From Neoantigens to Nanocarriers: Modern Methods and Modalities in Using Peptides for Cancer VaccinationClick to copy article linkArticle link copied!
- Aleah Harris TreitererAleah Harris TreitererDepartment of Chemical and Biomolecular Engineering, The Ohio State University, Columbus, Ohio 43210, United StatesMore by Aleah Harris Treiterer
- Blaise RobinsonBlaise RobinsonDepartment of Chemical and Biomolecular Engineering, The Ohio State University, Columbus, Ohio 43210, United StatesMore by Blaise Robinson
- Sean HugginsSean HugginsDepartment of Chemistry, The Ohio State University, Columbus, Ohio 43210, United StatesMore by Sean Huggins
- Blaise R. Kimmel*Blaise R. Kimmel*Email: [email protected]Department of Chemical and Biomolecular Engineering, The Ohio State University, Columbus, Ohio 43210, United StatesCenter for Cancer Engineering, Ohio State University Comprehensive Cancer Center, The Ohio State University, Columbus, Ohio 43210, United StatesPelotonia Institute for Immuno-Oncology, Ohio State University Comprehensive Cancer Center, The Ohio State University, Columbus, Ohio 43210, United StatesMore by Blaise R. Kimmel
Abstract
The success of cancer vaccination depends on the ability of therapeutics to sustain prolonged immune activation, leading to the destruction of tumor cells. However, only a few therapeutic cancer vaccines have been FDA-approved due to challenges in targeting and eliciting a sufficiently strong immune response. Peptides have emerged as promising drugs owing to their ability to interact with cell-surface receptors and their low manufacturing cost. Despite the peptides’ positive characteristics, additional research is needed to develop more effective methods for using peptides to stimulate the immune system for a sustained period to induce tumor cell regression. This review focuses on recent work in peptide-based vaccine design and development, aiming to determine the optimal formulation of peptide vaccines by identifying and isolating neoantigens for tumor targeting, thereby delivering peptide antigens to specific locations. The expansion of the current landscape of cancer treatments, including peptide vaccines and combination therapies, is revolutionizing the possibilities for patient care and treatment.
This publication is licensed under
License Summary*
You are free to share(copy and redistribute) this article in any medium or format and to adapt(remix, transform, and build upon) the material for any purpose, even commercially within the parameters below:
Creative Commons (CC): This is a Creative Commons license.
Attribution (BY): Credit must be given to the creator.
*Disclaimer
This summary highlights only some of the key features and terms of the actual license. It is not a license and has no legal value. Carefully review the actual license before using these materials.
License Summary*
You are free to share(copy and redistribute) this article in any medium or format and to adapt(remix, transform, and build upon) the material for any purpose, even commercially within the parameters below:
Creative Commons (CC): This is a Creative Commons license.
Attribution (BY): Credit must be given to the creator.
*Disclaimer
This summary highlights only some of the key features and terms of the actual license. It is not a license and has no legal value. Carefully review the actual license before using these materials.
License Summary*
You are free to share(copy and redistribute) this article in any medium or format and to adapt(remix, transform, and build upon) the material for any purpose, even commercially within the parameters below:
Creative Commons (CC): This is a Creative Commons license.
Attribution (BY): Credit must be given to the creator.
*Disclaimer
This summary highlights only some of the key features and terms of the actual license. It is not a license and has no legal value. Carefully review the actual license before using these materials.
Special Issue
Published as part of Biochemistry special issue “Chemistry and Biology of Peptides”.
Introduction
Figure 1
Figure 1. Comparison of different types of cancer vaccinations. (A) Cellular vaccines contain cells or parts of cells isolated from cancer patients, typically either cancerous cells or dendritic cells. These isolated cells are modified and then administered back to the patient to trigger an immune response against cancer cells. While these vaccinations can elicit an immune response that leads to tumor regression, isolating and processing patient cells is time-consuming and expensive. (B) Nucleic acid vaccines are made of either DNA or RNA. The nucleic acid is delivered to cells, allowing the genetic information to be processed and expressed as proteins. Currently, more focus has been placed on mRNA vaccines because the genetic information is not incorporated within the cell’s nucleus; therefore, both dividing and nondividing cells can express the protein of interest. The mRNA vaccines are cheaper and easier to manufacture than cellular vaccines. However, these technologies require a delivery system to ensure the material is successfully transported into the cell. (C) Peptide vaccines contain cancer antigens, designed to stimulate the immune system and induce cancer regression. While these vaccines have high binding affinity for cell receptors and can be combined with other molecules, this approach has known limitations, including difficulty in identifying immunogenic antigens and the potential for peptides to be easily degraded in the body.
Mechanisms of Cancer Vaccination
Figure 2
Figure 2. Overview of immune stimulation following the administration of a peptide vaccination. Following the administration of the peptide vaccination, the “drug load” or peptide antigens will be released into the body. The peptides within the body will be taken up by a dendritic cell, and processed either in the cytosol or an endosome, leading to both MHC Class I and Class II presentation. CD8+ and CD4+ T cells can then recognize antigens presented by MHC Class molecules, becoming mature T cells that can attack and destroy cancer cells.
| Cancer Types | Vaccine Used | Combination | Phase | Clinical Trail Number |
|---|---|---|---|---|
| Advanced solid tumor (wide range of types) | Personalized neoantigen with sargramostim or GM-CSF SC | Pembrolizumab, cyclophosphamide | I/II | NCT05269381 (27) |
| Colorectal and pancreatic cancer | KRAS-targeted long peptide vaccine | Poly-ILCL, nivolumab, ipilimumab | I | NCT04117087 (28) |
| Squamous nonsmall cell lung cancer and squamous cell carcinoma of head and neck | Adjusted peptide vaccine (PANDA-VAC) | Pembrolizumab | I | NCT04266730 (29) |
| Squamous nonsmall cell lung cancer, squamous cell carcinoma of head and neck, and urothelial bladder cancer | IDO and PD-L1 peptides (IO102-IO103) | Pembrolizumab | II | NCT05077709 (30) |
| Stage IIIC-IV melanoma | Personalized neoantigen vaccine | Poly-ICLC | I | NCT05098210 (31) |
| Hormone receptor positive HER2 negative breast cancer | ||||
| Stage III–IV nonsmall cell lung cancer | ||||
| Pancreatic cancer (nonmetastatic resectable pancreatic adenocarcinoma) | Autologous dendritic cells loaded with personalized peptides | Standard of care (SOC) adjuvant chemotherapy, nivolumab | Ib | NCT04627246 (32) |
| Ovarian cancer | Multineoepitope vaccine with relevant TAAs (OSE2101) | Pembrolizumab | II | NCT04713514 (33) |
| Neoantigenic peptides | Poly-ICLC, nivolumab | I | NCT04024878 (34) | |
| Nonsmall cell lung cancer | UCPVax – based on telomerase-derived helper peptides | Nivolumab | II | NCT04263051 (35) |
| Melonoma | NeoVax – personalized neoantigen | Poly-ICLC, CDX-301, nivolumab, pembrolizumab | I | NCT04930783 (36) |
| Liver cancer | DNAJB1-PKACA Peptide Vaccine | Nivolumab, ipilimumab | I | NCT04248569 (37) |
| Glioblastoma | EO2401 peptide vaccine | Nivolumab, bevacizumab | Ib/IIa | NCT04116658 (38) |
| Glioma | IDH1R132H peptide vaccine | Avelumab | I | NCT03893903 (39) |
| Gastric cancer | OTSGC-A24 peptide vaccine | Nivolumab, Ipilimumab | I | NCT03784040 (40) |
| Breast cancer | PVX-410 muli-peptide vaccine | Pembrolizumab, chemotherapy | II | NCT04634747 (41) |
| AE37 peptide vaccine | Pembrolizumab | II | NCT04024800 (42) |
All about Antigens
| Class/Category of Tumor Antigen | Description | Example Antigens | Example of Cancers Containing Antigen | |
|---|---|---|---|---|
| Tumor-associated antigens (TAA) | Overexpressed antigens (associated with oncogenes) | Antigen is expressed at higher levels in tumor cells than in normal cells | RAGE-1 | Pancreatic, (48) lung, (49) breast, prostate, colorectal, gastric, liver (49) |
| hTERT | Breast, skin, thyroid, (50) glioblastoma (51) | |||
| HER2 | Breast, gastric, gastroesophageal, nonsmall-cell, endometrial, ovarian (52) | |||
| Mesothelin | Mesothelioma, ovarian, pancreatic, lung, breast, cholangiocarcinoma, bile duct carcinoma, gastric cancer (53,54) | |||
| Differentiation antigens | Antigens are expressed in both tumor and normal cells, but only in specific normal cells | Tyrosinase | Melanoma, (55) neuroblastoma (56) | |
| gp100 | Melanoma (57) | |||
| MART-1 | Melanoma (58) | |||
| Prostate-specific antigen (PSA) | Prostate (59) | |||
| Tumor-specific antigens (TSA) | Oncogenic Viral Antigens | Abnormal expression of antigen due to viral infection | EBV LMP-1 | Nasopharyngeal carcinoma, gastric, lymphoma (60) |
| HPV-E6/E7 | Cervical, (61) head neck squamous cell carcinoma, anal, penile, vaginal, vulvar (62) | |||
| HTLV-1 | Adult T-cell leukemia/lymphoma (ATL), (63) endometrial (64) | |||
| Tumor-specific antigens | Antigens expressed due to mutations within tumor cells | KRAS | Nonsmall cell lung, colorectal, pancreatic (65) | |
| NRAS | Melanoma, lung adenocarcinoma, colon, pancreatic, leukemia (66) | |||
| ETV6 | Leukemia, (67) lymphoma (68) | |||
| NPM/ALK | Lymphoma, lung (69) |
Figure 3
Figure 3. Comparing in silico vs LC-MS neo-antigen screening methods. Both in silico and LC-MS neoantigen screening methods compare healthy cells to tumor samples to identify differences in the two genetic sequences. In silico analysis methods typically involve computer-based screening, whereas LC-MS methods involve eluting MHC peptide ligands prior to LC-MS to identify potential neoantigen candidates. Following the identification of potential neo-antigens, the candidates are screened in vitro to assess TIL activation.
| Computational Tool | Purpose | Link to Tool |
|---|---|---|
| timsTOF Prosit (81) | Trained to identify and predict immunopeptides and HLA-I peptides using PSM rescoring of MaxQuant results. | https://koina.proteomicsdb.org/ |
| AlphaPept (82) | Python based database to process large high resolution MS data sets, including features such as peptide identification and protein quantification. One drawback is the database only has functionality for DDA proteomics. | https://alphapept.org/ |
| RPEMHC (83) | Deep learning method to predict the binding affinity between peptides and MHC Class I and II molecules based on residue–residue pair encoding. | https://github.com/lennylv/RPEMHC |
| SNAF (84) | Computational tool to predict possible T cell and B cell antigens by identifying and interpreting classes of splicing neo-antigens. | N/A |
| TIMS2Rescore (85) | Tool that works with timsTOF model to assist with analyzing a sample’s proteome, including immunopeptidomics. | https://github.com/compomics/tims2rescore |
| UniPMT (86) | Computational tool that predicts binding of the peptide-MHC-TCR complex, the peptide-MHC complex, and the peptide-TCR complex. | N/A |
| VirusImmu (87) | Machine learning tool to predict B cell epitope immunogenicity. However, it is important to note that this tool is used for virus epitope predictions. | https://github.com/zhangjbig/VirusImmu |
| CNNeoPP (88) | Prediction model for neoantigen immunogenic classification. | https://github.com/AaronChen007/neoantigen |
| NeoPrecis-Immuno (89) | Neo-antigen immunogenicity prediction model, specifically shown to assist with predicting patient outcomes within Immune Checkpoint Inhibition treatment. | N/A |
Types and Deliveries of Peptide Vaccinations
Figure 4
Figure 4. How SLP and ROP vaccinations stimulate the immune system. Both SLP and ROP vaccinations stimulate the immune system by being processed by APCs, leading to the activation of CD4+ and CD8+ T cells. However, SLP vaccinations comprise a pool of peptides, each representing a different antigen epitope, whereas ROP vaccinations contain a single peptide with multiple epitopes encoded within the peptide.
Figure 5
Figure 5. Different vaccine delivery methods for peptide vaccinations. There have been significant advances in drug delivery technologies, and four of the main methods are detailed within the figure. Each specific method has its advantages and drawbacks, which are detailed in the bullet points below each icon.
| Type of Delivery Method | Application |
|---|---|
| Polymeric NP (109) | Vaccine formulation: Poly(lactic-co-glycolic) acid (PLGA) and dimethyl-dioctadecyl-ammonium bromide (DDAB) nanoparticle |
| Drug load: Model antigen (OVA) conjugated on the surface | |
| Immune Response: Generated CD4+ and CD8+ T cell response in addition to having successful nanoparticle delivery to the lymph nodes (109) | |
| Additional Information: Did not perform in vivo tumor studies, therefore further research is needed to determine the ability of the nanoparticle to slow cancer progression | |
| Lipid nanoparticles (110) | Vaccine formulation: Cationic liposomes |
| Drug load: OVA24 or OVA17 SLPs | |
| Immune Response: Vaccines were able to both activate T-cells and induce an immune response in vivo leading to tumor regression | |
| Additional Information: Cationic liposome vaccination was also compared to PLGA vaccination with same drug load, and the liposome was found to generate a more effective immune response (110) | |
| Nanogels (101,102) | Vaccine formulation: Nanogel made of cationic dextran |
| Drug load: SLPs that included cytotoxic C lymphocytes (CTL) and CD4+ T helpers | |
| Immune Response: Delivery in vivo lead to T cell activation (101) | |
| Additional Information: Preliminary study only, so further investigation within tumor models is needed to determine efficacy of treatment | |
| Vaccine Formulation: Nanogel made of monomer N-[(2,2-dimethyl-1,3-dioxolane)methyl]acrylamide (DMDOMA) | |
| Drug load: N/A, this was a study to only investigate delivery mechanism of nanogel | |
| Additional Information: Shown to completely degrade from hydrolysis within acidic environments, which was hypothesized to provide controlled drug release of anticancer drugs within acidic environments (102) | |
| Polymer-drug conjugates (107,111−114) | Vaccine Formulation: Hydrophilic polymers with cleavable linkers |
| Drug Load: Immunostimulatory payloads including targeted STING agonists | |
| Additional Information: Programmable pH responsive nanocarriers provided a tunable carrier architecture that can be combined with antigen delivery to reprogram the vascular-immune interface and substantially broaden responses to checkpoint inhibitors and adoptive cell therapies – self-assembling nanoparticles with peptide-TLR-7/8a conjugates was engineered to be able to self-assemble no matter the charge of peptide antigen used by conjugating a charge-modifying group and hydrophobic block to either end of the peptide of interest (114)-in vivo vaccination was shown to have uptake within APCs and elicit a significant T cell response | |
| Vaccine Formulation: Self-assembling nanoparticles with peptide-TLR-7/8a conjugates | |
| Drug Load: Variable peptide antigen | |
| Additional Information: Nanoparticles were engineered to self-assemble no matter the charge of peptide antigen. The in vivo vaccination was shown to have uptake within APCs and elicit a significant T cell response. |
Current Advancements in Peptide Vaccinations
Figure 6
Figure 6. How immune stimulants (DAMPs/PAMPs) can bolster the immune response from peptide vaccinations. Following the death of a cancer cell, immune stimulants such as DAMPs and PAMPs are released and can bind to pattern recognition receptors (PRRs). The binding of PRRs enables DAMPs/PAMPs to interact with dendritic cells, leading to antigen presentation and the activation of CD4+ and CD8+ T cells. The activated T cells are then able to target additional cancer cells, leading to the death of these cells and initiating the cycle again.
Roadblocks to Peptide Cancer Vaccinations
Figure 7
Figure 7. The different barriers that contribute to no peptide cancer vaccinations being FDA approved. TME is inter-related with immune suppression and evasion, and both work together to prevent tumor cells from being recognized and destroyed by the immune system. Tumor heterogeneity complicates the ability of cancer treatments to properly target and destroy all cancer cells, as the rapid mutations present can cause cancer cells to escape surveillance, leading to more proliferation and growth.
Conclusions
Acknowledgments
This work was supported in part by The Ohio State University Center for Cancer Engineering-Curing Cancer through Research in Engineering and Sciences. B.R.K. acknowledges financial support from the Prostate Cancer Foundation Young Investigator Award. We acknowledge the use of BioRender for the creation of all figures.
References
This article references 145 other publications.
- 1NCI Cancer Statistics; NCI. https://www.cancer.gov/about-cancer/understanding/statistics. (Accessed 16 October 2025).Google ScholarThere is no corresponding record for this reference.
- 2Hsu, J.Y.; Chang, C.J.; Cheng, J.S. Survival, treatment regimens and medical costs of women newly diagnosed with metastatic triple-negative breast cancer. Sci. Rep. 2022, 12 (1), 729, DOI: 10.1038/s41598-021-04316-2Google ScholarThere is no corresponding record for this reference.
- 3American Cancer Society Survival Rates for Pancreatic Cancer; American Cancer Society, https://www.cancer.org/cancer/types/pancreatic-cancer/detection-diagnosis-staging/survival-rates.html. (Accessed16 October 2025).Google ScholarThere is no corresponding record for this reference.
- 4Baskar, R.; Dai, J.; Wenlong, N.; Yeo, R.; Yeoh, K.-W. Biological Response of Cancer Cells to Radiation Treatment. Front. Mol. Biosci. 2014, 1, 24, DOI: 10.3389/fmolb.2014.00024Google ScholarThere is no corresponding record for this reference.
- 5Stephens, A. J.; Burgess-Brown, N. A.; Jiang, S. Beyond Just Peptide Antigens: The Complex World of Peptide-Based Cancer Vaccines. Front. Immunol. 2021, 12, 696791, DOI: 10.3389/fimmu.2021.696791Google ScholarThere is no corresponding record for this reference.
- 6O’Donnell, J.S.; Teng, M.W.; Smyth, M.J. Cancer immunoediting and resistance to T cell-based immunotherapy. Nat. Rev. Clin. Oncol. 2019, 16 (3), 151– 167, DOI: 10.1038/s41571-018-0142-8Google ScholarThere is no corresponding record for this reference.
- 7Wang, C.; Hao, P. Therapeutic Vaccine Strategies in Cancer: Advances, Challenges, and Future Directions. Clin. Cancer Bull. 2025, 4 (1), 10, DOI: 10.1007/s44272-025-00039-xGoogle ScholarThere is no corresponding record for this reference.
- 8Ma, L.; Diao, L.; Peng, Z.; Jia, Y.; Xie, H.; Li, B.; Ma, J.; Zhang, M.; Cheng, L.; Ding, D.; Zhang, X.; Chen, H.; Mo, F.; Jiang, H.; Xu, G.; Meng, F.; Zhong, Z.; Liu, M. Immunotherapy and Prevention of Cancer by Nanovaccines Loaded with Whole-Cell Components of Tumor Tissues or Cells. Adv. Mater. 2021, 33 (43), 2104849, DOI: 10.1002/adma.202104849Google ScholarThere is no corresponding record for this reference.
- 9Gao, C.; Luo, R.; Kwong, C. H. T.; Liu, J.; Tang, M.; Xie, B.; Duan, T.; Wang, R. Cancer Vaccine from Intracellularly Gelated Tumor Cells Functionalized with CD47 Blockage and Damage-Associated Molecular Pattern Exposure. Cell Rep. Med. 2025, 6 (5), 102092, DOI: 10.1016/j.xcrm.2025.102092Google ScholarThere is no corresponding record for this reference.
- 10Maeng, H. M.; Olkhanud, P. B.; Black, M.; Highfill, S. L.; Stroncek, D. F.; Berzofsky, J. A. Dendritic Cell Cancer Vaccines: Clinical Production for Cancer Immunotherapy. In In Cancer Vaccines: methods and Protocols, Padler-Karavani, V.; ed.; Springer: New York, NY, 2025; pp. 207– 221. DOI: DOI: 10.1007/978-1-0716-4542-0_16 .Google ScholarThere is no corresponding record for this reference.
- 11Bodey, B.; Bodey, B.; Siegel, S. E.; Kaiser, H. E. Failure of Cancer Vaccines: The Significant Limitations of This Approach to Immunotherapy. Anticancer Res. 2000, 20 (4), 2665– 2676Google ScholarThere is no corresponding record for this reference.
- 12Chen, J.; Tan, Q.; Yang, Z.; Chen, W.; Zhou, E.; Li, M.; Deng, J.; Wu, Y.; Liu, J.; Xu, J.; Guo, M.; Jin, Y. Dendritic Cell Derived-Extracellular Vesicles Engineered to Express Interleukin-12 and Anti-CTLA-4 on Their Surface for Combinational Cancer Immunotherapy. J. Extracell. Vesicles 2025, 14 (4), e70068 DOI: 10.1002/jev2.70068Google ScholarThere is no corresponding record for this reference.
- 13Liao, H.-C.; Liu, S.-J. Advances in Nucleic Acid-Based Cancer Vaccines. J. Biomed. Sci. 2025, 32 (1), 10, DOI: 10.1186/s12929-024-01102-wGoogle ScholarThere is no corresponding record for this reference.
- 14Kaczmarek, M.; Poznańska, J.; Fechner, F.; Michalska, N.; Paszkowska, S.; Napierała, A.; Mackiewicz, A. Cancer Vaccine Therapeutics: Limitations and Effectiveness─A Literature Review. Cells 2023, 12 (17), 2159, DOI: 10.3390/cells12172159Google ScholarThere is no corresponding record for this reference.
- 15Srivastava, R. Stimuli-Responsive Nanomaterials for the Precision Delivery of mRNA Cancer Vaccines. Nano Trends 2025, 11, 100147, DOI: 10.1016/j.nwnano.2025.100147Google ScholarThere is no corresponding record for this reference.
- 16Zhou, J.; Liu, Y.; Xu, W.; Bhatta, R.; Han, J.; Baskaran, D.; Devmal, S.; Leal, C.; Wang, H. Macroporous Hydrogel-Based mRNA Cancer Vaccine for in Situ Recruitment and Modulation of Dendritic Cells. Acta Biomater. 2025, 204, 234– 245, DOI: 10.1016/j.actbio.2025.08.005Google ScholarThere is no corresponding record for this reference.
- 17Kim, Y. A.; Jeong, H.; Kim, H.; Lee, S.; Kim, K. S.; Na, K. Lipid Nanoparticles with Prazole Adjuvant to Enhance the Efficacy of mRNA Cancer Vaccines. J. Controlled Release 2025, 383, 113756, DOI: 10.1016/j.jconrel.2025.113756Google ScholarThere is no corresponding record for this reference.
- 18Paula Fawzy, M.; Hassan, H. A. F. M.; Umair Amin, M.; Preis, E.; Bakowsky, U.; Ashraf Fahmy, S. Deploying Nucleic Acids-Loaded Plant-Derived Exosomes as Green Nano Gadget in Cancer Gene Therapy. Mater. Adv. 2025, 6 (4), 1230– 1261, DOI: 10.1039/D4MA01189AGoogle ScholarThere is no corresponding record for this reference.
- 19Chudasama, R.; Phung, Q.; Hsu, A.; Almhanna, K. Vaccines in Gastrointestinal Malignancies: From Prevention to Treatment. Vaccines 2021, 9 (6), 647, DOI: 10.3390/vaccines9060647Google ScholarThere is no corresponding record for this reference.
- 20Xu, F.; Yuan, Y.; Wang, Y.; Yin, Q. Emerging Peptide-Based Nanovaccines: From Design Synthesis to Defense against Cancer and Infection. Biomedecine Pharmacother 2023, 158, 114117, DOI: 10.1016/j.biopha.2022.114117Google ScholarThere is no corresponding record for this reference.
- 21Eiz-Vesper, B.; Schmetzer, H. M. Antigen-Presenting Cells: Potential of Proven Und New Players in Immune Therapies. Transfus. Med. Hemotherapy 2021, 47 (6), 429– 431, DOI: 10.1159/000512729Google ScholarThere is no corresponding record for this reference.
- 22Borst, J.; Ahrends, T.; Bąbała, N.; Melief, C. J. M.; Kastenmüller, W. CD4+ T Cell Help in Cancer Immunology and Immunotherapy. Nat. Rev. Immunol. 2018, 18 (10), 635– 647, DOI: 10.1038/s41577-018-0044-0Google ScholarThere is no corresponding record for this reference.
- 23Wu, L.; Yi, W.; Yao, S.; Xie, S.; Peng, R.; Zhang, J.; Tan, W. mRNA-Based Cancer Vaccines: Advancements and Prospects. Nano Lett. 2024, 24 (41), 12711– 12721, DOI: 10.1021/acs.nanolett.4c03296Google ScholarThere is no corresponding record for this reference.
- 24Abd-Aziz, N.; Poh, C. L. Development of Peptide-Based Vaccines for Cancer. J. Oncol. 2022, 2022, 9749363, DOI: 10.1155/2022/9749363Google ScholarThere is no corresponding record for this reference.
- 25Bacillus Calmette-Guerin (BCG) Treatment BCG Treatment: What It Is, Procedure & Side Effects. Bacillus Calmette-Guerin (BCG) Treatment, https://my.clevelandclinic.org/health/treatments/17908-bacillus-calmette-guerin-bcg-treatment. (Accessed 17 October 2025).Google ScholarThere is no corresponding record for this reference.
- 26Anassi, E.; Ndefo, U. A. Sipuleucel-T (Provenge) Injection. Pharm. Ther. 2011, 36 (4), 197– 202Google ScholarThere is no corresponding record for this reference.
- 27Mayo Clinic. A Phase I/II Study of Personalized Neoantigen Peptide-Based Vaccine in Combination With Pembrolizumab in Advanced Solid Tumors (PNeoVCA). Clinical trial registration NCT05269381; clinicaltrials.gov, 2024. https://clinicaltrials.gov/study/NCT05269381. (Accessed 05 November 2025).Google ScholarThere is no corresponding record for this reference.
- 28Sidney Kimmel Comprehensive Cancer Center at Johns Hopkins. Pooled Mutant KRAS-Targeted Long Peptide Vaccine Combined With Nivolumab and Ipilimumab for Patients With Resected MMR-p Colorectal and Pancreatic Cancer. Clinical trial registration NCT04117087; clinicaltrials.gov, 2025. https://clinicaltrials.gov/study/NCT04117087. (Accessed 05 November 2025).Google ScholarThere is no corresponding record for this reference.
- 29UNC Lineberger Comprehensive Cancer Center. Phase I Trial of a Personalized and Adaptive Neoantigen Dose-Adjusted Vaccine Administered Concurrently With Pembrolizumab. Clinical trial registration NCT04266730; clinicaltrials.gov, 2025. https://clinicaltrials.gov/study/NCT04266730. (Accessed 05 November 2025).Google ScholarThere is no corresponding record for this reference.
- 30IO Biotech. A Phase II Multi-Arm (Basket) Trial Investigating the Safety and Efficacy of IO102-IO103 in Combination With Pembrolizumab, as First-Line Treatment for Patients With Metastatic NSCLC, SCCHN, or Metastatic mUBC. Clinical trial registration NCT05077709; clinicaltrials.gov, 2024. https://clinicaltrials.gov/study/NCT05077709. (Accessed 05 November 2025).Google ScholarThere is no corresponding record for this reference.
- 31Fred Hutchinson Cancer Center. PNV21–001: A Phase I Study of a Personalized Multi-Peptide Neo-Antigen Vaccine in Breast Cancer, PD1/PD-L1 Inhibitor-Refractory Melanoma, and Pretreated Non-Small Cell Lung Cancer. Clinical trial registration NCT05098210; clinicaltrials.gov, 2025. https://clinicaltrials.gov/study/NCT05098210 (Accessed 05 November 2025).Google ScholarThere is no corresponding record for this reference.
- 32Digklia, A. A Phase Ib Study of the Combination of Personalized Autologous Dendritic Cell Vaccine and Standard of Care Adjuvant Chemotherapy Followed by Nivolumab for Resected Pancreatic Adenocarcinoma. Clinical trial registration NCT04627246; clinicaltrials.gov, 2025. https://clinicaltrials.gov/study/NCT04627246. (Accessed 05 November 2025).Google ScholarThere is no corresponding record for this reference.
- 33ARCAGY/GINECO GROUP. Randomized Phase II Study Comparing Neo-Epitope Based Vaccine OSE2101 (TEDOPI®) With or Without Anti-PD1 (Pembrolizumab) Versus Best Supportive Care as Maintenance Treatment in Platinum-Sensitive Recurrent Ovarian Cancer Patient With Controlled Disease After Platinum-Based Chemotherapy. Clinical trial registration NCT04713514; clinicaltrials.gov, 2024. https://clinicaltrials.gov/study/NCT04713514. (Accessed 05 November 2025).Google ScholarThere is no corresponding record for this reference.
- 34Ott, P. NeoVax With Nivolumab in Patients With Ovarian Cancer. Clinical trial registration NCT04024878; clinicaltrials.gov, 2025. https://clinicaltrials.gov/study/NCT04024878. (Accessed 05 November 2025).Google ScholarThere is no corresponding record for this reference.
- 35Centre Hospitalier Universitaire de Besancon. Evaluation of UCPVax Plus Nivolumab as Second Line Therapy in Advanced Non Small Cell Lung: A Randomized Non Comparative Phase II Trial; Clinical trial registration NCT04263051; clinicaltrials.gov, 2025. https://clinicaltrials.gov/study/NCT04263051 (Accessed 05 November 2025).Google ScholarThere is no corresponding record for this reference.
- 36Ott, P. A Phase Ib Study of NeoVax in Combination With CDX-301 and Nivolumab or Pembrolizumab and in Patients With Melanoma; Clinical trial registration NCT04930783; clinicaltrials.gov, 2025. https://clinicaltrials.gov/study/NCT04930783 (Accessed 05 November 2025).Google ScholarThere is no corresponding record for this reference.
- 37Sidney Kimmel Comprehensive Cancer Center at Johns Hopkins. A Pilot Study of a DNAJB1-PRKACA Fusion Kinase Peptide Vaccine Combined With Nivolumab and Ipilimumab for Patients With Fibrolamellar Hepatocellular Carcinoma; Clinical trial registration NCT04248569; clinicaltrials.gov, 2025. https://clinicaltrials.gov/study/NCT04248569 (Accessed 05 November 2025).Google ScholarThere is no corresponding record for this reference.
- 38Enterome. A Multicenter, Open-Label, First-in-Human, Phase 1b/2a Trial of EO2401, a Novel Multipeptide Therapeutic Vaccine, With and Without Check Point Inhibitor, Following Standard Treatment in Patients With Progressive Glioblastoma; Clinical trial registration NCT04116658; clinicaltrials.gov, 2024. https://clinicaltrials.gov/study/NCT04116658 (Accessed 05 November 2025).Google ScholarThere is no corresponding record for this reference.
- 39German Cancer Research Center. AMPLIFYing NEOepitope-Specific VACcine Responses in Progressive Diffuse Glioma - a Randomized, Open Label, 3 Arm Multicenter Phase I Trial to Assess Safety, Tolerability and Immunogenicity of IDH1R132H-Specific Peptide Vaccine in Combination with Checkpoint Inhibitor Avelumab; Clinical trial registration NCT03893903; clinicaltrials.gov, 2025. https://clinicaltrials.gov/study/NCT03893903 (Accessed 05 November 2025).Google ScholarThere is no corresponding record for this reference.
- 40National University Hospital, Singapore. Da VINci Study (OTSGC-A24 Therapeutic Peptide Vaccine + Ipilimumab + Nivolumab) Nivolumab, Ipilimumab and OTSGC-A24 Therapeutic Peptide Vaccine in Gastric Cancer - a Combination Immunotherapy Phase Ib Study.; Clinical trial registration NCT03784040; clinicaltrials.gov, 2019. https://clinicaltrials.gov/study/NCT03784040 (Accessed 05 November 2025).Google ScholarThere is no corresponding record for this reference.
- 41OncoPep, Inc. A Single-Arm, Open-Label, Multi-Center Phase II Study to Evaluate the Combination of PVX-410 + Pembrolizumab + Chemotherapy for Frontline Therapy of Metastatic, PD-L1+ Triple-Negative Breast Cancer (TNBC) in HLA-A2-Positive Patients; Clinical trial registration NCT04634747; clinicaltrials.gov, 2022. https://clinicaltrials.gov/study/NCT04634747 (Accessed 05 November 2025).Google ScholarThere is no corresponding record for this reference.
- 42NuGenerex Immuno-Oncology. A Phase II Clinical Trial of Pembrolizumab in Combination With the AE37 Peptide Vaccine in Patients With Metastatic Triple-Negative Breast Cancer; Clinical trial registration NCT04024800; clinicaltrials.gov, 2021. https://clinicaltrials.gov/study/NCT04024800 (Accessed 05 November 2025).Google ScholarThere is no corresponding record for this reference.
- 43Abd-Aziz, N.; Poh, C. L. Development of Peptide-Based Vaccines for Cancer. J. Oncol. 2022, 2022 (1), 9749363, DOI: 10.1155/2022/9749363Google ScholarThere is no corresponding record for this reference.
- 44Wang, L.; Wang, N.; Zhang, W.; Cheng, X.; Yan, Z.; Shao, G.; Wang, X.; Wang, R.; Fu, C. Therapeutic peptides: current applications and future directions. Signal Transduction Targeted Ther. 2022, 7, 48, DOI: 10.1038/s41392-022-00904-4Google ScholarThere is no corresponding record for this reference.
- 45Chen, Y.; Clay, N.; Phan, N.; Lothrop, E.; Culkins, C.; Robinson, B.; Stubblefield, A.; Ferguson, A.; Kimmel, B. R. Molecular Matchmakers: Bioconjugation Techniques Enhance Prodrug Potency for Immunotherapy. Mol. Pharmaceutics 2025, 22 (1), 58– 80, DOI: 10.1021/acs.molpharmaceut.4c00867Google ScholarThere is no corresponding record for this reference.
- 46Ott, P.A.; Hu, Z.; Keskin, D.B.; Shukla, S.A.; Sun, J.; Bozym, D.J.; Zhang, W.; Luoma, A.; Giobbie-Hurder, A.; Peter, L. An immunogenic personal neoantigen vaccine for patients with melanoma\. Nature 2017, 547, 217– 221, DOI: 10.1038/nature22991Google ScholarThere is no corresponding record for this reference.
- 47Sam-Yellowe, T. Y. Antigens. In Immunology: Overview and Laboratory Manual; Springer International Publishing: Cham, 2021; pp. 65– 72. DOI: DOI: 10.1007/978-3-030-64686-8_9 .Google ScholarThere is no corresponding record for this reference.
- 48Kang, R.; Loux, T.; Tang, D.; Schapiro, N. E.; Vernon, P.; Livesey, K. M.; Krasinskas, A.; Lotze, M. T.; Zeh, H. J. The Expression of the Receptor for Advanced Glycation Endproducts (RAGE) Is Permissive for Early Pancreatic Neoplasia. Proc. Natl. Acad. Sci. U. S. A. 2012, 109 (18), 7031– 7036, DOI: 10.1073/pnas.1113865109Google ScholarThere is no corresponding record for this reference.
- 49Chen, M.-C.; Chen, K.-C.; Chang, G.-C.; Lin, H.; Wu, C.-C.; Kao, W.-H.; Teng, C.-L. J.; Hsu, S.-L.; Yang, T.-Y. RAGE Acts as an Oncogenic Role and Promotes the Metastasis of Human Lung Cancer. Cell Death Dis. 2020, 11 (4), 265, DOI: 10.1038/s41419-020-2432-1Google ScholarThere is no corresponding record for this reference.
- 50Leão, R.; Apolónio, J. D.; Lee, D.; Figueiredo, A.; Tabori, U.; Castelo-Branco, P. Mechanisms of Human Telomerase Reverse Transcriptase (hTERT) Regulation: Clinical Impacts in Cancer. J. Biomed. Sci. 2018, 25 (1), 22, DOI: 10.1186/s12929-018-0422-8Google ScholarThere is no corresponding record for this reference.
- 51Sharma, S.; Chowdhury, S. Emerging Mechanisms of Telomerase Reactivation in Cancer. Trends Cancer 2022, 8 (8), 632– 641, DOI: 10.1016/j.trecan.2022.03.005Google ScholarThere is no corresponding record for this reference.
- 52Cheng, X. A Comprehensive Review of HER2 in Cancer Biology and Therapeutics. Genes 2024, 15 (7), 903, DOI: 10.3390/genes15070903Google ScholarThere is no corresponding record for this reference.
- 53NCI Antibody and Immunotoxin Treatments for Mesothelin-expressing Cancers; NCI, https://techtransfer.cancer.gov/available-technologies?abstract=TAB-4422. (Accessed 04 November 2025).Google ScholarThere is no corresponding record for this reference.
- 54Weidemann, S.; Gagelmann, P.; Gorbokon, N.; Lennartz, M.; Menz, A.; Luebke, A. M.; Kluth, M.; Hube-Magg, C.; Blessin, N. C.; Fraune, C.; Möller, K.; Bernreuther, C.; Lebok, P.; Clauditz, T. S.; Jacobsen, F.; Izbicki, J. R.; Jansen, K.; Sauter, G.; Uhlig, R.; Wilczak, W.; Steurer, S.; Minner, S.; Burandt, E.; Krech, R. H.; Dum, D.; Krech, T.; Marx, A. H.; Simon, R. Mesothelin Expression in Human Tumors: A Tissue Microarray Study on 12,679 Tumors. Biomedicines 2021, 9 (4), 397, DOI: 10.3390/biomedicines9040397Google ScholarThere is no corresponding record for this reference.
- 55Vargas, A. J.; Sittadjody, S.; Thangasamy, T.; Mendoza, E. E.; Limesand, K. H.; Burd, R. Exploiting Tyrosinase Expression and Activity in Melanocytic Tumors: Quercetin and the Central Role of p53. Integr. Cancer Ther 2011, 10 (4), 328– 340, DOI: 10.1177/1534735410391661Google ScholarThere is no corresponding record for this reference.
- 56Vargas, A. J.; Sittadjody, S.; Thangasamy, T.; Mendoza, E. E.; Limesand, K. H.; Burd, R. Exploiting Tyrosinase Expression and Activity in Melanocytic Tumors: Quercetin and the Central Role of P53. Integr. Cancer Ther. 2011, 10 (4), 328– 340, DOI: 10.1177/1534735410391661Google ScholarThere is no corresponding record for this reference.
- 57Mann, J. E.; Hasson, N.; Su, D. G.; Adeniran, A. J.; Smalley, K. S. M.; Djureinovic, D.; Jilaveanu, L. B.; Schoenfeld, D. A.; Kluger, H. M. GP100 Expression Is Variable in Intensity in Melanoma. Cancer Immunol. Immunother. 2024, 73 (10), 191, DOI: 10.1007/s00262-024-03776-5Google ScholarThere is no corresponding record for this reference.
- 58Wang, J.; Jia, R.; Yao, Y.; Cun, B.; Huang, X.; Gu, P.; Ge, S.; Fan, X. Differential Expression of Mart-1 in Human Uveal Melanoma Cells. Mol. Med. Rep. 2011, 4 (5), 799– 803, DOI: 10.3892/mmr.2011.504Google ScholarThere is no corresponding record for this reference.
- 59Mayo Clinic Types of prostate cancer: Common and rare forms. Diseases and Conditions; Mayo Clinic. https://www.mayoclinic.org/diseases-conditions/prostate-cancer/in-depth/prostate-cancer-types/art-20584938. (Accessed 05 November 2025).Google ScholarThere is no corresponding record for this reference.
- 60Shair, K. H. Y.; Bendt, K. M.; Edwards, R. H.; Nielsen, J. N.; Moore, D. T.; Raab-Traub, N. Epstein-Barr Virus-Encoded Latent Membrane Protein 1 (LMP1) and LMP2A Function Cooperatively To Promote Carcinoma Development in a Mouse Carcinogenesis Model. J. Virol. 2012, 86 (9), 5352– 5365, DOI: 10.1128/JVI.07035-11Google ScholarThere is no corresponding record for this reference.
- 61Pal, A.; Kundu, R. Human Papillomavirus E6 and E7: The Cervical Cancer Hallmarks and Targets for Therapy. Front Microbiol. 2020, 10, 3116, DOI: 10.3389/fmicb.2019.03116Google ScholarThere is no corresponding record for this reference.
- 62Baba, S. K.; Alblooshi, S. S. E.; Yaqoob, R.; Behl, S.; Al Saleem, M.; Rakha, E. A.; Malik, F.; Singh, M.; Macha, M. A.; Akhtar, M. K.; Houry, W. A.; Bhat, A. A.; Al Menhali, A.; Zheng, Z.-M.; Mirza, S. Human Papilloma Virus (HPV) Mediated Cancers: An Insightful Update. J. Transl. Med. 2025, 23 (1), 483, DOI: 10.1186/s12967-025-06470-xGoogle ScholarThere is no corresponding record for this reference.
- 63Valcarcel, B.; Enriquez-Vera, D.; De-la-Cruz-Ku, G.; Chambergo-Michilot, D.; Calderón-Huaycochea, H.; Malpica, L. Epidemiological Features and Outcomes of HTLV-1 Carriers Diagnosed With Cancer: A Retrospective Cohort Study in an Endemic Country. JCO Glob. Oncol. 2023, 9, e2200369 DOI: 10.1200/GO.22.00369Google ScholarThere is no corresponding record for this reference.
- 64Du, G.; Zhang, W.; Zhang, Z.; Zeng, M.; Wang, Y. HTLV-1-Associated Genes as Potential Biomarkers for Endometrial Cancer. Oncol Lett. 2019, 18 (1), 699– 705, DOI: 10.3892/ol.2019.10389Google ScholarThere is no corresponding record for this reference.
- 65Yang, Y.; Zhang, H.; Huang, S.; Chu, Q. KRAS Mutations in Solid Tumors: Characteristics, Current Therapeutic Strategy, and Potential Treatment Exploration. J. Clin. Med. 2023, 12 (2), 709, DOI: 10.3390/jcm12020709Google ScholarThere is no corresponding record for this reference.
- 66Johnson, D. B.; Puzanov, I. Treatment of NRAS-Mutant Melanoma. Curr. Treat Options Oncol. 2015, 16 (4), 15, DOI: 10.1007/s11864-015-0330-zGoogle ScholarThere is no corresponding record for this reference.
- 67Pencheva, B.; Di Paola, J.; Porter, C. C.Adam, M.; Feldman, J.; Mirzaa, G. M.; Pagon, R. A.; Wallace, S. E.; Amemiya, A. ETV6-Related Thrombocytopenia and Predisposition to Leukemia. In GeneReviews; University of Washington: Seattle: Seattle (WA, 1993.Google ScholarThere is no corresponding record for this reference.
- 68Biswas, A.; Rajesh, Y.; Mitra, P.; Mandal, M. ETV6 Gene Aberrations in Non-Haematological Malignancies: A Review Highlighting ETV6 Associated Fusion Genes in Solid Tumors. Biochim. Biophys. Acta BBA - Rev. Cancer 2020, 1874 (1), 188389, DOI: 10.1016/j.bbcan.2020.188389Google ScholarThere is no corresponding record for this reference.
- 69Andraos, E.; Dignac, J.; Meggetto, F. NPM-ALK: A Driver of Lymphoma Pathogenesis and a Therapeutic Target. Cancers 2021, 13 (1), 144, DOI: 10.3390/cancers13010144Google ScholarThere is no corresponding record for this reference.
- 70Buonaguro, L.; Tagliamonte, M. Selecting Target Antigens for Cancer Vaccine Development. Vaccines 2020, 8 (4), 615, DOI: 10.3390/vaccines8040615Google ScholarThere is no corresponding record for this reference.
- 71Xie, N.; Shen, G.; Gao, W.; Huang, Z.; Huang, C.; Fu, L. Neoantigens: promising targets for cancer therapy. Signal Transduction Targeted Ther. 2023, 8, 9, DOI: 10.1038/s41392-022-01270-xGoogle ScholarThere is no corresponding record for this reference.
- 72Verdon, D. J.; Jenkins, M. R. Identification and Targeting of Mutant Peptide Neoantigens in Cancer Immunotherapy. Cancers 2021, 13 (16), 4245, DOI: 10.3390/cancers13164245Google ScholarThere is no corresponding record for this reference.
- 73Chumakov, P.M. Function of the p53 Gene: Choice between Life and Death. Biochemistry 2000, 65, 34– 47Google ScholarThere is no corresponding record for this reference.
- 74Parikh, N.; Hilsenbeck, S.; Creighton, C.J.; Dayaram, T.; Shuck, R.; Shinbrot, E.; Xi, L.; Gibbs, R.A.; Wheeler, D.A.; Donehower, L.A. Effects of TP53 mutational status on gene expression patterns across 10 human cancer types - Parikh - 2014 - The Journal of Pathology - Wiley Online Library. cancer types. J. Pathol. 2014, 232, 522– 533, DOI: 10.1002/path.4321Google ScholarThere is no corresponding record for this reference.
- 75Chong, C.; Coukos, G.; Bassani-Sternberg, M. Identification of Tumor Antigens with Immunopeptidomics. Nat. Biotechnol. 2022, 40 (2), 175– 188, DOI: 10.1038/s41587-021-01038-8Google ScholarThere is no corresponding record for this reference.
- 76Tejaswi, L.; Ramesh, P.; Aditya, S.; Raju, R.; Prasad, T. S. K. Computational Neoantigen Prediction for Cancer Immunotherapy. Genes Immun. 2025, 26 (6), 531– 538, DOI: 10.1038/s41435-025-00365-zGoogle ScholarThere is no corresponding record for this reference.
- 77Manakongtreecheep, K.; Ctortecka, C.; Correa-Medero, L.O.; Zhu, T.; Lippincott, I.; Lawrence, G.M.; Howard, A.; Hernandez, G.M.; Forman, C.; Duggan, E.C. Sensitive detection of cancer antigens enabled by user-defined peptide libraries. Nature Biotechnol. 2026, 1– 13, DOI: 10.1038/s41587-026-03003-9Google ScholarThere is no corresponding record for this reference.
- 78Science Direct Topics Ligandome - an overview; Science Direct Topics, 2016.Google ScholarThere is no corresponding record for this reference.
- 79Verdon, D.J.; Jenkins, M.R. Identification and Targeting of Mutant Peptide Neoantigens in Cancer Immunotherapy. Cancers 2021, 13 (16), 4245, DOI: 10.3390/cancers13164245Google ScholarThere is no corresponding record for this reference.
- 80Shao, Y.; Ge, S.; Dong, R.; Ji, W.; Qin, C.; Wen, P. NeoTImmuML: A Machine Learning-Based Prediction Model for Human Tumor Neoantigen Immunogenicity. Front. Immunol. 2025, 16, 1681396, DOI: 10.3389/fimmu.2025.1681396Google ScholarThere is no corresponding record for this reference.
- 81Adams, C.; Gabriel, W.; Laukens, K.; Picciani, M.; Wilhelm, M.; Bittremieux, W.; Boonen, K. Fragment Ion Intensity Prediction Improves the Identification Rate of Non-Tryptic Peptides in timsTOF. Nat. Commun. 2024, 15 (1), 3956, DOI: 10.1038/s41467-024-48322-0Google ScholarThere is no corresponding record for this reference.
- 82Strauss, M. T.; Bludau, I.; Zeng, W.-F.; Voytik, E.; Ammar, C.; Schessner, J. P.; Ilango, R.; Gill, M.; Meier, F.; Willems, S.; Mann, M. AlphaPept: A Modern and Open Framework for MS-Based Proteomics. Nat. Commun. 2024, 15 (1), 2168, DOI: 10.1038/s41467-024-46485-4Google ScholarThere is no corresponding record for this reference.
- 83Wang, X.; Wu, T.; Jiang, Y.; Chen, T.; Pan, D.; Jin, Z.; Xie, J.; Quan, L.; Lyu, Q. RPEMHC: Improved Prediction of MHC–Peptide Binding Affinity by a Deep Learning Approach Based on Residue–Residue Pair Encoding. Bioinformatics 2024, 40 (1), btad785, DOI: 10.1093/bioinformatics/btad785Google ScholarThere is no corresponding record for this reference.
- 84Li, G.; Mahajan, S.; Ma, S.; Jeffery, E. D.; Zhang, X.; Bhattacharjee, A.; Venkatasubramanian, M.; Weirauch, M. T.; Miraldi, E. R.; Grimes, H. L.; Sheynkman, G. M.; Tilburgs, T.; Salomonis, N. Splicing Neoantigen Discovery with SNAF Reveals Shared Targets for Cancer Immunotherapy. Sci. Transl. Med. 2024, 16 (730), eade2886 DOI: 10.1126/scitranslmed.ade2886Google ScholarThere is no corresponding record for this reference.
- 85Declercq, A.; Devreese, R.; Scheid, J.; Jachmann, C.; Van Den Bossche, T.; Preikschat, A.; Gomez-Zepeda, D.; Rijal, J. B.; Hirschler, A.; Krieger, J. R.; Srikumar, T.; Rosenberger, G.; Martelli, C.; Trede, D.; Carapito, C.; Tenzer, S.; Walz, J. S.; Degroeve, S.; Bouwmeester, R.; Martens, L.; Gabriels, R. TIMS2Rescore: A Data Dependent Acquisition-Parallel Accumulation and Serial Fragmentation-Optimized Data-Driven Rescoring Pipeline Based on MS2Rescore. J. Proteome Res. 2025, 24 (3), 1067– 1076, DOI: 10.1021/acs.jproteome.4c00609Google ScholarThere is no corresponding record for this reference.
- 86Zhao, Y.; Yu, J.; Su, Y.; Shu, Y.; Ma, E.; Wang, J.; Jiang, S.; Wei, C.; Li, D.; Huang, Z.; Cheng, G.; Ren, H.; Feng, J. A Unified Deep Framework for Peptide–Major Histocompatibility Complex–T Cell Receptor Binding Prediction. Nat. Mach. Intell. 2025, 7 (4), 650– 660, DOI: 10.1038/s42256-025-01002-0Google ScholarThere is no corresponding record for this reference.
- 87Li, J.; Zhao, Z.; Tai, C.; Sun, T.; Tan, L.; Li, X.; He, W.; Li, H.; Zhang, J. VirusImmu: A Novel Ensemble Machine Learning Approach for Viral Immunogenicity Prediction. Brief Funct. Genomics 2025, 24, elaf008 DOI: 10.1093/bfgp/elaf008Google ScholarThere is no corresponding record for this reference.
- 88Cai, Y.; Chen, R.; Song, M.; Wang, L.; Huo, Z.; Yang, D.; Zhang, S.; Gao, S.; Hwang, S.; Bai, L.; Lv, Y.; Cui, Y.; Zhang, X. CNNeoPP: A Large Language Model-Enhanced Deep Learning Pipeline for Personalized Neoantigen Prediction and Liquid Biopsy Applications. Front. Immunol. 2026, 17, 1722117, DOI: 10.3389/fimmu.2026.1722117Google ScholarThere is no corresponding record for this reference.
- 89Lee, K.-H.; Sears, T. J.; Zanetti, M.; Carter, H. NeoPrecis: Enhancing Immunotherapy Response Prediction through Integration of Qualified Immunogenicity and Clonality-Aware Neoantigen Landscapes. Nat. Commun. 2026, 17 (1), 1966, DOI: 10.1038/s41467-026-68651-6Google ScholarThere is no corresponding record for this reference.
- 90Slaton, E. W.; Clay, N.; Phan, N.; Kimmel, B. R. Innovations in Yeast Synthetic Biology: Engineered Discovery Systems for Immunotherapy. ACS Synth. Biol. 2025, 14 (9), 3293– 3305, DOI: 10.1021/acssynbio.5c00321Google ScholarThere is no corresponding record for this reference.
- 91Calis, J. J. A.; Maybeno, M.; Greenbaum, J. A.; Weiskopf, D.; De Silva, A. D.; Sette, A.; Keşmir, C.; Peters, B. Properties of MHC Class I Presented Peptides That Enhance Immunogenicity. PLoS Comput. Biol. 2013, 9 (10), e1003266 DOI: 10.1371/journal.pcbi.1003266Google ScholarThere is no corresponding record for this reference.
- 92Capietto, A.H.; Jhunjhunwala, S.; Pollock, S.B.; Lupardus, P.; Wong, J.; Hänsch, L.; Cevallos, J.; Chestnut, Y.; Fernandez, A.; Lounsbury, N. Mutation position is an important determinant for predicting cancer neoantigens. J Exp Med. 2020, 217 (4), e20190179 DOI: 10.1084/jem.20190179Google ScholarThere is no corresponding record for this reference.
- 93Apostolopoulos, V.; Bojarska, J.; Chai, T.-T.; Elnagdy, S.; Kaczmarek, K.; Matsoukas, J.; New, R.; Parang, K.; Lopez, O. P.; Parhiz, H.; Perera, C. O.; Pickholz, M.; Remko, M.; Saviano, M.; Skwarczynski, M.; Tang, Y.; Wolf, W. M.; Yoshiya, T.; Zabrocki, J.; Zielenkiewicz, P.; AlKhazindar, M.; Barriga, V.; Kelaidonis, K.; Sarasia, E. M.; Toth, I. A Global Review on Short Peptides: Frontiers and Perspectives. Molecules 2021, 26 (2), 430, DOI: 10.3390/molecules26020430Google ScholarThere is no corresponding record for this reference.
- 94Li, W.; Joshi, M. D.; Singhania, S.; Ramsey, K. H.; Murthy, A. K. Peptide Vaccine: Progress and Challenges. Vaccines 2014, 2 (3), 515– 536, DOI: 10.3390/vaccines2030515Google ScholarThere is no corresponding record for this reference.
- 95Onodi, F.; Maherzi-Mechalikh, C.; Mougel, A.; Ben Hamouda, N.; Taboas, C.; Gueugnon, F.; Tran, T.; Nozach, H.; Marcon, E.; Gey, A. High Therapeutic Efficacy of a New Survivin LSP-Cancer Vaccine Containing CD4+ and CD8+ T-Cell Epitopes. Front. Oncol 2018, 8, 517, DOI: 10.3389/fonc.2018.00517Google ScholarThere is no corresponding record for this reference.
- 96Cai, L.; Zhang, J.; Zhu, R.; Shi, W.; Xia, X.; Edwards, M.; Finch, W.; Coombs, A.; Gao, J.; Chen, K.; Owen, S.; Jiang, S.; Lu, W. Protective Cellular Immunity Generated by Cross-Presenting Recombinant Overlapping Peptide Proteins. Oncotarget 2017, 8 (44), 76516– 76524, DOI: 10.18632/oncotarget.20407Google ScholarThere is no corresponding record for this reference.
- 97McDowell, S. H.; Gallaher, S. A.; Burden, R. E.; Scott, C. J. Leading the Invasion: The Role of Cathepsin S in the Tumour Microenvironment. Biochim. Biophys. Acta BBA - Mol. Cell Res. 2020, 1867 (10), 118781, DOI: 10.1016/j.bbamcr.2020.118781Google ScholarThere is no corresponding record for this reference.
- 98Zhang, H.; Hong, H.; Li, D.; Ma, S.; Di, Y.; Stoten, A.; Haig, N.; Gleria, K. D.; Yu, Z.; Xu, X.-N. Comparing Pooled Peptides with Intact Protein for Accessing Cross-Presentation Pathways for Protective CD8+ and CD4+ T Cells *. J. Biol. Chem. 2009, 284 (14), 9184– 9191, DOI: 10.1074/jbc.M809456200Google ScholarThere is no corresponding record for this reference.
- 99Wang, Z.; Yang, X.; Wei, X.; Shi, L.; Wang, X.; Zhuo, Z.; Su, H.; Wu, W.; Cao, Y. J. Designing Multifunctional Recombinant Vaccines: An Engineering Strategy Based on Innovative Epitope Prediction-Guided Splicing. Theranostics 2025, 15 (9), 3924– 3942, DOI: 10.7150/thno.103755Google ScholarThere is no corresponding record for this reference.
- 100Oxford Vacmedix UK Ltd A Phase 1, Multicentre, Open-Label, Nonrandomised, First-in-Human Study of OVM-200 as a Therapeutic Vaccine in Patients With Locally Advanced or Metastatic Non-Small Cell Lung Cancer, Ovarian Cancer, and Prostate Cancer; Clinical trial registration NCT05104515; clinicaltrials.gov, 2025. https://clinicaltrials.gov/study/NCT05104515. (Accessed 17 February 2026).Google ScholarThere is no corresponding record for this reference.
- 101Kordalivand, N.; Tondini, E.; Lau, C. Y. J.; Vermonden, T.; Mastrobattista, E.; Hennink, W. E.; Ossendorp, F.; Nostrum, C. F. Cationic Synthetic Long Peptides-Loaded Nanogels: An Efficient Therapeutic Vaccine Formulation for Induction of T-Cell Responses. J. Controlled Release 2019, 315, 114– 125, DOI: 10.1016/j.jconrel.2019.10.048Google ScholarThere is no corresponding record for this reference.
- 102Wei, P.; Gangapurwala, G.; Pretzel, D.; Leiske, M.N.; Wang, L.; Hoeppener, S.; Schubert, S.; Brendel, J.C.; Schubert, U.S. Smart pH-Sensitive Nanogels for Controlled Release in an Acidic Environment. Biomacromolecules 2019, 20, 130– 140, DOI: 10.1021/acs.biomac.8b01228Google ScholarThere is no corresponding record for this reference.
- 103Palmer, C.R.; Pastora, L.E.; Kimmel, B.R.; Pagendarm, H.M.; Kwiatkowski, A.J.; Stone, P.T.; Arora, K.; Francini, N.; Fedorova, O.; Pyle, A.M. Covalent Polymer-RNA Conjugates for Potent Activation of the RIG-I Pathway. Adv. Healthcare Mater. 2025, 14, 2303815, DOI: 10.1002/adhm.202303815Google ScholarThere is no corresponding record for this reference.
- 104Beach, M. A.; Nayanathara, U.; Gao, Y.; Zhang, C.; Xiong, Y.; Wang, Y.; Such, G. K. Polymeric Nanoparticles for Drug Delivery. Chem. Rev. 2024, 124 (9), 5505– 5616, DOI: 10.1021/acs.chemrev.3c00705Google ScholarThere is no corresponding record for this reference.
- 105Haim Zada, M.; Rottenberg, Y.; Domb, A. J. Peptide Loaded Polymeric Nanoparticles by Non-Aqueous Nanoprecipitation. J. Colloid Interface Sci. 2022, 622, 904– 913, DOI: 10.1016/j.jcis.2022.05.007Google ScholarThere is no corresponding record for this reference.
- 106Mehta, M.; Bui, T. A.; Yang, X.; Aksoy, Y.; Goldys, E. M.; Deng, W. Lipid-Based Nanoparticles for Drug/Gene Delivery: An Overview of the Production Techniques and Difficulties Encountered in Their Industrial Development. ACS Mater. Au 2023, 3 (6), 600– 619, DOI: 10.1021/acsmaterialsau.3c00032Google ScholarThere is no corresponding record for this reference.
- 107Xiao, W.; Jiang, W.; Chen, Z.; Huang, Y.; Mao, J.; Zheng, W.; Hu, Y.; Shi, J. Advance in Peptide-Based Drug Development: Delivery Platforms, Therapeutics and Vaccines. Signal Transduction Targeted Ther. 2025, 10 (1), 74, DOI: 10.1038/s41392-024-02107-5Google ScholarThere is no corresponding record for this reference.
- 108Pagendarm, H. M.; Stone, P. T.; Kimmel, B. R.; Baljon, J. J.; Aziz, M. H.; Pastora, L. E.; Hubert, L.; Roth, E. W.; Almunif, S.; Scott, E. A.; Wilson, J. T. Engineering Endosomolytic Nanocarriers of Diverse Morphologies Using Confined Impingement Jet Mixing. Nanoscale 2023, 15, 16016– 16029, DOI: 10.1039/D3NR02874GGoogle ScholarThere is no corresponding record for this reference.
- 109Han, S.; Ma, W.; Jiang, D.; Sutherlin, L.; Zhang, J.; Lu, Y.; Huo, N.; Chen, Z.; Engle, J.W.; Wang, Y. Intracellular signaling pathway in dendritic cells and antigen transport pathway in vivo mediated by an OVA@DDAB/PLGA nano-vaccine. J Nanobiotechnol. 2021, 19, 394, DOI: 10.1186/s12951-021-01116-8Google ScholarThere is no corresponding record for this reference.
- 110Varypataki, E. M.; Silva, A. L.; Barnier-Quer, C.; Collin, N.; Ossendorp, F.; Jiskoot, W. Synthetic long peptide-based vaccine formulations for induction of cell mediated immunity: A comparative study of cationic liposomes and PLGA nanoparticles. J. Controlled Release 2016, 226, 98– 106, DOI: 10.1016/j.jconrel.2016.02.018Google ScholarThere is no corresponding record for this reference.
- 111Sheehy, T. L.; Kwiatkowski, A. J.; Arora, K.; Kimmel, B. R.; Schulman, J. A.; Gibson-Corley, K. N.; Wilson, J. T. STING-Activating Polymer–Drug Conjugates for Cancer Immunotherapy. ACS Cent. Sci. 2024, 10 (9), 1765– 1781, DOI: 10.1021/acscentsci.4c00579Google ScholarThere is no corresponding record for this reference.
- 112Arora, K.; Sheehy, T. L.; Schulman, J. A.; Loken, J. R.; Lehmann, Z.; Kimmel, B. R.; McAtee, C.; Bharti, V.; Stone, P. T.; Weaver, A. M.; Tyska, M.; Pathak, R. K.; Wilson, J. T. Macromolecular Diamidobenzimidazole Conjugates Can Activate Stimulator of Interferon Genes. J. Am. Chem. Soc. 2025, 147 (38), 35149– 35163, DOI: 10.1021/jacs.5c13195Google ScholarThere is no corresponding record for this reference.
- 113Kimmel, B. R.; Arora, K.; Chada, N. C.; Bharti, V.; Kwiatkowski, A. J.; Finkelstein, J. E.; Hanna, A.; Arner, E. N.; Sheehy, T. L.; Pastora, L. E.; Yang, J.; Pagendarm, H. M.; Stone, P. T.; Hargrove-Wiley, E.; Taylor, B. C.; Hubert, L. A.; Fingleton, B. M.; Gibson-Corley, K. N.; May, J. C.; McLean, J. A.; Rathmell, J. C.; Richmond, A.; Rathmell, W. K.; Balko, J. M.; Wilson, J. T. Potentiating Cancer Immunotherapies with Modular Albumin-Hitchhiking Nanobody–STING Agonist Conjugates. Nat. Biomed. Eng. 2025, 9 (10), 1719– 1739, DOI: 10.1038/s41551-025-01400-0Google ScholarThere is no corresponding record for this reference.
- 114Lynn, G.M.; Sedlik, C.; Baharom, F.; Zhu, Y.; Ramirez-Valdez, R.A.; Coble, V.L.; Tobin, K.; Nichols, S.R.; Itzkowitz, Y.; Zaidi, N. Peptide–TLR-7/8a conjugate vaccines chemically programmed for nanoparticle self-assembly enhance CD8 T-cell immunity to tumor antigens. Nat Biotechnol. 2020, 38, 320– 332, DOI: 10.1038/s41587-019-0390-xGoogle ScholarThere is no corresponding record for this reference.
- 115Moon, J. H.; Phoo, M. T.; Kim, Y.; Ahn, J.; Kim, A.; Nam, J.; Lee, S.-H.; Moon, J. J.; Son, S. Polysaccharide Adjuvants as Innate Immune Trainers: Bridging Pattern Recognition Receptor (PRR) Activation and Metabolic Reprogramming for Synthetic Vaccine Design. Adv. Sci. 2025, 12 (48), e09022 DOI: 10.1002/advs.202509022Google ScholarThere is no corresponding record for this reference.
- 116Spunde, K.; Korotkaja, K.; Sominskaya, I.; Zajakina, A. Genetic Adjuvants: A Paradigm Shift in Vaccine Development and Immune Modulation. Mol. Ther Nucleic Acids 2025, 36 (2), 102536, DOI: 10.1016/j.omtn.2025.102536Google ScholarThere is no corresponding record for this reference.
- 117Wang-Bishop, L.; Wehbe, M.; Pastora, L. E.; Yang, J.; Kimmel, B. R.; Garland, K. M.; Becker, K. W.; Carson, C. S.; Roth, E. W.; Gibson-Corley, K. N.; Ulkoski, D.; Krishnamurthy, V.; Fedorova, O.; Richmond, A.; Pyle, A. M.; Wilson, J. T. Nanoparticle Retinoic Acid-Inducible Gene I Agonist for Cancer Immunotherapy. ACS Nano 2024, 18 (18), 11631– 11643, DOI: 10.1021/acsnano.3c06225Google ScholarThere is no corresponding record for this reference.
- 118Wang-Bishop, L.; Kimmel, B. R.; Ngwa, V. M.; Madden, M. Z.; Baljon, J. J.; Florian, D. C.; Hanna, A.; Pastora, L. E.; Sheehy, T. L.; Kwiatkowski, A. J.; Wehbe, M.; Wen, X.; Becker, K. W.; Garland, K. M.; Schulman, J. A.; Shae, D.; Edwards, D.; Wolf, M. M.; Delapp, R.; Christov, P. P.; Beckermann, K. E.; Balko, J. M.; Rathmell, W. K.; Rathmell, J. C.; Chen, J.; Wilson, J. T. STING-Activating Nanoparticles Normalize the Vascular-Immune Interface to Potentiate Cancer Immunotherapy. Sci. Immunol. 2023, 8 (83), eadd1153 DOI: 10.1126/sciimmunol.add1153Google ScholarThere is no corresponding record for this reference.
- 119Baljon, J. J.; Kwiatkowski, A. J.; Pagendarm, H. M.; Stone, P. T.; Kumar, A.; Bharti, V.; Schulman, J. A.; Becker, K. W.; Roth, E. W.; Christov, P. P.; Joyce, S.; Wilson, J. T. A Cancer Nanovaccine for Co-Delivery of Peptide Neoantigens and Optimized Combinations of STING and TLR4 Agonists. ACS Nano 2024, 18 (9), 6845– 6862, DOI: 10.1021/acsnano.3c04471Google ScholarThere is no corresponding record for this reference.
- 120Xi, Y.; Chen, L.; Tang, J.; Yu, B.; Shen, W.; Niu, X. Amplifying “Eat Me Signal” by Immunogenic Cell Death for Potentiating Cancer Immunotherapy. Immunol Rev. 2024, 321 (1), 94– 114, DOI: 10.1111/imr.13251Google ScholarThere is no corresponding record for this reference.
- 121Van Lint, S.; Renmans, D.; Broos, K.; Goethals, L.; Maenhout, S.; Benteyn, D.; Goyvaerts, C.; Du Four, S.; Van der Jeught, K.; Bialkowski, L. Intratumoral Delivery of TriMix mRNA Results in T-cell Activation by Cross-Presenting Dendritic Cells. Cancer Immunol Res. 2016, 4, 146– 156, DOI: 10.1158/2326-6066.CIR-15-0163Google ScholarThere is no corresponding record for this reference.
- 122Wijfjes, Z.; Tomillero, I. R.; Gall, C. M. L.; van Dinther, E. A. W.; Turlings, F.; Classens, R.; Manna, S.; van Dalen, D.; Peters, W. R. J. R.; Schouren, K. Co-Delivery of Antigen and Adjuvant by Site-Specific Conjugation to Dendritic Cell-Targeted Fab Fragments Potentiates T Cell Responses. RSC Chem. Biol. 2025, 6 (6), 948– 962, DOI: 10.1039/D5CB00014AGoogle ScholarThere is no corresponding record for this reference.
- 123Botelho, N. K.; Tschumi, B. O.; Hubbell, J. A.; Swartz, M. A.; Donda, A.; Romero, P. Combination of Synthetic Long Peptides and XCL1 Fusion Proteins Results in Superior Tumor Control. Front. Immunol 2019, 10, 294, DOI: 10.3389/fimmu.2019.00294Google ScholarThere is no corresponding record for this reference.
- 124Carreno, B. M.; Magrini, V.; Becker-Hapak, M.; Kaabinejadian, S.; Hundal, J.; Petti, A. A.; Ly, A.; Lie, W.-R.; Hildebrand, W. H.; Mardis, E. R.; Linette, G. P. A Dendritic Cell Vaccine Increases the Breadth and Diversity of Melanoma Neoantigen-Specific T Cells. Science 2015, 348 (6236), 803– 808, DOI: 10.1126/science.aaa3828Google ScholarThere is no corresponding record for this reference.
- 125Belnoue, E.; Mayol, J.-F.; Carboni, S.; Besson, W. D. B.; Dupuychaffray, E.; Nelde, A.; Stevanovic, S.; Santiago-Raber, M.-L.; Walker, P. R.; Derouazi, M. Targeting Self- and Neoepitopes with a Modular Self-Adjuvanting Cancer Vaccine. JCI Insight 2019, 4 (11), 127305, DOI: 10.1172/jci.insight.127305Google ScholarThere is no corresponding record for this reference.
- 126Bajiya, N.; Najrin, S.; Kumar, P.; Choudhury, S.; Tomer, R.; Raghava, G. P. S. CPPsite3: An Updated Large Repository of Experimentally Validated Cell-Penetrating Peptides. Drug Discovery Today 2025, 30 (8), 104421, DOI: 10.1016/j.drudis.2025.104421Google ScholarThere is no corresponding record for this reference.
- 127Gall, V.A.; Philips, A.V.; Qiao, N.A.; Clise-Dwyer, K.; Perakis, A.A.; Zhang, M.; Clifton, G.T.; Sukhumalchandra, P.; Ma, Q.; Reddy, S.M. Trastuzumab Increases HER2 Uptake and Cross-Presentation by Dendritic Cells. Cancer Res 2017, 77 (19), 5374– 5383, DOI: 10.1158/0008-5472.CAN-16-2774Google ScholarThere is no corresponding record for this reference.
- 128Gall, V.A.; Philips, A.V.; Qiao, N.A.; Clise-Dwyer, K.; Perakis, A.A.; Zhang, M.; Clifton, G.T.; Sukhumalchandra, P.; Ma, Q.; Reddy, S.M.; An open-label, multinational, multicenter study of G17DT vaccination combined with cisplatin and 5-fluorouracil in patients with untreated, advanced gastric or gastroesophageal cancer: The GC4 study - Ajani - 2006 - Cancer - Wiley Online Library. https://acsjournals.onlinelibrary.wiley.com/doi/full/10.1002/cncr.21814 (accessed 2025–10–17).Google ScholarThere is no corresponding record for this reference.
- 129Li, Q.; Lei, X.; Zhu, J.; Zhong, Y.; Yang, J.; Wang, J.; Tan, H. Radiotherapy/Chemotherapy-Immunotherapy for Cancer Management: From Mechanisms to Clinical Implications - Li - 2023 - Oxidative Medicine and Cellular Longevity - Wiley Online Library. Oxid. Med. Cell. Longevity 2023, 2023, 7530794, DOI: 10.1155/2023/7530794Google ScholarThere is no corresponding record for this reference.
- 130Lee, H.-M.; Lee, H.-J.; Chang, J.-E. Inflammatory Cytokine: An Attractive Target for Cancer Treatment. Biomedicines 2022, 10 (9), 2116, DOI: 10.3390/biomedicines10092116Google ScholarThere is no corresponding record for this reference.
- 131Tufail, M.; Jiang, C.-H.; Li, N. Immune Evasion in Cancer: Mechanisms and Cutting-Edge Therapeutic Approaches. Signal Transduction Targeted Ther. 2025, 10 (1), 227, DOI: 10.1038/s41392-025-02280-1Google ScholarThere is no corresponding record for this reference.
- 132Dagogo-Jack, I.; Shaw, A. T. Tumour Heterogeneity and Resistance to Cancer Therapies. Nat. Rev. Clin. Oncol. 2018, 15 (2), 81– 94, DOI: 10.1038/nrclinonc.2017.166Google ScholarThere is no corresponding record for this reference.
- 133Jing, H.; Gao, Y.; Sun, Z.; Liu, S. Recent Advances in Novel Tumor Immunotherapy Strategies Based on Regulating the Tumor Microenvironment and Immune Checkpoints. Front. Immunol. 2025, 16, 1529403, DOI: 10.3389/fimmu.2025.1529403Google ScholarThere is no corresponding record for this reference.
- 134Shah, D. D.; Chorawala, M. R.; Raghani, N. R.; Patel, R.; Fareed, M.; Kashid, V. A.; Prajapati, B. G. Tumor Microenvironment: Recent Advances in Understanding and Its Role in Modulating Cancer Therapies. Med. Oncol. 2025, 42 (4), 117, DOI: 10.1007/s12032-025-02641-4Google ScholarThere is no corresponding record for this reference.
- 135Emami Nejad, A.; Najafgholian, S.; Rostami, A.; Sistani, A.; Shojaeifar, S.; Esparvarinha, M.; Nedaeinia, R.; Haghjooy Javanmard, S.; Taherian, M.; Ahmadlou, M.; Salehi, R.; Sadeghi, B.; Manian, M. The Role of Hypoxia in the Tumor Microenvironment and Development of Cancer Stem Cell: A Novel Approach to Developing Treatment. Cancer Cell Int. 2021, 21 (1), 62, DOI: 10.1186/s12935-020-01719-5Google ScholarThere is no corresponding record for this reference.
- 136Basak, U.; Sarkar, T.; Mukherjee, S.; Chakraborty, S.; Dutta, A.; Dutta, S.; Nayak, D.; Kaushik, S.; Das, T.; Sa, G. Tumor-Associated Macrophages: An Effective Player of the Tumor Microenvironment. Front. Immunol. 2023, 14, 1295257, DOI: 10.3389/fimmu.2023.1295257Google ScholarThere is no corresponding record for this reference.
- 137Yu, Y.; Yao, X.; Wang, Q.; Yang, M.; Li, R.; Qin, J.; Zhuang, J.; Sun, C. T Cell Exhaustion in Cancer Immunotherapy: Heterogeneity, Mechanisms, and Therapeutic Opportunities. Adv. Sci. 2026, n/a (n/a), e20634 DOI: 10.1002/advs.202520634Google ScholarThere is no corresponding record for this reference.
- 138Singhaviranon, S.; Dempsey, J. P.; Hagymasi, A. T.; Mandoiu, I. I.; Srivastava, P. K. Low-Avidity T Cells Drive Endogenous Tumor Immunity in Mice and Humans. Nat. Immunol. 2025, 26 (2), 240– 251, DOI: 10.1038/s41590-024-02044-zGoogle ScholarThere is no corresponding record for this reference.
- 139Tagore, S.; Caprio, L.; Amin, A. D.; Bestak, K.; Luthria, K.; D’Souza, E.; Barrera, I.; Melms, J. C.; Wu, S.; Abuzaid, S.; Wang, Y.; Jakubikova, V.; Koch, P.; Brodtman, D. Z.; Bawa, B.; Deshmukh, S. K.; Ebel, L.; Ibarra-Arellano, M. A.; Jaiswal, A.; Gurjao, C.; Biermann, J.; Shaikh, N.; Ramaradj, P.; Georgis, Y.; Lagos, G. G.; Ehrlich, M. I.; Ho, P.; Walsh, Z. H.; Rogava, M.; Politis, M. G.; Biswas, D.; Cottarelli, A.; Rizvi, N.; Shu, C. A.; Herzberg, B.; Anandasabapathy, N.; Sledge, G.; Zorn, E.; Canoll, P.; Bruce, J. N.; Rizvi, N. A.; Taylor, A. M.; Saqi, A.; Hibshoosh, H.; Schwartz, G. K.; Henick, B. S.; Chen, F.; Schapiro, D.; Shah, P.; Izar, B. Single-Cell and Spatial Genomic Landscape of Non-Small Cell Lung Cancer Brain Metastases. Nat. Med. 2025, 31 (4), 1351– 1363, DOI: 10.1038/s41591-025-03530-zGoogle ScholarThere is no corresponding record for this reference.
- 140Le, J.; Dian, Y.; Zhao, D.; Guo, Z.; Luo, Z.; Chen, X.; Zeng, F.; Deng, G. Single-Cell Multi-Omics in Cancer Immunotherapy: From Tumor Heterogeneity to Personalized Precision Treatment. Mol. Cancer 2025, 24 (1), 221, DOI: 10.1186/s12943-025-02426-3Google ScholarThere is no corresponding record for this reference.
- 141Hamley, I. W. Peptides for Vaccine Development. ACS Appl. Bio Mater. 2022, 5 (3), 905– 944, DOI: 10.1021/acsabm.1c01238Google ScholarThere is no corresponding record for this reference.
- 142Culkins, C.; Adomanis, R.; Phan, N.; Robinson, B.; Slaton, E.; Lothrop, E.; Chen, Y.; Kimmel, B.R. Unlocking the Gates: Therapeutic Agents for Noninvasive Drug Delivery Across the Blood-Brain Barrier. Mol. Pharmaceutics 2024, 21, 5430– 5454, DOI: 10.1021/acs.molpharmaceut.4c00604Google ScholarThere is no corresponding record for this reference.
- 143Mansurov, A.; Hosseinchi, P.; Chang, K.; Lauterbach, A. L.; Gray, L. T.; Alpar, A. T.; Budina, E.; Slezak, A. J.; Kang, S.; Cao, S.; Solanki, A.; Gomes, S.; Williford, J.-M.; Swartz, M. A.; Mendoza, J. L.; Ishihara, J.; Hubbell, J. A. Masking the Immunotoxicity of Interleukin-12 by Fusing It with a Domain of Its Receptor via a Tumour-Protease-Cleavable Linker. Nat. Biomed. Eng. 2022, 6 (7), 819– 829, DOI: 10.1038/s41551-022-00888-0Google ScholarThere is no corresponding record for this reference.
- 144Kimmel, B. R.; Mrksich, M. Development of an Enzyme-Inhibitor Reaction Using Cellular Retinoic Acid Binding Protein II for One-Pot Megamolecule Assembly. Chem. – Eur. J. 2021, 27 (71), 17843– 17848, DOI: 10.1002/chem.202103059Google ScholarThere is no corresponding record for this reference.
- 145Metcalf, K. J.; Kimmel, B. R.; Sykora, D. J.; Modica, J. A.; Parker, K. A.; Berens, E.; Dai, R.; Dravid, V. P.; Werb, Z.; Mrksich, M. Synthetic Tuning of Domain Stoichiometry in Nanobody–Enzyme Megamolecules. Bioconjugate Chem. 2021, 32 (1), 143– 152, DOI: 10.1021/acs.bioconjchem.0c00578Google ScholarThere is no corresponding record for this reference.
Cited By
This article has not yet been cited by other publications.
Article Views
Altmetric
Citations
Article Views are the COUNTER-compliant sum of full text article downloads since November 2008 (both PDF and HTML) across all institutions and individuals. These metrics are regularly updated to reflect usage leading up to the last few days.
Citations are the number of other articles citing this article, calculated by Crossref and updated daily. Find more information about Crossref citation counts.
The Altmetric Attention Score is a quantitative measure of the attention that a research article has received online. Clicking on the donut icon will load a page at altmetric.com with additional details about the score and the social media presence for the given article. Find more information on the Altmetric Attention Score and how the score is calculated.
Recommended Articles
Abstract
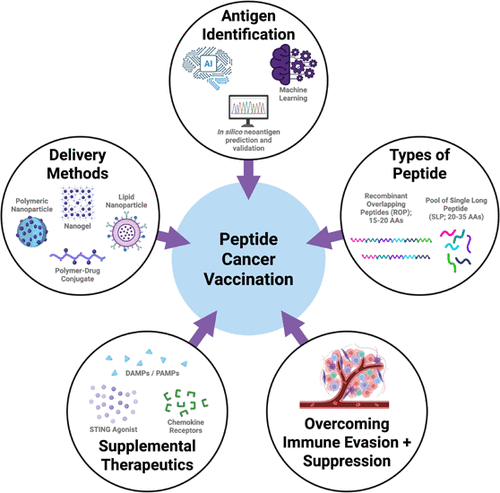
Figure 1

Figure 1. Comparison of different types of cancer vaccinations. (A) Cellular vaccines contain cells or parts of cells isolated from cancer patients, typically either cancerous cells or dendritic cells. These isolated cells are modified and then administered back to the patient to trigger an immune response against cancer cells. While these vaccinations can elicit an immune response that leads to tumor regression, isolating and processing patient cells is time-consuming and expensive. (B) Nucleic acid vaccines are made of either DNA or RNA. The nucleic acid is delivered to cells, allowing the genetic information to be processed and expressed as proteins. Currently, more focus has been placed on mRNA vaccines because the genetic information is not incorporated within the cell’s nucleus; therefore, both dividing and nondividing cells can express the protein of interest. The mRNA vaccines are cheaper and easier to manufacture than cellular vaccines. However, these technologies require a delivery system to ensure the material is successfully transported into the cell. (C) Peptide vaccines contain cancer antigens, designed to stimulate the immune system and induce cancer regression. While these vaccines have high binding affinity for cell receptors and can be combined with other molecules, this approach has known limitations, including difficulty in identifying immunogenic antigens and the potential for peptides to be easily degraded in the body.
Figure 2
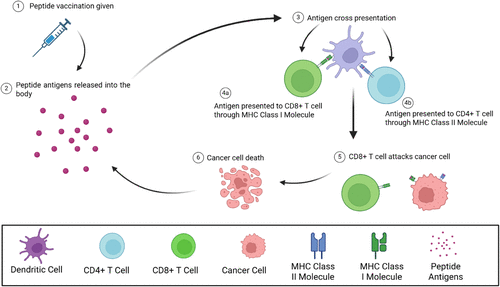
Figure 2. Overview of immune stimulation following the administration of a peptide vaccination. Following the administration of the peptide vaccination, the “drug load” or peptide antigens will be released into the body. The peptides within the body will be taken up by a dendritic cell, and processed either in the cytosol or an endosome, leading to both MHC Class I and Class II presentation. CD8+ and CD4+ T cells can then recognize antigens presented by MHC Class molecules, becoming mature T cells that can attack and destroy cancer cells.
Figure 3

Figure 3. Comparing in silico vs LC-MS neo-antigen screening methods. Both in silico and LC-MS neoantigen screening methods compare healthy cells to tumor samples to identify differences in the two genetic sequences. In silico analysis methods typically involve computer-based screening, whereas LC-MS methods involve eluting MHC peptide ligands prior to LC-MS to identify potential neoantigen candidates. Following the identification of potential neo-antigens, the candidates are screened in vitro to assess TIL activation.
Figure 4
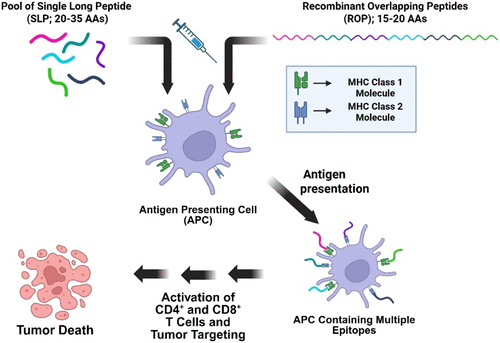
Figure 4. How SLP and ROP vaccinations stimulate the immune system. Both SLP and ROP vaccinations stimulate the immune system by being processed by APCs, leading to the activation of CD4+ and CD8+ T cells. However, SLP vaccinations comprise a pool of peptides, each representing a different antigen epitope, whereas ROP vaccinations contain a single peptide with multiple epitopes encoded within the peptide.
Figure 5

Figure 5. Different vaccine delivery methods for peptide vaccinations. There have been significant advances in drug delivery technologies, and four of the main methods are detailed within the figure. Each specific method has its advantages and drawbacks, which are detailed in the bullet points below each icon.
Figure 6

Figure 6. How immune stimulants (DAMPs/PAMPs) can bolster the immune response from peptide vaccinations. Following the death of a cancer cell, immune stimulants such as DAMPs and PAMPs are released and can bind to pattern recognition receptors (PRRs). The binding of PRRs enables DAMPs/PAMPs to interact with dendritic cells, leading to antigen presentation and the activation of CD4+ and CD8+ T cells. The activated T cells are then able to target additional cancer cells, leading to the death of these cells and initiating the cycle again.
Figure 7

Figure 7. The different barriers that contribute to no peptide cancer vaccinations being FDA approved. TME is inter-related with immune suppression and evasion, and both work together to prevent tumor cells from being recognized and destroyed by the immune system. Tumor heterogeneity complicates the ability of cancer treatments to properly target and destroy all cancer cells, as the rapid mutations present can cause cancer cells to escape surveillance, leading to more proliferation and growth.
References
This article references 145 other publications.
- 1NCI Cancer Statistics; NCI. https://www.cancer.gov/about-cancer/understanding/statistics. (Accessed 16 October 2025).There is no corresponding record for this reference.
- 2Hsu, J.Y.; Chang, C.J.; Cheng, J.S. Survival, treatment regimens and medical costs of women newly diagnosed with metastatic triple-negative breast cancer. Sci. Rep. 2022, 12 (1), 729, DOI: 10.1038/s41598-021-04316-2There is no corresponding record for this reference.
- 3American Cancer Society Survival Rates for Pancreatic Cancer; American Cancer Society, https://www.cancer.org/cancer/types/pancreatic-cancer/detection-diagnosis-staging/survival-rates.html. (Accessed16 October 2025).There is no corresponding record for this reference.
- 4Baskar, R.; Dai, J.; Wenlong, N.; Yeo, R.; Yeoh, K.-W. Biological Response of Cancer Cells to Radiation Treatment. Front. Mol. Biosci. 2014, 1, 24, DOI: 10.3389/fmolb.2014.00024There is no corresponding record for this reference.
- 5Stephens, A. J.; Burgess-Brown, N. A.; Jiang, S. Beyond Just Peptide Antigens: The Complex World of Peptide-Based Cancer Vaccines. Front. Immunol. 2021, 12, 696791, DOI: 10.3389/fimmu.2021.696791There is no corresponding record for this reference.
- 6O’Donnell, J.S.; Teng, M.W.; Smyth, M.J. Cancer immunoediting and resistance to T cell-based immunotherapy. Nat. Rev. Clin. Oncol. 2019, 16 (3), 151– 167, DOI: 10.1038/s41571-018-0142-8There is no corresponding record for this reference.
- 7Wang, C.; Hao, P. Therapeutic Vaccine Strategies in Cancer: Advances, Challenges, and Future Directions. Clin. Cancer Bull. 2025, 4 (1), 10, DOI: 10.1007/s44272-025-00039-xThere is no corresponding record for this reference.
- 8Ma, L.; Diao, L.; Peng, Z.; Jia, Y.; Xie, H.; Li, B.; Ma, J.; Zhang, M.; Cheng, L.; Ding, D.; Zhang, X.; Chen, H.; Mo, F.; Jiang, H.; Xu, G.; Meng, F.; Zhong, Z.; Liu, M. Immunotherapy and Prevention of Cancer by Nanovaccines Loaded with Whole-Cell Components of Tumor Tissues or Cells. Adv. Mater. 2021, 33 (43), 2104849, DOI: 10.1002/adma.202104849There is no corresponding record for this reference.
- 9Gao, C.; Luo, R.; Kwong, C. H. T.; Liu, J.; Tang, M.; Xie, B.; Duan, T.; Wang, R. Cancer Vaccine from Intracellularly Gelated Tumor Cells Functionalized with CD47 Blockage and Damage-Associated Molecular Pattern Exposure. Cell Rep. Med. 2025, 6 (5), 102092, DOI: 10.1016/j.xcrm.2025.102092There is no corresponding record for this reference.
- 10Maeng, H. M.; Olkhanud, P. B.; Black, M.; Highfill, S. L.; Stroncek, D. F.; Berzofsky, J. A. Dendritic Cell Cancer Vaccines: Clinical Production for Cancer Immunotherapy. In In Cancer Vaccines: methods and Protocols, Padler-Karavani, V.; ed.; Springer: New York, NY, 2025; pp. 207– 221. DOI: DOI: 10.1007/978-1-0716-4542-0_16 .There is no corresponding record for this reference.
- 11Bodey, B.; Bodey, B.; Siegel, S. E.; Kaiser, H. E. Failure of Cancer Vaccines: The Significant Limitations of This Approach to Immunotherapy. Anticancer Res. 2000, 20 (4), 2665– 2676There is no corresponding record for this reference.
- 12Chen, J.; Tan, Q.; Yang, Z.; Chen, W.; Zhou, E.; Li, M.; Deng, J.; Wu, Y.; Liu, J.; Xu, J.; Guo, M.; Jin, Y. Dendritic Cell Derived-Extracellular Vesicles Engineered to Express Interleukin-12 and Anti-CTLA-4 on Their Surface for Combinational Cancer Immunotherapy. J. Extracell. Vesicles 2025, 14 (4), e70068 DOI: 10.1002/jev2.70068There is no corresponding record for this reference.
- 13Liao, H.-C.; Liu, S.-J. Advances in Nucleic Acid-Based Cancer Vaccines. J. Biomed. Sci. 2025, 32 (1), 10, DOI: 10.1186/s12929-024-01102-wThere is no corresponding record for this reference.
- 14Kaczmarek, M.; Poznańska, J.; Fechner, F.; Michalska, N.; Paszkowska, S.; Napierała, A.; Mackiewicz, A. Cancer Vaccine Therapeutics: Limitations and Effectiveness─A Literature Review. Cells 2023, 12 (17), 2159, DOI: 10.3390/cells12172159There is no corresponding record for this reference.
- 15Srivastava, R. Stimuli-Responsive Nanomaterials for the Precision Delivery of mRNA Cancer Vaccines. Nano Trends 2025, 11, 100147, DOI: 10.1016/j.nwnano.2025.100147There is no corresponding record for this reference.
- 16Zhou, J.; Liu, Y.; Xu, W.; Bhatta, R.; Han, J.; Baskaran, D.; Devmal, S.; Leal, C.; Wang, H. Macroporous Hydrogel-Based mRNA Cancer Vaccine for in Situ Recruitment and Modulation of Dendritic Cells. Acta Biomater. 2025, 204, 234– 245, DOI: 10.1016/j.actbio.2025.08.005There is no corresponding record for this reference.
- 17Kim, Y. A.; Jeong, H.; Kim, H.; Lee, S.; Kim, K. S.; Na, K. Lipid Nanoparticles with Prazole Adjuvant to Enhance the Efficacy of mRNA Cancer Vaccines. J. Controlled Release 2025, 383, 113756, DOI: 10.1016/j.jconrel.2025.113756There is no corresponding record for this reference.
- 18Paula Fawzy, M.; Hassan, H. A. F. M.; Umair Amin, M.; Preis, E.; Bakowsky, U.; Ashraf Fahmy, S. Deploying Nucleic Acids-Loaded Plant-Derived Exosomes as Green Nano Gadget in Cancer Gene Therapy. Mater. Adv. 2025, 6 (4), 1230– 1261, DOI: 10.1039/D4MA01189AThere is no corresponding record for this reference.
- 19Chudasama, R.; Phung, Q.; Hsu, A.; Almhanna, K. Vaccines in Gastrointestinal Malignancies: From Prevention to Treatment. Vaccines 2021, 9 (6), 647, DOI: 10.3390/vaccines9060647There is no corresponding record for this reference.
- 20Xu, F.; Yuan, Y.; Wang, Y.; Yin, Q. Emerging Peptide-Based Nanovaccines: From Design Synthesis to Defense against Cancer and Infection. Biomedecine Pharmacother 2023, 158, 114117, DOI: 10.1016/j.biopha.2022.114117There is no corresponding record for this reference.
- 21Eiz-Vesper, B.; Schmetzer, H. M. Antigen-Presenting Cells: Potential of Proven Und New Players in Immune Therapies. Transfus. Med. Hemotherapy 2021, 47 (6), 429– 431, DOI: 10.1159/000512729There is no corresponding record for this reference.
- 22Borst, J.; Ahrends, T.; Bąbała, N.; Melief, C. J. M.; Kastenmüller, W. CD4+ T Cell Help in Cancer Immunology and Immunotherapy. Nat. Rev. Immunol. 2018, 18 (10), 635– 647, DOI: 10.1038/s41577-018-0044-0There is no corresponding record for this reference.
- 23Wu, L.; Yi, W.; Yao, S.; Xie, S.; Peng, R.; Zhang, J.; Tan, W. mRNA-Based Cancer Vaccines: Advancements and Prospects. Nano Lett. 2024, 24 (41), 12711– 12721, DOI: 10.1021/acs.nanolett.4c03296There is no corresponding record for this reference.
- 24Abd-Aziz, N.; Poh, C. L. Development of Peptide-Based Vaccines for Cancer. J. Oncol. 2022, 2022, 9749363, DOI: 10.1155/2022/9749363There is no corresponding record for this reference.
- 25Bacillus Calmette-Guerin (BCG) Treatment BCG Treatment: What It Is, Procedure & Side Effects. Bacillus Calmette-Guerin (BCG) Treatment, https://my.clevelandclinic.org/health/treatments/17908-bacillus-calmette-guerin-bcg-treatment. (Accessed 17 October 2025).There is no corresponding record for this reference.
- 26Anassi, E.; Ndefo, U. A. Sipuleucel-T (Provenge) Injection. Pharm. Ther. 2011, 36 (4), 197– 202There is no corresponding record for this reference.
- 27Mayo Clinic. A Phase I/II Study of Personalized Neoantigen Peptide-Based Vaccine in Combination With Pembrolizumab in Advanced Solid Tumors (PNeoVCA). Clinical trial registration NCT05269381; clinicaltrials.gov, 2024. https://clinicaltrials.gov/study/NCT05269381. (Accessed 05 November 2025).There is no corresponding record for this reference.
- 28Sidney Kimmel Comprehensive Cancer Center at Johns Hopkins. Pooled Mutant KRAS-Targeted Long Peptide Vaccine Combined With Nivolumab and Ipilimumab for Patients With Resected MMR-p Colorectal and Pancreatic Cancer. Clinical trial registration NCT04117087; clinicaltrials.gov, 2025. https://clinicaltrials.gov/study/NCT04117087. (Accessed 05 November 2025).There is no corresponding record for this reference.
- 29UNC Lineberger Comprehensive Cancer Center. Phase I Trial of a Personalized and Adaptive Neoantigen Dose-Adjusted Vaccine Administered Concurrently With Pembrolizumab. Clinical trial registration NCT04266730; clinicaltrials.gov, 2025. https://clinicaltrials.gov/study/NCT04266730. (Accessed 05 November 2025).There is no corresponding record for this reference.
- 30IO Biotech. A Phase II Multi-Arm (Basket) Trial Investigating the Safety and Efficacy of IO102-IO103 in Combination With Pembrolizumab, as First-Line Treatment for Patients With Metastatic NSCLC, SCCHN, or Metastatic mUBC. Clinical trial registration NCT05077709; clinicaltrials.gov, 2024. https://clinicaltrials.gov/study/NCT05077709. (Accessed 05 November 2025).There is no corresponding record for this reference.
- 31Fred Hutchinson Cancer Center. PNV21–001: A Phase I Study of a Personalized Multi-Peptide Neo-Antigen Vaccine in Breast Cancer, PD1/PD-L1 Inhibitor-Refractory Melanoma, and Pretreated Non-Small Cell Lung Cancer. Clinical trial registration NCT05098210; clinicaltrials.gov, 2025. https://clinicaltrials.gov/study/NCT05098210 (Accessed 05 November 2025).There is no corresponding record for this reference.
- 32Digklia, A. A Phase Ib Study of the Combination of Personalized Autologous Dendritic Cell Vaccine and Standard of Care Adjuvant Chemotherapy Followed by Nivolumab for Resected Pancreatic Adenocarcinoma. Clinical trial registration NCT04627246; clinicaltrials.gov, 2025. https://clinicaltrials.gov/study/NCT04627246. (Accessed 05 November 2025).There is no corresponding record for this reference.
- 33ARCAGY/GINECO GROUP. Randomized Phase II Study Comparing Neo-Epitope Based Vaccine OSE2101 (TEDOPI®) With or Without Anti-PD1 (Pembrolizumab) Versus Best Supportive Care as Maintenance Treatment in Platinum-Sensitive Recurrent Ovarian Cancer Patient With Controlled Disease After Platinum-Based Chemotherapy. Clinical trial registration NCT04713514; clinicaltrials.gov, 2024. https://clinicaltrials.gov/study/NCT04713514. (Accessed 05 November 2025).There is no corresponding record for this reference.
- 34Ott, P. NeoVax With Nivolumab in Patients With Ovarian Cancer. Clinical trial registration NCT04024878; clinicaltrials.gov, 2025. https://clinicaltrials.gov/study/NCT04024878. (Accessed 05 November 2025).There is no corresponding record for this reference.
- 35Centre Hospitalier Universitaire de Besancon. Evaluation of UCPVax Plus Nivolumab as Second Line Therapy in Advanced Non Small Cell Lung: A Randomized Non Comparative Phase II Trial; Clinical trial registration NCT04263051; clinicaltrials.gov, 2025. https://clinicaltrials.gov/study/NCT04263051 (Accessed 05 November 2025).There is no corresponding record for this reference.
- 36Ott, P. A Phase Ib Study of NeoVax in Combination With CDX-301 and Nivolumab or Pembrolizumab and in Patients With Melanoma; Clinical trial registration NCT04930783; clinicaltrials.gov, 2025. https://clinicaltrials.gov/study/NCT04930783 (Accessed 05 November 2025).There is no corresponding record for this reference.
- 37Sidney Kimmel Comprehensive Cancer Center at Johns Hopkins. A Pilot Study of a DNAJB1-PRKACA Fusion Kinase Peptide Vaccine Combined With Nivolumab and Ipilimumab for Patients With Fibrolamellar Hepatocellular Carcinoma; Clinical trial registration NCT04248569; clinicaltrials.gov, 2025. https://clinicaltrials.gov/study/NCT04248569 (Accessed 05 November 2025).There is no corresponding record for this reference.
- 38Enterome. A Multicenter, Open-Label, First-in-Human, Phase 1b/2a Trial of EO2401, a Novel Multipeptide Therapeutic Vaccine, With and Without Check Point Inhibitor, Following Standard Treatment in Patients With Progressive Glioblastoma; Clinical trial registration NCT04116658; clinicaltrials.gov, 2024. https://clinicaltrials.gov/study/NCT04116658 (Accessed 05 November 2025).There is no corresponding record for this reference.
- 39German Cancer Research Center. AMPLIFYing NEOepitope-Specific VACcine Responses in Progressive Diffuse Glioma - a Randomized, Open Label, 3 Arm Multicenter Phase I Trial to Assess Safety, Tolerability and Immunogenicity of IDH1R132H-Specific Peptide Vaccine in Combination with Checkpoint Inhibitor Avelumab; Clinical trial registration NCT03893903; clinicaltrials.gov, 2025. https://clinicaltrials.gov/study/NCT03893903 (Accessed 05 November 2025).There is no corresponding record for this reference.
- 40National University Hospital, Singapore. Da VINci Study (OTSGC-A24 Therapeutic Peptide Vaccine + Ipilimumab + Nivolumab) Nivolumab, Ipilimumab and OTSGC-A24 Therapeutic Peptide Vaccine in Gastric Cancer - a Combination Immunotherapy Phase Ib Study.; Clinical trial registration NCT03784040; clinicaltrials.gov, 2019. https://clinicaltrials.gov/study/NCT03784040 (Accessed 05 November 2025).There is no corresponding record for this reference.
- 41OncoPep, Inc. A Single-Arm, Open-Label, Multi-Center Phase II Study to Evaluate the Combination of PVX-410 + Pembrolizumab + Chemotherapy for Frontline Therapy of Metastatic, PD-L1+ Triple-Negative Breast Cancer (TNBC) in HLA-A2-Positive Patients; Clinical trial registration NCT04634747; clinicaltrials.gov, 2022. https://clinicaltrials.gov/study/NCT04634747 (Accessed 05 November 2025).There is no corresponding record for this reference.
- 42NuGenerex Immuno-Oncology. A Phase II Clinical Trial of Pembrolizumab in Combination With the AE37 Peptide Vaccine in Patients With Metastatic Triple-Negative Breast Cancer; Clinical trial registration NCT04024800; clinicaltrials.gov, 2021. https://clinicaltrials.gov/study/NCT04024800 (Accessed 05 November 2025).There is no corresponding record for this reference.
- 43Abd-Aziz, N.; Poh, C. L. Development of Peptide-Based Vaccines for Cancer. J. Oncol. 2022, 2022 (1), 9749363, DOI: 10.1155/2022/9749363There is no corresponding record for this reference.
- 44Wang, L.; Wang, N.; Zhang, W.; Cheng, X.; Yan, Z.; Shao, G.; Wang, X.; Wang, R.; Fu, C. Therapeutic peptides: current applications and future directions. Signal Transduction Targeted Ther. 2022, 7, 48, DOI: 10.1038/s41392-022-00904-4There is no corresponding record for this reference.
- 45Chen, Y.; Clay, N.; Phan, N.; Lothrop, E.; Culkins, C.; Robinson, B.; Stubblefield, A.; Ferguson, A.; Kimmel, B. R. Molecular Matchmakers: Bioconjugation Techniques Enhance Prodrug Potency for Immunotherapy. Mol. Pharmaceutics 2025, 22 (1), 58– 80, DOI: 10.1021/acs.molpharmaceut.4c00867There is no corresponding record for this reference.
- 46Ott, P.A.; Hu, Z.; Keskin, D.B.; Shukla, S.A.; Sun, J.; Bozym, D.J.; Zhang, W.; Luoma, A.; Giobbie-Hurder, A.; Peter, L. An immunogenic personal neoantigen vaccine for patients with melanoma\. Nature 2017, 547, 217– 221, DOI: 10.1038/nature22991There is no corresponding record for this reference.
- 47Sam-Yellowe, T. Y. Antigens. In Immunology: Overview and Laboratory Manual; Springer International Publishing: Cham, 2021; pp. 65– 72. DOI: DOI: 10.1007/978-3-030-64686-8_9 .There is no corresponding record for this reference.
- 48Kang, R.; Loux, T.; Tang, D.; Schapiro, N. E.; Vernon, P.; Livesey, K. M.; Krasinskas, A.; Lotze, M. T.; Zeh, H. J. The Expression of the Receptor for Advanced Glycation Endproducts (RAGE) Is Permissive for Early Pancreatic Neoplasia. Proc. Natl. Acad. Sci. U. S. A. 2012, 109 (18), 7031– 7036, DOI: 10.1073/pnas.1113865109There is no corresponding record for this reference.
- 49Chen, M.-C.; Chen, K.-C.; Chang, G.-C.; Lin, H.; Wu, C.-C.; Kao, W.-H.; Teng, C.-L. J.; Hsu, S.-L.; Yang, T.-Y. RAGE Acts as an Oncogenic Role and Promotes the Metastasis of Human Lung Cancer. Cell Death Dis. 2020, 11 (4), 265, DOI: 10.1038/s41419-020-2432-1There is no corresponding record for this reference.
- 50Leão, R.; Apolónio, J. D.; Lee, D.; Figueiredo, A.; Tabori, U.; Castelo-Branco, P. Mechanisms of Human Telomerase Reverse Transcriptase (hTERT) Regulation: Clinical Impacts in Cancer. J. Biomed. Sci. 2018, 25 (1), 22, DOI: 10.1186/s12929-018-0422-8There is no corresponding record for this reference.
- 51Sharma, S.; Chowdhury, S. Emerging Mechanisms of Telomerase Reactivation in Cancer. Trends Cancer 2022, 8 (8), 632– 641, DOI: 10.1016/j.trecan.2022.03.005There is no corresponding record for this reference.
- 52Cheng, X. A Comprehensive Review of HER2 in Cancer Biology and Therapeutics. Genes 2024, 15 (7), 903, DOI: 10.3390/genes15070903There is no corresponding record for this reference.
- 53NCI Antibody and Immunotoxin Treatments for Mesothelin-expressing Cancers; NCI, https://techtransfer.cancer.gov/available-technologies?abstract=TAB-4422. (Accessed 04 November 2025).There is no corresponding record for this reference.
- 54Weidemann, S.; Gagelmann, P.; Gorbokon, N.; Lennartz, M.; Menz, A.; Luebke, A. M.; Kluth, M.; Hube-Magg, C.; Blessin, N. C.; Fraune, C.; Möller, K.; Bernreuther, C.; Lebok, P.; Clauditz, T. S.; Jacobsen, F.; Izbicki, J. R.; Jansen, K.; Sauter, G.; Uhlig, R.; Wilczak, W.; Steurer, S.; Minner, S.; Burandt, E.; Krech, R. H.; Dum, D.; Krech, T.; Marx, A. H.; Simon, R. Mesothelin Expression in Human Tumors: A Tissue Microarray Study on 12,679 Tumors. Biomedicines 2021, 9 (4), 397, DOI: 10.3390/biomedicines9040397There is no corresponding record for this reference.
- 55Vargas, A. J.; Sittadjody, S.; Thangasamy, T.; Mendoza, E. E.; Limesand, K. H.; Burd, R. Exploiting Tyrosinase Expression and Activity in Melanocytic Tumors: Quercetin and the Central Role of p53. Integr. Cancer Ther 2011, 10 (4), 328– 340, DOI: 10.1177/1534735410391661There is no corresponding record for this reference.
- 56Vargas, A. J.; Sittadjody, S.; Thangasamy, T.; Mendoza, E. E.; Limesand, K. H.; Burd, R. Exploiting Tyrosinase Expression and Activity in Melanocytic Tumors: Quercetin and the Central Role of P53. Integr. Cancer Ther. 2011, 10 (4), 328– 340, DOI: 10.1177/1534735410391661There is no corresponding record for this reference.
- 57Mann, J. E.; Hasson, N.; Su, D. G.; Adeniran, A. J.; Smalley, K. S. M.; Djureinovic, D.; Jilaveanu, L. B.; Schoenfeld, D. A.; Kluger, H. M. GP100 Expression Is Variable in Intensity in Melanoma. Cancer Immunol. Immunother. 2024, 73 (10), 191, DOI: 10.1007/s00262-024-03776-5There is no corresponding record for this reference.
- 58Wang, J.; Jia, R.; Yao, Y.; Cun, B.; Huang, X.; Gu, P.; Ge, S.; Fan, X. Differential Expression of Mart-1 in Human Uveal Melanoma Cells. Mol. Med. Rep. 2011, 4 (5), 799– 803, DOI: 10.3892/mmr.2011.504There is no corresponding record for this reference.
- 59Mayo Clinic Types of prostate cancer: Common and rare forms. Diseases and Conditions; Mayo Clinic. https://www.mayoclinic.org/diseases-conditions/prostate-cancer/in-depth/prostate-cancer-types/art-20584938. (Accessed 05 November 2025).There is no corresponding record for this reference.
- 60Shair, K. H. Y.; Bendt, K. M.; Edwards, R. H.; Nielsen, J. N.; Moore, D. T.; Raab-Traub, N. Epstein-Barr Virus-Encoded Latent Membrane Protein 1 (LMP1) and LMP2A Function Cooperatively To Promote Carcinoma Development in a Mouse Carcinogenesis Model. J. Virol. 2012, 86 (9), 5352– 5365, DOI: 10.1128/JVI.07035-11There is no corresponding record for this reference.
- 61Pal, A.; Kundu, R. Human Papillomavirus E6 and E7: The Cervical Cancer Hallmarks and Targets for Therapy. Front Microbiol. 2020, 10, 3116, DOI: 10.3389/fmicb.2019.03116There is no corresponding record for this reference.
- 62Baba, S. K.; Alblooshi, S. S. E.; Yaqoob, R.; Behl, S.; Al Saleem, M.; Rakha, E. A.; Malik, F.; Singh, M.; Macha, M. A.; Akhtar, M. K.; Houry, W. A.; Bhat, A. A.; Al Menhali, A.; Zheng, Z.-M.; Mirza, S. Human Papilloma Virus (HPV) Mediated Cancers: An Insightful Update. J. Transl. Med. 2025, 23 (1), 483, DOI: 10.1186/s12967-025-06470-xThere is no corresponding record for this reference.
- 63Valcarcel, B.; Enriquez-Vera, D.; De-la-Cruz-Ku, G.; Chambergo-Michilot, D.; Calderón-Huaycochea, H.; Malpica, L. Epidemiological Features and Outcomes of HTLV-1 Carriers Diagnosed With Cancer: A Retrospective Cohort Study in an Endemic Country. JCO Glob. Oncol. 2023, 9, e2200369 DOI: 10.1200/GO.22.00369There is no corresponding record for this reference.
- 64Du, G.; Zhang, W.; Zhang, Z.; Zeng, M.; Wang, Y. HTLV-1-Associated Genes as Potential Biomarkers for Endometrial Cancer. Oncol Lett. 2019, 18 (1), 699– 705, DOI: 10.3892/ol.2019.10389There is no corresponding record for this reference.
- 65Yang, Y.; Zhang, H.; Huang, S.; Chu, Q. KRAS Mutations in Solid Tumors: Characteristics, Current Therapeutic Strategy, and Potential Treatment Exploration. J. Clin. Med. 2023, 12 (2), 709, DOI: 10.3390/jcm12020709There is no corresponding record for this reference.
- 66Johnson, D. B.; Puzanov, I. Treatment of NRAS-Mutant Melanoma. Curr. Treat Options Oncol. 2015, 16 (4), 15, DOI: 10.1007/s11864-015-0330-zThere is no corresponding record for this reference.
- 67Pencheva, B.; Di Paola, J.; Porter, C. C.Adam, M.; Feldman, J.; Mirzaa, G. M.; Pagon, R. A.; Wallace, S. E.; Amemiya, A. ETV6-Related Thrombocytopenia and Predisposition to Leukemia. In GeneReviews; University of Washington: Seattle: Seattle (WA, 1993.There is no corresponding record for this reference.
- 68Biswas, A.; Rajesh, Y.; Mitra, P.; Mandal, M. ETV6 Gene Aberrations in Non-Haematological Malignancies: A Review Highlighting ETV6 Associated Fusion Genes in Solid Tumors. Biochim. Biophys. Acta BBA - Rev. Cancer 2020, 1874 (1), 188389, DOI: 10.1016/j.bbcan.2020.188389There is no corresponding record for this reference.
- 69Andraos, E.; Dignac, J.; Meggetto, F. NPM-ALK: A Driver of Lymphoma Pathogenesis and a Therapeutic Target. Cancers 2021, 13 (1), 144, DOI: 10.3390/cancers13010144There is no corresponding record for this reference.
- 70Buonaguro, L.; Tagliamonte, M. Selecting Target Antigens for Cancer Vaccine Development. Vaccines 2020, 8 (4), 615, DOI: 10.3390/vaccines8040615There is no corresponding record for this reference.
- 71Xie, N.; Shen, G.; Gao, W.; Huang, Z.; Huang, C.; Fu, L. Neoantigens: promising targets for cancer therapy. Signal Transduction Targeted Ther. 2023, 8, 9, DOI: 10.1038/s41392-022-01270-xThere is no corresponding record for this reference.
- 72Verdon, D. J.; Jenkins, M. R. Identification and Targeting of Mutant Peptide Neoantigens in Cancer Immunotherapy. Cancers 2021, 13 (16), 4245, DOI: 10.3390/cancers13164245There is no corresponding record for this reference.
- 73Chumakov, P.M. Function of the p53 Gene: Choice between Life and Death. Biochemistry 2000, 65, 34– 47There is no corresponding record for this reference.
- 74Parikh, N.; Hilsenbeck, S.; Creighton, C.J.; Dayaram, T.; Shuck, R.; Shinbrot, E.; Xi, L.; Gibbs, R.A.; Wheeler, D.A.; Donehower, L.A. Effects of TP53 mutational status on gene expression patterns across 10 human cancer types - Parikh - 2014 - The Journal of Pathology - Wiley Online Library. cancer types. J. Pathol. 2014, 232, 522– 533, DOI: 10.1002/path.4321There is no corresponding record for this reference.
- 75Chong, C.; Coukos, G.; Bassani-Sternberg, M. Identification of Tumor Antigens with Immunopeptidomics. Nat. Biotechnol. 2022, 40 (2), 175– 188, DOI: 10.1038/s41587-021-01038-8There is no corresponding record for this reference.
- 76Tejaswi, L.; Ramesh, P.; Aditya, S.; Raju, R.; Prasad, T. S. K. Computational Neoantigen Prediction for Cancer Immunotherapy. Genes Immun. 2025, 26 (6), 531– 538, DOI: 10.1038/s41435-025-00365-zThere is no corresponding record for this reference.
- 77Manakongtreecheep, K.; Ctortecka, C.; Correa-Medero, L.O.; Zhu, T.; Lippincott, I.; Lawrence, G.M.; Howard, A.; Hernandez, G.M.; Forman, C.; Duggan, E.C. Sensitive detection of cancer antigens enabled by user-defined peptide libraries. Nature Biotechnol. 2026, 1– 13, DOI: 10.1038/s41587-026-03003-9There is no corresponding record for this reference.
- 78Science Direct Topics Ligandome - an overview; Science Direct Topics, 2016.There is no corresponding record for this reference.
- 79Verdon, D.J.; Jenkins, M.R. Identification and Targeting of Mutant Peptide Neoantigens in Cancer Immunotherapy. Cancers 2021, 13 (16), 4245, DOI: 10.3390/cancers13164245There is no corresponding record for this reference.
- 80Shao, Y.; Ge, S.; Dong, R.; Ji, W.; Qin, C.; Wen, P. NeoTImmuML: A Machine Learning-Based Prediction Model for Human Tumor Neoantigen Immunogenicity. Front. Immunol. 2025, 16, 1681396, DOI: 10.3389/fimmu.2025.1681396There is no corresponding record for this reference.
- 81Adams, C.; Gabriel, W.; Laukens, K.; Picciani, M.; Wilhelm, M.; Bittremieux, W.; Boonen, K. Fragment Ion Intensity Prediction Improves the Identification Rate of Non-Tryptic Peptides in timsTOF. Nat. Commun. 2024, 15 (1), 3956, DOI: 10.1038/s41467-024-48322-0There is no corresponding record for this reference.
- 82Strauss, M. T.; Bludau, I.; Zeng, W.-F.; Voytik, E.; Ammar, C.; Schessner, J. P.; Ilango, R.; Gill, M.; Meier, F.; Willems, S.; Mann, M. AlphaPept: A Modern and Open Framework for MS-Based Proteomics. Nat. Commun. 2024, 15 (1), 2168, DOI: 10.1038/s41467-024-46485-4There is no corresponding record for this reference.
- 83Wang, X.; Wu, T.; Jiang, Y.; Chen, T.; Pan, D.; Jin, Z.; Xie, J.; Quan, L.; Lyu, Q. RPEMHC: Improved Prediction of MHC–Peptide Binding Affinity by a Deep Learning Approach Based on Residue–Residue Pair Encoding. Bioinformatics 2024, 40 (1), btad785, DOI: 10.1093/bioinformatics/btad785There is no corresponding record for this reference.
- 84Li, G.; Mahajan, S.; Ma, S.; Jeffery, E. D.; Zhang, X.; Bhattacharjee, A.; Venkatasubramanian, M.; Weirauch, M. T.; Miraldi, E. R.; Grimes, H. L.; Sheynkman, G. M.; Tilburgs, T.; Salomonis, N. Splicing Neoantigen Discovery with SNAF Reveals Shared Targets for Cancer Immunotherapy. Sci. Transl. Med. 2024, 16 (730), eade2886 DOI: 10.1126/scitranslmed.ade2886There is no corresponding record for this reference.
- 85Declercq, A.; Devreese, R.; Scheid, J.; Jachmann, C.; Van Den Bossche, T.; Preikschat, A.; Gomez-Zepeda, D.; Rijal, J. B.; Hirschler, A.; Krieger, J. R.; Srikumar, T.; Rosenberger, G.; Martelli, C.; Trede, D.; Carapito, C.; Tenzer, S.; Walz, J. S.; Degroeve, S.; Bouwmeester, R.; Martens, L.; Gabriels, R. TIMS2Rescore: A Data Dependent Acquisition-Parallel Accumulation and Serial Fragmentation-Optimized Data-Driven Rescoring Pipeline Based on MS2Rescore. J. Proteome Res. 2025, 24 (3), 1067– 1076, DOI: 10.1021/acs.jproteome.4c00609There is no corresponding record for this reference.
- 86Zhao, Y.; Yu, J.; Su, Y.; Shu, Y.; Ma, E.; Wang, J.; Jiang, S.; Wei, C.; Li, D.; Huang, Z.; Cheng, G.; Ren, H.; Feng, J. A Unified Deep Framework for Peptide–Major Histocompatibility Complex–T Cell Receptor Binding Prediction. Nat. Mach. Intell. 2025, 7 (4), 650– 660, DOI: 10.1038/s42256-025-01002-0There is no corresponding record for this reference.
- 87Li, J.; Zhao, Z.; Tai, C.; Sun, T.; Tan, L.; Li, X.; He, W.; Li, H.; Zhang, J. VirusImmu: A Novel Ensemble Machine Learning Approach for Viral Immunogenicity Prediction. Brief Funct. Genomics 2025, 24, elaf008 DOI: 10.1093/bfgp/elaf008There is no corresponding record for this reference.
- 88Cai, Y.; Chen, R.; Song, M.; Wang, L.; Huo, Z.; Yang, D.; Zhang, S.; Gao, S.; Hwang, S.; Bai, L.; Lv, Y.; Cui, Y.; Zhang, X. CNNeoPP: A Large Language Model-Enhanced Deep Learning Pipeline for Personalized Neoantigen Prediction and Liquid Biopsy Applications. Front. Immunol. 2026, 17, 1722117, DOI: 10.3389/fimmu.2026.1722117There is no corresponding record for this reference.
- 89Lee, K.-H.; Sears, T. J.; Zanetti, M.; Carter, H. NeoPrecis: Enhancing Immunotherapy Response Prediction through Integration of Qualified Immunogenicity and Clonality-Aware Neoantigen Landscapes. Nat. Commun. 2026, 17 (1), 1966, DOI: 10.1038/s41467-026-68651-6There is no corresponding record for this reference.
- 90Slaton, E. W.; Clay, N.; Phan, N.; Kimmel, B. R. Innovations in Yeast Synthetic Biology: Engineered Discovery Systems for Immunotherapy. ACS Synth. Biol. 2025, 14 (9), 3293– 3305, DOI: 10.1021/acssynbio.5c00321There is no corresponding record for this reference.
- 91Calis, J. J. A.; Maybeno, M.; Greenbaum, J. A.; Weiskopf, D.; De Silva, A. D.; Sette, A.; Keşmir, C.; Peters, B. Properties of MHC Class I Presented Peptides That Enhance Immunogenicity. PLoS Comput. Biol. 2013, 9 (10), e1003266 DOI: 10.1371/journal.pcbi.1003266There is no corresponding record for this reference.
- 92Capietto, A.H.; Jhunjhunwala, S.; Pollock, S.B.; Lupardus, P.; Wong, J.; Hänsch, L.; Cevallos, J.; Chestnut, Y.; Fernandez, A.; Lounsbury, N. Mutation position is an important determinant for predicting cancer neoantigens. J Exp Med. 2020, 217 (4), e20190179 DOI: 10.1084/jem.20190179There is no corresponding record for this reference.
- 93Apostolopoulos, V.; Bojarska, J.; Chai, T.-T.; Elnagdy, S.; Kaczmarek, K.; Matsoukas, J.; New, R.; Parang, K.; Lopez, O. P.; Parhiz, H.; Perera, C. O.; Pickholz, M.; Remko, M.; Saviano, M.; Skwarczynski, M.; Tang, Y.; Wolf, W. M.; Yoshiya, T.; Zabrocki, J.; Zielenkiewicz, P.; AlKhazindar, M.; Barriga, V.; Kelaidonis, K.; Sarasia, E. M.; Toth, I. A Global Review on Short Peptides: Frontiers and Perspectives. Molecules 2021, 26 (2), 430, DOI: 10.3390/molecules26020430There is no corresponding record for this reference.
- 94Li, W.; Joshi, M. D.; Singhania, S.; Ramsey, K. H.; Murthy, A. K. Peptide Vaccine: Progress and Challenges. Vaccines 2014, 2 (3), 515– 536, DOI: 10.3390/vaccines2030515There is no corresponding record for this reference.
- 95Onodi, F.; Maherzi-Mechalikh, C.; Mougel, A.; Ben Hamouda, N.; Taboas, C.; Gueugnon, F.; Tran, T.; Nozach, H.; Marcon, E.; Gey, A. High Therapeutic Efficacy of a New Survivin LSP-Cancer Vaccine Containing CD4+ and CD8+ T-Cell Epitopes. Front. Oncol 2018, 8, 517, DOI: 10.3389/fonc.2018.00517There is no corresponding record for this reference.
- 96Cai, L.; Zhang, J.; Zhu, R.; Shi, W.; Xia, X.; Edwards, M.; Finch, W.; Coombs, A.; Gao, J.; Chen, K.; Owen, S.; Jiang, S.; Lu, W. Protective Cellular Immunity Generated by Cross-Presenting Recombinant Overlapping Peptide Proteins. Oncotarget 2017, 8 (44), 76516– 76524, DOI: 10.18632/oncotarget.20407There is no corresponding record for this reference.
- 97McDowell, S. H.; Gallaher, S. A.; Burden, R. E.; Scott, C. J. Leading the Invasion: The Role of Cathepsin S in the Tumour Microenvironment. Biochim. Biophys. Acta BBA - Mol. Cell Res. 2020, 1867 (10), 118781, DOI: 10.1016/j.bbamcr.2020.118781There is no corresponding record for this reference.
- 98Zhang, H.; Hong, H.; Li, D.; Ma, S.; Di, Y.; Stoten, A.; Haig, N.; Gleria, K. D.; Yu, Z.; Xu, X.-N. Comparing Pooled Peptides with Intact Protein for Accessing Cross-Presentation Pathways for Protective CD8+ and CD4+ T Cells *. J. Biol. Chem. 2009, 284 (14), 9184– 9191, DOI: 10.1074/jbc.M809456200There is no corresponding record for this reference.
- 99Wang, Z.; Yang, X.; Wei, X.; Shi, L.; Wang, X.; Zhuo, Z.; Su, H.; Wu, W.; Cao, Y. J. Designing Multifunctional Recombinant Vaccines: An Engineering Strategy Based on Innovative Epitope Prediction-Guided Splicing. Theranostics 2025, 15 (9), 3924– 3942, DOI: 10.7150/thno.103755There is no corresponding record for this reference.
- 100Oxford Vacmedix UK Ltd A Phase 1, Multicentre, Open-Label, Nonrandomised, First-in-Human Study of OVM-200 as a Therapeutic Vaccine in Patients With Locally Advanced or Metastatic Non-Small Cell Lung Cancer, Ovarian Cancer, and Prostate Cancer; Clinical trial registration NCT05104515; clinicaltrials.gov, 2025. https://clinicaltrials.gov/study/NCT05104515. (Accessed 17 February 2026).There is no corresponding record for this reference.
- 101Kordalivand, N.; Tondini, E.; Lau, C. Y. J.; Vermonden, T.; Mastrobattista, E.; Hennink, W. E.; Ossendorp, F.; Nostrum, C. F. Cationic Synthetic Long Peptides-Loaded Nanogels: An Efficient Therapeutic Vaccine Formulation for Induction of T-Cell Responses. J. Controlled Release 2019, 315, 114– 125, DOI: 10.1016/j.jconrel.2019.10.048There is no corresponding record for this reference.
- 102Wei, P.; Gangapurwala, G.; Pretzel, D.; Leiske, M.N.; Wang, L.; Hoeppener, S.; Schubert, S.; Brendel, J.C.; Schubert, U.S. Smart pH-Sensitive Nanogels for Controlled Release in an Acidic Environment. Biomacromolecules 2019, 20, 130– 140, DOI: 10.1021/acs.biomac.8b01228There is no corresponding record for this reference.
- 103Palmer, C.R.; Pastora, L.E.; Kimmel, B.R.; Pagendarm, H.M.; Kwiatkowski, A.J.; Stone, P.T.; Arora, K.; Francini, N.; Fedorova, O.; Pyle, A.M. Covalent Polymer-RNA Conjugates for Potent Activation of the RIG-I Pathway. Adv. Healthcare Mater. 2025, 14, 2303815, DOI: 10.1002/adhm.202303815There is no corresponding record for this reference.
- 104Beach, M. A.; Nayanathara, U.; Gao, Y.; Zhang, C.; Xiong, Y.; Wang, Y.; Such, G. K. Polymeric Nanoparticles for Drug Delivery. Chem. Rev. 2024, 124 (9), 5505– 5616, DOI: 10.1021/acs.chemrev.3c00705There is no corresponding record for this reference.
- 105Haim Zada, M.; Rottenberg, Y.; Domb, A. J. Peptide Loaded Polymeric Nanoparticles by Non-Aqueous Nanoprecipitation. J. Colloid Interface Sci. 2022, 622, 904– 913, DOI: 10.1016/j.jcis.2022.05.007There is no corresponding record for this reference.
- 106Mehta, M.; Bui, T. A.; Yang, X.; Aksoy, Y.; Goldys, E. M.; Deng, W. Lipid-Based Nanoparticles for Drug/Gene Delivery: An Overview of the Production Techniques and Difficulties Encountered in Their Industrial Development. ACS Mater. Au 2023, 3 (6), 600– 619, DOI: 10.1021/acsmaterialsau.3c00032There is no corresponding record for this reference.
- 107Xiao, W.; Jiang, W.; Chen, Z.; Huang, Y.; Mao, J.; Zheng, W.; Hu, Y.; Shi, J. Advance in Peptide-Based Drug Development: Delivery Platforms, Therapeutics and Vaccines. Signal Transduction Targeted Ther. 2025, 10 (1), 74, DOI: 10.1038/s41392-024-02107-5There is no corresponding record for this reference.
- 108Pagendarm, H. M.; Stone, P. T.; Kimmel, B. R.; Baljon, J. J.; Aziz, M. H.; Pastora, L. E.; Hubert, L.; Roth, E. W.; Almunif, S.; Scott, E. A.; Wilson, J. T. Engineering Endosomolytic Nanocarriers of Diverse Morphologies Using Confined Impingement Jet Mixing. Nanoscale 2023, 15, 16016– 16029, DOI: 10.1039/D3NR02874GThere is no corresponding record for this reference.
- 109Han, S.; Ma, W.; Jiang, D.; Sutherlin, L.; Zhang, J.; Lu, Y.; Huo, N.; Chen, Z.; Engle, J.W.; Wang, Y. Intracellular signaling pathway in dendritic cells and antigen transport pathway in vivo mediated by an OVA@DDAB/PLGA nano-vaccine. J Nanobiotechnol. 2021, 19, 394, DOI: 10.1186/s12951-021-01116-8There is no corresponding record for this reference.
- 110Varypataki, E. M.; Silva, A. L.; Barnier-Quer, C.; Collin, N.; Ossendorp, F.; Jiskoot, W. Synthetic long peptide-based vaccine formulations for induction of cell mediated immunity: A comparative study of cationic liposomes and PLGA nanoparticles. J. Controlled Release 2016, 226, 98– 106, DOI: 10.1016/j.jconrel.2016.02.018There is no corresponding record for this reference.
- 111Sheehy, T. L.; Kwiatkowski, A. J.; Arora, K.; Kimmel, B. R.; Schulman, J. A.; Gibson-Corley, K. N.; Wilson, J. T. STING-Activating Polymer–Drug Conjugates for Cancer Immunotherapy. ACS Cent. Sci. 2024, 10 (9), 1765– 1781, DOI: 10.1021/acscentsci.4c00579There is no corresponding record for this reference.
- 112Arora, K.; Sheehy, T. L.; Schulman, J. A.; Loken, J. R.; Lehmann, Z.; Kimmel, B. R.; McAtee, C.; Bharti, V.; Stone, P. T.; Weaver, A. M.; Tyska, M.; Pathak, R. K.; Wilson, J. T. Macromolecular Diamidobenzimidazole Conjugates Can Activate Stimulator of Interferon Genes. J. Am. Chem. Soc. 2025, 147 (38), 35149– 35163, DOI: 10.1021/jacs.5c13195There is no corresponding record for this reference.
- 113Kimmel, B. R.; Arora, K.; Chada, N. C.; Bharti, V.; Kwiatkowski, A. J.; Finkelstein, J. E.; Hanna, A.; Arner, E. N.; Sheehy, T. L.; Pastora, L. E.; Yang, J.; Pagendarm, H. M.; Stone, P. T.; Hargrove-Wiley, E.; Taylor, B. C.; Hubert, L. A.; Fingleton, B. M.; Gibson-Corley, K. N.; May, J. C.; McLean, J. A.; Rathmell, J. C.; Richmond, A.; Rathmell, W. K.; Balko, J. M.; Wilson, J. T. Potentiating Cancer Immunotherapies with Modular Albumin-Hitchhiking Nanobody–STING Agonist Conjugates. Nat. Biomed. Eng. 2025, 9 (10), 1719– 1739, DOI: 10.1038/s41551-025-01400-0There is no corresponding record for this reference.
- 114Lynn, G.M.; Sedlik, C.; Baharom, F.; Zhu, Y.; Ramirez-Valdez, R.A.; Coble, V.L.; Tobin, K.; Nichols, S.R.; Itzkowitz, Y.; Zaidi, N. Peptide–TLR-7/8a conjugate vaccines chemically programmed for nanoparticle self-assembly enhance CD8 T-cell immunity to tumor antigens. Nat Biotechnol. 2020, 38, 320– 332, DOI: 10.1038/s41587-019-0390-xThere is no corresponding record for this reference.
- 115Moon, J. H.; Phoo, M. T.; Kim, Y.; Ahn, J.; Kim, A.; Nam, J.; Lee, S.-H.; Moon, J. J.; Son, S. Polysaccharide Adjuvants as Innate Immune Trainers: Bridging Pattern Recognition Receptor (PRR) Activation and Metabolic Reprogramming for Synthetic Vaccine Design. Adv. Sci. 2025, 12 (48), e09022 DOI: 10.1002/advs.202509022There is no corresponding record for this reference.
- 116Spunde, K.; Korotkaja, K.; Sominskaya, I.; Zajakina, A. Genetic Adjuvants: A Paradigm Shift in Vaccine Development and Immune Modulation. Mol. Ther Nucleic Acids 2025, 36 (2), 102536, DOI: 10.1016/j.omtn.2025.102536There is no corresponding record for this reference.
- 117Wang-Bishop, L.; Wehbe, M.; Pastora, L. E.; Yang, J.; Kimmel, B. R.; Garland, K. M.; Becker, K. W.; Carson, C. S.; Roth, E. W.; Gibson-Corley, K. N.; Ulkoski, D.; Krishnamurthy, V.; Fedorova, O.; Richmond, A.; Pyle, A. M.; Wilson, J. T. Nanoparticle Retinoic Acid-Inducible Gene I Agonist for Cancer Immunotherapy. ACS Nano 2024, 18 (18), 11631– 11643, DOI: 10.1021/acsnano.3c06225There is no corresponding record for this reference.
- 118Wang-Bishop, L.; Kimmel, B. R.; Ngwa, V. M.; Madden, M. Z.; Baljon, J. J.; Florian, D. C.; Hanna, A.; Pastora, L. E.; Sheehy, T. L.; Kwiatkowski, A. J.; Wehbe, M.; Wen, X.; Becker, K. W.; Garland, K. M.; Schulman, J. A.; Shae, D.; Edwards, D.; Wolf, M. M.; Delapp, R.; Christov, P. P.; Beckermann, K. E.; Balko, J. M.; Rathmell, W. K.; Rathmell, J. C.; Chen, J.; Wilson, J. T. STING-Activating Nanoparticles Normalize the Vascular-Immune Interface to Potentiate Cancer Immunotherapy. Sci. Immunol. 2023, 8 (83), eadd1153 DOI: 10.1126/sciimmunol.add1153There is no corresponding record for this reference.
- 119Baljon, J. J.; Kwiatkowski, A. J.; Pagendarm, H. M.; Stone, P. T.; Kumar, A.; Bharti, V.; Schulman, J. A.; Becker, K. W.; Roth, E. W.; Christov, P. P.; Joyce, S.; Wilson, J. T. A Cancer Nanovaccine for Co-Delivery of Peptide Neoantigens and Optimized Combinations of STING and TLR4 Agonists. ACS Nano 2024, 18 (9), 6845– 6862, DOI: 10.1021/acsnano.3c04471There is no corresponding record for this reference.
- 120Xi, Y.; Chen, L.; Tang, J.; Yu, B.; Shen, W.; Niu, X. Amplifying “Eat Me Signal” by Immunogenic Cell Death for Potentiating Cancer Immunotherapy. Immunol Rev. 2024, 321 (1), 94– 114, DOI: 10.1111/imr.13251There is no corresponding record for this reference.
- 121Van Lint, S.; Renmans, D.; Broos, K.; Goethals, L.; Maenhout, S.; Benteyn, D.; Goyvaerts, C.; Du Four, S.; Van der Jeught, K.; Bialkowski, L. Intratumoral Delivery of TriMix mRNA Results in T-cell Activation by Cross-Presenting Dendritic Cells. Cancer Immunol Res. 2016, 4, 146– 156, DOI: 10.1158/2326-6066.CIR-15-0163There is no corresponding record for this reference.
- 122Wijfjes, Z.; Tomillero, I. R.; Gall, C. M. L.; van Dinther, E. A. W.; Turlings, F.; Classens, R.; Manna, S.; van Dalen, D.; Peters, W. R. J. R.; Schouren, K. Co-Delivery of Antigen and Adjuvant by Site-Specific Conjugation to Dendritic Cell-Targeted Fab Fragments Potentiates T Cell Responses. RSC Chem. Biol. 2025, 6 (6), 948– 962, DOI: 10.1039/D5CB00014AThere is no corresponding record for this reference.
- 123Botelho, N. K.; Tschumi, B. O.; Hubbell, J. A.; Swartz, M. A.; Donda, A.; Romero, P. Combination of Synthetic Long Peptides and XCL1 Fusion Proteins Results in Superior Tumor Control. Front. Immunol 2019, 10, 294, DOI: 10.3389/fimmu.2019.00294There is no corresponding record for this reference.
- 124Carreno, B. M.; Magrini, V.; Becker-Hapak, M.; Kaabinejadian, S.; Hundal, J.; Petti, A. A.; Ly, A.; Lie, W.-R.; Hildebrand, W. H.; Mardis, E. R.; Linette, G. P. A Dendritic Cell Vaccine Increases the Breadth and Diversity of Melanoma Neoantigen-Specific T Cells. Science 2015, 348 (6236), 803– 808, DOI: 10.1126/science.aaa3828There is no corresponding record for this reference.
- 125Belnoue, E.; Mayol, J.-F.; Carboni, S.; Besson, W. D. B.; Dupuychaffray, E.; Nelde, A.; Stevanovic, S.; Santiago-Raber, M.-L.; Walker, P. R.; Derouazi, M. Targeting Self- and Neoepitopes with a Modular Self-Adjuvanting Cancer Vaccine. JCI Insight 2019, 4 (11), 127305, DOI: 10.1172/jci.insight.127305There is no corresponding record for this reference.
- 126Bajiya, N.; Najrin, S.; Kumar, P.; Choudhury, S.; Tomer, R.; Raghava, G. P. S. CPPsite3: An Updated Large Repository of Experimentally Validated Cell-Penetrating Peptides. Drug Discovery Today 2025, 30 (8), 104421, DOI: 10.1016/j.drudis.2025.104421There is no corresponding record for this reference.
- 127Gall, V.A.; Philips, A.V.; Qiao, N.A.; Clise-Dwyer, K.; Perakis, A.A.; Zhang, M.; Clifton, G.T.; Sukhumalchandra, P.; Ma, Q.; Reddy, S.M. Trastuzumab Increases HER2 Uptake and Cross-Presentation by Dendritic Cells. Cancer Res 2017, 77 (19), 5374– 5383, DOI: 10.1158/0008-5472.CAN-16-2774There is no corresponding record for this reference.
- 128Gall, V.A.; Philips, A.V.; Qiao, N.A.; Clise-Dwyer, K.; Perakis, A.A.; Zhang, M.; Clifton, G.T.; Sukhumalchandra, P.; Ma, Q.; Reddy, S.M.; An open-label, multinational, multicenter study of G17DT vaccination combined with cisplatin and 5-fluorouracil in patients with untreated, advanced gastric or gastroesophageal cancer: The GC4 study - Ajani - 2006 - Cancer - Wiley Online Library. https://acsjournals.onlinelibrary.wiley.com/doi/full/10.1002/cncr.21814 (accessed 2025–10–17).There is no corresponding record for this reference.
- 129Li, Q.; Lei, X.; Zhu, J.; Zhong, Y.; Yang, J.; Wang, J.; Tan, H. Radiotherapy/Chemotherapy-Immunotherapy for Cancer Management: From Mechanisms to Clinical Implications - Li - 2023 - Oxidative Medicine and Cellular Longevity - Wiley Online Library. Oxid. Med. Cell. Longevity 2023, 2023, 7530794, DOI: 10.1155/2023/7530794There is no corresponding record for this reference.
- 130Lee, H.-M.; Lee, H.-J.; Chang, J.-E. Inflammatory Cytokine: An Attractive Target for Cancer Treatment. Biomedicines 2022, 10 (9), 2116, DOI: 10.3390/biomedicines10092116There is no corresponding record for this reference.
- 131Tufail, M.; Jiang, C.-H.; Li, N. Immune Evasion in Cancer: Mechanisms and Cutting-Edge Therapeutic Approaches. Signal Transduction Targeted Ther. 2025, 10 (1), 227, DOI: 10.1038/s41392-025-02280-1There is no corresponding record for this reference.
- 132Dagogo-Jack, I.; Shaw, A. T. Tumour Heterogeneity and Resistance to Cancer Therapies. Nat. Rev. Clin. Oncol. 2018, 15 (2), 81– 94, DOI: 10.1038/nrclinonc.2017.166There is no corresponding record for this reference.
- 133Jing, H.; Gao, Y.; Sun, Z.; Liu, S. Recent Advances in Novel Tumor Immunotherapy Strategies Based on Regulating the Tumor Microenvironment and Immune Checkpoints. Front. Immunol. 2025, 16, 1529403, DOI: 10.3389/fimmu.2025.1529403There is no corresponding record for this reference.
- 134Shah, D. D.; Chorawala, M. R.; Raghani, N. R.; Patel, R.; Fareed, M.; Kashid, V. A.; Prajapati, B. G. Tumor Microenvironment: Recent Advances in Understanding and Its Role in Modulating Cancer Therapies. Med. Oncol. 2025, 42 (4), 117, DOI: 10.1007/s12032-025-02641-4There is no corresponding record for this reference.
- 135Emami Nejad, A.; Najafgholian, S.; Rostami, A.; Sistani, A.; Shojaeifar, S.; Esparvarinha, M.; Nedaeinia, R.; Haghjooy Javanmard, S.; Taherian, M.; Ahmadlou, M.; Salehi, R.; Sadeghi, B.; Manian, M. The Role of Hypoxia in the Tumor Microenvironment and Development of Cancer Stem Cell: A Novel Approach to Developing Treatment. Cancer Cell Int. 2021, 21 (1), 62, DOI: 10.1186/s12935-020-01719-5There is no corresponding record for this reference.
- 136Basak, U.; Sarkar, T.; Mukherjee, S.; Chakraborty, S.; Dutta, A.; Dutta, S.; Nayak, D.; Kaushik, S.; Das, T.; Sa, G. Tumor-Associated Macrophages: An Effective Player of the Tumor Microenvironment. Front. Immunol. 2023, 14, 1295257, DOI: 10.3389/fimmu.2023.1295257There is no corresponding record for this reference.
- 137Yu, Y.; Yao, X.; Wang, Q.; Yang, M.; Li, R.; Qin, J.; Zhuang, J.; Sun, C. T Cell Exhaustion in Cancer Immunotherapy: Heterogeneity, Mechanisms, and Therapeutic Opportunities. Adv. Sci. 2026, n/a (n/a), e20634 DOI: 10.1002/advs.202520634There is no corresponding record for this reference.
- 138Singhaviranon, S.; Dempsey, J. P.; Hagymasi, A. T.; Mandoiu, I. I.; Srivastava, P. K. Low-Avidity T Cells Drive Endogenous Tumor Immunity in Mice and Humans. Nat. Immunol. 2025, 26 (2), 240– 251, DOI: 10.1038/s41590-024-02044-zThere is no corresponding record for this reference.
- 139Tagore, S.; Caprio, L.; Amin, A. D.; Bestak, K.; Luthria, K.; D’Souza, E.; Barrera, I.; Melms, J. C.; Wu, S.; Abuzaid, S.; Wang, Y.; Jakubikova, V.; Koch, P.; Brodtman, D. Z.; Bawa, B.; Deshmukh, S. K.; Ebel, L.; Ibarra-Arellano, M. A.; Jaiswal, A.; Gurjao, C.; Biermann, J.; Shaikh, N.; Ramaradj, P.; Georgis, Y.; Lagos, G. G.; Ehrlich, M. I.; Ho, P.; Walsh, Z. H.; Rogava, M.; Politis, M. G.; Biswas, D.; Cottarelli, A.; Rizvi, N.; Shu, C. A.; Herzberg, B.; Anandasabapathy, N.; Sledge, G.; Zorn, E.; Canoll, P.; Bruce, J. N.; Rizvi, N. A.; Taylor, A. M.; Saqi, A.; Hibshoosh, H.; Schwartz, G. K.; Henick, B. S.; Chen, F.; Schapiro, D.; Shah, P.; Izar, B. Single-Cell and Spatial Genomic Landscape of Non-Small Cell Lung Cancer Brain Metastases. Nat. Med. 2025, 31 (4), 1351– 1363, DOI: 10.1038/s41591-025-03530-zThere is no corresponding record for this reference.
- 140Le, J.; Dian, Y.; Zhao, D.; Guo, Z.; Luo, Z.; Chen, X.; Zeng, F.; Deng, G. Single-Cell Multi-Omics in Cancer Immunotherapy: From Tumor Heterogeneity to Personalized Precision Treatment. Mol. Cancer 2025, 24 (1), 221, DOI: 10.1186/s12943-025-02426-3There is no corresponding record for this reference.
- 141Hamley, I. W. Peptides for Vaccine Development. ACS Appl. Bio Mater. 2022, 5 (3), 905– 944, DOI: 10.1021/acsabm.1c01238There is no corresponding record for this reference.
- 142Culkins, C.; Adomanis, R.; Phan, N.; Robinson, B.; Slaton, E.; Lothrop, E.; Chen, Y.; Kimmel, B.R. Unlocking the Gates: Therapeutic Agents for Noninvasive Drug Delivery Across the Blood-Brain Barrier. Mol. Pharmaceutics 2024, 21, 5430– 5454, DOI: 10.1021/acs.molpharmaceut.4c00604There is no corresponding record for this reference.
- 143Mansurov, A.; Hosseinchi, P.; Chang, K.; Lauterbach, A. L.; Gray, L. T.; Alpar, A. T.; Budina, E.; Slezak, A. J.; Kang, S.; Cao, S.; Solanki, A.; Gomes, S.; Williford, J.-M.; Swartz, M. A.; Mendoza, J. L.; Ishihara, J.; Hubbell, J. A. Masking the Immunotoxicity of Interleukin-12 by Fusing It with a Domain of Its Receptor via a Tumour-Protease-Cleavable Linker. Nat. Biomed. Eng. 2022, 6 (7), 819– 829, DOI: 10.1038/s41551-022-00888-0There is no corresponding record for this reference.
- 144Kimmel, B. R.; Mrksich, M. Development of an Enzyme-Inhibitor Reaction Using Cellular Retinoic Acid Binding Protein II for One-Pot Megamolecule Assembly. Chem. – Eur. J. 2021, 27 (71), 17843– 17848, DOI: 10.1002/chem.202103059There is no corresponding record for this reference.
- 145Metcalf, K. J.; Kimmel, B. R.; Sykora, D. J.; Modica, J. A.; Parker, K. A.; Berens, E.; Dai, R.; Dravid, V. P.; Werb, Z.; Mrksich, M. Synthetic Tuning of Domain Stoichiometry in Nanobody–Enzyme Megamolecules. Bioconjugate Chem. 2021, 32 (1), 143– 152, DOI: 10.1021/acs.bioconjchem.0c00578There is no corresponding record for this reference.



