Idler Compounds: A Simple Protocol for Openly Sharing Fridge Contents for Cross-ScreeningClick to copy article linkArticle link copied!
- Rebecka IsakssonRebecka IsakssonSchool of Pharmacy and Structural Genomics Consortium, University College London, 29-39 Brunswick Square, London WC1N 1AX, U.K.More by Rebecka Isaksson
- Eve M. CarterEve M. CarterSchool of Pharmacy and Structural Genomics Consortium, University College London, 29-39 Brunswick Square, London WC1N 1AX, U.K.More by Eve M. Carter
- Charlotte K. HindCharlotte K. HindAntimicrobial Discovery, Development and Diagnostics, Countermeasures, Development, Evaluation and Preparedness, UK Health Security Agency, Porton Down, Salisbury SP4 0JG, U.K.More by Charlotte K. Hind
- J. Mark SuttonJ. Mark SuttonAntimicrobial Discovery, Development and Diagnostics, Countermeasures, Development, Evaluation and Preparedness, UK Health Security Agency, Porton Down, Salisbury SP4 0JG, U.K.More by J. Mark Sutton
- Hazel RudgyardHazel RudgyardDiscovery Group, Countermeasures, Development, Evaluation and Preparedness, UK Health Security Agency, Porton Down, Salisbury SP4 0JG, U.K.More by Hazel Rudgyard
- Adam H. RobertsAdam H. RobertsDiscovery Group, Countermeasures, Development, Evaluation and Preparedness, UK Health Security Agency, Porton Down, Salisbury SP4 0JG, U.K.More by Adam H. Roberts
- Christopher W. MoonChristopher W. MoonDiscovery Group, Countermeasures, Development, Evaluation and Preparedness, UK Health Security Agency, Porton Down, Salisbury SP4 0JG, U.K.More by Christopher W. Moon
- Yinuo WangYinuo WangSchool of Pharmacy and Structural Genomics Consortium, University College London, 29-39 Brunswick Square, London WC1N 1AX, U.K.More by Yinuo Wang
- Sandra CodonySandra CodonyInstitute of Biomedicine of the University of Barcelona (IBUB), Universitat de Barcelona, Barcelona 08028, SpainLaboratori de Química Farmacèutica, Facultat de Farmàcia i Ciències de l’Alimentació, Universitat de Barcelona, Av. Joan XXIII, 27-31, Barcelona 08028, SpainMore by Sandra Codony
- Antón L. MartínezAntón L. MartínezInnopharma Drug Screening Platform/Biofarma Research Group, CIMUS Research Center, Pharmacology Department, School of Pharmacy, University of Santiago de Compostela (USC) and Instituto de Investigación Sanitaria de Santiago de Compostela (IDIS), Santiago de Compostela 15782, SpainMore by Antón L. Martínez
- Joanna BaconJoanna BaconDiscovery Group, Countermeasures, Development, Evaluation and Preparedness, UK Health Security Agency, Porton Down, Salisbury SP4 0JG, U.K.More by Joanna Bacon
- Matthew H. Todd*Matthew H. Todd*Email: [email protected]School of Pharmacy and Structural Genomics Consortium, University College London, 29-39 Brunswick Square, London WC1N 1AX, U.K.More by Matthew H. Todd
Abstract
Academic drug discovery laboratories tend to accumulate collections of compounds with great potential value that merely reside in fridges and freezers. Cross screening these libraries against alternative targets holds significant potential for uncovering novel hits, but in the academic setting compound collections are rarely used, and shared, in this way. We present a short guide for collecting small molecules not being actively pursued in group projects (which we term “idlers”) to establish an open compound library. We describe how a diverse subset of this library was screened against a panel of pathogens, with the resulting data made publicly available. We hope to encourage other academic groups to develop and share their own libraries of idlers, thereby maximizing the utility of existing resources, enabling new insights, and catalyzing novel research directions through open science.
This publication is licensed under
License Summary*
You are free to share(copy and redistribute) this article in any medium or format and to adapt(remix, transform, and build upon) the material for any purpose, even commercially within the parameters below:
Creative Commons (CC): This is a Creative Commons license.
Attribution (BY): Credit must be given to the creator.
*Disclaimer
This summary highlights only some of the key features and terms of the actual license. It is not a license and has no legal value. Carefully review the actual license before using these materials.
License Summary*
You are free to share(copy and redistribute) this article in any medium or format and to adapt(remix, transform, and build upon) the material for any purpose, even commercially within the parameters below:
Creative Commons (CC): This is a Creative Commons license.
Attribution (BY): Credit must be given to the creator.
*Disclaimer
This summary highlights only some of the key features and terms of the actual license. It is not a license and has no legal value. Carefully review the actual license before using these materials.
License Summary*
You are free to share(copy and redistribute) this article in any medium or format and to adapt(remix, transform, and build upon) the material for any purpose, even commercially within the parameters below:
Creative Commons (CC): This is a Creative Commons license.
Attribution (BY): Credit must be given to the creator.
*Disclaimer
This summary highlights only some of the key features and terms of the actual license. It is not a license and has no legal value. Carefully review the actual license before using these materials.
1. Introduction
1.1. Library and Plate Design
Figure 1
Figure 1. Overview of library setup and the sharing process. Compounds were synthesized or purchased for a range of projects in the research group. Idler compounds were then prepared as stock solutions and added to an online library. A subset was selected, plated and shared. A detailed step-by-step guide is provided in the Supporting Information.
1.2. Building the Library and Online Resources
1.3. Sharing Compounds from the Library
Figure 2
Figure 2. Summary of the selection process for the 96-well plate to ensure compound diversity; researchers selected 56 compounds with the remaining 40 compounds selected computationally to ensure diversity as outlined in the Supporting Information.
1.4. Composition of the Compounds in the First Plate
Figure 3
Figure 3. Targets for which the compounds on plate MHT-0001 were originally intended. The shared plate contained compounds from various projects focusing on viral (60% of compounds), parasitic (25%), bacterial (9%), and human (6%) targets; most projects were target-based rather than phenotypic.
Figure 4
Figure 4. Compounds of plate MHT-0001 were selected to be structurally diverse. A range of functional groups is represented, and most compounds align with desirable values of descriptors commonly used in medicinal chemistry programmes.
1.5. Where to Send Idler Compounds
2. Results and Discussion
Figure 5
Figure 5. (A) Comparison of antimicrobial activity against A. baumannii (∓PMBN) and E. coli (∓PMBN). Compounds are reported as active when inhibition is >80% at 100 μM compound concentration. (B) Chemical structure and antimicrobial activity of WEL2.76.
Figure 6
Figure 6. (A) Antimicrobial activity against S. aureus, A. baumannii (∓PMBN), E. coli (∓PMBN), C. albicans, M. avium, and M. abscessus. Compounds are reported as active when inhibition is >80% at 100 μM compound concentration (S. aureus, A. baumannii (∓PMBN), E. coli (∓PMBN), C. albicans) and >50% inhibition at 10 μM compound concentration (M. avium, and M. abscessus). (B) Solubility of the compounds. (C) Chemical structures of the highlighted compounds.
3. Conclusions
4. Experimental Section
4.1. Assay to Test Activity against S. aureus, A. baumannii, and E. coli
4.2. Assay to Test Activity against C. albicans
4.3. Assay to Test Activity against Mycobacterium abscessus and Mycobacterium avium
4.4. Hemolysis
4.5. Solubility
Supporting Information
The Supporting Information is available free of charge at https://pubs.acs.org/doi/10.1021/acs.jmedchem.5c02354.
A guide of library generation, links to additional online material, chemistry synthesis information and supporting biological protocols (PDF)
Compound ID names and SMILES (CSV)
The assay results including compound details (SMILES, molecular weight), the structural motif analysis of the selected compounds, and the 2D and 3D descriptors of the compounds (XLSX)
Terms & Conditions
Most electronic Supporting Information files are available without a subscription to ACS Web Editions. Such files may be downloaded by article for research use (if there is a public use license linked to the relevant article, that license may permit other uses). Permission may be obtained from ACS for other uses through requests via the RightsLink permission system: http://pubs.acs.org/page/copyright/permissions.html.
Acknowledgments
The authors would like to extend our heartfelt thanks to Prof. Dr Santi Vázquez at the Laboratorio de Química Farmacéutica e Instituto de Biomedicina (IBUB) at Universidad de Barcelona, and Dr Chris Southan for fruitful discussions. The UKHSA Open Innovation Platform would like to thank Grant-in-aid funding (Project 113361). Funding for compounds evaluated as direct acting antivirals was provided by NIH grant 1U19AI171292-01 (READDI-AViDD Center). Other compounds were developed with funding from The Maplethorpe Fellowship, BIO Ventures for Global Health and the National Council for Scientific and Technological Development (CNPq, grant number 142488/2020-8 to Wellington da Silva) and the São Paulo Research Foundation (FAPESP, grant numbers 2022/03731-5, and 2023/03295-3 to Wellington da Silva; 2023/00383-9 to Carlos Roque Duarte Correia). We would like to acknowledge the support of the University College London School of Pharmacy Nuclear Magnetic Resonance Core Facility (RRID: SCR_027123). This work, integrated into the framework of PERTE for Cutting-edge Health, has been cofinanced by the Spanish Ministry of Science, Innovation and Universities with funds from the European Union NextGenerationEU, from the Recovery, Transformation and Resilience Plan (PRTR-C17.I1) and from the Autonomous Community of Galicia within the framework of the Biotechnology Plan Applied to Health. The Todd Group Researchers who contributed to this work are (in alphabetical order): Mohsen Alamoudi, Hadia Almahli, Guilherme Fernandes, Kangping Liu, Álvaro F. Magalhães, Evans Mainsah, Xin Qiu, Rahman Shah Zaib Saleem, Wellington da Silva, and Yuhang Wang.
| CAMHB | cation-adjusted Mueller Hinton Broth |
| CC-BY | Creative Commons Attribution License |
| CFU | colony-forming unit |
| CO-ADD | community for open antimicrobial drug discovery |
| CRO | Contract Research Organization |
| DPBS | Dulbecco’s phosphate-buffered saline |
| ELF | European Lead Factory |
| ELN | electronic laboratory notebook |
| EUCAST | European Committee on Antimicrobial Susceptibility Testing |
| FAIR | findable, accessible, interoperable and reusable |
| GSH | glutathione |
| NGO | nongovernmental organization |
| Nsp | nonstructural protein |
| OADC | oleic acid, albumin, dextrose, and catalase |
| PMBN | polymyxin b nonapeptide |
| SARS-CoV-2 | severe acute respiratory syndrome coronavirus 2 |
| TPSA | topological polar surface area |
| TSA | tryptic soy agar |
| UKHSA | UK Health Security Agency |
References
This article references 49 other publications.
- 1Duffy, S.; Sykes, M. L.; Jones, A. J.; Shelper, T. B.; Simpson, M.; Lang, R.; Poulsen, S.-A.; Sleebs, B. E.; Avery, V. M. Screening the Medicines for Malaria Venture Pathogen Box across Multiple Pathogens Reclassifies Starting Points for Open-Source Drug Discovery. Antimicrob. Agents Chemother. 2017, 61 (9), e0037917 DOI: 10.1128/AAC.00379-17Google ScholarThere is no corresponding record for this reference.
- 2Spangenberg, T.; Burrows, J. N.; Kowalczyk, P.; McDonald, S.; Wells, T. N. C.; Willis, P. The Open Access Malaria Box: A Drug Discovery Catalyst for Neglected Diseases. PLoS One 2013, 8 (6), e62906 DOI: 10.1371/journal.pone.0062906Google ScholarThere is no corresponding record for this reference.
- 3Adam, A.; Besson, D.; Bryant, R.; Rees, S.; Willis, P. A.; Burrows, J. N.; Hooft van Huisjduijnen, R.; Laleu, B.; Norton, L.; Canan, S.; Hawryluk, N.; Robinson, D.; Palmer, M.; Samby, K. K. Global Health Priority Box–Proactive Pandemic Preparedness. ACS Infect. Dis. 2024, 10 (12), 4030– 4039, DOI: 10.1021/acsinfecdis.4c00700Google ScholarThere is no corresponding record for this reference.
- 4Tredup, C.; Ndreshkjana, B.; Schneider, N. S.; Tjaden, A.; Kemas, A. M.; Youhanna, S.; Lauschke, V. M.; Berger, B.-T.; Krämer, A.; Berger, L. M.; Röhm, S.; Knapp, S.; Farin, H. F.; Müller, S. Deep Annotation of Donated Chemical Probes (DCP) in Organotypic Human Liver Cultures and Patient-Derived Organoids from Tumor and Normal Colorectum. ACS Chem. Biol. 2023, 18 (4), 822– 836, DOI: 10.1021/acschembio.2c00877Google ScholarThere is no corresponding record for this reference.
- 5Müller, S.; Ackloo, S.; Arrowsmith, C. H.; Bauser, M.; Baryza, J. L.; Blagg, J.; Böttcher, J.; Bountra, C.; Brown, P. J.; Bunnage, M. E.; Carter, A. J.; Damerell, D.; Dötsch, V.; Drewry, D. H.; Edwards, A. M.; Edwards, J.; Elkins, J. M.; Fischer, C.; Frye, S. V.; Gollner, A.; Grimshaw, C. E.; IJzerman, A.; Hanke, T.; Hartung, I. V.; Hitchcock, S.; Howe, T.; Hughes, T. V.; Laufer, S.; Li, V. M.; Liras, S.; Marsden, B. D.; Matsui, H.; Mathias, J.; O’Hagan, R. C.; Owen, D. R.; Pande, V.; Rauh, D.; Rosenberg, S. H.; Roth, B. L.; Schneider, N. S.; Scholten, C.; Singh Saikatendu, K.; Simeonov, A.; Takizawa, M.; Tse, C.; Thompson, P. R.; Treiber, D. K.; Viana, A. Y.; Wells, C. I.; Willson, T. M.; Zuercher, W. J.; Knapp, S.; Mueller-Fahrnow, A. Donated Chemical Probes for Open Science. Elife 2018, 7, e34311 DOI: 10.7554/eLife.34311Google ScholarThere is no corresponding record for this reference.
- 6Wells, C. I.; Al-Ali, H.; Andrews, D. M.; Asquith, C. R. M.; Axtman, A. D.; Dikic, I.; Ebner, D.; Ettmayer, P.; Fischer, C.; Frederiksen, M.; Futrell, R. E.; Gray, N. S.; Hatch, S. B.; Knapp, S.; Lücking, U.; Michaelides, M.; Mills, C. E.; Müller, S.; Owen, D.; Picado, A.; Saikatendu, K. S.; Schröder, M.; Stolz, A.; Tellechea, M.; Turunen, B. J.; Vilar, S.; Wang, J.; Zuercher, W. J.; Willson, T. M.; Drewry, D. H. The Kinase Chemogenomic Set (KCGS): An Open Science Resource for Kinase Vulnerability Identification. Int. J. Mol. Sci. 2021, 22 (2), 566, DOI: 10.3390/ijms22020566Google ScholarThere is no corresponding record for this reference.
- 7AstraZeneca. Open Innovation. https://openinnovation.astrazeneca.com/(accessed March 4, 2025).Google ScholarThere is no corresponding record for this reference.
- 8European Lead Factory. https://www.europeanleadfactory.eu/(accessed April 10, 2025).Google ScholarThere is no corresponding record for this reference.
- 9Griffith University. Compounds Australia. https://www.griffith.edu.au/griffith-sciences/compounds-australia (accessed June 12, 2025).Google ScholarThere is no corresponding record for this reference.
- 10Todd Group Library Spreadsheet. https://tinyurl.com/yynht8mu (accessed June 12, 2025).Google ScholarThere is no corresponding record for this reference.
- 11Todd Group Library LabArchive. https://tinyurl.com/3jf88u32 (accessed June 12, 2025).Google ScholarThere is no corresponding record for this reference.
- 12LabArchives. https://www.labarchives.com/(accessed April 10, 2025).Google ScholarThere is no corresponding record for this reference.
- 13Wilkinson, M. D.; Dumontier, M.; Aalbersberg, Ij. J.; Appleton, G.; Axton, M.; Baak, A.; Blomberg, N.; Boiten, J.-W.; da Silva Santos, L. B.; Bourne, P. E.; Bouwman, J.; Brookes, A. J.; Clark, T.; Crosas, M.; Dillo, I.; Dumon, O.; Edmunds, S.; Evelo, C. T.; Finkers, R.; Gonzalez-Beltran, A.; Gray, A. J. G.; Groth, P.; Goble, C.; Grethe, J. S.; Heringa, J.; ’t Hoen, P. A.; Hooft, R.; Kuhn, T.; Kok, R.; Kok, J.; Lusher, S. J.; Martone, M. E.; Mons, A.; Packer, A. L.; Persson, B.; Rocca-Serra, P.; Roos, M.; van Schaik, R.; Sansone, S.-A.; Schultes, E.; Sengstag, T.; Slater, T.; Strawn, G.; Swertz, M. A.; Thompson, M.; van der Lei, J.; van Mulligen, E.; Velterop, J.; Waagmeester, A.; Wittenburg, P.; Wolstencroft, K.; Zhao, J.; Mons, B. The FAIR Guiding Principles for Scientific Data Management and Stewardship. Sci. Data 2016, 3 (1), 160018, DOI: 10.1038/sdata.2016.18Google ScholarThere is no corresponding record for this reference.
- 14Creative Commons. Attribution 4.0 International. https://creativecommons.org/licenses/by/4.0/(accessed May 13, 2025).Google ScholarThere is no corresponding record for this reference.
- 15Bloodworth, S.; Willoughby, C.; Coles, S. J. Data Accessibility in the Chemical Sciences: An Analysis of Recent Practice in Organic Chemistry Journals. Beilstein J. Org. Chem. 2025, 21 (1), 864– 876, DOI: 10.3762/bjoc.21.70Google ScholarThere is no corresponding record for this reference.
- 16Xie, S. C.; Wang, Y.; Morton, C. J.; Metcalfe, R. D.; Dogovski, C.; Pasaje, C. F. A.; Dunn, E.; Luth, M. R.; Kumpornsin, K.; Istvan, E. S.; Park, J. S.; Fairhurst, K. J.; Ketprasit, N.; Yeo, T.; Yildirim, O.; Bhebhe, M. N.; Klug, D. M.; Rutledge, P. J.; Godoy, L. C.; Dey, S.; De Souza, M. L.; Siqueira-Neto, J. L.; Du, Y.; Puhalovich, T.; Amini, M.; Shami, G.; Loesbanluechai, D.; Nie, S.; Williamson, N.; Jana, G. P.; Maity, B. C.; Thomson, P.; Foley, T.; Tan, D. S.; Niles, J. C.; Han, B. W.; Goldberg, D. E.; Burrows, J.; Fidock, D. A.; Lee, M. C. S.; Winzeler, E. A.; Griffin, M. D. W.; Todd, M. H.; Tilley, L. Reaction Hijacking Inhibition of Plasmodium Falciparum Asparagine TRNA Synthetase. Nat. Commun. 2024, 15 (1), 937, DOI: 10.1038/s41467-024-45224-zGoogle ScholarThere is no corresponding record for this reference.
- 17Lipinski, C. A.; Lombardo, F.; Dominy, B. W.; Feeney, P. J. Experimental and Computational Approaches to Estimate Solubility and Permeability in Drug Discovery and Development Settings. Adv. Drug Delivery Rev. 2001, 46 (1–3), 3– 26Google ScholarThere is no corresponding record for this reference.
- 18Lipinski, C. A. Lead- and Drug-like Compounds: The Rule-of-Five Revolution. Drug Discovery Today: Technol. 2004, 1 (4), 337– 341, DOI: 10.1016/j.ddtec.2004.11.007Google ScholarThere is no corresponding record for this reference.
- 19Lovering, F.; Bikker, J.; Humblet, C. Escape from Flatland: Increasing Saturation as an Approach to Improving Clinical Success. J. Med. Chem. 2009, 52 (21), 6752– 6756, DOI: 10.1021/jm901241eGoogle ScholarThere is no corresponding record for this reference.
- 20Kombo, D. C.; Tallapragada, K.; Jain, R.; Chewning, J.; Mazurov, A. A.; Speake, J. D.; Hauser, T. A.; Toler, S. 3D Molecular Descriptors Important for Clinical Success. J. Chem. Inf. Model. 2013, 53 (2), 327– 342, DOI: 10.1021/ci300445eGoogle ScholarThere is no corresponding record for this reference.
- 21Churcher, I.; Newbold, S.; Murray, C. W. Return to Flatland. Nat. Rev. Chem. 2025, 9 (3), 140– 141, DOI: 10.1038/s41570-025-00688-5Google ScholarThere is no corresponding record for this reference.
- 22Daina, A.; Michielin, O.; Zoete, V. SwissADME: A Free Web Tool to Evaluate Pharmacokinetics, Drug-Likeness and Medicinal Chemistry Friendliness of Small Molecules. Sci. Rep. 2017, 7 (1), 42717, DOI: 10.1038/srep42717Google ScholarThere is no corresponding record for this reference.
- 23CNP18-DENVRdrp-VirtualScreen. https://github.com/StructuralGenomicsConsortium/CNP18-DENVRdrp-VirtualScreen/wiki (accessed May 13, 2025).Google ScholarThere is no corresponding record for this reference.
- 24CNP11-DENVRdrp-LifeChem. https://github.com/StructuralGenomicsConsortium/CNP11-DENVRdrp-LifeChem/wiki (accessed April 10, 2025).Google ScholarThere is no corresponding record for this reference.
- 25NCI Division of Cancer Treatment & Diagnosis. https://dtp.cancer.gov/discovery_development/nci-60/(accessed April 10, 2025).Google ScholarThere is no corresponding record for this reference.
- 26CO-ADD. https://www.co-add.org/?q=content/the-project (accessed March 19, 2025).Google ScholarThere is no corresponding record for this reference.
- 27EU-OPENSCREEN. https://www.eu-openscreen.eu/index.html (accessed May 13, 2025).Google ScholarThere is no corresponding record for this reference.
- 28EXSCALATE4CoV. https://www.exscalate4cov.eu/(accessed April 10, 2025).Google ScholarThere is no corresponding record for this reference.
- 29UKHSA. Spotlight on Antimicrobial Resistance. https://research.ukhsa.gov.uk/our-research/antimicrobial-resistance/(accessed May 13, 2025).Google ScholarThere is no corresponding record for this reference.
- 30Llompart, P.; Minoletti, C.; Baybekov, S.; Horvath, D.; Marcou, G.; Varnek, A. Will We Ever Be Able to Accurately Predict Solubility?. Sci. Data 2024, 11 (1), 303, DOI: 10.1038/s41597-024-03105-6Google ScholarThere is no corresponding record for this reference.
- 31Glenn, I. S.; Hall, L. N.; Khalid, M. M.; Ott, M.; Shoichet, B. K. Colloidal Aggregation Confounds Cell-Based Covid-19 Antiviral Screens. J. Med. Chem. 2024, 67 (12), 10263– 10274, DOI: 10.1021/acs.jmedchem.4c00597Google ScholarThere is no corresponding record for this reference.
- 32Hind, C.; Clifford, M.; Woolley, C.; Harmer, J.; McGee, L. M. C.; Tyson-Hirst, I.; Tait, H. J.; Brooke, D. P.; Dancer, S. J.; Hunter, I. S.; Suckling, C. J.; Beveridge, R.; Parkinson, J. A.; Sutton, J. M.; Scott, F. J. Insights into the Spectrum of Activity and Mechanism of Action of MGB-BP-3. ACS Infect. Dis. 2022, 8 (12), 2552– 2563, DOI: 10.1021/acsinfecdis.2c00445Google ScholarThere is no corresponding record for this reference.
- 33OSA. Murligases. https://github.com/opensourceantibiotics/murligase/wiki (accessed April 10, 2025).Google ScholarThere is no corresponding record for this reference.
- 34OSM. Series 6. https://github.com/OpenSourceMalaria/Series6/wiki (accessed April 10, 2025).Google ScholarThere is no corresponding record for this reference.
- 35CNP16-iminoquinone-SarsRdRp-mechanistic-study. https://github.com/StructuralGenomicsConsortium/CNP16-iminoquinone-SarsRdRp-mechanstic-study/wiki (accessed April 10, 2025).Google ScholarThere is no corresponding record for this reference.
- 36CNP21-CHIKVhelicase-Asinex. https://github.com/StructuralGenomicsConsortium/CNP21-CHIKVhelicase-Asinex/wiki/The-Story-So-Far (accessed April 10, 2025).Google ScholarThere is no corresponding record for this reference.
- 37OSM. Series 3. https://github.com/OpenSourceMalaria/Series3/wiki/The-Data-on-the-Original-Hit (accessed April 10, 2025).Google ScholarThere is no corresponding record for this reference.
- 38Todd, M.; Carter, E.; Isaksson, R. Todd Group Compound Library - Idler Plate MHT0001; Zenodo. https://doi.org/10.5281/zenodo.15409476 (accessed May 17, 2025).Google ScholarThere is no corresponding record for this reference.
- 39National Center for Biotechnology Information. Pubchem. Bioassay Record for AID 2061108, First Todd Group Plate of Idler Compounds. Source: Research Group of Professor Matthew Todd, School of Pharmacy; University College London, 2025. https://pubchem.ncbi.nlm.nih.gov/bioassay/2061108.Google ScholarThere is no corresponding record for this reference.
- 40Todd, M.; Carter, E.; Isaksson, R. Todd Group Library ELN Snapshot May 13th 2025, Dataset; University College London, 2025.Google ScholarThere is no corresponding record for this reference.
- 41Todd, M.; Carter, E.; Isaksson, R. Todd Group Idler Library - Plate MHT0001 Results, Dataset; University College London, 2025.Google ScholarThere is no corresponding record for this reference.
- 42Zdrazil, B.; Felix, E.; Hunter, F.; Manners, E. J.; Blackshaw, J.; Corbett, S.; de Veij, M.; Ioannidis, H.; Lopez, D. M.; Mosquera, J. F.; Magarinos, M. P.; Bosc, N.; Arcila, R.; Kizilören, T.; Gaulton, A.; Bento, A. P.; Adasme, M. F.; Monecke, P.; Landrum, G. A.; Leach, A. R. The ChEMBL Database in 2023: A Drug Discovery Platform Spanning Multiple Bioactivity Data Types and Time Periods. Nucleic Acids Res. 2024, 52 (D1), D1180– D1192, DOI: 10.1093/nar/gkad1004Google ScholarThere is no corresponding record for this reference.
- 43ChEMBL Data Deposition Guide. https://chembl.gitbook.io/chembl-data-deposition-guide (accessed June 12, 2025).Google ScholarThere is no corresponding record for this reference.
- 44BioAssays - PubChem. https://pubchem.ncbi.nlm.nih.gov/docs/bioassays (accessed June 12, 2025).Google ScholarThere is no corresponding record for this reference.
- 45EUCAST MIC determination of non-fastidious and fastidious organisms. https://www.eucast.org/ast_of_bacteria/mic_determination (accessed May 13, 2025).Google ScholarThere is no corresponding record for this reference.
- 46EUCAST Methods in fungi. https://www.eucast.org/astoffungi/methodsinantifungalsusceptibilitytesting (accessed May 13, 2025).Google ScholarThere is no corresponding record for this reference.
- 47Dupont, C.; Viljoen, A.; Thomas, S.; Roquet-Banères, F.; Herrmann, J.-L.; Pethe, K.; Kremer, L. Bedaquiline Inhibits the ATP Synthase in Mycobacterium Abscessus and Is Effective in Infected Zebrafish. Antimicrob. Agents Chemother. 2017, 61 (11), e0122517 DOI: 10.1128/AAC.01225-17Google ScholarThere is no corresponding record for this reference.
- 48Litvinov, V.; Makarova, M.; Kudlay, D.; Nikolenko, N.; Mikhailova, J. In Vitro Activity of Bedaquiline against Mycobacterium Avium Complex. J. Med. Microbiol. 2021, 70 (10), 001439, DOI: 10.1099/jmm.0.001439Google ScholarThere is no corresponding record for this reference.
- 49Kim, S.-H.; Hind, C. K.; Fernandes, G. F. S.; Wu, J.; Semenya, D.; Clifford, M.; Marsh, C.; Anselmi, S.; Mason, A. J.; Bruce, K. D.; Sutton, J. M.; Castagnolo, D. Development of Novel Membrane Disrupting Lipoguanidine Compounds Sensitizing Gram-Negative Bacteria to Antibiotics. ACS Med. Chem. Lett. 2024, 15 (2), 239– 249, DOI: 10.1021/acsmedchemlett.3c00460Google ScholarThere is no corresponding record for this reference.
Cited By
This article has not yet been cited by other publications.
Article Views
Altmetric
Citations
Article Views are the COUNTER-compliant sum of full text article downloads since November 2008 (both PDF and HTML) across all institutions and individuals. These metrics are regularly updated to reflect usage leading up to the last few days.
Citations are the number of other articles citing this article, calculated by Crossref and updated daily. Find more information about Crossref citation counts.
The Altmetric Attention Score is a quantitative measure of the attention that a research article has received online. Clicking on the donut icon will load a page at altmetric.com with additional details about the score and the social media presence for the given article. Find more information on the Altmetric Attention Score and how the score is calculated.
Recommended Articles
Abstract
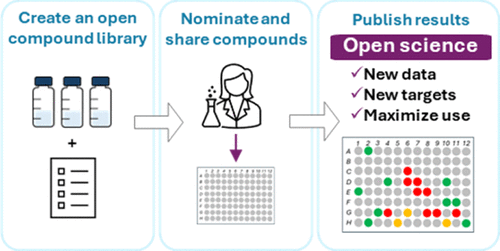
Figure 1

Figure 1. Overview of library setup and the sharing process. Compounds were synthesized or purchased for a range of projects in the research group. Idler compounds were then prepared as stock solutions and added to an online library. A subset was selected, plated and shared. A detailed step-by-step guide is provided in the Supporting Information.
Figure 2

Figure 2. Summary of the selection process for the 96-well plate to ensure compound diversity; researchers selected 56 compounds with the remaining 40 compounds selected computationally to ensure diversity as outlined in the Supporting Information.
Figure 3

Figure 3. Targets for which the compounds on plate MHT-0001 were originally intended. The shared plate contained compounds from various projects focusing on viral (60% of compounds), parasitic (25%), bacterial (9%), and human (6%) targets; most projects were target-based rather than phenotypic.
Figure 4

Figure 4. Compounds of plate MHT-0001 were selected to be structurally diverse. A range of functional groups is represented, and most compounds align with desirable values of descriptors commonly used in medicinal chemistry programmes.
Figure 5

Figure 5. (A) Comparison of antimicrobial activity against A. baumannii (∓PMBN) and E. coli (∓PMBN). Compounds are reported as active when inhibition is >80% at 100 μM compound concentration. (B) Chemical structure and antimicrobial activity of WEL2.76.
Figure 6

Figure 6. (A) Antimicrobial activity against S. aureus, A. baumannii (∓PMBN), E. coli (∓PMBN), C. albicans, M. avium, and M. abscessus. Compounds are reported as active when inhibition is >80% at 100 μM compound concentration (S. aureus, A. baumannii (∓PMBN), E. coli (∓PMBN), C. albicans) and >50% inhibition at 10 μM compound concentration (M. avium, and M. abscessus). (B) Solubility of the compounds. (C) Chemical structures of the highlighted compounds.
References
This article references 49 other publications.
- 1Duffy, S.; Sykes, M. L.; Jones, A. J.; Shelper, T. B.; Simpson, M.; Lang, R.; Poulsen, S.-A.; Sleebs, B. E.; Avery, V. M. Screening the Medicines for Malaria Venture Pathogen Box across Multiple Pathogens Reclassifies Starting Points for Open-Source Drug Discovery. Antimicrob. Agents Chemother. 2017, 61 (9), e0037917 DOI: 10.1128/AAC.00379-17There is no corresponding record for this reference.
- 2Spangenberg, T.; Burrows, J. N.; Kowalczyk, P.; McDonald, S.; Wells, T. N. C.; Willis, P. The Open Access Malaria Box: A Drug Discovery Catalyst for Neglected Diseases. PLoS One 2013, 8 (6), e62906 DOI: 10.1371/journal.pone.0062906There is no corresponding record for this reference.
- 3Adam, A.; Besson, D.; Bryant, R.; Rees, S.; Willis, P. A.; Burrows, J. N.; Hooft van Huisjduijnen, R.; Laleu, B.; Norton, L.; Canan, S.; Hawryluk, N.; Robinson, D.; Palmer, M.; Samby, K. K. Global Health Priority Box–Proactive Pandemic Preparedness. ACS Infect. Dis. 2024, 10 (12), 4030– 4039, DOI: 10.1021/acsinfecdis.4c00700There is no corresponding record for this reference.
- 4Tredup, C.; Ndreshkjana, B.; Schneider, N. S.; Tjaden, A.; Kemas, A. M.; Youhanna, S.; Lauschke, V. M.; Berger, B.-T.; Krämer, A.; Berger, L. M.; Röhm, S.; Knapp, S.; Farin, H. F.; Müller, S. Deep Annotation of Donated Chemical Probes (DCP) in Organotypic Human Liver Cultures and Patient-Derived Organoids from Tumor and Normal Colorectum. ACS Chem. Biol. 2023, 18 (4), 822– 836, DOI: 10.1021/acschembio.2c00877There is no corresponding record for this reference.
- 5Müller, S.; Ackloo, S.; Arrowsmith, C. H.; Bauser, M.; Baryza, J. L.; Blagg, J.; Böttcher, J.; Bountra, C.; Brown, P. J.; Bunnage, M. E.; Carter, A. J.; Damerell, D.; Dötsch, V.; Drewry, D. H.; Edwards, A. M.; Edwards, J.; Elkins, J. M.; Fischer, C.; Frye, S. V.; Gollner, A.; Grimshaw, C. E.; IJzerman, A.; Hanke, T.; Hartung, I. V.; Hitchcock, S.; Howe, T.; Hughes, T. V.; Laufer, S.; Li, V. M.; Liras, S.; Marsden, B. D.; Matsui, H.; Mathias, J.; O’Hagan, R. C.; Owen, D. R.; Pande, V.; Rauh, D.; Rosenberg, S. H.; Roth, B. L.; Schneider, N. S.; Scholten, C.; Singh Saikatendu, K.; Simeonov, A.; Takizawa, M.; Tse, C.; Thompson, P. R.; Treiber, D. K.; Viana, A. Y.; Wells, C. I.; Willson, T. M.; Zuercher, W. J.; Knapp, S.; Mueller-Fahrnow, A. Donated Chemical Probes for Open Science. Elife 2018, 7, e34311 DOI: 10.7554/eLife.34311There is no corresponding record for this reference.
- 6Wells, C. I.; Al-Ali, H.; Andrews, D. M.; Asquith, C. R. M.; Axtman, A. D.; Dikic, I.; Ebner, D.; Ettmayer, P.; Fischer, C.; Frederiksen, M.; Futrell, R. E.; Gray, N. S.; Hatch, S. B.; Knapp, S.; Lücking, U.; Michaelides, M.; Mills, C. E.; Müller, S.; Owen, D.; Picado, A.; Saikatendu, K. S.; Schröder, M.; Stolz, A.; Tellechea, M.; Turunen, B. J.; Vilar, S.; Wang, J.; Zuercher, W. J.; Willson, T. M.; Drewry, D. H. The Kinase Chemogenomic Set (KCGS): An Open Science Resource for Kinase Vulnerability Identification. Int. J. Mol. Sci. 2021, 22 (2), 566, DOI: 10.3390/ijms22020566There is no corresponding record for this reference.
- 7AstraZeneca. Open Innovation. https://openinnovation.astrazeneca.com/(accessed March 4, 2025).There is no corresponding record for this reference.
- 8European Lead Factory. https://www.europeanleadfactory.eu/(accessed April 10, 2025).There is no corresponding record for this reference.
- 9Griffith University. Compounds Australia. https://www.griffith.edu.au/griffith-sciences/compounds-australia (accessed June 12, 2025).There is no corresponding record for this reference.
- 10Todd Group Library Spreadsheet. https://tinyurl.com/yynht8mu (accessed June 12, 2025).There is no corresponding record for this reference.
- 11Todd Group Library LabArchive. https://tinyurl.com/3jf88u32 (accessed June 12, 2025).There is no corresponding record for this reference.
- 12LabArchives. https://www.labarchives.com/(accessed April 10, 2025).There is no corresponding record for this reference.
- 13Wilkinson, M. D.; Dumontier, M.; Aalbersberg, Ij. J.; Appleton, G.; Axton, M.; Baak, A.; Blomberg, N.; Boiten, J.-W.; da Silva Santos, L. B.; Bourne, P. E.; Bouwman, J.; Brookes, A. J.; Clark, T.; Crosas, M.; Dillo, I.; Dumon, O.; Edmunds, S.; Evelo, C. T.; Finkers, R.; Gonzalez-Beltran, A.; Gray, A. J. G.; Groth, P.; Goble, C.; Grethe, J. S.; Heringa, J.; ’t Hoen, P. A.; Hooft, R.; Kuhn, T.; Kok, R.; Kok, J.; Lusher, S. J.; Martone, M. E.; Mons, A.; Packer, A. L.; Persson, B.; Rocca-Serra, P.; Roos, M.; van Schaik, R.; Sansone, S.-A.; Schultes, E.; Sengstag, T.; Slater, T.; Strawn, G.; Swertz, M. A.; Thompson, M.; van der Lei, J.; van Mulligen, E.; Velterop, J.; Waagmeester, A.; Wittenburg, P.; Wolstencroft, K.; Zhao, J.; Mons, B. The FAIR Guiding Principles for Scientific Data Management and Stewardship. Sci. Data 2016, 3 (1), 160018, DOI: 10.1038/sdata.2016.18There is no corresponding record for this reference.
- 14Creative Commons. Attribution 4.0 International. https://creativecommons.org/licenses/by/4.0/(accessed May 13, 2025).There is no corresponding record for this reference.
- 15Bloodworth, S.; Willoughby, C.; Coles, S. J. Data Accessibility in the Chemical Sciences: An Analysis of Recent Practice in Organic Chemistry Journals. Beilstein J. Org. Chem. 2025, 21 (1), 864– 876, DOI: 10.3762/bjoc.21.70There is no corresponding record for this reference.
- 16Xie, S. C.; Wang, Y.; Morton, C. J.; Metcalfe, R. D.; Dogovski, C.; Pasaje, C. F. A.; Dunn, E.; Luth, M. R.; Kumpornsin, K.; Istvan, E. S.; Park, J. S.; Fairhurst, K. J.; Ketprasit, N.; Yeo, T.; Yildirim, O.; Bhebhe, M. N.; Klug, D. M.; Rutledge, P. J.; Godoy, L. C.; Dey, S.; De Souza, M. L.; Siqueira-Neto, J. L.; Du, Y.; Puhalovich, T.; Amini, M.; Shami, G.; Loesbanluechai, D.; Nie, S.; Williamson, N.; Jana, G. P.; Maity, B. C.; Thomson, P.; Foley, T.; Tan, D. S.; Niles, J. C.; Han, B. W.; Goldberg, D. E.; Burrows, J.; Fidock, D. A.; Lee, M. C. S.; Winzeler, E. A.; Griffin, M. D. W.; Todd, M. H.; Tilley, L. Reaction Hijacking Inhibition of Plasmodium Falciparum Asparagine TRNA Synthetase. Nat. Commun. 2024, 15 (1), 937, DOI: 10.1038/s41467-024-45224-zThere is no corresponding record for this reference.
- 17Lipinski, C. A.; Lombardo, F.; Dominy, B. W.; Feeney, P. J. Experimental and Computational Approaches to Estimate Solubility and Permeability in Drug Discovery and Development Settings. Adv. Drug Delivery Rev. 2001, 46 (1–3), 3– 26There is no corresponding record for this reference.
- 18Lipinski, C. A. Lead- and Drug-like Compounds: The Rule-of-Five Revolution. Drug Discovery Today: Technol. 2004, 1 (4), 337– 341, DOI: 10.1016/j.ddtec.2004.11.007There is no corresponding record for this reference.
- 19Lovering, F.; Bikker, J.; Humblet, C. Escape from Flatland: Increasing Saturation as an Approach to Improving Clinical Success. J. Med. Chem. 2009, 52 (21), 6752– 6756, DOI: 10.1021/jm901241eThere is no corresponding record for this reference.
- 20Kombo, D. C.; Tallapragada, K.; Jain, R.; Chewning, J.; Mazurov, A. A.; Speake, J. D.; Hauser, T. A.; Toler, S. 3D Molecular Descriptors Important for Clinical Success. J. Chem. Inf. Model. 2013, 53 (2), 327– 342, DOI: 10.1021/ci300445eThere is no corresponding record for this reference.
- 21Churcher, I.; Newbold, S.; Murray, C. W. Return to Flatland. Nat. Rev. Chem. 2025, 9 (3), 140– 141, DOI: 10.1038/s41570-025-00688-5There is no corresponding record for this reference.
- 22Daina, A.; Michielin, O.; Zoete, V. SwissADME: A Free Web Tool to Evaluate Pharmacokinetics, Drug-Likeness and Medicinal Chemistry Friendliness of Small Molecules. Sci. Rep. 2017, 7 (1), 42717, DOI: 10.1038/srep42717There is no corresponding record for this reference.
- 23CNP18-DENVRdrp-VirtualScreen. https://github.com/StructuralGenomicsConsortium/CNP18-DENVRdrp-VirtualScreen/wiki (accessed May 13, 2025).There is no corresponding record for this reference.
- 24CNP11-DENVRdrp-LifeChem. https://github.com/StructuralGenomicsConsortium/CNP11-DENVRdrp-LifeChem/wiki (accessed April 10, 2025).There is no corresponding record for this reference.
- 25NCI Division of Cancer Treatment & Diagnosis. https://dtp.cancer.gov/discovery_development/nci-60/(accessed April 10, 2025).There is no corresponding record for this reference.
- 26CO-ADD. https://www.co-add.org/?q=content/the-project (accessed March 19, 2025).There is no corresponding record for this reference.
- 27EU-OPENSCREEN. https://www.eu-openscreen.eu/index.html (accessed May 13, 2025).There is no corresponding record for this reference.
- 28EXSCALATE4CoV. https://www.exscalate4cov.eu/(accessed April 10, 2025).There is no corresponding record for this reference.
- 29UKHSA. Spotlight on Antimicrobial Resistance. https://research.ukhsa.gov.uk/our-research/antimicrobial-resistance/(accessed May 13, 2025).There is no corresponding record for this reference.
- 30Llompart, P.; Minoletti, C.; Baybekov, S.; Horvath, D.; Marcou, G.; Varnek, A. Will We Ever Be Able to Accurately Predict Solubility?. Sci. Data 2024, 11 (1), 303, DOI: 10.1038/s41597-024-03105-6There is no corresponding record for this reference.
- 31Glenn, I. S.; Hall, L. N.; Khalid, M. M.; Ott, M.; Shoichet, B. K. Colloidal Aggregation Confounds Cell-Based Covid-19 Antiviral Screens. J. Med. Chem. 2024, 67 (12), 10263– 10274, DOI: 10.1021/acs.jmedchem.4c00597There is no corresponding record for this reference.
- 32Hind, C.; Clifford, M.; Woolley, C.; Harmer, J.; McGee, L. M. C.; Tyson-Hirst, I.; Tait, H. J.; Brooke, D. P.; Dancer, S. J.; Hunter, I. S.; Suckling, C. J.; Beveridge, R.; Parkinson, J. A.; Sutton, J. M.; Scott, F. J. Insights into the Spectrum of Activity and Mechanism of Action of MGB-BP-3. ACS Infect. Dis. 2022, 8 (12), 2552– 2563, DOI: 10.1021/acsinfecdis.2c00445There is no corresponding record for this reference.
- 33OSA. Murligases. https://github.com/opensourceantibiotics/murligase/wiki (accessed April 10, 2025).There is no corresponding record for this reference.
- 34OSM. Series 6. https://github.com/OpenSourceMalaria/Series6/wiki (accessed April 10, 2025).There is no corresponding record for this reference.
- 35CNP16-iminoquinone-SarsRdRp-mechanistic-study. https://github.com/StructuralGenomicsConsortium/CNP16-iminoquinone-SarsRdRp-mechanstic-study/wiki (accessed April 10, 2025).There is no corresponding record for this reference.
- 36CNP21-CHIKVhelicase-Asinex. https://github.com/StructuralGenomicsConsortium/CNP21-CHIKVhelicase-Asinex/wiki/The-Story-So-Far (accessed April 10, 2025).There is no corresponding record for this reference.
- 37OSM. Series 3. https://github.com/OpenSourceMalaria/Series3/wiki/The-Data-on-the-Original-Hit (accessed April 10, 2025).There is no corresponding record for this reference.
- 38Todd, M.; Carter, E.; Isaksson, R. Todd Group Compound Library - Idler Plate MHT0001; Zenodo. https://doi.org/10.5281/zenodo.15409476 (accessed May 17, 2025).There is no corresponding record for this reference.
- 39National Center for Biotechnology Information. Pubchem. Bioassay Record for AID 2061108, First Todd Group Plate of Idler Compounds. Source: Research Group of Professor Matthew Todd, School of Pharmacy; University College London, 2025. https://pubchem.ncbi.nlm.nih.gov/bioassay/2061108.There is no corresponding record for this reference.
- 40Todd, M.; Carter, E.; Isaksson, R. Todd Group Library ELN Snapshot May 13th 2025, Dataset; University College London, 2025.There is no corresponding record for this reference.
- 41Todd, M.; Carter, E.; Isaksson, R. Todd Group Idler Library - Plate MHT0001 Results, Dataset; University College London, 2025.There is no corresponding record for this reference.
- 42Zdrazil, B.; Felix, E.; Hunter, F.; Manners, E. J.; Blackshaw, J.; Corbett, S.; de Veij, M.; Ioannidis, H.; Lopez, D. M.; Mosquera, J. F.; Magarinos, M. P.; Bosc, N.; Arcila, R.; Kizilören, T.; Gaulton, A.; Bento, A. P.; Adasme, M. F.; Monecke, P.; Landrum, G. A.; Leach, A. R. The ChEMBL Database in 2023: A Drug Discovery Platform Spanning Multiple Bioactivity Data Types and Time Periods. Nucleic Acids Res. 2024, 52 (D1), D1180– D1192, DOI: 10.1093/nar/gkad1004There is no corresponding record for this reference.
- 43ChEMBL Data Deposition Guide. https://chembl.gitbook.io/chembl-data-deposition-guide (accessed June 12, 2025).There is no corresponding record for this reference.
- 44BioAssays - PubChem. https://pubchem.ncbi.nlm.nih.gov/docs/bioassays (accessed June 12, 2025).There is no corresponding record for this reference.
- 45EUCAST MIC determination of non-fastidious and fastidious organisms. https://www.eucast.org/ast_of_bacteria/mic_determination (accessed May 13, 2025).There is no corresponding record for this reference.
- 46EUCAST Methods in fungi. https://www.eucast.org/astoffungi/methodsinantifungalsusceptibilitytesting (accessed May 13, 2025).There is no corresponding record for this reference.
- 47Dupont, C.; Viljoen, A.; Thomas, S.; Roquet-Banères, F.; Herrmann, J.-L.; Pethe, K.; Kremer, L. Bedaquiline Inhibits the ATP Synthase in Mycobacterium Abscessus and Is Effective in Infected Zebrafish. Antimicrob. Agents Chemother. 2017, 61 (11), e0122517 DOI: 10.1128/AAC.01225-17There is no corresponding record for this reference.
- 48Litvinov, V.; Makarova, M.; Kudlay, D.; Nikolenko, N.; Mikhailova, J. In Vitro Activity of Bedaquiline against Mycobacterium Avium Complex. J. Med. Microbiol. 2021, 70 (10), 001439, DOI: 10.1099/jmm.0.001439There is no corresponding record for this reference.
- 49Kim, S.-H.; Hind, C. K.; Fernandes, G. F. S.; Wu, J.; Semenya, D.; Clifford, M.; Marsh, C.; Anselmi, S.; Mason, A. J.; Bruce, K. D.; Sutton, J. M.; Castagnolo, D. Development of Novel Membrane Disrupting Lipoguanidine Compounds Sensitizing Gram-Negative Bacteria to Antibiotics. ACS Med. Chem. Lett. 2024, 15 (2), 239– 249, DOI: 10.1021/acsmedchemlett.3c00460There is no corresponding record for this reference.
Supporting Information
Supporting Information
The Supporting Information is available free of charge at https://pubs.acs.org/doi/10.1021/acs.jmedchem.5c02354.
A guide of library generation, links to additional online material, chemistry synthesis information and supporting biological protocols (PDF)
Compound ID names and SMILES (CSV)
The assay results including compound details (SMILES, molecular weight), the structural motif analysis of the selected compounds, and the 2D and 3D descriptors of the compounds (XLSX)
Terms & Conditions
Most electronic Supporting Information files are available without a subscription to ACS Web Editions. Such files may be downloaded by article for research use (if there is a public use license linked to the relevant article, that license may permit other uses). Permission may be obtained from ACS for other uses through requests via the RightsLink permission system: http://pubs.acs.org/page/copyright/permissions.html.



