; Mirkin, C. A. Large-Area, Highly Crystalline DNA-Assembled Metasurfaces Exhibiting Widely Tunable Epsilon-Near-Zero Behavior. ACS Nano 2021, 15 (11), 18289– 18296, DOI: 10.1021/acsnano.1c07496Google ScholarThere is no corresponding record for this reference.
- 246Sun, L.; Lin, H.; Park, D. J.; Bourgeois, M. R.; Ross, M. B.; Ku, J. C.; Schatz, G. C.; Mirkin, C. A. Polarization-Dependent Optical Response in Anisotropic Nanoparticle-DNA Superlattices. Nano Lett. 2017, 17 (4), 2313– 2318, DOI: 10.1021/acs.nanolett.6b05101Google ScholarThere is no corresponding record for this reference.
- 247Li, Y.; Tanriover, I.; Zhou, W.; Hadibrata, W.; Dereshgi, S. A.; Samanta, D.; Aydin, K.; Mirkin, C. A. Monolayer Plasmonic Nanoframes as Large-Area, Broadband Metasurface Absorbers. Small 2022, 18 (33), 2201171, DOI: 10.1002/smll.202201171Google ScholarThere is no corresponding record for this reference.
- 248Li, Y.; Jin, H.; Zhou, W.; Wang, Z.; Lin, Z.; Mirkin, C. A.; Espinosa, H. D. Ultrastrong Colloidal Crystal Metamaterials Engineered with DNA. Sci. Adv. 2023, 9 (39), eadj8103 DOI: 10.1126/sciadv.adj8103Google ScholarThere is no corresponding record for this reference.
- 249Petrosko, S. H.; Coleman, B. D.; Drout, R. J.; Schultz, J. D.; Mirkin, C. A. Spherical Nucleic Acids: Integrating Nanotechnology Concepts into General Chemistry Curricula. J. Chem. Educ. 2021, 98 (10), 3090– 3099, DOI: 10.1021/acs.jchemed.1c00441Google ScholarThere is no corresponding record for this reference.
- 250Johnson, G. T.; Autin, L.; Goodsell, D. S.; Sanner, M. F.; Olson, A. J. ePMV Embeds Molecular Modeling into Professional Animation Software Environments. Structure 2011, 19 (3), 293– 303, DOI: 10.1016/j.str.2010.12.023Google ScholarThere is no corresponding record for this reference.
Cited By
This article has not yet been cited by other publications.
Article Views
Altmetric
Citations
Article Views are the COUNTER-compliant sum of full text article downloads since November 2008 (both PDF and HTML) across all institutions and individuals. These metrics are regularly updated to reflect usage leading up to the last few days.
Citations are the number of other articles citing this article, calculated by Crossref and updated daily. Find more information about Crossref citation counts.
The Altmetric Attention Score is a quantitative measure of the attention that a research article has received online. Clicking on the donut icon will load a page at altmetric.com with additional details about the score and the social media presence for the given article. Find more information on the Altmetric Attention Score and how the score is calculated.
Recommended Articles
Abstract

Figure 1
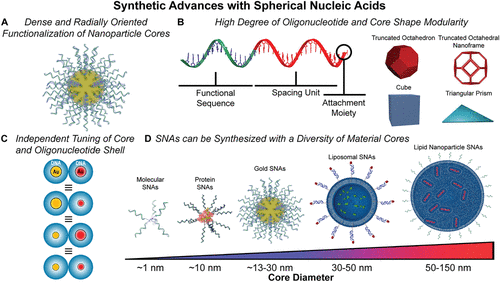
Figure 1. Synthetic advances with SNAs. (A) Schematic shows a gold-core SNA radially functionalized with strands of oligonucleotides. (B) SNAs exhibit a high degree of oligonucleotide and core shape modularity. Functional oligonucleotides typically contain three segments: a functional oligonucleotide sequence, a spacer segment, and an attachment group for nanoparticle surface binding. SNA-like properties can be achieved using anisotropic nanoparticle cores including truncated octahedron, truncated octahedral nanoframes, cubes, and triangular prisms. Adapted with permission from ref (206). Copyright 2022 Springer Nature Ltd. (C) SNA diameter is dependent on the core size and oligonucleotide shell length. Both parameters can be tuned independently. SNAs of the same diameter can be prepared using different core sizes and DNA shell lengths. Adapted with permission from ref (10). Copyright 2011 American Association for the Advancement of Science. (D) To date, SNAs have been synthesized using a variety of core materials that allow access to vastly different physicochemical properties. Core materials have included both inorganic and organic cores, and they endow the SNA structures with specific properties (e.g., plasmonic and enzymatic) based on composition. Pictured are a selection of different SNAs with a diversity of core materials, including molecular, protein, gold, liposomal, and lipid nanoparticle cores ranging in size from ∼1 to >150 nm. The representation of a CRISPR SNA is adapted from PDB ID 8UZA. ePMV software was used in the generation of panels A, B, and D, ref (250).
Figure 2

Figure 2. Fundamental research and engineering development with SNA-based probes. (A) Solutions of gold nanoparticle-core SNAs before (left; dispersed particles are red) and after the addition of a target DNA sequence links the SNAs (right; aggregated particles are purple). (B) Melting curves show free DNA and the same DNA attached to a gold-core SNA and a core-free polyvalent nucleic acid nanostructure (PNAN) (left); enthalpies and entropies of DNA hybridization to linear DNA and SNAs are derived from a concentration-dependent fluorescence hybridization assay (right). Panel B, left: Adapted with permission from ref (59). Copyright 2011 American Chemical Society. Panel B, right: Reprinted with permission from ref (131). Copyright 2018 American Chemical Society. (C) Example shows polynucleotide target detection using the colorimetric Northwestern Spot Test (left). Solutions of SNA probes were mixed with a fully complementary target strand (Target), without the target (No Target) and with a target contaning a single base pair mismatch (Mismatch Target) and heated to the indicated temperature. Note the high degree of discrimination between the fully complementary and mismatched targets, allowing for the detection of single nucleotide mismatches. Example shows the first SNA-based scanometric detection system wherein oligonucleotide arrays were treated with target strands (either fully complementary [A] or with a single base pair mismatch, substituting A for G, T, or C) and SNA probes were developed at the indicated temperatures (right). Panel C, left: Adapted with permission from ref (112). Copyright 1997 American Association for the Advancement of Science. Panel C, right: Reprinted with permission from ref (126). Copyright 2000 American Association for the Advancement of Science. (D) Chip-based multiplexed scanometric detection of barcode-DNA strands are released from SNA probes after incubation with DNA targets derived from the human immunodeficiency virus (HIV), Ebola virus (EV), variola virus (VV), and hepatitis B virus (HBV), or a mixture of all four (Mix). Adapted with permission from ref (139). Copyright 2006 WILEY-VCH Verlag GmbH & Co. KGaA, Weinheim. ePMV software was used in the generation of panel A, ref (250).
Figure 3
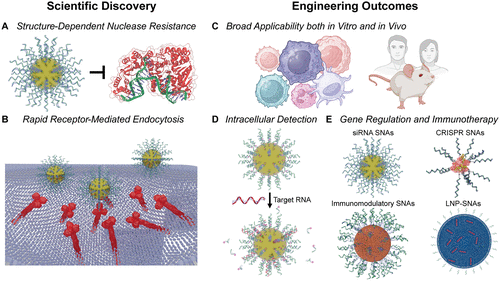
Figure 3. Scientific discoveries and engineering outcomes of SNAs in nanomedicine research. (A) The dense 3D DNA shell of SNAs confers enhanced resistance to nuclease degradation relative to linear oligonucleotides. This protection arises from slower enzymatic hydrolysis, as the negatively charged and sterically bulky shell inhibits nuclease activity. The protein corona that forms on SNAs in biological media is not depicted. FAN1 Nuclease PDB ID 8S5A. (B) Compared to linear or transfected oligonucleotides, SNAs exhibit high cellular uptake due to their multivalent interactions with class A scavenger receptors, which facilitate endocytosis via a lipid-raft-dependent, caveolae-mediated pathway. The representation of scavenger receptors is adapted from PDB IDs 2QIH and 7DPX. (C) Their high uptake efficiency, stability, and limited activation of host immune responses enable SNAs to function broadly as tools in both in vitro and in vivo systems. This versatility supports their use in intracellular detection, imaging, gene regulation, and immunotherapy. (D) Detection probes, including nanoflares, sticky-flares, and FIT-flares, have been engineered to sense intracellular RNA, ions, small molecules, and diseased tissue in vivo, as well as to perform live-cell genetic and metabolic analysis. These probes incorporate fluorophores that are either released or activated upon target binding, generating sensitive and responsive readouts. (E) As gene regulation agents, SNAs can be constructed from DNA or RNA to modulate expression through antisense or RNA interference pathways. As immunotherapies, they stimulate both innate and adaptive immune responses and have shown promise as nanomedicines for treating diverse cancers and infectious disease. The representation of a CRISPR SNA is adapted from PDB ID 8UZA. ePMV software was used in the generation of panels A, B, D, and E, ref (250).
Figure 4

Figure 4. Fundamental research and engineering development with SNA-based immunotherapeutics. Key scientific discoveries such as (A) the multivalent display of CpG DNA on SNAs potently stimulating TLR9 have pioneered the new field of structural immunotherapy. The representation of TLR9 is adapted from PDB IDs 5Y3M and 6US8. (B) Compositionally identical yet structurally distinct SNAs suppress tumor growth differentially, demonstrating the importance of nanoarchitecture in next-generation therapeutic development. Adapted with permission from ref (6). Copyright 2019 National Academy of Sciences. (C) These structure–function relationships extend to more complex therapeutics. The differential placements of MHC-I- and MHC-II-restricted antigens in DA-SNA 1 and 2 substantially alter tumor suppression. Adapted with permission from ref (22). Copyright 2023 Springer Nature Ltd. (D) The oligonucleotide shell of SNAs dictates their circulation and biodistribution, leading to the discovery of new agents that cross the blood–brain barrier (BBB). (E) Gold SNAs bearing Bcl2L12 siRNA labeled with Gd(III) localize in glioma tumor tissue following intracranial injection as shown in magnetic resonance (MR) images (top), hematoxylin and eosin-stained brain sections (middle), and the 3D reconstruction of MR data (bottom). Reprinted with permission from ref (195). Copyright 2013 American Association for the Advancement of Science (AAAS). (F) β-galactosidase-based ProSNA functionalized with transferrin aptamer (upper left) successfully accumulates within the brain (lower left), quantified by radiant efficiency (right). Reprinted with permission from ref (41). Copyright 2022 American Chemical Society. ePMV software was used in the generation of panels A, B, and C, ref (250).
Figure 5

Figure 5. Scientific discoveries, materials design, and engineering outcomes through the crystallization of SNAs and other types of programmable atom equivalents (PAEs). (A) The colloidal crystal design space arises from the synthetic advances in SNA preparation, where enthalpy-dominated DNA interactions direct crystallization. Metal nanoparticle core (shape and size) and DNA design determine the lattice symmetry and crystal habit. Scale bars = 125, 75, and 1000 nm from left to right. (B) Enthalpic DNA bonding interactions are maximized with facet registry between anisotropic nanoparticle cores, such as cubes. Scale bar = 500 nm. (C) DNA bonding is maximized along the edges of nanoframes, forming a crystalline symmetry from a non-space-filling polyhedral. Scale bar = 1 μm. Reprinted with permission from ref (206). Copyright 2022 Springer Nature Ltd. (D) Thin films and Winterbottom constructions can be formed from heterogeneous crystal growth on substrates. Reprinted with permission from ref (226). Copyright 2017 American Chemical Society. (E) The refractive index of colloidal crystals can be designed as a function of NP shape, NP size, volume fraction, and symmetry. Reprinted with permission from ref (206). Copyright 2022 Springer Nature Ltd. (F) Colloidal crystals components can be stabilized and used as heterogeneous catalysts. (G) Deterministic placement is shown for single PAEs to couple photonic response (determined by spacing, P) and plasmonic response (distance between Au cube and Au substrate). Reprinted with permission from ref (243). Copyright 2015 American Chemical Society. (H) Colloidal crystals exhibit shape memory upon dehydration and rehydration. The mechanical response of the colloidal crystal due to a reversible change in refractive index results in tunable optical response. ePMV software was used in the generation of panel A, ref (250).
References
This article references 250 other publications.
- 1Mirkin, C. A.; Letsinger, R. L.; Mucic, R. C.; Storhoff, J. J. A DNA-Based Method for Rationally Assembling Nanoparticles into Macroscopic Materials. Nature 1996, 382 (6592), 607– 609, DOI: 10.1038/382607a0There is no corresponding record for this reference.
- 2Cutler, J. I.; Auyeung, E.; Mirkin, C. A. Spherical Nucleic Acids. J. Am. Chem. Soc. 2012, 134 (3), 1376– 1391, DOI: 10.1021/ja209351uThere is no corresponding record for this reference.
- 3Mirkin, C. A.; Petrosko, S. H. Inspired Beyond Nature: Three Decades of Spherical Nucleic Acids and Colloidal Crystal Engineering with DNA. ACS Nano 2023, 17 (17), 16291– 16307, DOI: 10.1021/acsnano.3c06564There is no corresponding record for this reference.
- 4Macfarlane, R. J.; O’Brien, M. N.; Petrosko, S. H.; Mirkin, C. A. Nucleic Acid-Modified Nanostructures as Programmable Atom Equivalents: Forging a New ″Table of Elements″. Angew. Chem., Int. Ed. 2013, 52 (22), 5688– 5698, DOI: 10.1002/anie.201209336There is no corresponding record for this reference.
- 5Samanta, D.; Zhou, W.; Ebrahimi, S. B.; Petrosko, S. H.; Mirkin, C. A. Programmable Matter: The Nanoparticle Atom and DNA Bond. Adv. Mater. 2022, 34 (12), 2107875, DOI: 10.1002/adma.202107875There is no corresponding record for this reference.
- 6Wang, S.; Qin, L.; Yamankurt, G.; Skakuj, K.; Huang, Z.; Chen, P.-C.; Dominguez, D.; Lee, A.; Zhang, B.; Mirkin, C. A. Rational Vaccinology with Spherical Nucleic Acids. Proc. Natl. Acad. Soc. U. S. A. 2019, 116 (21), 10473– 10481, DOI: 10.1073/pnas.1902805116There is no corresponding record for this reference.
- 7Rosi, N. L.; Giljohann, D. A.; Thaxton, C. S.; Lytton-Jean, A. K. R.; Han, M. S.; Mirkin, C. A. Oligonucleotide-Modified Gold Nanoparticles for Intracellular Gene Regulation. Science 2006, 312 (5776), 1027– 1030, DOI: 10.1126/science.1125559There is no corresponding record for this reference.
- 8Mirkin, C. A.; Langer, R.; Mrksich, M.; Margolin, A. A.; Petrosko, S. H.; Artzi, N. Blueprints for Better Drugs: The Structural Revolution in Nanomedicine. ACS Nano 2025, 19 (20), 18889– 18901, DOI: 10.1021/acsnano.5c06380There is no corresponding record for this reference.
- 9Mirkin, C. A.; Mrksich, M.; Artzi, N. The Emerging Era of Structural Nanomedicine. Nat. Rev. Bioeng. 2025, 3 (7), 526– 528, DOI: 10.1038/s44222-025-00306-5There is no corresponding record for this reference.
- 10Macfarlane, R. J.; Lee, B.; Jones, M. R.; Harris, N.; Schatz, G. C.; Mirkin, C. A. Nanoparticle Superlattice Engineering with DNA. Science 2011, 334 (6053), 204– 208, DOI: 10.1126/science.1210493There is no corresponding record for this reference.
- 11Park, J.; Evangelopoulos, M.; Vasher, M. K.; Kudruk, S.; Ramani, N.; Mayer, V.; Solivan, A. C.; Lee, A.; Mirkin, C. A. Enhancing Endosomal Escape and Gene Regulation Activity for Spherical Nucleic Acids. Small 2024, 20 (11), 2306902, DOI: 10.1002/smll.202306902There is no corresponding record for this reference.
- 12Samanta, D.; Ebrahimi, S. B.; Kusmierz, C. D.; Cheng, H. F.; Mirkin, C. A. Protein Spherical Nucleic Acids for Live-Cell Chemical Analysis. J. Am. Chem. Soc. 2020, 142 (31), 13350– 13355, DOI: 10.1021/jacs.0c06866There is no corresponding record for this reference.
- 13Seferos, D. S.; Giljohann, D. A.; Hill, H. D.; Prigodich, A. E.; Mirkin, C. A. Nano-Flares: Probes for Transfection and mRNA Detection in Living Cells. J. Am. Chem. Soc. 2007, 129 (50), 15477– 15479, DOI: 10.1021/ja0776529There is no corresponding record for this reference.
- 14Briley, W. E.; Bondy, M. H.; Randeria, P. S.; Dupper, T. J.; Mirkin, C. A. Quantification and Real-Time Tracking of RNA in Live Cells Using Sticky-Flares. Proc. Natl. Acad. Soc. U. S. A. 2015, 112 (31), 9591– 9595, DOI: 10.1073/pnas.1510581112There is no corresponding record for this reference.
- 15Yamankurt, G.; Stawicki, R. J.; Posadas, D. M.; Nguyen, J. Q.; Carthew, R. W.; Mirkin, C. A. The Effector Mechanism of siRNA Spherical Nucleic Acids. Proc. Natl. Acad. Soc. U. S. A. 2020, 117 (3), 1312– 1320, DOI: 10.1073/pnas.1915907117There is no corresponding record for this reference.
- 16Li, J.; Mao, X.; Zhao, T.; Fang, W.; Jin, Y.; Liu, M.; Fan, C.; Tian, Y. Tetrahedral DNA Framework-Based Spherical Nucleic Acids for Efficient siRNA Delivery. Angew. Chem., Int. Ed. 2025, 64 (5), e202416988 DOI: 10.1002/anie.202416988There is no corresponding record for this reference.
- 17Randeria, P. S.; Seeger, M. A.; Wang, X.-Q.; Wilson, H.; Shipp, D.; Mirkin, C. A.; Paller, A. S. siRNA-Based Spherical Nucleic Acids Reverse Impaired Wound Healing in Diabetic Mice by Ganglioside GM3 Synthase Knockdown. Proc. Natl. Acad. Soc. U. S. A. 2015, 112 (18), 5573– 5578, DOI: 10.1073/pnas.1505951112There is no corresponding record for this reference.
- 18Kumthekar, P.; Ko, C. H.; Paunesku, T.; Dixit, K.; Sonabend, A. M.; Bloch, O.; Tate, M.; Schwartz, M.; Zuckerman, L.; Lezon, R. A First-in-Human Phase 0 Clinical Study of RNA Interference-Based Spherical Nucleic Acids in Patients with Recurrent Glioblastoma. Sci. Transl. Med. 2021, 13 (584), eabb3945 DOI: 10.1126/scitranslmed.abb3945There is no corresponding record for this reference.
- 19Huang, C.; Han, Z.; Evangelopoulos, M.; Mirkin, C. A. CRISPR Spherical Nucleic Acids. J. Am. Chem. Soc. 2022, 144 (41), 18756– 18760, DOI: 10.1021/jacs.2c07913There is no corresponding record for this reference.
- 20Han, Z.; Huang, C.; Luo, T.; Mirkin, C. A. A General Genome Editing Strategy Using CRISPR Lipid Nanoparticle Spherical Nucleic Acids. Proc. Natl. Acad. Soc. U. S. A. 2025, 122 (36), e2426094122 DOI: 10.1073/pnas.2426094122There is no corresponding record for this reference.
- 21Qin, L.; Wang, S.; Dominguez, D.; Long, A.; Chen, S.; Fan, J.; Ahn, J.; Skakuj, K.; Huang, Z.; Lee, A. Development of Spherical Nucleic Acids for Prostate Cancer Immunotherapy. Front. Immunol. 2020, 11, 1333, DOI: 10.3389/fimmu.2020.01333There is no corresponding record for this reference.
- 22Teplensky, M. H.; Evangelopoulos, M.; Dittmar, J. W.; Forsyth, C. M.; Sinegra, A. J.; Wang, S.; Mirkin, C. A. Multi-Antigen Spherical Nucleic Acid Cancer Vaccines. Nat. Biomed. Eng. 2023, 7 (7), 911– 927, DOI: 10.1038/s41551-022-01000-2There is no corresponding record for this reference.
- 23Hwang, J.; Dittmar, J. W.; Kang, J.; Ocampo, T.; Evangelopoulos, M.; Han, Z.; Kudruk, S.; Lorch, J.; Mirkin, C. A. DNA Anchoring Strength Directly Correlates with Spherical Nucleic Acid-Based HPV E7 Cancer Vaccine Potency. Nano Lett. 2024, 24 (25), 7629– 7636, DOI: 10.1021/acs.nanolett.4c01392There is no corresponding record for this reference.
- 24Auyeung, E.; Morris, W.; Mondloch, J. E.; Hupp, J. T.; Farha, O. K.; Mirkin, C. A. Controlling Structure and Porosity in Catalytic Nanoparticle Superlattices with DNA. J. Am. Chem. Soc. 2015, 137 (4), 1658– 1662, DOI: 10.1021/ja512116pThere is no corresponding record for this reference.
- 25Sun, L.; Lin, H.; Kohlstedt, K. L.; Schatz, G. C.; Mirkin, C. A. Design Principles for Photonic Crystals Based on Plasmonic Nanoparticle Superlattices. Proc. Natl. Acad. Soc. U. S. A. 2018, 115 (28), 7242– 7247, DOI: 10.1073/pnas.1800106115There is no corresponding record for this reference.
- 26Lee, S.; Calcaterra, H. A.; Lee, S.; Hadibrata, W.; Lee, B.; Oh, E.; Aydin, K.; Glotzer, S. C.; Mirkin, C. A. Shape Memory in Self-Adapting Colloidal Crystals. Nature 2022, 610 (7933), 674– 679, DOI: 10.1038/s41586-022-05232-9There is no corresponding record for this reference.
- 27Michelson, A.; Shani, L.; Kahn, J. S.; Redeker, D. C.; Lee, W.-I.; DeOlivares, K. R.; Kisslinger, K.; Tiwale, N.; Yan, H.; Pattammattel, A. Scalable Fabrication of Chip-Integrated 3D-Nanostructured Electronic Devices via DNA-Programmable Assembly. Sci. Adv. 2025, 11 (13), eadt5620 DOI: 10.1126/sciadv.adt5620There is no corresponding record for this reference.
- 28Daniel, W. L.; Lorch, U.; Mix, S.; Bexon, A. S. A First-in-Human Phase 1 Study of Cavrotolimod, a TLR9 Agonist Spherical Nucleic Acid, in Healthy Participants: Evidence of Immune Activation. Front. Immunol. 2022, 13, 1073777, DOI: 10.3389/fimmu.2022.1073777There is no corresponding record for this reference.
- 29Milhem, M. M.; Wise-Draper, T. M.; Chandra, S.; Hanna, G. J.; Laux, D. E.; Medina, T. M.; Ansstas, G.; Daud, A.; Kelly, C. M.; O’Day, S. J. Phase 1b/2 Study Evaluating Safety, Efficacy and Immune Effects of TLR9 Agonist Cavrotolimod with Anti-PD-1 Antibodies Among Patients with Advanced Solid Tumors. J. Immunother. Cancer 2025, 13 (7), e011651 DOI: 10.1136/jitc-2025-011651There is no corresponding record for this reference.
- 30Zhang, C.; Macfarlane, R. J.; Young, K. L.; Choi, C. H. J.; Hao, L.; Auyeung, E.; Liu, G.; Zhou, X.; Mirkin, C. A. A General Approach to DNA-Programmable Atom Equivalents. Nat. Mater. 2013, 12 (8), 741– 746, DOI: 10.1038/nmat3647There is no corresponding record for this reference.
- 31Mucic, R. C.; Storhoff, J. J.; Mirkin, C. A.; Letsinger, R. L. DNA-Directed Synthesis of Binary Nanoparticle Network Materials. J. Am. Chem. Soc. 1998, 120 (48), 12674– 12675, DOI: 10.1021/ja982721sThere is no corresponding record for this reference.
- 32Callmann, C. E.; Vasher, M. K.; Das, A.; Kusmierz, C. D.; Mirkin, C. A. In Vivo Behavior of Ultrasmall Spherical Nucleic Acids. Small 2023, 19 (24), 2300097, DOI: 10.1002/smll.202300097There is no corresponding record for this reference.
- 33Hill, H. D.; Millstone, J. E.; Banholzer, M. J.; Mirkin, C. A. The Role Radius of Curvature Plays in Thiolated Oligonucleotide Loading on Gold Nanoparticles. ACS Nano 2009, 3 (2), 418– 424, DOI: 10.1021/nn800726eThere is no corresponding record for this reference.
- 34PELCO Gold Nanoparticles─Gold Colloids; Ted Pella. https://www.tedpella.com/gold_html/pelco-gold-nanoparticles.aspx#Passive (accessed 2025-03-22).There is no corresponding record for this reference.
- 35Frens, G. Controlled Nucleation for the Regulation of the Particle Size in Monodisperse Gold Suspensions. Nat. Phys. Sci. 1973, 241 (105), 20– 22, DOI: 10.1038/physci241020a0There is no corresponding record for this reference.
- 36Hayat, M. A. Colloidal Gold: Principles, Methods, and Applications; Academic Press: New York, 1989.There is no corresponding record for this reference.
- 37Hill, H. D.; Macfarlane, R. J.; Senesi, A. J.; Lee, B.; Park, S. Y.; Mirkin, C. A. Controlling the Lattice Parameters of Gold Nanoparticle FCC Crystals with Duplex DNA Linkers. Nano Lett. 2008, 8 (8), 2341– 2344, DOI: 10.1021/nl8011787There is no corresponding record for this reference.
- 38Wang, M. D. Ruler of Life. Nat. Phys. 2021, 17 (8), 976– 976, DOI: 10.1038/s41567-021-01300-5There is no corresponding record for this reference.
- 39Banga, R. J.; Chernyak, N.; Narayan, S. P.; Nguyen, S. T.; Mirkin, C. A. Liposomal Spherical Nucleic Acids. J. Am. Chem. Soc. 2014, 136 (28), 9866– 9869, DOI: 10.1021/ja504845fThere is no corresponding record for this reference.
- 40Yamankurt, G.; Berns, E. J.; Xue, A.; Lee, A.; Bagheri, N.; Mrksich, M.; Mirkin, C. A. Exploration of the Nanomedicine-Design Space with High-Throughput Screening and Machine Learning. Nat. Biomed. Eng. 2019, 3 (4), 318– 327, DOI: 10.1038/s41551-019-0351-1There is no corresponding record for this reference.
- 41Kusmierz, C. D.; Callmann, C. E.; Kudruk, S.; Distler, M. E.; Mirkin, C. A. Transferrin Aptamers Increase the In Vivo Blood-Brain Barrier Targeting of Protein Spherical Nucleic Acids. Bioconjugate Chem. 2022, 33 (10), 1803– 1810, DOI: 10.1021/acs.bioconjchem.2c00389There is no corresponding record for this reference.
- 42Ebrahimi, S. B.; Samanta, D.; Kusmierz, C. D.; Mirkin, C. A. Protein Transfection via Spherical Nucleic Acids. Nat. Protoc. 2022, 17 (2), 327– 357, DOI: 10.1038/s41596-021-00642-xThere is no corresponding record for this reference.
- 43Girard, M.; Wang, S.; Du, J. S.; Das, A.; Huang, Z.; Dravid, V. P.; Lee, B.; Mirkin, C. A.; Olvera de la Cruz, M. Particle Analogs of Electrons in Colloidal Crystals. Science 2019, 364 (6446), 1174– 1178, DOI: 10.1126/science.aaw8237There is no corresponding record for this reference.
- 44Daniel, M.-C.; Astruc, D. Gold Nanoparticles: Assembly, Supramolecular Chemistry, Quantum-Size-Related Properties, and Applications toward Biology, Catalysis, and Nanotechnology. Chem. Rev. 2004, 104 (1), 293– 346, DOI: 10.1021/cr030698+There is no corresponding record for this reference.
- 45Lee, J.-S.; Lytton-Jean, A. K. R.; Hurst, S. J.; Mirkin, C. A. Silver Nanoparticle-Oligonucleotide Conjugates Based on DNA with Triple Cyclic Disulfide Moieties. Nano Lett. 2007, 7 (7), 2112– 2115, DOI: 10.1021/nl071108gThere is no corresponding record for this reference.
- 46Xue, C.; Chen, X.; Hurst, S. J.; Mirkin, C. A. Self-Assembled Monolayer Mediated Silica Coating of Silver Triangular Nanoprisms. Adv. Mater. 2007, 19 (22), 4071– 4074, DOI: 10.1002/adma.200701506There is no corresponding record for this reference.
- 47Young, K. L.; Scott, A. W.; Hao, L.; Mirkin, S. E.; Liu, G.; Mirkin, C. A. Hollow Spherical Nucleic Acids for Intracellular Gene Regulation Based upon Biocompatible Silica Shells. Nano Lett. 2012, 12 (7), 3867– 3871, DOI: 10.1021/nl3020846There is no corresponding record for this reference.
- 48Cutler, J. I.; Zheng, D.; Xu, X.; Giljohann, D. A.; Mirkin, C. A. Polyvalent Oligonucleotide Iron Oxide Nanoparticle “Click” Conjugates. Nano Lett. 2010, 10 (4), 1477– 1480, DOI: 10.1021/nl100477mThere is no corresponding record for this reference.
- 49Mitchell, G. P.; Mirkin, C. A.; Letsinger, R. L. Programmed Assembly of DNA Functionalized Quantum Dots. J. Am. Chem. Soc. 1999, 121 (35), 8122– 8123, DOI: 10.1021/ja991662vThere is no corresponding record for this reference.
- 50Sun, D.; Gang, O. DNA-Functionalized Quantum Dots: Fabrication, Structural, and Physicochemical Properties. Langmuir 2013, 29 (23), 7038– 7046, DOI: 10.1021/la4000186There is no corresponding record for this reference.
- 51Mulvaney, P. Surface Plasmon Spectroscopy of Nanosized Metal Particles. Langmuir 1996, 12 (3), 788– 800, DOI: 10.1021/la9502711There is no corresponding record for this reference.
- 52Gao, C.; Hu, Y.; Wang, M.; Chi, M.; Yin, Y. Fully Alloyed Ag/Au Nanospheres: Combining the Plasmonic Property of Ag with the Stability of Au. J. Am. Chem. Soc. 2014, 136 (20), 7474– 7479, DOI: 10.1021/ja502890cThere is no corresponding record for this reference.
- 53Banga, R. J.; Meckes, B.; Narayan, S. P.; Sprangers, A. J.; Nguyen, S. T.; Mirkin, C. A. Cross-Linked Micellar Spherical Nucleic Acids from Thermoresponsive Templates. J. Am. Chem. Soc. 2017, 139 (12), 4278– 4281, DOI: 10.1021/jacs.6b13359There is no corresponding record for this reference.
- 54Zhang, C.; Hao, L.; Calabrese, C. M.; Zhou, Y.; Choi, C. H. J.; Xing, H.; Mirkin, C. A. Biodegradable DNA-Brush Block Copolymer Spherical Nucleic Acids Enable Transfection Agent-Free Intracellular Gene Regulation. Small 2015, 11 (40), 5360– 5368, DOI: 10.1002/smll.201501573There is no corresponding record for this reference.
- 55Chen, H.; Ding, Q.; Li, L.; Wei, P.; Niu, Z.; Kong, T.; Fu, P.; Wang, Y.; Li, J.; Wang, K. Extracellular Vesicle Spherical Nucleic Acids. JACS Au 2024, 4 (6), 2381– 2392, DOI: 10.1021/jacsau.4c00338There is no corresponding record for this reference.
- 56Zhu, S.; Xing, H.; Gordiichuk, P.; Park, J.; Mirkin, C. A. PLGA Spherical Nucleic Acids. Adv. Mater. 2018, 30 (22), 1707113, DOI: 10.1002/adma.201707113There is no corresponding record for this reference.
- 57Brodin, J. D.; Sprangers, A. J.; McMillan, J. R.; Mirkin, C. A. DNA-Mediated Cellular Delivery of Functional Enzymes. J. Am. Chem. Soc. 2015, 137 (47), 14838– 14841, DOI: 10.1021/jacs.5b09711There is no corresponding record for this reference.
- 58Sinegra, A. J.; Evangelopoulos, M.; Park, J.; Huang, Z.; Mirkin, C. A. Lipid Nanoparticle Spherical Nucleic Acids for Intracellular DNA and RNA Delivery. Nano Lett. 2021, 21 (15), 6584– 6591, DOI: 10.1021/acs.nanolett.1c01973There is no corresponding record for this reference.
- 59Cutler, J. I.; Zhang, K.; Zheng, D.; Auyeung, E.; Prigodich, A. E.; Mirkin, C. A. Polyvalent Nucleic Acid Nanostructures. J. Am. Chem. Soc. 2011, 133 (24), 9254– 9257, DOI: 10.1021/ja203375nThere is no corresponding record for this reference.
- 60Auyeung, E.; Cutler, J. I.; Macfarlane, R. J.; Jones, M. R.; Wu, J.; Liu, G.; Zhang, K.; Osberg, K. D.; Mirkin, C. A. Synthetically Programmable Nanoparticle Superlattices Using a Hollow Three-Dimensional Spacer Approach. Nat. Nanotechnol. 2012, 7 (1), 24– 28, DOI: 10.1038/nnano.2011.222There is no corresponding record for this reference.
- 61Teplensky, M. H.; Distler, M. E.; Kusmierz, C. D.; Evangelopoulos, M.; Gula, H.; Elli, D.; Tomatsidou, A.; Nicolaescu, V.; Gelarden, I.; Yeldandi, A. Spherical Nucleic Acids as an Infectious Disease Vaccine Platform. Proc. Natl. Acad. Sci. U. S. A. 2022, 119 (14), e2119093119 DOI: 10.1073/pnas.2119093119There is no corresponding record for this reference.
- 62Giljohann, D. A.; Seferos, D. S.; Prigodich, A. E.; Patel, P. C.; Mirkin, C. A. Gene Regulation with Polyvalent siRNA-Nanoparticle Conjugates. J. Am. Chem. Soc. 2009, 131 (6), 2072– 2073, DOI: 10.1021/ja808719pThere is no corresponding record for this reference.
- 63Liu, L.; Lu, H.; Shi, R.; Peng, X.-X.; Xiang, Q.; Wang, B.; Wan, Q.-Q.; Sun, Y.; Yang, F.; Zhang, G.-J. Synergy of Peptide-Nucleic Acid and Spherical Nucleic Acid Enabled Quantitative and Specific Detection of Tumor Exosomal MicroRNA. Anal. Chem. 2019, 91 (20), 13198– 13205, DOI: 10.1021/acs.analchem.9b03622There is no corresponding record for this reference.
- 64Lytton-Jean, A. K. R.; Gibbs-Davis, J. M.; Long, H.; Schatz, G. C.; Mirkin, C. A.; Nguyen, S. T. Highly Cooperative Behavior of Peptide Nucleic Acid-Linked DNA-Modified Gold-Nanoparticle and Comb-Polymer Aggregates. Adv. Mater. 2009, 21 (6), 706– 709, DOI: 10.1002/adma.200801724There is no corresponding record for this reference.
- 65McKenzie, F.; Faulds, K.; Graham, D. Sequence-Specific DNA Detection Using High-Affinity LNA-Functionalized Gold Nanoparticles. Small 2007, 3 (11), 1866– 1868, DOI: 10.1002/smll.200700225There is no corresponding record for this reference.
- 66Seferos, D. S.; Giljohann, D. A.; Rosi, N. L.; Mirkin, C. A. Locked Nucleic Acid-Nanoparticle Conjugates. ChemBioChem. 2007, 8 (11), 1230– 1232, DOI: 10.1002/cbic.200700262There is no corresponding record for this reference.
- 67Dittmar, J. W.; Teplensky, M. H.; Evangelopoulos, M.; Qin, L.; Zhang, B.; Mirkin, C. A. Tuning DNA Dissociation from Spherical Nucleic Acids for Enhanced Immunostimulation. ACS Nano 2023, 17 (18), 17996– 18007, DOI: 10.1021/acsnano.3c04333There is no corresponding record for this reference.
- 68Zhang, K.; Hao, L.; Hurst, S. J.; Mirkin, C. A. Antibody-Linked Spherical Nucleic Acids for Cellular Targeting. J. Am. Chem. Soc. 2012, 134 (40), 16488– 16491, DOI: 10.1021/ja306854dThere is no corresponding record for this reference.
- 69Zhang, W.; Callmann, C. E.; Meckes, B.; Mirkin, C. A. Tumor-Associated Enzyme-Activatable Spherical Nucleic Acids. ACS Nano 2022, 16 (7), 10931– 10942, DOI: 10.1021/acsnano.2c03323There is no corresponding record for this reference.
- 70Choi, C. H. J.; Hao, L.; Narayan, S. P.; Auyeung, E.; Mirkin, C. A. Mechanism for the Endocytosis of Spherical Nucleic Acid Nanoparticle Conjugates. Proc. Natl. Acad. Sci. U. S. A. 2013, 110 (19), 7625– 7630, DOI: 10.1073/pnas.1305804110There is no corresponding record for this reference.
- 71Wu, X. A.; Choi, C. H. J.; Zhang, C.; Hao, L.; Mirkin, C. A. Intracellular Fate of Spherical Nucleic Acid Nanoparticle Conjugates. J. Am. Chem. Soc. 2014, 136 (21), 7726– 7733, DOI: 10.1021/ja503010aThere is no corresponding record for this reference.
- 72Han, Z.; Guo, A. X.; Luo, T.; Cai, T.; Mirkin, C. A. Biomineralization of Semiconductor Quantum Dots Using DNA-Functionalized Protein Nanoreactors. Sci. Adv. 2025, 11 (19), eadv6906 DOI: 10.1126/sciadv.adv6906There is no corresponding record for this reference.
- 73Hsiao, J. C.; Buryska, T.; Kim, E.; Howes, P. D.; deMello, A. J. Tuning DNA-Nanoparticle Conjugate Properties Allows Modulation of Nuclease Activity. Nanoscale 2021, 13 (9), 4956– 4970, DOI: 10.1039/D0NR08668AThere is no corresponding record for this reference.
- 74Seferos, D. S.; Prigodich, A. E.; Giljohann, D. A.; Patel, P. C.; Mirkin, C. A. Polyvalent DNA Nanoparticle Conjugates Stabilize Nucleic Acids. Nano Lett. 2009, 9 (1), 308– 311, DOI: 10.1021/nl802958fThere is no corresponding record for this reference.
- 75Kyriazi, M.-E.; El-Sagheer, A. H.; Medintz, I. L.; Brown, T.; Kanaras, A. G. An Investigation into the Resistance of Spherical Nucleic Acids against DNA Enzymatic Degradation. Bioconjugate Chem. 2022, 33 (1), 219– 225, DOI: 10.1021/acs.bioconjchem.1c00540There is no corresponding record for this reference.
- 76Giljohann, D. A.; Seferos, D. S.; Patel, P. C.; Millstone, J. E.; Rosi, N. L.; Mirkin, C. A. Oligonucleotide Loading Determines Cellular Uptake of DNA-Modified Gold Nanoparticles. Nano Lett. 2007, 7 (12), 3818– 3821, DOI: 10.1021/nl072471qThere is no corresponding record for this reference.
- 77Hurst, S. J.; Lytton-Jean, A. K. R.; Mirkin, C. A. Maximizing DNA Loading on a Range of Gold Nanoparticle Sizes. Anal. Chem. 2006, 78 (24), 8313– 8318, DOI: 10.1021/ac0613582There is no corresponding record for this reference.
- 78Liu, B.; Liu, J. Freezing-Driven DNA Adsorption on Gold Nanoparticles: Tolerating Extremely Low Salt Concentration but Requiring High DNA Concentration. Langmuir 2019, 35 (19), 6476– 6482, DOI: 10.1021/acs.langmuir.9b00746There is no corresponding record for this reference.
- 79Hao, Y.; Li, Y.; Song, L.; Deng, Z. Flash Synthesis of Spherical Nucleic Acids with Record DNA Density. J. Am. Chem. Soc. 2021, 143 (8), 3065– 3069, DOI: 10.1021/jacs.1c00568There is no corresponding record for this reference.
- 80Ye, Y.; Hao, Y.; Ye, M.; Song, X.; Deng, Z. Evaporative Drying: A General and Readily Scalable Route to Spherical Nucleic Acids with Quantitative, Fully Tunable, and Record-High DNA Loading. Small 2022, 18 (24), 2202458, DOI: 10.1002/smll.202202458There is no corresponding record for this reference.
- 81Liu, B.; Liu, J. Freezing Directed Construction of Bio/Nano Interfaces: Reagentless Conjugation, Denser Spherical Nucleic Acids, and Better Nanoflares. J. Am. Chem. Soc. 2017, 139 (28), 9471– 9474, DOI: 10.1021/jacs.7b04885There is no corresponding record for this reference.
- 82Eller, M. J.; Chandra, K.; Coughlin, E. E.; Odom, T. W.; Schweikert, E. A. Label Free Particle-by-Particle Quantification of DNA Loading on Sorted Gold Nanostars. Anal. Chem. 2019, 91 (9), 5566– 5572, DOI: 10.1021/acs.analchem.8b03715There is no corresponding record for this reference.
- 83Lee, J.; Lee, S. Non-Invasive, Reliable, and Fast Quantification of DNA Loading on Gold Nanoparticles by a One-Step Optical Measurement. Anal. Chem. 2023, 95 (3), 1856– 1866, DOI: 10.1021/acs.analchem.2c03378There is no corresponding record for this reference.
- 84Lee, S.; Pandit, S.; Gilman, G.; Bhattacharya, A.; Samanta, D. Label-Free Quantification of DNA Loading on Centrifugation-Resistant Spherical Nucleic Acids. Anal. Chem. 2025, 97 (23), 12267– 12275, DOI: 10.1021/acs.analchem.5c01112There is no corresponding record for this reference.
- 85Li, H.; Zhang, B.; Lu, X.; Tan, X.; Jia, F.; Xiao, Y.; Cheng, Z.; Li, Y.; Silva, D. O.; Schrekker, H. S. Molecular Spherical Nucleic Acids. Proc. Natl. Acad. Sci. U. S. A. 2018, 115 (17), 4340– 4344, DOI: 10.1073/pnas.1801836115There is no corresponding record for this reference.
- 86Cavaliere, J. P.; Forsyth, C. M.; Guo, A. X.; Kang, J.; Ngo, K. H.; Mirkin, C. A. Amphiphilic Polymeric Micelle Spherical Nucleic Acids (SNAs) as Drug Delivery Vehicles. Bioconjugate Chem. 2026, 37, 83, DOI: 10.1021/acs.bioconjchem.5c00507There is no corresponding record for this reference.
- 87Distler, M. E.; Teplensky, M. H.; Bujold, K. E.; Kusmierz, C. D.; Evangelopoulos, M.; Mirkin, C. A. DNA Dendrons as Agents for Intracellular Delivery. J. Am. Chem. Soc. 2021, 143 (34), 13513– 13518, DOI: 10.1021/jacs.1c07240There is no corresponding record for this reference.
- 88Chinen, A. B.; Guan, C. M.; Ko, C. H.; Mirkin, C. A. The Impact of Protein Corona Formation on the Macrophage Cellular Uptake and Biodistribution of Spherical Nucleic Acids. Small 2017, 13 (16), 1603847, DOI: 10.1002/smll.201603847There is no corresponding record for this reference.
- 89Mahajan, A. S.; Stegh, A. H. Spherical Nucleic Acids as Precision Therapeutics for the Treatment of Cancer-From Bench to Bedside. Cancers 2022, 14 (7), 1615, DOI: 10.3390/cancers14071615There is no corresponding record for this reference.
- 90Yan, R.; Chen, J.; Wang, J.; Rao, J.; Du, X.; Liu, Y.; Zhang, L.; Qiu, L.; Liu, B.; Zhao, Y.-D. A NanoFlare-Based Strategy for In Situ Tumor Margin Demarcation and Neoadjuvant Gene/Photothermal Therapy. Small 2018, 14 (50), 1802745, DOI: 10.1002/smll.201802745There is no corresponding record for this reference.
- 91Radovic-Moreno, A. F.; Chernyak, N.; Mader, C. C.; Nallagatla, S.; Kang, R. S.; Hao, L.; Walker, D. A.; Halo, T. L.; Merkel, T. J.; Rische, C. H. Immunomodulatory Spherical Nucleic Acids. Proc. Natl. Acad. Sci. U. S. A. 2015, 112 (13), 3892– 3897, DOI: 10.1073/pnas.1502850112There is no corresponding record for this reference.
- 92O’Brien, M. N.; Girard, M.; Lin, H.-X.; Millan, J. A.; Olvera de la Cruz, M.; Lee, B.; Mirkin, C. A. Exploring the Zone of Anisotropy and Broken Symmetries in DNA-Mediated Nanoparticle Crystallization. Proc. Natl. Acad. Sci. U. S. A. 2016, 113 (38), 10485– 10490, DOI: 10.1073/pnas.1611808113There is no corresponding record for this reference.
- 93Jones, M. R.; Macfarlane, R. J.; Lee, B.; Zhang, J.; Young, K. L.; Senesi, A. J.; Mirkin, C. A. DNA-Nanoparticle Superlattices Formed from Anisotropic Building Blocks. Nat. Mater. 2010, 9 (11), 913– 917, DOI: 10.1038/nmat2870There is no corresponding record for this reference.
- 94O’Brien, M. N.; Lin, H.-X.; Girard, M.; Olvera de la Cruz, M.; Mirkin, C. A. Programming Colloidal Crystal Habit with Anisotropic Nanoparticle Building Blocks and DNA Bonds. J. Am. Chem. Soc. 2016, 138 (44), 14562– 14565, DOI: 10.1021/jacs.6b09704There is no corresponding record for this reference.
- 95O’Brien, M. N.; Jones, M. R.; Lee, B.; Mirkin, C. A. Anisotropic Nanoparticle Complementarity in DNA-Mediated Co-Crystallization. Nat. Mater. 2015, 14 (8), 833– 839, DOI: 10.1038/nmat4293There is no corresponding record for this reference.
- 96Chang, C.-C.; Wu, H.-L.; Kuo, C.-H.; Huang, M. H. Hydrothermal Synthesis of Monodispersed Octahedral Gold Nanocrystals with Five Different Size Ranges and Their Self-Assembled Structures. Chem. Mater. 2008, 20 (24), 7570– 7574, DOI: 10.1021/cm8021984There is no corresponding record for this reference.
- 97Jin, R.; Cao, Y.; Mirkin, C. A.; Kelly, K. L.; Schatz, G. C.; Zheng, J. G. Photoinduced Conversion of Silver Nanospheres to Nanoprisms. Science 2001, 294 (5548), 1901– 1903, DOI: 10.1126/science.1066541There is no corresponding record for this reference.
- 98Jana, N. R.; Gearheart, L.; Murphy, C. J. Wet Chemical Synthesis of High Aspect Ratio Cylindrical Gold Nanorods. J. Phys. Chem. B 2001, 105 (19), 4065– 4067, DOI: 10.1021/jp0107964There is no corresponding record for this reference.
- 99O’Brien, M. N.; Jones, M. R.; Brown, K. A.; Mirkin, C. A. Universal Noble Metal Nanoparticle Seeds Realized Through Iterative Reductive Growth and Oxidative Dissolution Reactions. J. Am. Chem. Soc. 2014, 136 (21), 7603– 7606, DOI: 10.1021/ja503509kThere is no corresponding record for this reference.
- 100Akbulut, O.; Mace, C. R.; Martinez, R. V.; Kumar, A. A.; Nie, Z.; Patton, M. R.; Whitesides, G. M. Separation of Nanoparticles in Aqueous Multiphase Systems through Centrifugation. Nano Lett. 2012, 12 (8), 4060– 4064, DOI: 10.1021/nl301452xThere is no corresponding record for this reference.
- 101Jana, N. R. Nanorod Shape Separation Using Surfactant Assisted Self-Assembly. Chem. Commun. 2003, (15), 1950– 1951, DOI: 10.1039/b303103aThere is no corresponding record for this reference.
- 102Scarabelli, L.; Coronado-Puchau, M.; Giner-Casares, J. J.; Langer, J.; Liz-Marzán, L. M. Monodisperse Gold Nanotriangles: Size Control, Large-Scale Self-Assembly, and Performance in Surface-Enhanced Raman Scattering. ACS Nano 2014, 8 (6), 5833– 5842, DOI: 10.1021/nn500727wThere is no corresponding record for this reference.
- 103Cheng, Z.; Jones, M. R. Assembly of Planar Chiral Superlattices from Achiral Building Blocks. Nat. Commun. 2022, 13 (1), 4207, DOI: 10.1038/s41467-022-31868-2There is no corresponding record for this reference.
- 104Millstone, J. E.; Georganopoulou, D. G.; Xu, X.; Wei, W.; Li, S.; Mirkin, C. A. DNA-Gold Triangular Nanoprism Conjugates. Small 2008, 4 (12), 2176– 2180, DOI: 10.1002/smll.200800931There is no corresponding record for this reference.
- 105Zhou, W.; Li, Y.; Je, K.; Vo, T.; Lin, H.; Partridge, B. E.; Huang, Z.; Glotzer, S. C.; Mirkin, C. A. Space-Tiled Colloidal Crystals from DNA-Forced Shape-Complementary Polyhedra Pairing. Science 2024, 383 (6680), 312– 319, DOI: 10.1126/science.adj1021There is no corresponding record for this reference.
- 106Li, Y.; Lin, H.; Zhou, W.; Sun, L.; Samanta, D.; Mirkin, C. A. Corner-, Edge-, and Facet-Controlled Growth of Nanocrystals. Sci. Adv. 2021, 7 (3), eabf1410 DOI: 10.1126/sciadv.abf1410There is no corresponding record for this reference.
- 107Brodin, J. D.; Auyeung, E.; Mirkin, C. A. DNA-Mediated Engineering of Multicomponent Enzyme Crystals. Proc. Nat. Acad. Sci. U. S. A. 2015, 112 (15), 4564– 4569, DOI: 10.1073/pnas.1503533112There is no corresponding record for this reference.
- 108Wang, Y.; Wang, Y.; Breed, D. R.; Manoharan, V. N.; Feng, L.; Hollingsworth, A. D.; Weck, M.; Pine, D. J. Colloids with Valence and Specific Directional Bonding. Nature 2012, 491 (7422), 51– 55, DOI: 10.1038/nature11564There is no corresponding record for this reference.
- 109Chen, G.; Gibson, K. J.; Liu, D.; Rees, H. C.; Lee, J.-H.; Xia, W.; Lin, R.; Xin, H. L.; Gang, O.; Weizmann, Y. Regioselective Surface Encoding of Nanoparticles for Programmable Self-Assembly. Nat. Mater. 2019, 18 (2), 169– 174, DOI: 10.1038/s41563-018-0231-1There is no corresponding record for this reference.
- 110Tian, Y.; Zhang, Y.; Wang, T.; Xin, H. L.; Li, H.; Gang, O. Lattice Engineering through Nanoparticle-DNA Frameworks. Nat. Mater. 2016, 15 (6), 654– 661, DOI: 10.1038/nmat4571There is no corresponding record for this reference.
- 111Papavassiliou, G. C. Optical Properties of Small Inorganic and Organic Metal Particles. Prog. Solid State Chem. 1979, 12 (3), 185– 271, DOI: 10.1016/0079-6786(79)90001-3There is no corresponding record for this reference.
- 112Elghanian, R.; Storhoff, J. J.; Mucic, R. C.; Letsinger, R. L.; Mirkin, C. A. Selective Colorimetric Detection of Polynucleotides Based on the Distance-Dependent Optical Properties of Gold Nanoparticles. Science 1997, 277 (5329), 1078– 1081, DOI: 10.1126/science.277.5329.1078There is no corresponding record for this reference.
- 113Storhoff, J. J.; Lazarides, A. A.; Mucic, R. C.; Mirkin, C. A.; Letsinger, R. L.; Schatz, G. C. What Controls the Optical Properties of DNA-Linked Gold Nanoparticle Assemblies?. J. Am. Chem. Soc. 2000, 122 (19), 4640– 4650, DOI: 10.1021/ja993825lThere is no corresponding record for this reference.
- 114Xia, C.; Cheng, H.; Hou, X.; Zhang, Y.; Zhou, X.; Yan, Q.; Cao, S. Spherical Nucleic Acids for Biomedical Applications. Adv. Sens. Energy Mater. 2024, 3 (4), 100117, DOI: 10.1016/j.asems.2024.100117There is no corresponding record for this reference.
- 115Lee, J.-S.; Han, M. S.; Mirkin, C. A. Colorimetric Detection of Mercuric Ion (Hg2+) in Aqueous Media Using DNA-Functionalized Gold Nanoparticles. Angew. Chem., Int. Ed. 2007, 46 (22), 4093– 4096, DOI: 10.1002/anie.200700269There is no corresponding record for this reference.
- 116Lee, J.-S.; Mirkin, C. A. Chip-Based Scanometric Detection of Mercuric Ion Using DNA-Functionalized Gold Nanoparticles. Anal. Chem. 2008, 80 (17), 6805– 6808, DOI: 10.1021/ac801046aThere is no corresponding record for this reference.
- 117Xu, X.; Daniel, W. L.; Wei, W.; Mirkin, C. A. Colorimetric Cu2+ Detection Using DNA-Modified Gold-Nanoparticle Aggregates as Probes and Click Chemistry. Small 2010, 6 (5), 623– 626, DOI: 10.1002/smll.200901691There is no corresponding record for this reference.
- 118Han, M. S.; Lytton-Jean, A. K. R.; Oh, B.-K.; Heo, J.; Mirkin, C. A. Colorimetric Screening of DNA-Binding Molecules with Gold Nanoparticle Probes. Angew. Chem., Int. Ed. 2006, 45 (11), 1807– 1810, DOI: 10.1002/anie.200504277There is no corresponding record for this reference.
- 119Nam, J.-M.; Thaxton, C. S.; Mirkin, C. A. Nanoparticle-Based Bio-Bar Codes for the Ultrasensitive Detection of Proteins. Science 2003, 301 (5641), 1884– 1886, DOI: 10.1126/science.1088755There is no corresponding record for this reference.
- 120Pavlov, V.; Xiao, Y.; Shlyahovsky, B.; Willner, I. Aptamer-Functionalized Au Nanoparticles for the Amplified Optical Detection of Thrombin. J. Am. Chem. Soc. 2004, 126 (38), 11768– 11769, DOI: 10.1021/ja046970uThere is no corresponding record for this reference.
- 121Kim, D.; Daniel, W. L.; Mirkin, C. A. Microarray-Based Multiplexed Scanometric Immunoassay for Protein Cancer Markers Using Gold Nanoparticle Probes. Anal. Chem. 2009, 81 (21), 9183– 9187, DOI: 10.1021/ac9018389There is no corresponding record for this reference.
- 122Xu, X.; Han, M. S.; Mirkin, C. A. A Gold-Nanoparticle-Based Real-Time Colorimetric Screening Method for Endonuclease Activity and Inhibition. Angew. Chem., Int. Ed. 2007, 46 (19), 3468– 3470, DOI: 10.1002/anie.200605249There is no corresponding record for this reference.
- 123Zheng, G.; Daniel, W. L.; Mirkin, C. A. A New Approach to Amplified Telomerase Detection with Polyvalent Oligonucleotide Nanoparticle Conjugates. J. Am. Chem. Soc. 2008, 130 (30), 9644– 9645, DOI: 10.1021/ja803035pThere is no corresponding record for this reference.
- 124Chen, L.; Yuan, A.; Zhang, D.; Xie, W.; Peng, H. Fluorescence and Colorimetric Analysis of β-Estradiol Based on Aptamer Assembled Spherical Nucleic Acids. Anal. Methods 2024, 16 (37), 6356– 6363, DOI: 10.1039/D4AY01283FThere is no corresponding record for this reference.
- 125Storhoff, J. J.; Elghanian, R.; Mucic, R. C.; Mirkin, C. A.; Letsinger, R. L. One-Pot Colorimetric Differentiation of Polynucleotides with Single Base Imperfections Using Gold Nanoparticle Probes. J. Am. Chem. Soc. 1998, 120 (9), 1959– 1964, DOI: 10.1021/ja972332iThere is no corresponding record for this reference.
- 126Taton, T. A.; Mirkin, C. A.; Letsinger, R. L. Scanometric DNA Array Detection with Nanoparticle Probes. Science 2000, 289 (5485), 1757– 1760, DOI: 10.1126/science.289.5485.1757There is no corresponding record for this reference.
- 127Alhasan, A. H.; Kim, D. Y.; Daniel, W. L.; Watson, E.; Meeks, J. J.; Thaxton, C. S.; Mirkin, C. A. Scanometric MicroRNA Array Profiling of Prostate Cancer Markers Using Spherical Nucleic Acid-Gold Nanoparticle Conjugates. Anal. Chem. 2012, 84 (9), 4153– 4160, DOI: 10.1021/ac3004055There is no corresponding record for this reference.
- 128The Verigene System; Diasorin. https://us.diasorin.com/en/molecular-diagnostics/tools/verigene-system?f%5B0%5D=mdx_disease%3A249 (accessed 2025-03-24).There is no corresponding record for this reference.
- 129Hurst, S. J.; Hill, H. D.; Mirkin, C. A. “Three-Dimensional Hybridization” with Polyvalent DNA-Gold Nanoparticle Conjugates. J. Am. Chem. Soc. 2008, 130 (36), 12192– 12200, DOI: 10.1021/ja804266jThere is no corresponding record for this reference.
- 130Lytton-Jean, A. K. R.; Mirkin, C. A. A Thermodynamic Investigation into the Binding Properties of DNA Functionalized Gold Nanoparticle Probes and Molecular Fluorophore Probes. J. Am. Chem. Soc. 2005, 127 (37), 12754– 12755, DOI: 10.1021/ja052255oThere is no corresponding record for this reference.
- 131Fong, L.-K.; Wang, Z.; Schatz, G. C.; Luijten, E.; Mirkin, C. A. The Role of Structural Enthalpy in Spherical Nucleic Acid Hybridization. J. Am. Chem. Soc. 2018, 140 (20), 6226– 6230, DOI: 10.1021/jacs.8b03459There is no corresponding record for this reference.
- 132Randeria, P. S.; Jones, M. R.; Kohlstedt, K. L.; Banga, R. J.; Olvera de la Cruz, M.; Schatz, G. C.; Mirkin, C. A. What Controls the Hybridization Thermodynamics of Spherical Nucleic Acids?. J. Am. Chem. Soc. 2015, 137 (10), 3486– 3489, DOI: 10.1021/jacs.5b00670There is no corresponding record for this reference.
- 133Wang, X.; Huang, K.; Cheng, N.; Liu, J. Thermal Drying Synthesized Density-Tunable and High Stability Nonthiolated Spherical Nucleic Acids for Split Aptamer and Lateral Flow Biosensors. Biosens. Bioelectron. 2025, 289, 117863, DOI: 10.1016/j.bios.2025.117863There is no corresponding record for this reference.
- 134Nam, J.-M.; Park, S.-J.; Mirkin, C. A. Bio-Barcodes Based on Oligonucleotide-Modified Nanoparticles. J. Am. Chem. Soc. 2002, 124 (15), 3820– 3821, DOI: 10.1021/ja0178766There is no corresponding record for this reference.
- 135Nam, J.-M.; Stoeva, S. I.; Mirkin, C. A. Bio-Bar-Code-Based DNA Detection with PCR-like Sensitivity. J. Am. Chem. Soc. 2004, 126 (19), 5932– 5933, DOI: 10.1021/ja049384+There is no corresponding record for this reference.
- 136Thaxton, C. S.; Hill, H. D.; Georganopoulou, D. G.; Stoeva, S. I.; Mirkin, C. A. A Bio-Bar-Code Assay Based upon Dithiothreitol-Induced Oligonucleotide Release. Anal. Chem. 2005, 77 (24), 8174– 8178, DOI: 10.1021/ac0514265There is no corresponding record for this reference.
- 137Hill, H. D.; Mirkin, C. A. The Bio-Barcode Assay for the Detection of Protein and Nucleic Acid Targets Using DTT-Induced Ligand Exchange. Nat. Protoc. 2006, 1 (1), 324– 336, DOI: 10.1038/nprot.2006.51There is no corresponding record for this reference.
- 138Goluch, E. D.; Savka, S. I.; Georganopoulou, D. G.; Nam, J. M.; Shaikh, K. A.; Ryu, K. S.; Chiesl, T. N.; Barron, A. E.; Mirkin, C. A.; Liu, C. Chip-Based High-Sensitivity Detection of Multiple Disease Biomarkers. 19th IEEE International Conference on Micro Electro Mechanical Systems, Istanbul, Turkey; IEEE, 2006; pp 478– 481. DOI: 10.1109/memsys.2006.1627840 .There is no corresponding record for this reference.
- 139Stoeva, S. I.; Lee, J.-S.; Thaxton, C. S.; Mirkin, C. A. Multiplexed DNA Detection with Biobarcoded Nanoparticle Probes. Angew. Chem., Int. Ed. 2006, 45 (20), 3303– 3306, DOI: 10.1002/anie.200600124There is no corresponding record for this reference.
- 140Stoeva, S. I.; Lee, J.-S.; Smith, J. E.; Rosen, S. T.; Mirkin, C. A. Multiplexed Detection of Protein Cancer Markers with Biobarcoded Nanoparticle Probes. J. Am. Chem. Soc. 2006, 128 (26), 8378– 8379, DOI: 10.1021/ja0613106There is no corresponding record for this reference.
- 141Thaxton, C. S.; Elghanian, R.; Thomas, A. D.; Stoeva, S. I.; Lee, J.-S.; Smith, N. D.; Schaeffer, A. J.; Klocker, H.; Horninger, W.; Bartsch, G. Nanoparticle-Based Bio-Barcode Assay Redefines “Undetectable” PSA and Biochemical Recurrence after Radical Prostatectomy. Proc. Natl. Acad. Sci. U. S. A. 2009, 106 (44), 18437– 18442, DOI: 10.1073/pnas.0904719106There is no corresponding record for this reference.
- 142Lee, J. H.; Wang, Z.; Liu, J.; Lu, Y. Highly Sensitive and Selective Colorimetric Sensors for Uranyl (UO22+): Development and Comparison of Labeled and Label-Free DNAzyme-Gold Nanoparticle Systems. J. Am. Chem. Soc. 2008, 130 (43), 14217– 14226, DOI: 10.1021/ja803607zThere is no corresponding record for this reference.
- 143Wu, P.; Hwang, K.; Lan, T.; Lu, Y. A DNAzyme-Gold Nanoparticle Probe for Uranyl Ion in Living Cells. J. Am. Chem. Soc. 2013, 135 (14), 5254– 5257, DOI: 10.1021/ja400150vThere is no corresponding record for this reference.
- 144Li, Y.; Liu, M.-L.; Liang, W.-B.; Zhuo, Y.; He, X.-J. Spherical Nucleic Acid Enzyme Programmed Network to Accelerate CRISPR Assays for Electrochemiluminescence Biosensing Applications. Biosens. Bioelectron. 2023, 238, 115589, DOI: 10.1016/j.bios.2023.115589There is no corresponding record for this reference.
- 145Yang, X.; Yuan, L.; Xu, Y.; He, B. Target-Catalyzed Self-Assembled Spherical G-Quadruplex/Hemin DNAzymes for Highly Sensitive Colorimetric Detection of MicroRNA in Serum. Anal. Chim. Acta 2023, 1247, 340879, DOI: 10.1016/j.aca.2023.340879There is no corresponding record for this reference.
- 146Sun, Y.; Shi, L.; Wang, Q.; Mi, L.; Li, T. Spherical Nucleic Acid Enzyme (SNAzyme) Boosted Chemiluminescence miRNA Imaging Using a Smartphone. Anal. Chem. 2019, 91 (5), 3652– 3658, DOI: 10.1021/acs.analchem.8b05696There is no corresponding record for this reference.
- 147Shi, L.; Sun, Y.; Mi, L.; Li, T. Target-Catalyzed Self-Growing Spherical Nucleic Acid Enzyme (SNAzyme) as a Double Amplifier for Ultrasensitive Chemiluminescence MicroRNA Detection. ACS Sens. 2019, 4 (12), 3219– 3226, DOI: 10.1021/acssensors.9b01655There is no corresponding record for this reference.
- 148Shi, L.; Liu, C.; Wang, H.; Zheng, J.; Wang, Q.; Shi, L.; Li, T. Framework and Spherical Nucleic Acids Synergistically Enhanced Electrochemiluminescence Nanosensors for Rapidly Diagnosing Acute Myocardial Infarction Based on Circulating MicroRNA Levels. Anal. Chem. 2022, 94 (41), 14394– 14401, DOI: 10.1021/acs.analchem.2c03144There is no corresponding record for this reference.
- 149Wang, H.; Shi, L.; Wang, Q.; Shi, L.; Li, T. Robust Noncovalent Spherical Nucleic Acid Enzymes (SNAzymes) for Ultrasensitive Unamplified Electrochemiluminescence Detection of Endogenous Myocardial MicroRNAs. Biosens. Bioelectron. 2023, 241, 115687, DOI: 10.1016/j.bios.2023.115687There is no corresponding record for this reference.
- 150Qu, S.; Shi, L.; Li, H.; Li, T. A Label-Free Silicon-Based Spherical Nucleic Acid Enzyme (SNAzyme) for Ultrasensitive Chemiluminescence Detection of Acute Myocardial Infarction-Related Nucleic Acids. Sens. Diagn. 2023, 2 (1), 194– 202, DOI: 10.1039/D2SD00170EThere is no corresponding record for this reference.
- 151Wang, K.; Zhu, W.; Li, X.; Ling, G.; Zhang, P. Programmable DNAzyme Walkers on Gold Nanoplatforms: From Rational Design to Bioimaging/Biosensing Breakthroughs. Acta Biomater. 2025, 206, 74– 87, DOI: 10.1016/j.actbio.2025.09.020There is no corresponding record for this reference.
- 152Zhang, K.; Cutler, J. I.; Zhang, J.; Zheng, D.; Auyeung, E.; Mirkin, C. A. Nanopod Formation through Gold Nanoparticle Templated and Catalyzed Cross-linking of Polymers Bearing Pendant Propargyl Ethers. J. Am. Chem. Soc. 2010, 132 (43), 15151– 15153, DOI: 10.1021/ja107224sThere is no corresponding record for this reference.
- 153Zhang, K.; Zheng, D.; Hao, L.; Cutler, J. I.; Auyeung, E.; Mirkin, C. A. ImmunoPods: Polymer Shells with Native Antibody Cross-Links. Angew. Chem., Int. Ed. 2012, 51 (5), 1169– 1172, DOI: 10.1002/anie.201106313There is no corresponding record for this reference.
- 154Massich, M. D.; Giljohann, D. A.; Seferos, D. S.; Ludlow, L. E.; Horvath, C. M.; Mirkin, C. A. Regulating Immune Response Using Polyvalent Nucleic Acid-Gold Nanoparticle Conjugates. Mol. Pharmaceutics 2009, 6 (6), 1934– 1940, DOI: 10.1021/mp900172mThere is no corresponding record for this reference.
- 155Massich, M. D.; Giljohann, D. A.; Schmucker, A. L.; Patel, P. C.; Mirkin, C. A. Cellular Response of Polyvalent Oligonucleotide-Gold Nanoparticle Conjugates. ACS Nano 2010, 4 (10), 5641– 5646, DOI: 10.1021/nn102228sThere is no corresponding record for this reference.
- 156Zhao, J.; He, J.; Ding, X.; Zhou, Y.; Liu, M.; Chen, X.; Quan, W.; Hua, D.; Tong, J.; Li, J. DENV Peptides Delivered as Spherical Nucleic Acid Constructs Enhance Antigen Presentation and Immunogenicity in Vitro and in Vivo. Int. J. Nanomedicine 2024, 19, 9757– 9770, DOI: 10.2147/IJN.S467427There is no corresponding record for this reference.
- 157Sparwasser, T.; Hültner, L.; Koch, E. S.; Luz, A.; Lipford, G. B.; Wagner, H. Immunostimulatory CpG-Oligodeoxynucleotides Cause Extramedullary Murine Hemopoiesis. J. Immunol. 1999, 162 (4), 2368– 2374, DOI: 10.4049/jimmunol.162.4.2368There is no corresponding record for this reference.
- 158Sparwasser, T.; Miethke, T.; Lipford, G.; Borschert, K.; Häcker, H.; Heeg, K.; Wagner, H. Bacterial DNA Causes Septic Shock. Nature 1997, 386 (6623), 336– 337, DOI: 10.1038/386336a0There is no corresponding record for this reference.
- 159Ferrer, J. R.; Sinegra, A. J.; Ivancic, D.; Yeap, X. Y.; Qiu, L.; Wang, J.-J.; Zhang, Z. J.; Wertheim, J. A.; Mirkin, C. A. Structure-Dependent Biodistribution of Liposomal Spherical Nucleic Acids. ACS Nano 2020, 14 (2), 1682– 1693, DOI: 10.1021/acsnano.9b07254There is no corresponding record for this reference.
- 160Asohan, J.; Fakih, H. H.; Das, T.; Sleiman, H. F. Control of the Assembly and Disassembly of Spherical Nucleic Acids Is Critical for Enhanced Gene Silencing. ACS Nano 2024, 18 (5), 3996– 4007, DOI: 10.1021/acsnano.3c05940There is no corresponding record for this reference.
- 161Meckes, B.; Banga, R. J.; Nguyen, S. T.; Mirkin, C. A. Enhancing the Stability and Immunomodulatory Activity of Liposomal Spherical Nucleic Acids through Lipid-Tail DNA Modifications. Small 2018, 14 (5), 1702909, DOI: 10.1002/smll.201702909There is no corresponding record for this reference.
- 162Callmann, C. E.; Kusmierz, C. D.; Dittmar, J. W.; Broger, L.; Mirkin, C. A. Impact of Liposomal Spherical Nucleic Acid Structure on Immunotherapeutic Function. ACS Cent. Sci. 2021, 7 (5), 892– 899, DOI: 10.1021/acscentsci.1c00181There is no corresponding record for this reference.
- 163Huang, Z. N.; Cole, L. E.; Callmann, C. E.; Wang, S.; Mirkin, C. A. Sequence Multiplicity within Spherical Nucleic Acids. ACS Nano 2020, 14 (1), 1084– 1092, DOI: 10.1021/acsnano.9b08750There is no corresponding record for this reference.
- 164Liu, H.; Moynihan, K. D.; Zheng, Y.; Szeto, G. L.; Li, A. V.; Huang, B.; Van Egeren, D. S.; Park, C.; Irvine, D. J. Structure-Based Programming of Lymph-Node Targeting in Molecular Vaccines. Nature 2014, 507 (7493), 519– 522, DOI: 10.1038/nature12978There is no corresponding record for this reference.
- 165Smith, A. A. A.; Gale, E. C.; Roth, G. A.; Maikawa, C. L.; Correa, S.; Yu, A. C.; Appel, E. A. Nanoparticles Presenting Potent TLR7/8 Agonists Enhance Anti-PD-L1 Immunotherapy in Cancer Treatment. Biomacromolecules 2020, 21 (9), 3704– 3712, DOI: 10.1021/acs.biomac.0c00812There is no corresponding record for this reference.
- 166Oyler-Yaniv, A.; Oyler-Yaniv, J.; Whitlock, B. M.; Liu, Z.; Germain, R. N.; Huse, M.; Altan-Bonnet, G.; Krichevsky, O. A Tunable Diffusion-Consumption Mechanism of Cytokine Propagation Enables Plasticity in Cell-to-Cell Communication in the Immune System. Immunity 2017, 46 (4), 609– 620, DOI: 10.1016/j.immuni.2017.03.011There is no corresponding record for this reference.
- 167Kusmierz, C. D.; Bujold, K. E.; Callmann, C. E.; Mirkin, C. A. Defining the Design Parameters for in Vivo Enzyme Delivery Through Protein Spherical Nucleic Acids. ACS Cent. Sci. 2020, 6 (5), 815– 822, DOI: 10.1021/acscentsci.0c00313There is no corresponding record for this reference.
- 168Narayan, S. P.; Choi, C. H. J.; Hao, L.; Calabrese, C. M.; Auyeung, E.; Zhang, C.; Goor, O. J. G. M.; Mirkin, C. A. The Sequence-Specific Cellular Uptake of Spherical Nucleic Acid Nanoparticle Conjugates. Small 2015, 11 (33), 4173– 4182, DOI: 10.1002/smll.201500027There is no corresponding record for this reference.
- 169Pearson, A. M.; Rich, A.; Krieger, M. Polynucleotide Binding to Macrophage Scavenger Receptors Depends on the Formation of Base-Quartet-Stabilized Four-Stranded Helices. J. Biol. Chem. 1993, 268 (5), 3546– 3554, DOI: 10.1016/S0021-9258(18)53729-7There is no corresponding record for this reference.
- 170Paranandi, K. S.; Lee, A.; Stanic, N.; Mirkin, C. A. Cellular Export Fate of Liposomal Spherical Nucleic Acids. ACS Nano 2023, 17 (19), 19000– 19010, DOI: 10.1021/acsnano.3c04608There is no corresponding record for this reference.
- 171Wiraja, C.; Mori, Y.; Ichimura, T.; Hwang, J.; Xu, C.; Bonventre, J. V. Nephrotoxicity Assessment with Human Kidney Tubuloids Using Spherical Nucleic Acid-Based mRNA Nanoflares. Nano Lett. 2021, 21 (13), 5850– 5858, DOI: 10.1021/acs.nanolett.1c01840There is no corresponding record for this reference.
- 172Jayagopal, A.; Halfpenny, K. C.; Perez, J. W.; Wright, D. W. Hairpin DNA-Functionalized Gold Colloids for the Imaging of mRNA in Live Cells. J. Am. Chem. Soc. 2010, 132 (28), 9789– 9796, DOI: 10.1021/ja102585vThere is no corresponding record for this reference.
- 173Liu, L.; Li, N.; Huang, Z.-M.; Tang, L.-J.; Ying, Z.-M.; Jiang, J.-H. Gold Nanoflares with Computing Function as Smart Diagnostic Automata for Multi-miRNA Patterns in Living Cells. Anal. Chem. 2020, 92 (16), 10925– 10929, DOI: 10.1021/acs.analchem.0c02325There is no corresponding record for this reference.
- 174Li, L.; Feng, J.; Fan, Y.; Tang, B. Simultaneous Imaging of Zn2+ and Cu2+ in Living Cells Based on DNAzyme Modified Gold Nanoparticle. Anal. Chem. 2015, 87 (9), 4829– 4835, DOI: 10.1021/acs.analchem.5b00204There is no corresponding record for this reference.
- 175Zheng, D.; Seferos, D. S.; Giljohann, D. A.; Patel, P. C.; Mirkin, C. A. Aptamer Nano-flares for Molecular Detection in Living Cells. Nano Lett. 2009, 9 (9), 3258– 3261, DOI: 10.1021/nl901517bThere is no corresponding record for this reference.
- 176Chen, L.; Chao, J.; Qu, X.; Zhang, H.; Zhu, D.; Su, S.; Aldalbahi, A.; Wang, L.; Pei, H. Probing Cellular Molecules with PolyA-Based Engineered Aptamer Nanobeacon. ACS Appl. Mater. Interfaces 2017, 9 (9), 8014– 8020, DOI: 10.1021/acsami.6b16764There is no corresponding record for this reference.
- 177Yeo, D. C.; Wiraja, C.; Paller, A. S.; Mirkin, C. A.; Xu, C. Abnormal Scar Identification with Spherical-Nucleic-Acid Technology. Nat. Biomed. Eng. 2018, 2 (4), 227– 238, DOI: 10.1038/s41551-018-0218-xThere is no corresponding record for this reference.
- 178Liu, Z.; Zhao, J.; Zhang, R.; Han, G.; Zhang, C.; Liu, B.; Zhang, Z.; Han, M.-Y.; Gao, X. Cross-Platform Cancer Cell Identification Using Telomerase-Specific Spherical Nucleic Acids. ACS Nano 2018, 12 (4), 3629– 3637, DOI: 10.1021/acsnano.8b00743There is no corresponding record for this reference.
- 179Conde, J.; Oliva, N.; Artzi, N. Implantable Hydrogel Embedded Dark-Gold Nanoswitch as a Theranostic Probe to Sense and Overcome Cancer Multidrug Resistance. Proc. Natl. Acad. Sci. U. S. A. 2015, 112 (11), E1278-E1287 DOI: 10.1073/pnas.1421229112There is no corresponding record for this reference.
- 180Halo, T. L.; McMahon, K. M.; Angeloni, N. L.; Xu, Y.; Wang, W.; Chinen, A. B.; Malin, D.; Strekalova, E.; Cryns, V. L.; Cheng, C. NanoFlares for the Detection, Isolation, and Culture of Live Tumor Cells from Human Blood. Proc. Natl. Acad. Sci. U. S. A. 2014, 111 (48), 17104– 17109, DOI: 10.1073/pnas.1418637111There is no corresponding record for this reference.
- 181Zhu, D.; Li, X.; Zhu, Y.; Wei, Q.; Hu, Y.; Su, S.; Chao, J.; Wang, L.; Weng, L. Spatiotemporal Monitoring of Subcellular mRNAs In Situ via Polyadenine-Mediated Dual-Color Sticky Flares. ACS Appl. Mater. Interfaces 2023, 15 (12), 15250– 15259, DOI: 10.1021/acsami.3c01242There is no corresponding record for this reference.
- 182Lin, M.; Yi, X.; Huang, F.; Ma, X.; Zuo, X.; Xia, F. Photoactivated Nanoflares for mRNA Detection in Single Living Cells. Anal. Chem. 2019, 91 (3), 2021– 2027, DOI: 10.1021/acs.analchem.8b04434There is no corresponding record for this reference.
- 183Song, S.; Liang, Z.; Zhang, J.; Wang, L.; Li, G.; Fan, C. Gold-Nanoparticle-Based Multicolor Nanobeacons for Sequence-Specific DNA Analysis. Angew. Chem., Int. Ed. 2009, 48 (46), 8670– 8674, DOI: 10.1002/anie.200901887There is no corresponding record for this reference.
- 184Pan, W.; Zhang, T.; Yang, H.; Diao, W.; Li, N.; Tang, B. Multiplexed Detection and Imaging of Intracellular mRNAs Using a Four-Color Nanoprobe. Anal. Chem. 2013, 85 (21), 10581– 10588, DOI: 10.1021/ac402700sThere is no corresponding record for this reference.
- 185Melnychuk, N.; Klymchenko, A. S. DNA-Functionalized Dye-Loaded Polymeric Nanoparticles: Ultrabright FRET Platform for Amplified Detection of Nucleic Acids. J. Am. Chem. Soc. 2018, 140 (34), 10856– 10865, DOI: 10.1021/jacs.8b05840There is no corresponding record for this reference.
- 186Zhao, Y.; Zhao, J.; Zhang, J.; Sun, Y.; Li, L.; Li, Z.; Li, M. Enzymatically Controlled Nanoflares for Specific Molecular Recognition and Biosensing. Anal. Chem. 2022, 94 (25), 8883– 8889, DOI: 10.1021/acs.analchem.2c00166There is no corresponding record for this reference.
- 187Chen, M.; Duan, R.; Xu, S.; Duan, Z.; Yuan, Q.; Xia, F.; Huang, F. Photoactivated DNA Walker Based on DNA Nanoflares for Signal-Amplified MicroRNA Imaging in Single Living Cells. Anal. Chem. 2021, 93 (48), 16264– 16272, DOI: 10.1021/acs.analchem.1c04505There is no corresponding record for this reference.
- 188Li, J.; Liu, S.; Wang, J.; Liu, R.; Yang, X.; Wang, K.; Huang, J. Photocaged Amplified FRET Nanoflares: Spatiotemporal Controllable of mRNA-Powered Nanomachines for Precise and Sensitive MicroRNA Imaging in Live Cells. Nucleic Acids Res. 2022, 50 (7), e40 DOI: 10.1093/nar/gkab1258There is no corresponding record for this reference.
- 189Yang, Y.; Huang, J.; Yang, X.; Quan, K.; Wang, H.; Ying, L.; Xie, N.; Ou, M.; Wang, K. FRET Nanoflares for Intracellular mRNA Detection: Avoiding False Positive Signals and Minimizing Effects of System Fluctuations. J. Am. Chem. Soc. 2015, 137 (26), 8340– 8343, DOI: 10.1021/jacs.5b04007There is no corresponding record for this reference.
- 190Wang, N.; Song, L.; Xing, H.; Zhang, K.; Yang, R.; Li, J. A Spherical Nucleic Acid-Based Two-Photon Nanoprobe for RNase H Activity Assay in Living Cells and Tissues. Nanoscale 2019, 11 (17), 8133– 8137, DOI: 10.1039/C9NR00880BThere is no corresponding record for this reference.
- 191Zeng, T.; Fang, J.; Jiang, Y.; Xing, C.; Lu, C.; Yang, H. Spherical Nucleic Acid Probe Based on 2′-Fluorinated DNA Functionalization for High-Fidelity Intracellular Sensing. Anal. Chem. 2022, 94 (51), 18009– 18016, DOI: 10.1021/acs.analchem.2c04294There is no corresponding record for this reference.
- 192Xiao, F.; Lin, L.; Chao, Z.; Shao, C.; Chen, Z.; Wei, Z.; Lu, J.; Huang, Y.; Li, L.; Liu, Q. Organic Spherical Nucleic Acids for the Transport of a NIR-II-Emitting Dye Across the Blood-Brain Barrier. Angew. Chem., Int. Ed. 2020, 59 (24), 9702– 9710, DOI: 10.1002/anie.202002312There is no corresponding record for this reference.
- 193Fakih, H. H.; Katolik, A.; Malek-Adamian, E.; Fakhoury, J. J.; Kaviani, S.; Damha, M. J.; Sleiman, H. F. Design and Enhanced Gene Silencing Activity of Spherical 2′-Fluoroarabinose Nucleic Acids (FANA-SNAs). Chem. Sci. 2021, 12 (8), 2993– 3003, DOI: 10.1039/D0SC06645AThere is no corresponding record for this reference.
- 194Chen, P.; Wang, D.; Wang, Y.; Zhang, L.; Wang, Q.; Liu, L.; Li, J.; Sun, X.; Ren, M.; Wang, R. Maximizing TLR9 Activation in Cancer Immunotherapy with Dual-Adjuvanted Spherical Nucleic Acids. Nano Lett. 2022, 22 (10), 4058– 4066, DOI: 10.1021/acs.nanolett.2c00723There is no corresponding record for this reference.
- 195Jensen, S. A.; Day, E. S.; Ko, C. H.; Hurley, L. A.; Luciano, J. P.; Kouri, F. M.; Merkel, T. J.; Luthi, A. J.; Patel, P. C.; Cutler, J. I. Spherical Nucleic Acid Nanoparticle Conjugates as an RNAi-Based Therapy for Glioblastoma. Sci. Transl. Med. 2013, 5 (209), 209ra152, DOI: 10.1126/scitranslmed.3006839There is no corresponding record for this reference.
- 196Zheng, D.; Giljohann, D. A.; Chen, D. L.; Massich, M. D.; Wang, X.-Q.; Iordanov, H.; Mirkin, C. A.; Paller, A. S. Topical Delivery of siRNA-Based Spherical Nucleic Acid Nanoparticle Conjugates for Gene Regulation. Proc. Natl. Acad. Sci. U. S. A. 2012, 109 (30), 11975– 11980, DOI: 10.1073/pnas.1118425109There is no corresponding record for this reference.
- 197Fang, Y.; Lu, X.; Wang, D.; Cai, J.; Wang, Y.; Chen, P.; Ren, M.; Lu, H.; Union, J.; Zhang, L. Spherical Nucleic Acids for Topical Treatment of Hyperpigmentation. J. Am. Chem. Soc. 2021, 143 (3), 1296– 1300, DOI: 10.1021/jacs.0c12044There is no corresponding record for this reference.
- 198Huang, Z.; Callmann, C. E.; Wang, S.; Vasher, M. K.; Evangelopoulos, M.; Petrosko, S. H.; Mirkin, C. A. Rational Vaccinology: Harnessing Nanoscale Chemical Design for Cancer Immunotherapy. ACS Cent. Sci. 2022, 8 (6), 692– 704, DOI: 10.1021/acscentsci.2c00227There is no corresponding record for this reference.
- 199Skakuj, K.; Wang, S.; Qin, L.; Lee, A.; Zhang, B.; Mirkin, C. A. Conjugation Chemistry-Dependent T-Cell Activation with Spherical Nucleic Acids. J. Am. Chem. Soc. 2018, 140 (4), 1227– 1230, DOI: 10.1021/jacs.7b12579There is no corresponding record for this reference.
- 200Skakuj, K.; Teplensky, M. H.; Wang, S.; Dittmar, J. W.; Mirkin, C. A. Chemically Tuning the Antigen Release Kinetics from Spherical Nucleic Acids Maximizes Immune Stimulation. ACS Cent. Sci. 2021, 7 (11), 1838– 1846, DOI: 10.1021/acscentsci.1c00779There is no corresponding record for this reference.
- 201Teplensky, M. H.; Dittmar, J. W.; Qin, L.; Wang, S.; Evangelopoulos, M.; Zhang, B.; Mirkin, C. A. Spherical Nucleic Acid Vaccine Structure Markedly Influences Adaptive Immune Responses of Clinically Utilized Prostate Cancer Targets. Adv. Healthcare Mater. 2021, 10 (22), 2101262, DOI: 10.1002/adhm.202101262There is no corresponding record for this reference.
- 202Chinen, A. B.; Ferrer, J. R.; Merkel, T. J.; Mirkin, C. A. Relationships between Poly(ethylene glycol) Modifications on RNA-Spherical Nucleic Acid Conjugates and Cellular Uptake and Circulation Time. Bioconjugate Chem. 2016, 27 (11), 2715– 2721, DOI: 10.1021/acs.bioconjchem.6b00483There is no corresponding record for this reference.
- 203Park, S. Y.; Lytton-Jean, A. K. R.; Lee, B.; Weigand, S.; Schatz, G. C.; Mirkin, C. A. DNA-Programmable Nanoparticle Crystallization. Nature 2008, 451 (7178), 553– 556, DOI: 10.1038/nature06508There is no corresponding record for this reference.
- 204Laramy, C. R.; O’Brien, M. N.; Mirkin, C. A. Crystal Engineering with DNA. Nat. Rev. Mater. 2019, 4 (3), 201– 224, DOI: 10.1038/s41578-019-0087-2There is no corresponding record for this reference.
- 205Wang, S.; Lee, S.; Du, J. S.; Partridge, B. E.; Cheng, H. F.; Zhou, W.; Dravid, V. P.; Lee, B.; Glotzer, S. C.; Mirkin, C. A. The Emergence of Valency in Colloidal Crystals through Electron Equivalents. Nat. Mater. 2022, 21 (5), 580– 587, DOI: 10.1038/s41563-021-01170-5There is no corresponding record for this reference.
- 206Li, Y.; Zhou, W.; Tanriover, I.; Hadibrata, W.; Partridge, B. E.; Lin, H.; Hu, X.; Lee, B.; Liu, J.; Dravid, V. P. Open-Channel Metal Particle Superlattices. Nature 2022, 611 (7937), 695– 701, DOI: 10.1038/s41586-022-05291-yThere is no corresponding record for this reference.
- 207Cheng, H. F.; Distler, M. E.; Lee, B.; Zhou, W.; Weigand, S.; Mirkin, C. A. Nanoparticle Superlattices through Template-Encoded DNA Dendrimers. J. Am. Chem. Soc. 2021, 143 (41), 17170– 17179, DOI: 10.1021/jacs.1c07858There is no corresponding record for this reference.
- 208Distler, M. E.; Landy, K. M.; Gibson, K. J.; Lee, B.; Weigand, S.; Mirkin, C. A. Symmetry-Breaking Dendrimer Synthons in Colloidal Crystal Engineering with DNA. J. Am. Chem. Soc. 2023, 145 (2), 841– 850, DOI: 10.1021/jacs.2c08599There is no corresponding record for this reference.
- 209Li, Y.; Zhou, W.; Zhou, Y.; Cheng, H. F.; Lee, B.; Hu, X.; Roth, E. W.; Dravid, V. P.; Glotzer, S. C.; Mirkin, C. A. Cocrystals Combining Order and Correlated Disorder via Colloidal Crystal Engineering with DNA. Sci. Adv. 2025, 11 (16), eadu4919 DOI: 10.1126/sciadv.adu4919There is no corresponding record for this reference.
- 210Liu, H.; Matthies, M.; Russo, J.; Rovigatti, L.; Narayanan, R. P.; Diep, T.; McKeen, D.; Gang, O.; Stephanopoulos, N.; Sciortino, F. Inverse Design of a Pyrochlore Lattice of DNA Origami through Model-Driven Experiments. Science 2024, 384 (6697), 776– 781, DOI: 10.1126/science.adl5549There is no corresponding record for this reference.
- 211Kahn, J. S.; Minevich, B.; Michelson, A.; Emamy, H.; Wu, J.; Ji, H.; Yun, A.; Kisslinger, K.; Xiang, S.; Yu, N. Encoding Hierarchical 3D Architecture through Inverse Design of Programmable Bonds. Nat. Mater. 2025, 24 (8), 1273– 1282, DOI: 10.1038/s41563-025-02263-1There is no corresponding record for this reference.
- 212He, M.; Gales, J. P.; Ducrot, É.; Gong, Z.; Yi, G.-R.; Sacanna, S.; Pine, D. J. Colloidal Diamond. Nature 2020, 585 (7826), 524– 529, DOI: 10.1038/s41586-020-2718-6There is no corresponding record for this reference.
- 213Lin, H.; Lee, S.; Sun, L.; Spellings, M.; Engel, M.; Glotzer, S. C.; Mirkin, C. A. Clathrate Colloidal Crystals. Science 2017, 355 (6328), 931– 935, DOI: 10.1126/science.aal3919There is no corresponding record for this reference.
- 214Zhou, W.; Lim, Y.; Lin, H.; Lee, S.; Li, Y.; Huang, Z.; Du, J. S.; Lee, B.; Wang, S.; Sánchez-Iglesias, A. Colloidal Quasicrystals Engineered with DNA. Nat. Mater. 2024, 23 (3), 424– 428, DOI: 10.1038/s41563-023-01706-xThere is no corresponding record for this reference.
- 215Liu, W.; Tagawa, M.; Xin, H. L.; Wang, T.; Emamy, H.; Li, H.; Yager, K. G.; Starr, F. W.; Tkachenko, A. V.; Gang, O. Diamond Family of Nanoparticle Superlattices. Science 2016, 351 (6273), 582– 586, DOI: 10.1126/science.aad2080There is no corresponding record for this reference.
- 216Kim, Y.; Macfarlane, R. J.; Jones, M. R.; Mirkin, C. A. Transmutable Nanoparticles with Reconfigurable Surface Ligands. Science 2016, 351 (6273), 579– 582, DOI: 10.1126/science.aad2212There is no corresponding record for this reference.
- 217Rogers, W. B.; Manoharan, V. N. Programming Colloidal Phase Transitions with DNA Strand Displacement. Science 2015, 347 (6222), 639– 642, DOI: 10.1126/science.1259762There is no corresponding record for this reference.
- 218Oh, J. S.; Yi, G.-R.; Pine, D. J. Reconfigurable Self-Assembly and Kinetic Control of Multiprogrammed DNA-Coated Particles. ACS Nano 2020, 14 (4), 4595– 4600, DOI: 10.1021/acsnano.0c00164There is no corresponding record for this reference.
- 219Zhu, J.; Lin, H.; Kim, Y.; Yang, M.; Skakuj, K.; Du, J. S.; Lee, B.; Schatz, G. C.; Van Duyne, R. P.; Mirkin, C. A. Light-Responsive Colloidal Crystals Engineered with DNA. Adv. Mater. 2020, 32 (8), 1906600, DOI: 10.1002/adma.201906600There is no corresponding record for this reference.
- 220Zhu, J.; Kim, Y.; Lin, H.; Wang, S.; Mirkin, C. A. pH-Responsive Nanoparticle Superlattices with Tunable DNA Bonds. J. Am. Chem. Soc. 2018, 140 (15), 5061– 5064, DOI: 10.1021/jacs.8b02793There is no corresponding record for this reference.
- 221Mason, J. A.; Laramy, C. R.; Lai, C.-T.; O’Brien, M. N.; Lin, Q.-Y.; Dravid, V. P.; Schatz, G. C.; Mirkin, C. A. Contraction and Expansion of Stimuli-Responsive DNA Bonds in Flexible Colloidal Crystals. J. Am. Chem. Soc. 2016, 138 (28), 8722– 8725, DOI: 10.1021/jacs.6b05430There is no corresponding record for this reference.
- 222Hensley, A.; Jacobs, W. M.; Rogers, W. B. Self-Assembly of Photonic Crystals by Controlling the Nucleation and Growth of DNA-Coated Colloids. Proc. Natl. Acad. Sci. U. S. A. 2022, 119 (1), e2114050118 DOI: 10.1073/pnas.2114050118There is no corresponding record for this reference.
- 223Hensley, A.; Videbæk, T. E.; Seyforth, H.; Jacobs, W. M.; Rogers, W. B. Macroscopic Photonic Single Crystals via Seeded Growth of DNA-Coated Colloids. Nat. Commun. 2023, 14 (1), 4237, DOI: 10.1038/s41467-023-39992-3There is no corresponding record for this reference.
- 224Auyeung, E.; Li, T. I. N. G.; Senesi, A. J.; Schmucker, A. L.; Pals, B. C.; de la Cruz, M. O.; Mirkin, C. A. DNA-Mediated Nanoparticle Crystallization into Wulff Polyhedra. Nature 2014, 505 (7481), 73– 77, DOI: 10.1038/nature12739There is no corresponding record for this reference.
- 225Hellstrom, S. L.; Kim, Y.; Fakonas, J. S.; Senesi, A. J.; Macfarlane, R. J.; Mirkin, C. A.; Atwater, H. A. Epitaxial Growth of DNA-Assembled Nanoparticle Superlattices on Patterned Substrates. Nano Lett. 2013, 13 (12), 6084– 6090, DOI: 10.1021/nl4033654There is no corresponding record for this reference.
- 226Wang, M. X.; Seo, S. E.; Gabrys, P. A.; Fleischman, D.; Lee, B.; Kim, Y.; Atwater, H. A.; Macfarlane, R. J.; Mirkin, C. A. Epitaxy: Programmable Atom Equivalents Versus Atoms. ACS Nano 2017, 11 (1), 180– 185, DOI: 10.1021/acsnano.6b06584There is no corresponding record for this reference.
- 227Lewis, D. J.; Zornberg, L. Z.; Carter, D. J. D.; Macfarlane, R. J. Single-Crystal Winterbottom Constructions of Nanoparticle Superlattices. Nat. Mater. 2020, 19 (7), 719– 724, DOI: 10.1038/s41563-020-0643-6There is no corresponding record for this reference.
- 228Senesi, A. J.; Eichelsdoerfer, D. J.; Macfarlane, R. J.; Jones, M. R.; Auyeung, E.; Lee, B.; Mirkin, C. A. Stepwise Evolution of DNA-Programmable Nanoparticle Superlattices. Angew. Chem., Int. Ed. 2013, 52 (26), 6624– 6628, DOI: 10.1002/anie.201301936There is no corresponding record for this reference.
- 229Sun, L.; Lin, H.; Li, Y.; Zhou, W.; Du, J. S.; Mirkin, C. A. Position- and Orientation-Controlled Growth of Wulff-Shaped Colloidal Crystals Engineered with DNA. Adv. Mater. 2020, 32 (47), 2005316, DOI: 10.1002/adma.202005316There is no corresponding record for this reference.
- 230Gabrys, P. A.; Seo, S. E.; Wang, M. X.; Oh, E.; Macfarlane, R. J.; Mirkin, C. A. Lattice Mismatch in Crystalline Nanoparticle Thin Films. Nano Lett. 2018, 18 (1), 579– 585, DOI: 10.1021/acs.nanolett.7b04737There is no corresponding record for this reference.
- 231Young, K. L.; Ross, M. B.; Blaber, M. G.; Rycenga, M.; Jones, M. R.; Zhang, C.; Senesi, A. J.; Lee, B.; Schatz, G. C.; Mirkin, C. A. Using DNA to Design Plasmonic Metamaterials with Tunable Optical Properties. Adv. Mater. 2014, 26 (4), 653– 659, DOI: 10.1002/adma.201302938There is no corresponding record for this reference.
- 232Zornberg, L. Z.; Lewis, D. J.; Mertiri, A.; Hueckel, T.; Carter, D. J. D.; Macfarlane, R. J. Self-Assembling Systems for Optical Out-of-Plane Coupling Devices. ACS Nano 2023, 17 (4), 3394– 3400, DOI: 10.1021/acsnano.2c08344There is no corresponding record for this reference.
- 233Lewis, D. J.; Carter, D. J. D.; Macfarlane, R. J. Using DNA to Control the Mechanical Response of Nanoparticle Superlattices. J. Am. Chem. Soc. 2020, 142 (45), 19181– 19188, DOI: 10.1021/jacs.0c08790There is no corresponding record for this reference.
- 234Kim, S.; Svetlizky, I.; Weitz, D. A.; Spaepen, F. Work Hardening in Colloidal Crystals. Nature 2024, 630 (8017), 648– 653, DOI: 10.1038/s41586-024-07453-6There is no corresponding record for this reference.
- 235Seo, S. E.; Wang, M. X.; Shade, C. M.; Rouge, J. L.; Brown, K. A.; Mirkin, C. A. Modulating the Bond Strength of DNA-Nanoparticle Superlattices. ACS Nano 2016, 10 (2), 1771– 1779, DOI: 10.1021/acsnano.5b07103There is no corresponding record for this reference.
- 236Samanta, D.; Iscen, A.; Laramy, C. R.; Ebrahimi, S. B.; Bujold, K. E.; Schatz, G. C.; Mirkin, C. A. Multivalent Cation-Induced Actuation of DNA-Mediated Colloidal Superlattices. J. Am. Chem. Soc. 2019, 141 (51), 19973– 19977, DOI: 10.1021/jacs.9b09900There is no corresponding record for this reference.
- 237Michelson, A.; Subramanian, A.; Kisslinger, K.; Tiwale, N.; Xiang, S.; Shen, E.; Kahn, J. S.; Nykypanchuk, D.; Yan, H.; Nam, C.-Y. Three-Dimensional Nanoscale Metal, Metal Oxide, and Semiconductor Frameworks through DNA-Programmable Assembly and Templating. Sci. Adv. 2024, 10 (2), eadl0604 DOI: 10.1126/sciadv.adl0604There is no corresponding record for this reference.
- 238Auyeung, E.; Macfarlane, R. J.; Choi, C. H. J.; Cutler, J. I.; Mirkin, C. A. Transitioning DNA-Engineered Nanoparticle Superlattices from Solution to the Solid State. Adv. Mater. 2012, 24 (38), 5181– 5186, DOI: 10.1002/adma.201202069There is no corresponding record for this reference.
- 239Wang, S.; Park, S. S.; Buru, C. T.; Lin, H.; Chen, P.-C.; Roth, E. W.; Farha, O. K.; Mirkin, C. A. Colloidal Crystal Engineering with Metal-Organic Framework Nanoparticles and DNA. Nat. Commun. 2020, 11 (1), 2495, DOI: 10.1038/s41467-020-16339-wThere is no corresponding record for this reference.
- 240Kim, S.; Zheng, C. Y.; Schatz, G. C.; Aydin, K.; Kim, K.-H.; Mirkin, C. A. Mie-Resonant Three-Dimensional Metacrystals. Nano Lett. 2020, 20 (11), 8096– 8101, DOI: 10.1021/acs.nanolett.0c03089There is no corresponding record for this reference.
- 241Zhang, Y.; Xu, D. D.; Tanriover, I.; Zhou, W.; Li, Y.; López-Arteaga, R.; Aydin, K.; Mirkin, C. A. Nonlinear Optical Colloidal Metacrystals. Nat. Photonics 2025, 19 (1), 20– 27, DOI: 10.1038/s41566-024-01558-0There is no corresponding record for this reference.
- 242Li, Z.; Lim, Y.; Tanriover, I.; Zhou, W.; Li, Y.; Zhang, Y.; Aydin, K.; Glotzer, S. C.; Mirkin, C. A. DNA-Mediated Assembly of Au Bipyramids into Anisotropic Light Emitting Kagome Superlattices. Sci. Adv. 2024, 10 (29), eadp3756 DOI: 10.1126/sciadv.adp3756There is no corresponding record for this reference.
- 243Lin, Q.-Y.; Li, Z.; Brown, K. A.; O’Brien, M. N.; Ross, M. B.; Zhou, Y.; Butun, S.; Chen, P.-C.; Schatz, G. C.; Dravid, V. P. Strong Coupling between Plasmonic Gap Modes and Photonic Lattice Modes in DNA-Assembled Gold Nanocube Arrays. Nano Lett. 2015, 15 (7), 4699– 4703, DOI: 10.1021/acs.nanolett.5b01548There is no corresponding record for this reference.
- 244Zheng, C. Y.; Palacios, E.; Zhou, W.; Hadibrata, W.; Sun, L.; Huang, Z.; Schatz, G. C.; Aydin, K.; Mirkin, C. A. Tunable Fluorescence from Dye-Modified DNA-Assembled Plasmonic Nanocube Arrays. Adv. Mater. 2019, 31 (41), 1904448, DOI: 10.1002/adma.201904448There is no corresponding record for this reference.
- 245Zheng, C. Y.; Hadibrata, W.; Kim, S.; Schatz, G. C.; Aydin, K.; Mirkin, C. A. Large-Area, Highly Crystalline DNA-Assembled Metasurfaces Exhibiting Widely Tunable Epsilon-Near-Zero Behavior. ACS Nano 2021, 15 (11), 18289– 18296, DOI: 10.1021/acsnano.1c07496There is no corresponding record for this reference.
- 246Sun, L.; Lin, H.; Park, D. J.; Bourgeois, M. R.; Ross, M. B.; Ku, J. C.; Schatz, G. C.; Mirkin, C. A. Polarization-Dependent Optical Response in Anisotropic Nanoparticle-DNA Superlattices. Nano Lett. 2017, 17 (4), 2313– 2318, DOI: 10.1021/acs.nanolett.6b05101There is no corresponding record for this reference.
- 247Li, Y.; Tanriover, I.; Zhou, W.; Hadibrata, W.; Dereshgi, S. A.; Samanta, D.; Aydin, K.; Mirkin, C. A. Monolayer Plasmonic Nanoframes as Large-Area, Broadband Metasurface Absorbers. Small 2022, 18 (33), 2201171, DOI: 10.1002/smll.202201171There is no corresponding record for this reference.
- 248Li, Y.; Jin, H.; Zhou, W.; Wang, Z.; Lin, Z.; Mirkin, C. A.; Espinosa, H. D. Ultrastrong Colloidal Crystal Metamaterials Engineered with DNA. Sci. Adv. 2023, 9 (39), eadj8103 DOI: 10.1126/sciadv.adj8103There is no corresponding record for this reference.
- 249Petrosko, S. H.; Coleman, B. D.; Drout, R. J.; Schultz, J. D.; Mirkin, C. A. Spherical Nucleic Acids: Integrating Nanotechnology Concepts into General Chemistry Curricula. J. Chem. Educ. 2021, 98 (10), 3090– 3099, DOI: 10.1021/acs.jchemed.1c00441There is no corresponding record for this reference.
- 250Johnson, G. T.; Autin, L.; Goodsell, D. S.; Sanner, M. F.; Olson, A. J. ePMV Embeds Molecular Modeling into Professional Animation Software Environments. Structure 2011, 19 (3), 293– 303, DOI: 10.1016/j.str.2010.12.023There is no corresponding record for this reference.



