NanoBondy Reaction through NeissLock Anhydride Allows Covalent Immune Cell DecorationClick to copy article linkArticle link copied!
- Lasya R. VankayalaLasya R. VankayalaDepartment of Biochemistry, University of Oxford, South Parks Road, Oxford OX1 3QU, U.K.Department of Pharmacology, University of Cambridge, Tennis Court Road, Cambridge CB2 1PD, U.K.Engineering Biology Interdisciplinary Research Centre, University of Cambridge, Cambridge CB2 1GA, U.K.Sir William Dunn School of Pathology, University of Oxford, South Parks Road, Oxford OX1 3RE, U.K.More by Lasya R. Vankayala
- Kish R. AdoniKish R. AdoniInstitute of Structural and Molecular Biology, Division of Biosciences, University College London, London WC1E 6BT, U.K.Institute of Structural and Molecular Biology, Birkbeck College, University of London, London WC1E 6BT, U.K.More by Kish R. Adoni
- Sheryl Y. T. LimSheryl Y. T. LimDepartment of Biochemistry, University of Oxford, South Parks Road, Oxford OX1 3QU, U.K.More by Sheryl Y. T. Lim
- Tommy DamTommy DamSir William Dunn School of Pathology, University of Oxford, South Parks Road, Oxford OX1 3RE, U.K.More by Tommy Dam
- Omer DushekOmer DushekSir William Dunn School of Pathology, University of Oxford, South Parks Road, Oxford OX1 3RE, U.K.More by Omer Dushek
- Konstantinos ThalassinosKonstantinos ThalassinosInstitute of Structural and Molecular Biology, Division of Biosciences, University College London, London WC1E 6BT, U.K.Institute of Structural and Molecular Biology, Birkbeck College, University of London, London WC1E 6BT, U.K.More by Konstantinos Thalassinos
- Mark R. Howarth*Mark R. Howarth*E-mail: [email protected]Department of Pharmacology, University of Cambridge, Tennis Court Road, Cambridge CB2 1PD, U.K.Engineering Biology Interdisciplinary Research Centre, University of Cambridge, Cambridge CB2 1GA, U.K.More by Mark R. Howarth
Abstract
Cell-surface conjugation has enormous therapeutic and research potential. Existing technologies for cell-surface modification are usually reversible, nonspecific, or rely on genetic editing of target cells. Here, we present the NanoBondy, a nanobody modified for covalent ligation to an unmodified protein target at the cell surface. The NanoBondy utilizes the 20 naturally occurring amino acids, harnessing NeissLock chemistry engineered from Neisseria meningitidis. We evaluated the binding and specificity of a panel of nanobodies to CD45, a long-lived surface marker of nucleated hematopoietic cells. We demonstrated the conversion of existing nanobodies to covalently reacting NanoBondies using a disulfide clamp to position the self-processing module of FrpA close to the nanobody antigen-binding site. The addition of calcium induces anhydride formation at the NanoBondy C-terminus, enabling proximity-directed ligation to surface amines on CD45. We optimized the NanoBondy reaction by fine-tuning linkers and disulfide clamp sites to modulate anhydride positioning. Tandem mass spectrometry mapped reaction sites between NanoBondy and CD45. NanoBondy ligation was robust to buffer, pH, and temperature and was detected within 2 minutes. We established the reaction specificity of NanoBondies to endogenous CD45 at the surface of NK cells and T cells. NanoBondy technology provides a modular approach for targeted, inducible, and covalent cell-surface modification of immune cells without their genetic modification.
This publication is licensed under
License Summary*
You are free to share(copy and redistribute) this article in any medium or format and to adapt(remix, transform, and build upon) the material for any purpose, even commercially within the parameters below:
Creative Commons (CC): This is a Creative Commons license.
Attribution (BY): Credit must be given to the creator.
*Disclaimer
This summary highlights only some of the key features and terms of the actual license. It is not a license and has no legal value. Carefully review the actual license before using these materials.
License Summary*
You are free to share(copy and redistribute) this article in any medium or format and to adapt(remix, transform, and build upon) the material for any purpose, even commercially within the parameters below:
Creative Commons (CC): This is a Creative Commons license.
Attribution (BY): Credit must be given to the creator.
*Disclaimer
This summary highlights only some of the key features and terms of the actual license. It is not a license and has no legal value. Carefully review the actual license before using these materials.
License Summary*
You are free to share(copy and redistribute) this article in any medium or format and to adapt(remix, transform, and build upon) the material for any purpose, even commercially within the parameters below:
Creative Commons (CC): This is a Creative Commons license.
Attribution (BY): Credit must be given to the creator.
*Disclaimer
This summary highlights only some of the key features and terms of the actual license. It is not a license and has no legal value. Carefully review the actual license before using these materials.
Introduction
Figure 1
Figure 1. NanoBondy principle. (A) NeissLock chemistry. Upon the addition of calcium, the self-processing module (SPM) activates autoproteolysis at the aspartate–proline bond. This step generates a highly reactive aspartyl anhydride, which undergoes nucleophilic attack by a nearby nucleophilic amino acid or water. Fusing SPM to a binder (purple) thereby allows inducible covalent coupling to a target protein (green). (B) NanoBondy design. A nanobody (purple) employs complementarity-determining regions (CDRs) close to the N-terminus to bind its target (green). A regular nanobody can be engineered into a covalently reacting NanoBondy by inclusion of a flexible linker and disulfide clamp (magenta) to hold the reactive D (cyan) of SPM (orange) near the target, allowing anhydride-mediated covalent conjugation to the target after calcium activation.
Results
Selected Nanobody Candidates Demonstrate Specific Binding to CD45
Figure 2
Figure 2. Characterization of anti-CD45 nanobodies. (A) Purification of anti-CD45 nanobodies. Nanobodies were expressed in E. coli and purified by SpySwitch affinity chromatography, followed by SDS-PAGE ± dithiothreitol (DTT) and Coomassie staining to assess disulfide bond formation. The experiment was conducted once. (B) Schematic of the organization of CD45, including extracellular domains d1–d4. (C) Binding of nanobodies to purified CD45. Nanobodies were coated on a plate and incubated with the indicated concentration of biotinylated human CD45 domains 1 and 2 (CD45d1d2), followed by colorimetric ELISA detection (absorbance at 450 nm). Anti-HER2 nanobody was used as a negative control. Each triplicate data point is shown with a line connecting the mean. Representative ELISA data were obtained from two independent experiments. (D) Binding of anti-CD45 nanobodies at the cell surface by flow cytometry. Anti-CD45 nanobodies were incubated with Expi293F, NK92, or YTS cells. Nanobody binding was detected using anti-VHH-Alexa Fluor 647. Anti-CD45 antibody was used as a positive control, with anti-HER2 nanobodies or unstained (no binder) cells as negative controls. Representative flow cytometry data were obtained from two independent experiments for Expi293F and YTS and one experiment with all three cell lines.
Designed NanoBondies Demonstrate Specific, Inducible Coupling to Purified CD45
Figure 3
Figure 3. NanoBondy covalent conjugation to recombinant CD45. (A) AlphaFold prediction of 2H5 nanobody (purple) interaction with CD45d1d2 (green). Magenta represents the site for a disulfide clamp, with lysines on CD45d1d2 colored pink and the terminal aspartate in cyan. (B) MBP fusion improved the CD45 gel-based analysis. CD45d consists of MBP fused to domains 1 and 2 of CD45. PNGase F digestion decreased heterogeneous mobility of CD45d1d2 and CD45d upon SDS-PAGE with Coomassie staining. (C) Individual protein components for the conjugation assay in (D). CD45d, sfGFP, ODC, and PNGase F were validated by SDS-PAGE/Coomassie staining. (D) Specificity of the NanoBondy reaction with recombinant CD45. Anti-CD45 2H5 R72C or anti-IgG NanoBondy-SPM was incubated with CD45d, each at 10.5 μM, for 1 h at 37 °C in HBS ± calcium, followed by SDS-PAGE with Coomassie staining. ODC, sfGFP, and anti-IgG NanoBondy were used as negative controls for reaction specificity. Hydroxylamine was used as a competing nucleophile to block reactivity. Colon represents a covalent conjugate. The experiment was conducted once.
The NanoBondy Clamp Site and Linker Length Alter Conjugation Yield
Figure 4
Figure 4. Optimization of NanoBondy clamp site and linker length. (A) AlphaFold prediction of 2H5 NanoBondy (purple) bound to CD45d1d2 (green). The reactive anhydride is shown in cyan, alternative clamp sites are shown in magenta, and linkers are shown in orange. (B) Clamp-site variant reactivity. 2H5 NanoBondy variants were incubated with CD45d at 10.5 μM each for 1 h at 37 °C in HBS ± Ca2+, followed by SDS-PAGE with Coomassie staining. Anti-IgG NanoBondy was used as a negative control. The leftmost lane represents CD45d without PNGase F treatment. The experiment was conducted once. (C) Linker variant reactivity for 2H5 R72C anti-CD45 NanoBondy, analyzed as in (B). Representative gels were obtained from two independent experiments.
The NanoBondy Retains Reactivity across Various Conditions
Figure 5
Figure 5. Condition-dependence of NanoBondy reaction. (A) Buffer- and temperature-dependence of NanoBondy reaction. 2H5 R72C anti-CD45 NanoBondy was incubated with CD45d, each at 10.5 μM, in the indicated buffer and temperature before SDS-PAGE with Coomassie staining. Ctrl refers to the lane containing anti-IgG NanoBondy with CD45d for 120 min. Representative gel from two independent experiments. (B) pH-dependence of NanoBondy reaction. The reaction was evaluated as in (A) with HBS-MES buffer at the indicated pH at 37 °C. Ctrl refers to the lane containing anti-IgG NanoBondy with CD45d for 120 min. The experiment was conducted once.
Cross-Linking MS/MS Identifies Sites of NanoBondy-CD45d Cross-Linking
Figure 6
Figure 6. Mass spectrometry analysis of covalent conjugate between the NanoBondy and CD45d. (A) Dominant cross-linking sites. 2H5 R72C anti-CD45 NanoBondy was incubated with CD45d before cross-linking MS/MS. The number of identified cross-linked spectral matches for each NanoBondy cross-linking site on CD45 is shown. (B) Higher energy collision-induced dissociation (HCD) fragmentation spectrum of identified cross-link precursor ions corresponding to D173 (NanoBondy) to K131 (CD45d). Fragment ions matching fragmented cross-link (with cross-linker still intact) are annotated in bold, while peaks corresponding to fragments post-cross-link fragmentation are annotated with nonbold lines. “Car” indicates carbamidomethylation of cysteine. (C) Mapping of cross-link sites. AlphaFold prediction of NanoBondy (purple) bound to CD45d1d2 domains (green), highlighting cross-linking sites identified from the reactive aspartate (cyan) of the NanoBondy to target lysines (pink) or to serine (dark blue). Samples were run in technical triplicate. The cross-linking MS experiment was conducted twice.
NanoBondy Demonstrates Targeted Covalent Coupling at the Cell Surface
Figure 7
Figure 7. NanoBondy covalent conjugation at the cell surface. (A) Western blotting of NanoBondy reaction at the cell surface. Anti-CD45 NanoBondy at 5 μM was incubated with YTS cells or Expi293F cells for 1 h at 37 °C ± calcium. Covalent conjugation was evaluated by Western blot using an anti-VHH polyclonal antibody to detect the NanoBondy. Anti-IgG NanoBondy or hydroxylamine to react with the anhydride provided negative controls. Western blot to glyceraldehyde-3-phosphate dehydrogenase (GAPDH) was the loading control. The complete GAPDH blot is presented in Figure S6A. The experiment was conducted once. (B) YTS cells were stained as in (A), except with 25 μM anti-CD45 NanoBondy, and analyzed by Western blot using an anti-CD45 antibody. The CD45 band demonstrates an upward shift upon covalent conjugation with the a-CD45 NanoBondy. The full-length GAPDH blot is presented in Figure S6B. The experiment was conducted once. (C) Western blotting of the DuoBondy reaction on CD8+ T cells. DuoBondy (WT) or DuoBondy (DA) at 1 μM was incubated with CD8+ T cells for 40 min at 37 °C ± calcium. Covalent conjugation was evaluated by Western blot using an anti-VHH polyclonal antibody to detect the DuoBondy. Representative blot from two independent experiments. (D) DuoBondy consists of a nanobody binder (Nb102c3) to PD-1 (dark blue) fused N-terminally to the established covalently reacting anti-CD45 NanoBondy (purple) with SPM in orange.
Discussion
Supporting Information
The Supporting Information is available free of charge at https://pubs.acs.org/doi/10.1021/acs.bioconjchem.5c00519.
Comprehensive methods and supplementary results, including amino acid sequence, MS, structure prediction, and fluorescence microscopy (PDF)
Terms & Conditions
Most electronic Supporting Information files are available without a subscription to ACS Web Editions. Such files may be downloaded by article for research use (if there is a public use license linked to the relevant article, that license may permit other uses). Permission may be obtained from ACS for other uses through requests via the RightsLink permission system: http://pubs.acs.org/page/copyright/permissions.html.
Acknowledgments
L.R.V. was funded by MSD. M.R.H. was funded by the Engineering and Physical Sciences Research Council (EPSRC EP/W01565X/1). K.A. is funded by a Wellcome Collaborative Award in Science (209250/Z/17/Z) to K.T. The mass spectrometer used for cross-linking was funded by a Wellcome Trust Multiuser Equipment grant (221521/Z/20/Z) to K.T. S.Y.T.L. was funded by an A*STAR studentship. This work was supported by the Wellcome Trust (207537/Z/17/Z, 301534/Z/23/Z). We thank Dr. Anthony Tumber (University of Oxford, Department of Chemistry) for assistance with intact protein mass spectrometry, supported by the Biotechnology and Biological Sciences Research Council (BBSRC BB/R000344/1). We thank the flow cytometry facility staff at the University of Cambridge School of the Biological Sciences for the technical expertise provided. AlphaFold2-multimer docking was performed using resources provided by the Cambridge Service for Data Driven Discovery (CSD3). CSD3 is operated by the University of Cambridge Research Computing Service, provided by Dell EMC and Intel, using Tier-2 funding from the EPSRC (capital grant EP/T022159/1) and DiRAC funding from the Science and Technology Facilities Council. We would like to thank Dr. Alan Wainman and the Dunn School Bioimaging Facility.
References
This article references 71 other publications.
- 1Keeble, A. H.; Howarth, M. Power to the Protein: Enhancing and Combining Activities Using the Spy Toolbox. Chem. Sci. 2020, 11 (28), 7281– 7291, DOI: 10.1039/D0SC01878CGoogle ScholarThere is no corresponding record for this reference.
- 2Oostindie, S. C.; Lazar, G. A.; Schuurman, J.; Parren, P. W. H. I. Avidity in Antibody Effector Functions and Biotherapeutic Drug Design. Nat. Rev. Drug Discovery 2022, 21 (10), 715– 735, DOI: 10.1038/s41573-022-00501-8Google ScholarThere is no corresponding record for this reference.
- 3Khan, S. H.; Choi, Y.; Veena, M.; Lee, J. K.; Shin, D. S. Advances in CAR T Cell Therapy: Antigen Selection, Modifications, and Current Trials for Solid Tumors. Front. Immunol. 2025, 15, 1489827, DOI: 10.3389/fimmu.2024.1489827Google ScholarThere is no corresponding record for this reference.
- 4Brenner, J. S.; Mitragotri, S.; Muzykantov, V. R. Red Blood Cell Hitchhiking: A Novel Approach for Vascular Delivery of Nanocarriers. Annu. Rev. Biomed. Eng. 2021, 23 (1), 225– 248, DOI: 10.1146/annurev-bioeng-121219-024239Google ScholarThere is no corresponding record for this reference.
- 5Albelda, S. M. CAR T Cell Therapy for Patients with Solid Tumours: Key Lessons to Learn and Unlearn. Nat. Rev. Clin Oncol. 2024, 21 (1), 47– 66, DOI: 10.1038/s41571-023-00832-4Google ScholarThere is no corresponding record for this reference.
- 6D’Aloia, M. M.; Zizzari, I. G.; Sacchetti, B.; Pierelli, L.; Alimandi, M. CAR-T Cells: The Long and Winding Road to Solid Tumors. Cell Death Dis. 2018, 9 (3), 282, DOI: 10.1038/s41419-018-0278-6Google ScholarThere is no corresponding record for this reference.
- 7Guzman, G.; Reed, M. R.; Bielamowicz, K.; Koss, B.; Rodriguez, A. CAR-T Therapies in Solid Tumors: Opportunities and Challenges. Curr. Oncol. Rep. 2023, 25 (5), 479– 489, DOI: 10.1007/s11912-023-01380-xGoogle ScholarThere is no corresponding record for this reference.
- 8Hurton, L. V.; Singh, H.; Najjar, A. M.; Switzer, K. C.; Mi, T.; Maiti, S.; Olivares, S.; Rabinovich, B.; Huls, H.; Forget, M.-A. Tethered IL-15 Augments Antitumor Activity and Promotes a Stem-Cell Memory Subset in Tumor-Specific T Cells. Proc. Natl. Acad. Sci. U.S.A. 2016, 113 (48), E7788–E7797 DOI: 10.1073/pnas.1610544113Google ScholarThere is no corresponding record for this reference.
- 9Jones, D. S.; Nardozzi, J. D.; Sackton, K. L.; Ahmad, G.; Christensen, E.; Ringgaard, L.; Chang, D.-K.; Jaehger, D. E.; Konakondla, J. V.; Wiinberg, M.; Stokes, K. L.; Pratama, A.; Sauer, K.; Andresen, T. L. Cell Surface–Tethered IL-12 Repolarizes the Tumor Immune Microenvironment to Enhance the Efficacy of Adoptive T Cell Therapy. Sci. Adv. 2022, 8 (17), eabi8075 DOI: 10.1126/sciadv.abi8075Google ScholarThere is no corresponding record for this reference.
- 10Zhao, Y.; Dong, Y.; Yang, S.; Tu, Y.; Wang, C.; Li, J.; Yuan, Y.; Lian, Z. Bioorthogonal Equipping CAR-T Cells with Hyaluronidase and Checkpoint Blocking Antibody for Enhanced Solid Tumor Immunotherapy. ACS Cent. Sci. 2022, 8 (5), 603– 614, DOI: 10.1021/acscentsci.2c00163Google ScholarThere is no corresponding record for this reference.
- 11Stephan, M.; Stephan, S.; Bak, P.; Chen, J.; Irvine, D. J. Synapse-Directed Delivery of Immunomodulators Using T-Cell-Conjugated Nanoparticles. Biomaterials 2012, 33 (23), 5776– 5787, DOI: 10.1016/j.biomaterials.2012.04.029Google ScholarThere is no corresponding record for this reference.
- 12Luo, Y.; Chen, Z.; Sun, M.; Li, B.; Pan, F.; Ma, A.; Liao, J.; Yin, T.; Tang, X.; Huang, G.; Zhang, B.; Pan, H.; Zheng, M.; Cai, L. IL-12 Nanochaperone-Engineered CAR T Cell for Robust Tumor-Immunotherapy. Biomaterials 2022, 281, 121341, DOI: 10.1016/j.biomaterials.2021.121341Google ScholarThere is no corresponding record for this reference.
- 13Liu, Y.; Adu-Berchie, K.; Brockman, J. M.; Pezone, M.; Zhang, D. K. Y.; Zhou, J.; Pyrdol, J. W.; Wang, H.; Wucherpfennig, K. W.; Mooney, D. J. Cytokine Conjugation to Enhance T Cell Therapy. Proc. Natl. Acad. Sci. U.S.A. 2023, 120 (1), e2213222120 DOI: 10.1073/pnas.2213222120Google ScholarThere is no corresponding record for this reference.
- 14Zheng, Y.; Tang, L.; Mabardi, L.; Kumari, S.; Irvine, D. J. Enhancing Adoptive Cell Therapy of Cancer through Targeted Delivery of Small-Molecule Immunomodulators to Internalizing or Noninternalizing Receptors. ACS Nano 2017, 11 (3), 3089– 3100, DOI: 10.1021/acsnano.7b00078Google ScholarThere is no corresponding record for this reference.
- 15Stephan, M.; Moon, J.; Um, S. H.; Bershteyn, A.; Irvine, D. Therapeutic Cell Engineering with Surface-Conjugated Synthetic Nanoparticles. Nat. Med. 2010, 16 (9), 1035– 1041, DOI: 10.1038/nm.2198Google ScholarThere is no corresponding record for this reference.
- 16Tang, L.; Zheng, Y.; Melo, M. B.; Mabardi, L.; Castaño, A. P.; Xie, Y.-Q.; Li, N.; Kudchodkar, S. B.; Wong, H. C.; Jeng, E. K.; Maus, M. V.; Irvine, D. J. Enhancing T Cell Therapy through TCR-Signaling-Responsive Nanoparticle Drug Delivery. Nat. Biotechnol. 2018, 36 (8), 707– 716, DOI: 10.1038/nbt.4181Google ScholarThere is no corresponding record for this reference.
- 17Los, G. V.; Encell, L. P.; McDougall, M. G.; Hartzell, D. D.; Karassina, N.; Zimprich, C.; Wood, M. G.; Learish, R.; Ohana, R. F.; Urh, M.; Simpson, D.; Mendez, J.; Zimmerman, K.; Otto, P.; Vidugiris, G.; Zhu, J.; Darzins, A.; Klaubert, D. H.; Bulleit, R. F.; Wood, K. V. HaloTag: A Novel Protein Labeling Technology for Cell Imaging and Protein Analysis. ACS Chem. Biol. 2008, 3 (6), 373– 382, DOI: 10.1021/cb800025kGoogle ScholarThere is no corresponding record for this reference.
- 18Keppler, A.; Gendreizig, S.; Gronemeyer, T.; Pick, H.; Vogel, H.; Johnsson, K. A General Method for the Covalent Labeling of Fusion Proteins with Small Molecules in Vivo. Nat. Biotechnol. 2003, 21 (1), 86– 89, DOI: 10.1038/nbt765Google ScholarThere is no corresponding record for this reference.
- 19Sun, F.; Zhang, W.-B. Genetically Encoded Click Chemistry. Chin. J. Chem. 2020, 38 (8), 894– 896, DOI: 10.1002/cjoc.202000101Google ScholarThere is no corresponding record for this reference.
- 20Humberg, C.; Yilmaz, Z.; Fitzian, K.; Dörner, W.; Kümmel, D.; Mootz, H. D. A Cysteine-Less and Ultra-Fast Split Intein Rationally Engineered from Being Aggregation-Prone to Highly Efficient in Protein Trans-Splicing. Nat. Commun. 2025, 16 (1), 2723, DOI: 10.1038/s41467-025-57596-xGoogle ScholarThere is no corresponding record for this reference.
- 21Mao, H.; Hart, S. A.; Schink, A.; Pollok, B. A. Sortase-Mediated Protein Ligation: A New Method for Protein Engineering. J. Am. Chem. Soc. 2004, 126 (9), 2670– 2671, DOI: 10.1021/ja039915eGoogle ScholarThere is no corresponding record for this reference.
- 22Liu, X.; Wen, J.; Yi, H.; Hou, X.; Yin, Y.; Ye, G.; Wu, X.; Jiang, X. Split Chimeric Antigen Receptor-Modified T Cells Targeting Glypican-3 Suppress Hepatocellular Carcinoma Growth with Reduced Cytokine Release. Ther. Adv. Med. Oncol. 2020, 12, 175883592091034, DOI: 10.1177/1758835920910347Google ScholarThere is no corresponding record for this reference.
- 23Sutherland, A. R.; Owens, M. N.; Geyer, C. R. Modular Chimeric Antigen Receptor Systems for Universal CAR T Cell Retargeting. Int. J. Mol. Sci. 2020, 21 (19), 7222, DOI: 10.3390/ijms21197222Google ScholarThere is no corresponding record for this reference.
- 24Minutolo, N. G.; Sharma, P.; Poussin, M.; Shaw, L. C.; Brown, D. P.; Hollander, E. E.; Smole, A.; Rodriguez-Garcia, A.; Hui, J. Z.; Zappala, F.; Tsourkas, A.; Powell, D. J. Quantitative Control of Gene-Engineered T-Cell Activity through the Covalent Attachment of Targeting Ligands to a Universal Immune Receptor. J. Am. Chem. Soc. 2020, 142 (14), 6554– 6568, DOI: 10.1021/jacs.9b11622Google ScholarThere is no corresponding record for this reference.
- 25Rahikainen, R.; Rijal, P.; Tan, T. K.; Wu, H.; Andersson, A. C.; Barrett, J. R.; Bowden, T. A.; Draper, S. J.; Townsend, A. R.; Howarth, M. Overcoming Symmetry Mismatch in Vaccine Nanoassembly through Spontaneous Amidation. Angew. Chem. Int. Ed. 2021, 60 (1), 321– 330, DOI: 10.1002/anie.202009663Google ScholarThere is no corresponding record for this reference.
- 26Bishani, A.; Chernolovskaya, E. L. Activation of Innate Immunity by Therapeutic Nucleic Acids. Int. J. Mol. Sci. 2021, 22 (24), 13360, DOI: 10.3390/ijms222413360Google ScholarThere is no corresponding record for this reference.
- 27Bonifant, C. L.; Jackson, H. J.; Brentjens, R. J.; Curran, K. J. Toxicity and Management in CAR T-Cell Therapy. Mol. Ther. Oncolytics 2016, 3, 16011, DOI: 10.1038/mto.2016.11Google ScholarThere is no corresponding record for this reference.
- 28Hamdy, N.; Goustin, A. S.; Desaulniers, J.-P.; Li, M.; Chow, C. S.; Al-Katib, A. Sheep Red Blood Cells Armed with Anti-CD20 Single-Chain Variable Fragments (scFvs) Fused to a Glycosylphosphatidylinositol (GPI) Anchor: A Strategy to Target CD20-Positive Tumor Cells. J. Immunol. Methods 2005, 297 (1–2), 109– 124, DOI: 10.1016/j.jim.2004.12.003Google ScholarThere is no corresponding record for this reference.
- 29White, S.; Vonheijne, G. Transmembrane Helices before, during, and after Insertion. Curr. Opin. Struct. Biol. 2005, 15 (4), 378– 386, DOI: 10.1016/j.sbi.2005.07.004Google ScholarThere is no corresponding record for this reference.
- 30Reinisch, K. M.; Prinz, W. A. Mechanisms of Nonvesicular Lipid Transport. J. Cell Biol. 2021, 220 (3), e202012058 DOI: 10.1083/jcb.202012058Google ScholarThere is no corresponding record for this reference.
- 31Yu, B.; Li, S.; Tabata, T.; Wang, N.; Cao, L.; Kumar, G. R.; Sun, W.; Liu, J.; Ott, M.; Wang, L. Accelerating PERx Reaction Enables Covalent Nanobodies for Potent Neutralization of SARS-CoV-2 and Variants. Chem 2022, 8 (10), 2766– 2783, DOI: 10.1016/j.chempr.2022.07.012Google ScholarThere is no corresponding record for this reference.
- 32Yu, B.; Cao, L.; Li, S.; Klauser, P. C.; Wang, L. The Proximity-Enabled Sulfur Fluoride Exchange Reaction in the Protein Context. Chem. Sci. 2023, 14 (29), 7913– 7921, DOI: 10.1039/D3SC01921GGoogle ScholarThere is no corresponding record for this reference.
- 33Li, Q.; Chen, Q.; Klauser, P. C.; Li, M.; Zheng, F.; Wang, N.; Li, X.; Zhang, Q.; Fu, X.; Wang, Q. Developing Covalent Protein Drugs via Proximity-Enabled Reactive Therapeutics. Cell 2020, 182 (1), P85– 97.E16, DOI: 10.1016/j.cell.2020.05.028Google ScholarThere is no corresponding record for this reference.
- 34De Bank, P. A.; Kellam, B.; Kendall, D. A.; Shakesheff, K. M. Surface Engineering of Living Myoblasts via Selective Periodate Oxidation. Biotechnol. Bioeng. 2003, 81 (7), 800– 808, DOI: 10.1002/bit.10525Google ScholarThere is no corresponding record for this reference.
- 35Glassman, P. M.; Hood, E. D.; Ferguson, L. T.; Zhao, Z.; Siegel, D. L.; Mitragotri, S.; Brenner, J. S.; Muzykantov, V. R. Red Blood Cells: The Metamorphosis of a Neglected Carrier into the Natural Mothership for Artificial Nanocarriers. Adv. Drug Delivery Rev. 2021, 178, 113992, DOI: 10.1016/j.addr.2021.113992Google ScholarThere is no corresponding record for this reference.
- 36Bao, G.; Tang, M.; Zhao, J.; Zhu, X. Nanobody: A Promising Toolkit for Molecular Imaging and Disease Therapy. EJNMMI Res. 2021, 11 (1), 6, DOI: 10.1186/s13550-021-00750-5Google ScholarThere is no corresponding record for this reference.
- 37Asaadi, Y.; Jouneghani, F. F.; Janani, S.; Rahbarizadeh, F. A Comprehensive Comparison between Camelid Nanobodies and Single Chain Variable Fragments. Biomark. Res. 2021, 9 (1), 87, DOI: 10.1186/s40364-021-00332-6Google ScholarThere is no corresponding record for this reference.
- 38Tack, L.; Miret-Casals, L.; Zwaenepoel, O.; Cadoni, E.; Van Troys, M.; Gettemans, J.; Madder, A. Development of “Furan Warhead”-Equipped Antagonistic Nanobodies for Covalent Cross-Linking to the Epidermal Growth Factor Receptor. Bioconjugate Chem. 2025, 36 (8), 1649– 1660, DOI: 10.1021/acs.bioconjchem.5c00144Google ScholarThere is no corresponding record for this reference.
- 39Gao, W.; Cho, E.; Liu, Y.; Lu, Y. Advances and Challenges in Cell-Free Incorporation of Unnatural Amino Acids Into Proteins. Front. Pharmacol. 2019, 10, 611, DOI: 10.3389/fphar.2019.00611Google ScholarThere is no corresponding record for this reference.
- 40Scheu, A.; Lim, S. Y. T.; Metzner, F. J.; Mohammed, S.; Howarth, M. NeissLock Provides an Inducible Protein Anhydride for Covalent Targeting of Endogenous Proteins. Nat. Commun. 2021, 12 (1), 717, DOI: 10.1038/s41467-021-20963-5Google ScholarThere is no corresponding record for this reference.
- 41Lim, S.; Keeble, A. H.; Howarth, M. R. Split NeissLock with Spy-Acceleration Arms Mammalian Proteins for Anhydride-Mediated Cell Ligation. ACS Chem. Biol. 2025, 20, 2475, DOI: 10.1021/acschembio.5c00515Google ScholarThere is no corresponding record for this reference.
- 42Sviridova, E.; Rezacova, P.; Bondar, A.; Veverka, V.; Novak, P.; Schenk, G.; Svergun, D. I.; Kuta Smatanova, I.; Bumba, L. Structural Basis of the Interaction between the Putative Adhesion-Involved and Iron-Regulated FrpD and FrpC Proteins of Neisseria Meningitidis. Sci. Rep. 2017, 7 (1), 40408, DOI: 10.1038/srep40408Google ScholarThere is no corresponding record for this reference.
- 43Borowska, M. T.; Liu, L. D.; Caveney, N. A.; Jude, K. M.; Kim, W.-J.; Masubuchi, T.; Hui, E.; Majzner, R. G.; Garcia, K. C. Orientation-Dependent CD45 Inhibition with Viral and Engineered Ligands. Sci. Immunol. 2024, 9 (100), eadp0707 DOI: 10.1126/sciimmunol.adp0707Google ScholarThere is no corresponding record for this reference.
- 44Rokkam, D.; Lupardus, P. J. Discovery and Characterization of Llama VHH Targeting the RO Form of Human CD45. bioRxiv . 2020. DOI: 10.1101/2020.09.01.278853Google ScholarThere is no corresponding record for this reference.
- 45Keeble, A. H.; Turkki, P.; Stokes, S.; Khairil Anuar, I. N. A.; Rahikainen, R.; Hytönen, V. P.; Howarth, M. Approaching Infinite Affinity through Engineering of Peptide–Protein Interaction. Proc. Natl. Acad. Sci. U.S.A. 2019, 116 (52), 26523– 26533, DOI: 10.1073/pnas.1909653116Google ScholarThere is no corresponding record for this reference.
- 46Vester, S. K.; Rahikainen, R.; Khairil Anuar, I. N. A.; Hills, R. A.; Tan, T. K.; Howarth, M. SpySwitch Enables pH- or Heat-Responsive Capture and Release for Plug-and-Display Nanoassembly. Nat. Commun. 2022, 13 (1), 3714, DOI: 10.1038/s41467-022-31193-8Google ScholarThere is no corresponding record for this reference.
- 47Bryant, P.; Pozzati, G.; Elofsson, A. Improved Prediction of Protein-Protein Interactions Using AlphaFold2. Nat. Commun. 2022, 13 (1), 1265, DOI: 10.1038/s41467-022-28865-wGoogle ScholarThere is no corresponding record for this reference.
- 48Evans, R.; O’Neill, M.; Pritzel, A.; Antropova, N.; Senior, A.; Green, T.; Žídek, A.; Bates, R.; Blackwell, S.; Yim, J. Protein Complex Prediction With AlphaFold-Multimer. bioRxiv . 2022. DOI: 10.1101/2021.10.04.463034Google ScholarThere is no corresponding record for this reference.
- 49Zhong, B.; Su, X.; Wen, M.; Zuo, S.; Hong, L.; Lin, J. ParaFold: Paralleling AlphaFold For Large-Scale Predictions. arXiv . 2021. DOI: 10.48550/arXiv.2111.06340Google ScholarThere is no corresponding record for this reference.
- 50Breitwieser, G. E. Extracellular Calcium as an Integrator of Tissue Function. Int. J. Biochem. Cell Biol. 2008, 40 (8), 1467– 1480, DOI: 10.1016/j.biocel.2008.01.019Google ScholarThere is no corresponding record for this reference.
- 51Hermiston, M. L.; Xu, Z.; Weiss, A. CD45: A Critical Regulator of Signaling Thresholds in Immune Cells. Annu. Rev. Immunol. 2003, 21 (1), 107– 137, DOI: 10.1146/annurev.immunol.21.120601.140946Google ScholarThere is no corresponding record for this reference.
- 52Savchenko, A. A.; Tikhonova, E.; Kudryavtsev, I.; Kudlay, D.; Korsunsky, I.; Beleniuk, V.; Borisov, A. TREC/KREC Levels and T and B Lymphocyte Subpopulations in COVID-19 Patients at Different Stages of the Disease. Viruses 2022, 14 (3), 646, DOI: 10.3390/v14030646Google ScholarThere is no corresponding record for this reference.
- 53Kalinin, R. S.; Ukrainskaya, V. M.; Chumakov, S. P.; Moysenovich, A. M.; Tereshchuk, V. M.; Volkov, D. V.; Pershin, D. S.; Maksimov, E. G.; Zhang, H.; Maschan, M. A.; Rubtsov, Y. P.; Stepanov, A. V. Engineered Removal of PD-1 From the Surface of CD19 CAR-T Cells Results in Increased Activation and Diminished Survival. Front. Mol. Biosci. 2021, 8, 745286, DOI: 10.3389/fmolb.2021.745286Google ScholarThere is no corresponding record for this reference.
- 54Fernandes, R. A.; Su, L.; Nishiga, Y.; Ren, J.; Bhuiyan, A. M.; Cheng, N.; Kuo, C. J.; Picton, L. K.; Ohtsuki, S.; Majzner, R. G.; Rietberg, S. P.; Mackall, C. L.; Yin, Q.; Ali, L. R.; Yang, X.; Savvides, C. S.; Sage, J.; Dougan, M.; Garcia, K. C. Immune Receptor Inhibition through Enforced Phosphatase Recruitment. Nature 2020, 586 (7831), 779– 784, DOI: 10.1038/s41586-020-2851-2Google ScholarThere is no corresponding record for this reference.
- 55Fernández-Quintero, M. L.; Kokot, J.; Waibl, F.; Fischer, A.-L. M.; Quoika, P. K.; Deane, C. M.; Liedl, K. R. Challenges in Antibody Structure Prediction. mAbs 2023, 15 (1), 2175319, DOI: 10.1080/19420862.2023.2175319Google ScholarThere is no corresponding record for this reference.
- 56Erasmus, M. F.; Spector, L.; Ferrara, F.; DiNiro, R.; Pohl, T. J.; Perea-Schmittle, K.; Wang, W.; Tessier, P. M.; Richardson, C.; Turner, L.; Kumar, S.; Bedinger, D.; Sormanni, P.; Fernández-Quintero, M. L.; Ward, A. B.; Loeffler, J. R.; Swanson, O. M.; Deane, C. M.; Raybould, M. I. J.; Evers, A.; Sellmann, C.; Bachas, S.; Ruffolo, J.; Nastri, H. G.; Ramesh, K.; Sørensen, J.; Croasdale-Wood, R.; Hijano, O.; Leal-Lopes, C.; Shahsavarian, M.; Qiu, Y.; Marcatili, P.; Vernet, E.; Akbar, R.; Friedensohn, S.; Wagner, R.; Kurella, V. B.; Malhotra, S.; Kumar, S.; Kidger, P.; Almagro, J. C.; Furfine, E.; Stanton, M.; Graff, C. P.; Villalba, S. D.; Tomszak, F.; Teixeira, A. A. R.; Hopkins, E.; Dovner, M.; D’Angelo, S.; Bradbury, A. R. M. AIntibody: An Experimentally Validated in Silico Antibody Discovery Design Challenge. Nat. Biotechnol. 2024, 42 (11), 1637– 1642, DOI: 10.1038/s41587-024-02469-9Google ScholarThere is no corresponding record for this reference.
- 57Holm, L.; Moody, P.; Howarth, M. Electrophilic Affibodies Forming Covalent Bonds to Protein Targets. J. Biol. Chem. 2009, 284 (47), 32906– 32913, DOI: 10.1074/jbc.M109.034322Google ScholarThere is no corresponding record for this reference.
- 58De La Torre, D.; Chin, J. W. Reprogramming the Genetic Code. Nat. Rev. Genet. 2021, 22 (3), 169– 184, DOI: 10.1038/s41576-020-00307-7Google ScholarThere is no corresponding record for this reference.
- 59Zhang, H.; Han, Y.; Yang, Y.; Lin, F.; Li, K.; Kong, L.; Liu, H.; Dang, Y.; Lin, J.; Chen, P. R. Covalently Engineered Nanobody Chimeras for Targeted Membrane Protein Degradation. J. Am. Chem. Soc. 2021, 143 (40), 16377– 16382, DOI: 10.1021/jacs.1c08521Google ScholarThere is no corresponding record for this reference.
- 60Tyrrell, R. M. Induction of Pyrimidine Dimers In Bacterial DNA by 365 nm Radiation. Photochem. Photobiol. 1973, 17 (1), 69– 73, DOI: 10.1111/j.1751-1097.1973.tb06334.xGoogle ScholarThere is no corresponding record for this reference.
- 61Andley, U. P.; Lewis, R. M.; Reddan, J. R.; Kochevar, I. E. Action Spectrum for Cytotoxicity in the UVA- and UVB-Wavelength Region in Cultured Lens Epithelial Cells. Invest. Ophthalmol. Vis. Sci. 1994, 35 (2), 367– 373,Google ScholarThere is no corresponding record for this reference.
- 62Veggiani, G.; Nakamura, T.; Brenner, M. D.; Gayet, R. V.; Yan, J.; Robinson, C. V.; Howarth, M. Programmable Polyproteams Built Using Twin Peptide Superglues. Proc. Natl. Acad. Sci. U.S.A. 2016, 113 (5), 1202– 1207, DOI: 10.1073/pnas.1519214113Google ScholarThere is no corresponding record for this reference.
- 63Zimmermann, I.; Egloff, P.; Hutter, C. A. J.; Kuhn, B. T.; Bräuer, P.; Newstead, S.; Dawson, R. J. P.; Geertsma, E. R.; Seeger, M. A. Generation of Synthetic Nanobodies against Delicate Proteins. Nat. Protoc. 2020, 15 (5), 1707– 1741, DOI: 10.1038/s41596-020-0304-xGoogle ScholarThere is no corresponding record for this reference.
- 64Kunz, P.; Zinner, K.; Mücke, N.; Bartoschik, T.; Muyldermans, S.; Hoheisel, J. D. The Structural Basis of Nanobody Unfolding Reversibility and Thermoresistance. Sci. Rep. 2018, 8 (1), 7934, DOI: 10.1038/s41598-018-26338-zGoogle ScholarThere is no corresponding record for this reference.
- 65Berkmen, M. Production of Disulfide-Bonded Proteins in Escherichia Coli. Protein Expression Purif. 2012, 82 (1), 240– 251, DOI: 10.1016/j.pep.2011.10.009Google ScholarThere is no corresponding record for this reference.
- 66Gebauer, M.; Skerra, A. Engineered Protein Scaffolds as Next-Generation Therapeutics. Annu. Rev. Pharmacol. Toxicol. 2020, 60 (1), 391– 415, DOI: 10.1146/annurev-pharmtox-010818-021118Google ScholarThere is no corresponding record for this reference.
- 67Liang, Y.; Xu, Q.; Gao, Q. Advancing CAR-Based Immunotherapies in Solid Tumors: CAR- Macrophages and Neutrophils. Front. Immunol. 2023, 14, 1291619, DOI: 10.3389/fimmu.2023.1291619Google ScholarThere is no corresponding record for this reference.
- 68Wang, X.-W.; Zhang, W.-B. Chemical Topology and Complexity of Protein Architectures. Trends Biochem. Sci. 2018, 43 (10), 806– 817, DOI: 10.1016/j.tibs.2018.07.001Google ScholarThere is no corresponding record for this reference.
- 69Pedram, K.; Shon, D. J.; Tender, G. S.; Mantuano, N. R.; Northey, J. J.; Metcalf, K. J.; Wisnovsky, S. P.; Riley, N. M.; Forcina, G. C.; Malaker, S. A.; Kuo, A.; George, B. M.; Miller, C. L.; Casey, K. M.; Vilches-Moure, J. G.; Ferracane, M. J.; Weaver, V. M.; Läubli, H.; Bertozzi, C. R. Design of a Mucin-Selective Protease for Targeted Degradation of Cancer-Associated Mucins. Nat. Biotechnol. 2024, 42 (4), 597– 607, DOI: 10.1038/s41587-023-01840-6Google ScholarThere is no corresponding record for this reference.
- 70Eichhoff, A. M.; Börner, K.; Albrecht, B.; Schäfer, W.; Baum, N.; Haag, F.; Körbelin, J.; Trepel, M.; Braren, I.; Grimm, D. Nanobody-Enhanced Targeting of AAV Gene Therapy Vectors. Mol. Ther.–Methods Clin. Dev. 2019, 15, P211– 220, DOI: 10.1016/j.omtm.2019.09.003Google ScholarThere is no corresponding record for this reference.
- 71Bocancia-Mateescu, L.-A.; Stan, D.; Mirica, A.-C.; Ghita, M. G.; Stan, D.; Ruta, L. L. Nanobodies as Diagnostic and Therapeutic Tools for Cardiovascular Diseases (CVDs). Pharmaceuticals 2023, 16 (6), 863, DOI: 10.3390/ph16060863Google ScholarThere is no corresponding record for this reference.
Cited By
This article has not yet been cited by other publications.
Article Views
Altmetric
Citations
Article Views are the COUNTER-compliant sum of full text article downloads since November 2008 (both PDF and HTML) across all institutions and individuals. These metrics are regularly updated to reflect usage leading up to the last few days.
Citations are the number of other articles citing this article, calculated by Crossref and updated daily. Find more information about Crossref citation counts.
The Altmetric Attention Score is a quantitative measure of the attention that a research article has received online. Clicking on the donut icon will load a page at altmetric.com with additional details about the score and the social media presence for the given article. Find more information on the Altmetric Attention Score and how the score is calculated.
Recommended Articles
Abstract

Figure 1

Figure 1. NanoBondy principle. (A) NeissLock chemistry. Upon the addition of calcium, the self-processing module (SPM) activates autoproteolysis at the aspartate–proline bond. This step generates a highly reactive aspartyl anhydride, which undergoes nucleophilic attack by a nearby nucleophilic amino acid or water. Fusing SPM to a binder (purple) thereby allows inducible covalent coupling to a target protein (green). (B) NanoBondy design. A nanobody (purple) employs complementarity-determining regions (CDRs) close to the N-terminus to bind its target (green). A regular nanobody can be engineered into a covalently reacting NanoBondy by inclusion of a flexible linker and disulfide clamp (magenta) to hold the reactive D (cyan) of SPM (orange) near the target, allowing anhydride-mediated covalent conjugation to the target after calcium activation.
Figure 2
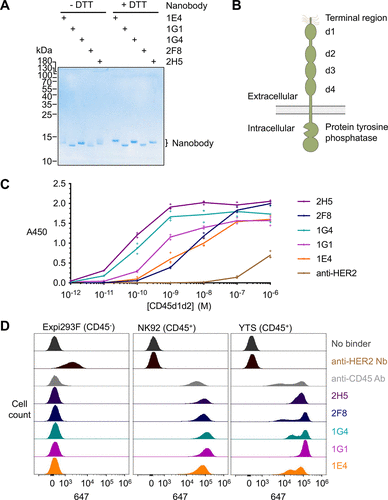
Figure 2. Characterization of anti-CD45 nanobodies. (A) Purification of anti-CD45 nanobodies. Nanobodies were expressed in E. coli and purified by SpySwitch affinity chromatography, followed by SDS-PAGE ± dithiothreitol (DTT) and Coomassie staining to assess disulfide bond formation. The experiment was conducted once. (B) Schematic of the organization of CD45, including extracellular domains d1–d4. (C) Binding of nanobodies to purified CD45. Nanobodies were coated on a plate and incubated with the indicated concentration of biotinylated human CD45 domains 1 and 2 (CD45d1d2), followed by colorimetric ELISA detection (absorbance at 450 nm). Anti-HER2 nanobody was used as a negative control. Each triplicate data point is shown with a line connecting the mean. Representative ELISA data were obtained from two independent experiments. (D) Binding of anti-CD45 nanobodies at the cell surface by flow cytometry. Anti-CD45 nanobodies were incubated with Expi293F, NK92, or YTS cells. Nanobody binding was detected using anti-VHH-Alexa Fluor 647. Anti-CD45 antibody was used as a positive control, with anti-HER2 nanobodies or unstained (no binder) cells as negative controls. Representative flow cytometry data were obtained from two independent experiments for Expi293F and YTS and one experiment with all three cell lines.
Figure 3

Figure 3. NanoBondy covalent conjugation to recombinant CD45. (A) AlphaFold prediction of 2H5 nanobody (purple) interaction with CD45d1d2 (green). Magenta represents the site for a disulfide clamp, with lysines on CD45d1d2 colored pink and the terminal aspartate in cyan. (B) MBP fusion improved the CD45 gel-based analysis. CD45d consists of MBP fused to domains 1 and 2 of CD45. PNGase F digestion decreased heterogeneous mobility of CD45d1d2 and CD45d upon SDS-PAGE with Coomassie staining. (C) Individual protein components for the conjugation assay in (D). CD45d, sfGFP, ODC, and PNGase F were validated by SDS-PAGE/Coomassie staining. (D) Specificity of the NanoBondy reaction with recombinant CD45. Anti-CD45 2H5 R72C or anti-IgG NanoBondy-SPM was incubated with CD45d, each at 10.5 μM, for 1 h at 37 °C in HBS ± calcium, followed by SDS-PAGE with Coomassie staining. ODC, sfGFP, and anti-IgG NanoBondy were used as negative controls for reaction specificity. Hydroxylamine was used as a competing nucleophile to block reactivity. Colon represents a covalent conjugate. The experiment was conducted once.
Figure 4

Figure 4. Optimization of NanoBondy clamp site and linker length. (A) AlphaFold prediction of 2H5 NanoBondy (purple) bound to CD45d1d2 (green). The reactive anhydride is shown in cyan, alternative clamp sites are shown in magenta, and linkers are shown in orange. (B) Clamp-site variant reactivity. 2H5 NanoBondy variants were incubated with CD45d at 10.5 μM each for 1 h at 37 °C in HBS ± Ca2+, followed by SDS-PAGE with Coomassie staining. Anti-IgG NanoBondy was used as a negative control. The leftmost lane represents CD45d without PNGase F treatment. The experiment was conducted once. (C) Linker variant reactivity for 2H5 R72C anti-CD45 NanoBondy, analyzed as in (B). Representative gels were obtained from two independent experiments.
Figure 5
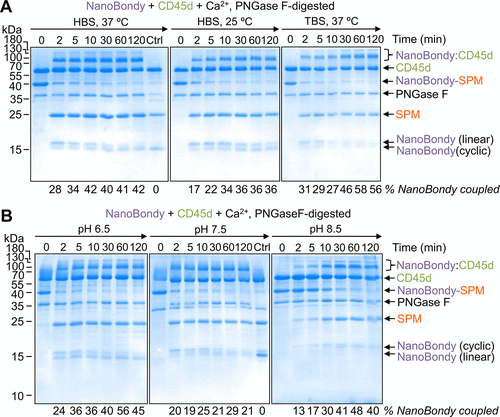
Figure 5. Condition-dependence of NanoBondy reaction. (A) Buffer- and temperature-dependence of NanoBondy reaction. 2H5 R72C anti-CD45 NanoBondy was incubated with CD45d, each at 10.5 μM, in the indicated buffer and temperature before SDS-PAGE with Coomassie staining. Ctrl refers to the lane containing anti-IgG NanoBondy with CD45d for 120 min. Representative gel from two independent experiments. (B) pH-dependence of NanoBondy reaction. The reaction was evaluated as in (A) with HBS-MES buffer at the indicated pH at 37 °C. Ctrl refers to the lane containing anti-IgG NanoBondy with CD45d for 120 min. The experiment was conducted once.
Figure 6

Figure 6. Mass spectrometry analysis of covalent conjugate between the NanoBondy and CD45d. (A) Dominant cross-linking sites. 2H5 R72C anti-CD45 NanoBondy was incubated with CD45d before cross-linking MS/MS. The number of identified cross-linked spectral matches for each NanoBondy cross-linking site on CD45 is shown. (B) Higher energy collision-induced dissociation (HCD) fragmentation spectrum of identified cross-link precursor ions corresponding to D173 (NanoBondy) to K131 (CD45d). Fragment ions matching fragmented cross-link (with cross-linker still intact) are annotated in bold, while peaks corresponding to fragments post-cross-link fragmentation are annotated with nonbold lines. “Car” indicates carbamidomethylation of cysteine. (C) Mapping of cross-link sites. AlphaFold prediction of NanoBondy (purple) bound to CD45d1d2 domains (green), highlighting cross-linking sites identified from the reactive aspartate (cyan) of the NanoBondy to target lysines (pink) or to serine (dark blue). Samples were run in technical triplicate. The cross-linking MS experiment was conducted twice.
Figure 7

Figure 7. NanoBondy covalent conjugation at the cell surface. (A) Western blotting of NanoBondy reaction at the cell surface. Anti-CD45 NanoBondy at 5 μM was incubated with YTS cells or Expi293F cells for 1 h at 37 °C ± calcium. Covalent conjugation was evaluated by Western blot using an anti-VHH polyclonal antibody to detect the NanoBondy. Anti-IgG NanoBondy or hydroxylamine to react with the anhydride provided negative controls. Western blot to glyceraldehyde-3-phosphate dehydrogenase (GAPDH) was the loading control. The complete GAPDH blot is presented in Figure S6A. The experiment was conducted once. (B) YTS cells were stained as in (A), except with 25 μM anti-CD45 NanoBondy, and analyzed by Western blot using an anti-CD45 antibody. The CD45 band demonstrates an upward shift upon covalent conjugation with the a-CD45 NanoBondy. The full-length GAPDH blot is presented in Figure S6B. The experiment was conducted once. (C) Western blotting of the DuoBondy reaction on CD8+ T cells. DuoBondy (WT) or DuoBondy (DA) at 1 μM was incubated with CD8+ T cells for 40 min at 37 °C ± calcium. Covalent conjugation was evaluated by Western blot using an anti-VHH polyclonal antibody to detect the DuoBondy. Representative blot from two independent experiments. (D) DuoBondy consists of a nanobody binder (Nb102c3) to PD-1 (dark blue) fused N-terminally to the established covalently reacting anti-CD45 NanoBondy (purple) with SPM in orange.
References
This article references 71 other publications.
- 1Keeble, A. H.; Howarth, M. Power to the Protein: Enhancing and Combining Activities Using the Spy Toolbox. Chem. Sci. 2020, 11 (28), 7281– 7291, DOI: 10.1039/D0SC01878CThere is no corresponding record for this reference.
- 2Oostindie, S. C.; Lazar, G. A.; Schuurman, J.; Parren, P. W. H. I. Avidity in Antibody Effector Functions and Biotherapeutic Drug Design. Nat. Rev. Drug Discovery 2022, 21 (10), 715– 735, DOI: 10.1038/s41573-022-00501-8There is no corresponding record for this reference.
- 3Khan, S. H.; Choi, Y.; Veena, M.; Lee, J. K.; Shin, D. S. Advances in CAR T Cell Therapy: Antigen Selection, Modifications, and Current Trials for Solid Tumors. Front. Immunol. 2025, 15, 1489827, DOI: 10.3389/fimmu.2024.1489827There is no corresponding record for this reference.
- 4Brenner, J. S.; Mitragotri, S.; Muzykantov, V. R. Red Blood Cell Hitchhiking: A Novel Approach for Vascular Delivery of Nanocarriers. Annu. Rev. Biomed. Eng. 2021, 23 (1), 225– 248, DOI: 10.1146/annurev-bioeng-121219-024239There is no corresponding record for this reference.
- 5Albelda, S. M. CAR T Cell Therapy for Patients with Solid Tumours: Key Lessons to Learn and Unlearn. Nat. Rev. Clin Oncol. 2024, 21 (1), 47– 66, DOI: 10.1038/s41571-023-00832-4There is no corresponding record for this reference.
- 6D’Aloia, M. M.; Zizzari, I. G.; Sacchetti, B.; Pierelli, L.; Alimandi, M. CAR-T Cells: The Long and Winding Road to Solid Tumors. Cell Death Dis. 2018, 9 (3), 282, DOI: 10.1038/s41419-018-0278-6There is no corresponding record for this reference.
- 7Guzman, G.; Reed, M. R.; Bielamowicz, K.; Koss, B.; Rodriguez, A. CAR-T Therapies in Solid Tumors: Opportunities and Challenges. Curr. Oncol. Rep. 2023, 25 (5), 479– 489, DOI: 10.1007/s11912-023-01380-xThere is no corresponding record for this reference.
- 8Hurton, L. V.; Singh, H.; Najjar, A. M.; Switzer, K. C.; Mi, T.; Maiti, S.; Olivares, S.; Rabinovich, B.; Huls, H.; Forget, M.-A. Tethered IL-15 Augments Antitumor Activity and Promotes a Stem-Cell Memory Subset in Tumor-Specific T Cells. Proc. Natl. Acad. Sci. U.S.A. 2016, 113 (48), E7788–E7797 DOI: 10.1073/pnas.1610544113There is no corresponding record for this reference.
- 9Jones, D. S.; Nardozzi, J. D.; Sackton, K. L.; Ahmad, G.; Christensen, E.; Ringgaard, L.; Chang, D.-K.; Jaehger, D. E.; Konakondla, J. V.; Wiinberg, M.; Stokes, K. L.; Pratama, A.; Sauer, K.; Andresen, T. L. Cell Surface–Tethered IL-12 Repolarizes the Tumor Immune Microenvironment to Enhance the Efficacy of Adoptive T Cell Therapy. Sci. Adv. 2022, 8 (17), eabi8075 DOI: 10.1126/sciadv.abi8075There is no corresponding record for this reference.
- 10Zhao, Y.; Dong, Y.; Yang, S.; Tu, Y.; Wang, C.; Li, J.; Yuan, Y.; Lian, Z. Bioorthogonal Equipping CAR-T Cells with Hyaluronidase and Checkpoint Blocking Antibody for Enhanced Solid Tumor Immunotherapy. ACS Cent. Sci. 2022, 8 (5), 603– 614, DOI: 10.1021/acscentsci.2c00163There is no corresponding record for this reference.
- 11Stephan, M.; Stephan, S.; Bak, P.; Chen, J.; Irvine, D. J. Synapse-Directed Delivery of Immunomodulators Using T-Cell-Conjugated Nanoparticles. Biomaterials 2012, 33 (23), 5776– 5787, DOI: 10.1016/j.biomaterials.2012.04.029There is no corresponding record for this reference.
- 12Luo, Y.; Chen, Z.; Sun, M.; Li, B.; Pan, F.; Ma, A.; Liao, J.; Yin, T.; Tang, X.; Huang, G.; Zhang, B.; Pan, H.; Zheng, M.; Cai, L. IL-12 Nanochaperone-Engineered CAR T Cell for Robust Tumor-Immunotherapy. Biomaterials 2022, 281, 121341, DOI: 10.1016/j.biomaterials.2021.121341There is no corresponding record for this reference.
- 13Liu, Y.; Adu-Berchie, K.; Brockman, J. M.; Pezone, M.; Zhang, D. K. Y.; Zhou, J.; Pyrdol, J. W.; Wang, H.; Wucherpfennig, K. W.; Mooney, D. J. Cytokine Conjugation to Enhance T Cell Therapy. Proc. Natl. Acad. Sci. U.S.A. 2023, 120 (1), e2213222120 DOI: 10.1073/pnas.2213222120There is no corresponding record for this reference.
- 14Zheng, Y.; Tang, L.; Mabardi, L.; Kumari, S.; Irvine, D. J. Enhancing Adoptive Cell Therapy of Cancer through Targeted Delivery of Small-Molecule Immunomodulators to Internalizing or Noninternalizing Receptors. ACS Nano 2017, 11 (3), 3089– 3100, DOI: 10.1021/acsnano.7b00078There is no corresponding record for this reference.
- 15Stephan, M.; Moon, J.; Um, S. H.; Bershteyn, A.; Irvine, D. Therapeutic Cell Engineering with Surface-Conjugated Synthetic Nanoparticles. Nat. Med. 2010, 16 (9), 1035– 1041, DOI: 10.1038/nm.2198There is no corresponding record for this reference.
- 16Tang, L.; Zheng, Y.; Melo, M. B.; Mabardi, L.; Castaño, A. P.; Xie, Y.-Q.; Li, N.; Kudchodkar, S. B.; Wong, H. C.; Jeng, E. K.; Maus, M. V.; Irvine, D. J. Enhancing T Cell Therapy through TCR-Signaling-Responsive Nanoparticle Drug Delivery. Nat. Biotechnol. 2018, 36 (8), 707– 716, DOI: 10.1038/nbt.4181There is no corresponding record for this reference.
- 17Los, G. V.; Encell, L. P.; McDougall, M. G.; Hartzell, D. D.; Karassina, N.; Zimprich, C.; Wood, M. G.; Learish, R.; Ohana, R. F.; Urh, M.; Simpson, D.; Mendez, J.; Zimmerman, K.; Otto, P.; Vidugiris, G.; Zhu, J.; Darzins, A.; Klaubert, D. H.; Bulleit, R. F.; Wood, K. V. HaloTag: A Novel Protein Labeling Technology for Cell Imaging and Protein Analysis. ACS Chem. Biol. 2008, 3 (6), 373– 382, DOI: 10.1021/cb800025kThere is no corresponding record for this reference.
- 18Keppler, A.; Gendreizig, S.; Gronemeyer, T.; Pick, H.; Vogel, H.; Johnsson, K. A General Method for the Covalent Labeling of Fusion Proteins with Small Molecules in Vivo. Nat. Biotechnol. 2003, 21 (1), 86– 89, DOI: 10.1038/nbt765There is no corresponding record for this reference.
- 19Sun, F.; Zhang, W.-B. Genetically Encoded Click Chemistry. Chin. J. Chem. 2020, 38 (8), 894– 896, DOI: 10.1002/cjoc.202000101There is no corresponding record for this reference.
- 20Humberg, C.; Yilmaz, Z.; Fitzian, K.; Dörner, W.; Kümmel, D.; Mootz, H. D. A Cysteine-Less and Ultra-Fast Split Intein Rationally Engineered from Being Aggregation-Prone to Highly Efficient in Protein Trans-Splicing. Nat. Commun. 2025, 16 (1), 2723, DOI: 10.1038/s41467-025-57596-xThere is no corresponding record for this reference.
- 21Mao, H.; Hart, S. A.; Schink, A.; Pollok, B. A. Sortase-Mediated Protein Ligation: A New Method for Protein Engineering. J. Am. Chem. Soc. 2004, 126 (9), 2670– 2671, DOI: 10.1021/ja039915eThere is no corresponding record for this reference.
- 22Liu, X.; Wen, J.; Yi, H.; Hou, X.; Yin, Y.; Ye, G.; Wu, X.; Jiang, X. Split Chimeric Antigen Receptor-Modified T Cells Targeting Glypican-3 Suppress Hepatocellular Carcinoma Growth with Reduced Cytokine Release. Ther. Adv. Med. Oncol. 2020, 12, 175883592091034, DOI: 10.1177/1758835920910347There is no corresponding record for this reference.
- 23Sutherland, A. R.; Owens, M. N.; Geyer, C. R. Modular Chimeric Antigen Receptor Systems for Universal CAR T Cell Retargeting. Int. J. Mol. Sci. 2020, 21 (19), 7222, DOI: 10.3390/ijms21197222There is no corresponding record for this reference.
- 24Minutolo, N. G.; Sharma, P.; Poussin, M.; Shaw, L. C.; Brown, D. P.; Hollander, E. E.; Smole, A.; Rodriguez-Garcia, A.; Hui, J. Z.; Zappala, F.; Tsourkas, A.; Powell, D. J. Quantitative Control of Gene-Engineered T-Cell Activity through the Covalent Attachment of Targeting Ligands to a Universal Immune Receptor. J. Am. Chem. Soc. 2020, 142 (14), 6554– 6568, DOI: 10.1021/jacs.9b11622There is no corresponding record for this reference.
- 25Rahikainen, R.; Rijal, P.; Tan, T. K.; Wu, H.; Andersson, A. C.; Barrett, J. R.; Bowden, T. A.; Draper, S. J.; Townsend, A. R.; Howarth, M. Overcoming Symmetry Mismatch in Vaccine Nanoassembly through Spontaneous Amidation. Angew. Chem. Int. Ed. 2021, 60 (1), 321– 330, DOI: 10.1002/anie.202009663There is no corresponding record for this reference.
- 26Bishani, A.; Chernolovskaya, E. L. Activation of Innate Immunity by Therapeutic Nucleic Acids. Int. J. Mol. Sci. 2021, 22 (24), 13360, DOI: 10.3390/ijms222413360There is no corresponding record for this reference.
- 27Bonifant, C. L.; Jackson, H. J.; Brentjens, R. J.; Curran, K. J. Toxicity and Management in CAR T-Cell Therapy. Mol. Ther. Oncolytics 2016, 3, 16011, DOI: 10.1038/mto.2016.11There is no corresponding record for this reference.
- 28Hamdy, N.; Goustin, A. S.; Desaulniers, J.-P.; Li, M.; Chow, C. S.; Al-Katib, A. Sheep Red Blood Cells Armed with Anti-CD20 Single-Chain Variable Fragments (scFvs) Fused to a Glycosylphosphatidylinositol (GPI) Anchor: A Strategy to Target CD20-Positive Tumor Cells. J. Immunol. Methods 2005, 297 (1–2), 109– 124, DOI: 10.1016/j.jim.2004.12.003There is no corresponding record for this reference.
- 29White, S.; Vonheijne, G. Transmembrane Helices before, during, and after Insertion. Curr. Opin. Struct. Biol. 2005, 15 (4), 378– 386, DOI: 10.1016/j.sbi.2005.07.004There is no corresponding record for this reference.
- 30Reinisch, K. M.; Prinz, W. A. Mechanisms of Nonvesicular Lipid Transport. J. Cell Biol. 2021, 220 (3), e202012058 DOI: 10.1083/jcb.202012058There is no corresponding record for this reference.
- 31Yu, B.; Li, S.; Tabata, T.; Wang, N.; Cao, L.; Kumar, G. R.; Sun, W.; Liu, J.; Ott, M.; Wang, L. Accelerating PERx Reaction Enables Covalent Nanobodies for Potent Neutralization of SARS-CoV-2 and Variants. Chem 2022, 8 (10), 2766– 2783, DOI: 10.1016/j.chempr.2022.07.012There is no corresponding record for this reference.
- 32Yu, B.; Cao, L.; Li, S.; Klauser, P. C.; Wang, L. The Proximity-Enabled Sulfur Fluoride Exchange Reaction in the Protein Context. Chem. Sci. 2023, 14 (29), 7913– 7921, DOI: 10.1039/D3SC01921GThere is no corresponding record for this reference.
- 33Li, Q.; Chen, Q.; Klauser, P. C.; Li, M.; Zheng, F.; Wang, N.; Li, X.; Zhang, Q.; Fu, X.; Wang, Q. Developing Covalent Protein Drugs via Proximity-Enabled Reactive Therapeutics. Cell 2020, 182 (1), P85– 97.E16, DOI: 10.1016/j.cell.2020.05.028There is no corresponding record for this reference.
- 34De Bank, P. A.; Kellam, B.; Kendall, D. A.; Shakesheff, K. M. Surface Engineering of Living Myoblasts via Selective Periodate Oxidation. Biotechnol. Bioeng. 2003, 81 (7), 800– 808, DOI: 10.1002/bit.10525There is no corresponding record for this reference.
- 35Glassman, P. M.; Hood, E. D.; Ferguson, L. T.; Zhao, Z.; Siegel, D. L.; Mitragotri, S.; Brenner, J. S.; Muzykantov, V. R. Red Blood Cells: The Metamorphosis of a Neglected Carrier into the Natural Mothership for Artificial Nanocarriers. Adv. Drug Delivery Rev. 2021, 178, 113992, DOI: 10.1016/j.addr.2021.113992There is no corresponding record for this reference.
- 36Bao, G.; Tang, M.; Zhao, J.; Zhu, X. Nanobody: A Promising Toolkit for Molecular Imaging and Disease Therapy. EJNMMI Res. 2021, 11 (1), 6, DOI: 10.1186/s13550-021-00750-5There is no corresponding record for this reference.
- 37Asaadi, Y.; Jouneghani, F. F.; Janani, S.; Rahbarizadeh, F. A Comprehensive Comparison between Camelid Nanobodies and Single Chain Variable Fragments. Biomark. Res. 2021, 9 (1), 87, DOI: 10.1186/s40364-021-00332-6There is no corresponding record for this reference.
- 38Tack, L.; Miret-Casals, L.; Zwaenepoel, O.; Cadoni, E.; Van Troys, M.; Gettemans, J.; Madder, A. Development of “Furan Warhead”-Equipped Antagonistic Nanobodies for Covalent Cross-Linking to the Epidermal Growth Factor Receptor. Bioconjugate Chem. 2025, 36 (8), 1649– 1660, DOI: 10.1021/acs.bioconjchem.5c00144There is no corresponding record for this reference.
- 39Gao, W.; Cho, E.; Liu, Y.; Lu, Y. Advances and Challenges in Cell-Free Incorporation of Unnatural Amino Acids Into Proteins. Front. Pharmacol. 2019, 10, 611, DOI: 10.3389/fphar.2019.00611There is no corresponding record for this reference.
- 40Scheu, A.; Lim, S. Y. T.; Metzner, F. J.; Mohammed, S.; Howarth, M. NeissLock Provides an Inducible Protein Anhydride for Covalent Targeting of Endogenous Proteins. Nat. Commun. 2021, 12 (1), 717, DOI: 10.1038/s41467-021-20963-5There is no corresponding record for this reference.
- 41Lim, S.; Keeble, A. H.; Howarth, M. R. Split NeissLock with Spy-Acceleration Arms Mammalian Proteins for Anhydride-Mediated Cell Ligation. ACS Chem. Biol. 2025, 20, 2475, DOI: 10.1021/acschembio.5c00515There is no corresponding record for this reference.
- 42Sviridova, E.; Rezacova, P.; Bondar, A.; Veverka, V.; Novak, P.; Schenk, G.; Svergun, D. I.; Kuta Smatanova, I.; Bumba, L. Structural Basis of the Interaction between the Putative Adhesion-Involved and Iron-Regulated FrpD and FrpC Proteins of Neisseria Meningitidis. Sci. Rep. 2017, 7 (1), 40408, DOI: 10.1038/srep40408There is no corresponding record for this reference.
- 43Borowska, M. T.; Liu, L. D.; Caveney, N. A.; Jude, K. M.; Kim, W.-J.; Masubuchi, T.; Hui, E.; Majzner, R. G.; Garcia, K. C. Orientation-Dependent CD45 Inhibition with Viral and Engineered Ligands. Sci. Immunol. 2024, 9 (100), eadp0707 DOI: 10.1126/sciimmunol.adp0707There is no corresponding record for this reference.
- 44Rokkam, D.; Lupardus, P. J. Discovery and Characterization of Llama VHH Targeting the RO Form of Human CD45. bioRxiv . 2020. DOI: 10.1101/2020.09.01.278853There is no corresponding record for this reference.
- 45Keeble, A. H.; Turkki, P.; Stokes, S.; Khairil Anuar, I. N. A.; Rahikainen, R.; Hytönen, V. P.; Howarth, M. Approaching Infinite Affinity through Engineering of Peptide–Protein Interaction. Proc. Natl. Acad. Sci. U.S.A. 2019, 116 (52), 26523– 26533, DOI: 10.1073/pnas.1909653116There is no corresponding record for this reference.
- 46Vester, S. K.; Rahikainen, R.; Khairil Anuar, I. N. A.; Hills, R. A.; Tan, T. K.; Howarth, M. SpySwitch Enables pH- or Heat-Responsive Capture and Release for Plug-and-Display Nanoassembly. Nat. Commun. 2022, 13 (1), 3714, DOI: 10.1038/s41467-022-31193-8There is no corresponding record for this reference.
- 47Bryant, P.; Pozzati, G.; Elofsson, A. Improved Prediction of Protein-Protein Interactions Using AlphaFold2. Nat. Commun. 2022, 13 (1), 1265, DOI: 10.1038/s41467-022-28865-wThere is no corresponding record for this reference.
- 48Evans, R.; O’Neill, M.; Pritzel, A.; Antropova, N.; Senior, A.; Green, T.; Žídek, A.; Bates, R.; Blackwell, S.; Yim, J. Protein Complex Prediction With AlphaFold-Multimer. bioRxiv . 2022. DOI: 10.1101/2021.10.04.463034There is no corresponding record for this reference.
- 49Zhong, B.; Su, X.; Wen, M.; Zuo, S.; Hong, L.; Lin, J. ParaFold: Paralleling AlphaFold For Large-Scale Predictions. arXiv . 2021. DOI: 10.48550/arXiv.2111.06340There is no corresponding record for this reference.
- 50Breitwieser, G. E. Extracellular Calcium as an Integrator of Tissue Function. Int. J. Biochem. Cell Biol. 2008, 40 (8), 1467– 1480, DOI: 10.1016/j.biocel.2008.01.019There is no corresponding record for this reference.
- 51Hermiston, M. L.; Xu, Z.; Weiss, A. CD45: A Critical Regulator of Signaling Thresholds in Immune Cells. Annu. Rev. Immunol. 2003, 21 (1), 107– 137, DOI: 10.1146/annurev.immunol.21.120601.140946There is no corresponding record for this reference.
- 52Savchenko, A. A.; Tikhonova, E.; Kudryavtsev, I.; Kudlay, D.; Korsunsky, I.; Beleniuk, V.; Borisov, A. TREC/KREC Levels and T and B Lymphocyte Subpopulations in COVID-19 Patients at Different Stages of the Disease. Viruses 2022, 14 (3), 646, DOI: 10.3390/v14030646There is no corresponding record for this reference.
- 53Kalinin, R. S.; Ukrainskaya, V. M.; Chumakov, S. P.; Moysenovich, A. M.; Tereshchuk, V. M.; Volkov, D. V.; Pershin, D. S.; Maksimov, E. G.; Zhang, H.; Maschan, M. A.; Rubtsov, Y. P.; Stepanov, A. V. Engineered Removal of PD-1 From the Surface of CD19 CAR-T Cells Results in Increased Activation and Diminished Survival. Front. Mol. Biosci. 2021, 8, 745286, DOI: 10.3389/fmolb.2021.745286There is no corresponding record for this reference.
- 54Fernandes, R. A.; Su, L.; Nishiga, Y.; Ren, J.; Bhuiyan, A. M.; Cheng, N.; Kuo, C. J.; Picton, L. K.; Ohtsuki, S.; Majzner, R. G.; Rietberg, S. P.; Mackall, C. L.; Yin, Q.; Ali, L. R.; Yang, X.; Savvides, C. S.; Sage, J.; Dougan, M.; Garcia, K. C. Immune Receptor Inhibition through Enforced Phosphatase Recruitment. Nature 2020, 586 (7831), 779– 784, DOI: 10.1038/s41586-020-2851-2There is no corresponding record for this reference.
- 55Fernández-Quintero, M. L.; Kokot, J.; Waibl, F.; Fischer, A.-L. M.; Quoika, P. K.; Deane, C. M.; Liedl, K. R. Challenges in Antibody Structure Prediction. mAbs 2023, 15 (1), 2175319, DOI: 10.1080/19420862.2023.2175319There is no corresponding record for this reference.
- 56Erasmus, M. F.; Spector, L.; Ferrara, F.; DiNiro, R.; Pohl, T. J.; Perea-Schmittle, K.; Wang, W.; Tessier, P. M.; Richardson, C.; Turner, L.; Kumar, S.; Bedinger, D.; Sormanni, P.; Fernández-Quintero, M. L.; Ward, A. B.; Loeffler, J. R.; Swanson, O. M.; Deane, C. M.; Raybould, M. I. J.; Evers, A.; Sellmann, C.; Bachas, S.; Ruffolo, J.; Nastri, H. G.; Ramesh, K.; Sørensen, J.; Croasdale-Wood, R.; Hijano, O.; Leal-Lopes, C.; Shahsavarian, M.; Qiu, Y.; Marcatili, P.; Vernet, E.; Akbar, R.; Friedensohn, S.; Wagner, R.; Kurella, V. B.; Malhotra, S.; Kumar, S.; Kidger, P.; Almagro, J. C.; Furfine, E.; Stanton, M.; Graff, C. P.; Villalba, S. D.; Tomszak, F.; Teixeira, A. A. R.; Hopkins, E.; Dovner, M.; D’Angelo, S.; Bradbury, A. R. M. AIntibody: An Experimentally Validated in Silico Antibody Discovery Design Challenge. Nat. Biotechnol. 2024, 42 (11), 1637– 1642, DOI: 10.1038/s41587-024-02469-9There is no corresponding record for this reference.
- 57Holm, L.; Moody, P.; Howarth, M. Electrophilic Affibodies Forming Covalent Bonds to Protein Targets. J. Biol. Chem. 2009, 284 (47), 32906– 32913, DOI: 10.1074/jbc.M109.034322There is no corresponding record for this reference.
- 58De La Torre, D.; Chin, J. W. Reprogramming the Genetic Code. Nat. Rev. Genet. 2021, 22 (3), 169– 184, DOI: 10.1038/s41576-020-00307-7There is no corresponding record for this reference.
- 59Zhang, H.; Han, Y.; Yang, Y.; Lin, F.; Li, K.; Kong, L.; Liu, H.; Dang, Y.; Lin, J.; Chen, P. R. Covalently Engineered Nanobody Chimeras for Targeted Membrane Protein Degradation. J. Am. Chem. Soc. 2021, 143 (40), 16377– 16382, DOI: 10.1021/jacs.1c08521There is no corresponding record for this reference.
- 60Tyrrell, R. M. Induction of Pyrimidine Dimers In Bacterial DNA by 365 nm Radiation. Photochem. Photobiol. 1973, 17 (1), 69– 73, DOI: 10.1111/j.1751-1097.1973.tb06334.xThere is no corresponding record for this reference.
- 61Andley, U. P.; Lewis, R. M.; Reddan, J. R.; Kochevar, I. E. Action Spectrum for Cytotoxicity in the UVA- and UVB-Wavelength Region in Cultured Lens Epithelial Cells. Invest. Ophthalmol. Vis. Sci. 1994, 35 (2), 367– 373,There is no corresponding record for this reference.
- 62Veggiani, G.; Nakamura, T.; Brenner, M. D.; Gayet, R. V.; Yan, J.; Robinson, C. V.; Howarth, M. Programmable Polyproteams Built Using Twin Peptide Superglues. Proc. Natl. Acad. Sci. U.S.A. 2016, 113 (5), 1202– 1207, DOI: 10.1073/pnas.1519214113There is no corresponding record for this reference.
- 63Zimmermann, I.; Egloff, P.; Hutter, C. A. J.; Kuhn, B. T.; Bräuer, P.; Newstead, S.; Dawson, R. J. P.; Geertsma, E. R.; Seeger, M. A. Generation of Synthetic Nanobodies against Delicate Proteins. Nat. Protoc. 2020, 15 (5), 1707– 1741, DOI: 10.1038/s41596-020-0304-xThere is no corresponding record for this reference.
- 64Kunz, P.; Zinner, K.; Mücke, N.; Bartoschik, T.; Muyldermans, S.; Hoheisel, J. D. The Structural Basis of Nanobody Unfolding Reversibility and Thermoresistance. Sci. Rep. 2018, 8 (1), 7934, DOI: 10.1038/s41598-018-26338-zThere is no corresponding record for this reference.
- 65Berkmen, M. Production of Disulfide-Bonded Proteins in Escherichia Coli. Protein Expression Purif. 2012, 82 (1), 240– 251, DOI: 10.1016/j.pep.2011.10.009There is no corresponding record for this reference.
- 66Gebauer, M.; Skerra, A. Engineered Protein Scaffolds as Next-Generation Therapeutics. Annu. Rev. Pharmacol. Toxicol. 2020, 60 (1), 391– 415, DOI: 10.1146/annurev-pharmtox-010818-021118There is no corresponding record for this reference.
- 67Liang, Y.; Xu, Q.; Gao, Q. Advancing CAR-Based Immunotherapies in Solid Tumors: CAR- Macrophages and Neutrophils. Front. Immunol. 2023, 14, 1291619, DOI: 10.3389/fimmu.2023.1291619There is no corresponding record for this reference.
- 68Wang, X.-W.; Zhang, W.-B. Chemical Topology and Complexity of Protein Architectures. Trends Biochem. Sci. 2018, 43 (10), 806– 817, DOI: 10.1016/j.tibs.2018.07.001There is no corresponding record for this reference.
- 69Pedram, K.; Shon, D. J.; Tender, G. S.; Mantuano, N. R.; Northey, J. J.; Metcalf, K. J.; Wisnovsky, S. P.; Riley, N. M.; Forcina, G. C.; Malaker, S. A.; Kuo, A.; George, B. M.; Miller, C. L.; Casey, K. M.; Vilches-Moure, J. G.; Ferracane, M. J.; Weaver, V. M.; Läubli, H.; Bertozzi, C. R. Design of a Mucin-Selective Protease for Targeted Degradation of Cancer-Associated Mucins. Nat. Biotechnol. 2024, 42 (4), 597– 607, DOI: 10.1038/s41587-023-01840-6There is no corresponding record for this reference.
- 70Eichhoff, A. M.; Börner, K.; Albrecht, B.; Schäfer, W.; Baum, N.; Haag, F.; Körbelin, J.; Trepel, M.; Braren, I.; Grimm, D. Nanobody-Enhanced Targeting of AAV Gene Therapy Vectors. Mol. Ther.–Methods Clin. Dev. 2019, 15, P211– 220, DOI: 10.1016/j.omtm.2019.09.003There is no corresponding record for this reference.
- 71Bocancia-Mateescu, L.-A.; Stan, D.; Mirica, A.-C.; Ghita, M. G.; Stan, D.; Ruta, L. L. Nanobodies as Diagnostic and Therapeutic Tools for Cardiovascular Diseases (CVDs). Pharmaceuticals 2023, 16 (6), 863, DOI: 10.3390/ph16060863There is no corresponding record for this reference.
Supporting Information
Supporting Information
The Supporting Information is available free of charge at https://pubs.acs.org/doi/10.1021/acs.bioconjchem.5c00519.
Comprehensive methods and supplementary results, including amino acid sequence, MS, structure prediction, and fluorescence microscopy (PDF)
Terms & Conditions
Most electronic Supporting Information files are available without a subscription to ACS Web Editions. Such files may be downloaded by article for research use (if there is a public use license linked to the relevant article, that license may permit other uses). Permission may be obtained from ACS for other uses through requests via the RightsLink permission system: http://pubs.acs.org/page/copyright/permissions.html.



