
Building a Foundation SERS Model for Lipids through Fatty Acid Pretraining for Annotation across Chemical SpacesClick to copy article linkArticle link copied!
- Rui HanRui HanKey Laboratory of Synthetic and Biological Colloids, Ministry of Education, International Joint Research Laboratory for Nano Energy Composites, School of Chemical and Material Engineering, Jiangnan University, Wuxi, P. R. China 214122More by Rui Han
- Emily Xi Tan*Emily Xi Tan*[email protected]Key Laboratory of Synthetic and Biological Colloids, Ministry of Education, International Joint Research Laboratory for Nano Energy Composites, School of Chemical and Material Engineering, Jiangnan University, Wuxi, P. R. China 214122School of Chemistry, Chemical Engineering and Biotechnology, Nanyang Technological University, 21 Nanyang Link, Singapore 637371Institute for Digital Molecular Analytics and Science (IDMxS), Nanyang Technological University, 59 Nanyang Drive, Singapore 636921More by Emily Xi Tan
- Yangcenzi XieYangcenzi XieSchool of Chemistry, Chemical Engineering and Biotechnology, Nanyang Technological University, 21 Nanyang Link, Singapore 637371More by Yangcenzi Xie
- Hong Sheng ChengHong Sheng ChengLee Kong Chian School of Medicine, Nanyang Technological University, 59 Nanyang Drive, Singapore 636921More by Hong Sheng Cheng
- Nguan Soon TanNguan Soon TanLee Kong Chian School of Medicine, Nanyang Technological University, 59 Nanyang Drive, Singapore 636921School of Biological Sciences, Nanyang Technological University, Singapore 637551More by Nguan Soon Tan
- Yan LvYan LvKey Laboratory of Synthetic and Biological Colloids, Ministry of Education, International Joint Research Laboratory for Nano Energy Composites, School of Chemical and Material Engineering, Jiangnan University, Wuxi, P. R. China 214122More by Yan Lv
- In Yee Phang*In Yee Phang*[email protected]Key Laboratory of Synthetic and Biological Colloids, Ministry of Education, International Joint Research Laboratory for Nano Energy Composites, School of Chemical and Material Engineering, Jiangnan University, Wuxi, P. R. China 214122More by In Yee Phang
- Xing Yi Ling*Xing Yi Ling*[email protected]Key Laboratory of Synthetic and Biological Colloids, Ministry of Education, International Joint Research Laboratory for Nano Energy Composites, School of Chemical and Material Engineering, Jiangnan University, Wuxi, P. R. China 214122School of Chemistry, Chemical Engineering and Biotechnology, Nanyang Technological University, 21 Nanyang Link, Singapore 637371Institute for Digital Molecular Analytics and Science (IDMxS), Nanyang Technological University, 59 Nanyang Drive, Singapore 636921School of Biological Sciences, Nanyang Technological University, Singapore 637551More by Xing Yi Ling
Abstract
Machine learning analysis of vibrational spectra is often closed-set, performing well for known molecular classes but degrading sharply when test molecules fall outside the training library. Herein, we introduce a SERS-based domain-informed foundation model for fatty acid-derived lipids that replaces categorical assignments with vector matching. The model is trained exclusively on primitive single-chain free fatty acids, with each structural attribute encoded by five orthogonal molecular vectors in a multidimensional space: (1) carbon number, (2) number of C═C bonds, (3) C═C position, (4) C═C geometry, and (5) number of carbon chains. This modular ensemble enables zero-shot prediction of all five vectors in parallel from previously unseen spectra, allowing untargeted molecular reconstruction by proximity in this space rather than by class labels. Despite being trained only on free fatty acids, the model generalizes to complex, multichain lipids absent from the training set, achieving accuracies of 91.7% for withheld free fatty acids, 85.5% for fatty acid esters of hydroxy fatty acids, and 80.0% for triglycerides. To enhance interpretability, we utilize density functional theory to provide a mechanistic basis for the spectral features underlying each prediction. We further demonstrate matrix-tolerant multiplex quantitation in artificial sweat and urine, recovering mixture ratios with 2–9% error across broad composition ranges. Collectively, this strategy enables extrapolative, interpretable spectral-to-structure prediction from SERS across adjacent chemical spaces.
This publication is licensed for personal use by The American Chemical Society.
Introduction
Figure 1
Figure 1. SERS foundation model for fatty-acid-derived lipids enables extrapolative lipid structure elucidation across related chemical spaces. The foundation model is trained on free fatty acids and tested on chemically adjacent but unseen lipid classes, including untested free fatty acids, FAHFAs, and triglycerides. Untested SERS spectra from different lipid classes exhibit conserved vibrational features (τCH2, δCH2, νCC, and νCO) that reflect shared molecular substructures rather than lipid class identity. The SERS lipid foundation model learns quantitative relationships between these spectral features and molecular vectors: (1) number of carbon atoms, (2) number of C═C bonds, positions of C═C bonds, cis/trans geometry of C═C, and total number of carbon chains corresponding directly to standard lipid structural notation. By inferring molecular vector values from untested spectra, the model predicts complete lipid structures outside the training set, exemplified by the fatty acid γ-linolenic acid, the FAHFA 5-SAHSA, and the triglyceride triolein.
Results and Discussion
Creating a Free Fatty Acid Benchmark Data Set
Figure 2
Figure 2. SERS fingerprints encode lipid molecular vectors governing chain length, unsaturation, and isomerism. (A) Lipid structures are encoded using five chemically interpretable molecular vectors: (1) number of carbon atoms, (2) number of C═C bonds, (3) positions of C═C bonds, (4) cis–trans geometry of C═C, and (5) total number of carbon chains. (B) A curated data set of various free fatty acids is shown alongside molecular structures and corresponding SERS spectra. Systematic and reproducible changes in vibrational features accompany variation of individual molecular vectors, indicating that SERS directly encodes lipid structural chemistry. (C) Increasing carbon number (C12–C22) yields a monotonic increase in the δCH2/νCO intensity ratio. (D) Increasing the number of C═C bonds (0–3) systematically modulates the νCC/νCH2 intensity ratio with increasing unsaturation. (E) Positional isomers exhibit correlated shifts in τCH2 and δCH2 modes as a function of C═C bond location, enabling discrimination of double-bond position. (F) Cis–trans geometry of C═C is resolved by a characteristic blue shift of the νCC. Symbols: ν, stretching vibration; δ, deformation (bending) vibration; τ, twisting vibration; ω, wagging vibration (out-of-plane bending).
Creating a Foundation Model for SERS Lipid Annotation
Figure 3
Figure 3. Architecture and interpretability of the SERS lipid foundation model for fatty-acid-derived lipids. (A) The foundation model comprises an ensemble of parallel regressors and classifiers trained exclusively on free fatty acid SERS spectra to predict five lipid molecular vectors independently. (B) Feature-importance analysis for each model reveals distinctive vibrational modes (δCH2, νCC, and νCO) associated with each molecular vector, showing that the models learn physically meaningful spectral–structure relationships from SERS spectra for robust lipid structure elucidation. Symbols: ν, stretching vibration; δ, deformation (bending) vibration; τ, twisting vibration; ω, wagging vibration (out-of-plane bending).
Foundation Model Extrapolation to Complex Lipids
Figure 4
Figure 4. SERS foundation model for vector-based lipid structure elucidation and cross-class generalization. (A) Schematics of the prediction workflow in which feature-matching highlights informative regions in an unknown SERS spectrum, an ensemble infers five molecular vectors, and the vector set is assembled to reconstruct the complete lipid structure. (B) Cross-class spectral comparison showing conserved fatty acid-derived bands alongside added contributions in complex lipids; FAHFAs and triglycerides exhibit broadened CH2 fingerprints, ester features, and glycerol signatures superimposed on fatty-acid modes. (C) Blind-test results across lipid classes demonstrate stable prediction beyond the single-chain FFA training chemical space, achieving overall elucidation accuracies of 91.7% (FFAs), 85.5% (FAHFAs), and 80.0% (triglycerides), with representative reconstructed structures shown for each class. Symbols: ν, stretching vibration; δ, deformation (bending) vibration; τ, twisting vibration; ω, wagging vibration (out-of-plane bending).
Multiplex Quantitation of Lipid Mixtures in Biomatrices
Figure 5
Figure 5. Multiplex quantitative lipid detection in complex biofluids. SERS combined with the support vector machine regressor (SVM-R) enables multiplex quantification of three different binary lipid mixtures in (A) artificial sweat and (B) artificial urine. The blind-test-predicted concentrations closely match experimental values across multiple lipid pairs with high coefficients of determination and low prediction errors, demonstrating matrix-tolerant multiplex analysis in biologically relevant environments and applicability in real-life scenarios.
Conclusions
Experimental Section
Chemicals
Synthesis and Purification of Fatty Acid Esters of Hydroxy Fatty Acids (FAHFA) Derivatives
Synthesis and Purification of Silver Nanocubes
SEM and UV–Vis Characterization of Ag Nanocubes
Preparation of SERS Substrates
SERS Measurement of Lipids
Density Functional Theory (DFT) Simulations
Chemometrics Analysis
Artificial Biofluid Preparation
Supporting Information
The Supporting Information is available free of charge at https://pubs.acs.org/doi/10.1021/acsami.6c00759.
Materials and methods, AgNC platform characterization, SERS measurements of lipids, density functional theory calculations, machine learning modeling (PDF)
Terms & Conditions
Most electronic Supporting Information files are available without a subscription to ACS Web Editions. Such files may be downloaded by article for research use (if there is a public use license linked to the relevant article, that license may permit other uses). Permission may be obtained from ACS for other uses through requests via the RightsLink permission system: http://pubs.acs.org/page/copyright/permissions.html.
| Ag | Silver |
| DFT | Density Functional Theory |
| FFA | Free Fatty Acid |
| FAHFA | Fatty Acid Ester of Hydroxy Fatty Acid |
| LA | Linoleic Acid |
| ML | Machine Learning |
| NMR | Nuclear Magnetic Resonance |
| OA | Oleic Acid |
| PsA | Petroselinic Acid |
| RF | Random Forest |
| RMSEP | Root Mean Square Error of Prediction |
| SERS | Surface-Enhanced Raman Scattering |
| SVM | Support Vector Machine |
| VA | Vaccenic Acid |
| α-LA | α-Linolenic Acid |
References
This article references 30 other publications.
- 1Guo, H.; Zhi, S.; Zhao, Z.; Gao, T.; Ma, H.; Luo, S.-H.; Zhang, W.; Guo, P.; Ren, B.; Tian, Z.-Q. Rapid SERS Analysis: From Laboratory to Real Sample. ACS Appl. Mater. Interfaces 2025, 17, 33318, DOI: 10.1021/acsami.5c05699Google ScholarThere is no corresponding record for this reference.
- 2Tan, E. X.; Zhong, Q.-Z.; Ting Chen, J. R.; Leong, Y. X.; Leon, G. K.; Tran, C. T.; Phang, I. Y.; Ling, X. Y. Surface-Enhanced Raman Scattering-Based Multimodal Techniques: Advances and Perspectives. ACS Nano 2024, 18 (47), 32315– 32334, DOI: 10.1021/acsnano.4c12996Google ScholarThere is no corresponding record for this reference.
- 3Tan, E. X.; Nguyen, L. B. T.; Jin, Y.; Lv, Y.; Phang, I. Y.; Ling, X. Y. SERS Cheminformatics: Opportunities for Data-Driven Discovery and Applications. ACS Central Science 2025, 11 (11), 2034– 2052, DOI: 10.1021/acscentsci.5c00785Google ScholarThere is no corresponding record for this reference.
- 4Tan, E. X.; Leong, Y. X.; Lim, S. H.; Chng, M. W. K.; Phang, I. Y.; Ling, X. Y. Chemistry-informed recommender system to predict optimal molecular receptors in SERS nanosensors. Nat. Commun. 2025, 16 (1), 7095, DOI: 10.1038/s41467-025-62519-xGoogle ScholarThere is no corresponding record for this reference.
- 5Tan, E. X.; Leong, S. X.; Liew, W. A.; Phang, I. Y.; Ng, J. Y.; Tan, N. S.; Lee, Y. H.; Ling, X. Y. Forward-predictive SERS-based chemical taxonomy for untargeted structural elucidation of epimeric cerebrosides. Nat. Commun. 2024, 15 (1), 2582, DOI: 10.1038/s41467-024-46838-zGoogle ScholarThere is no corresponding record for this reference.
- 6Tan, E. X.; Chen, J. R. T.; Pang, D. W. C.; Tan, N. S.; Phang, I. Y.; Ling, X. Y. Transfer Learning-Assisted SERS: Predicting Molecular Identity and Concentration in Mixtures Using Pure Compound Spectra. Angew Chem Int Ed 2025, 64, e202508717 DOI: 10.1002/anie.202508717Google ScholarThere is no corresponding record for this reference.
- 7Nguyen, L. B. T.; Tan, E. X.; Leong, S. X.; Koh, C. S. L.; Madhumita, M.; Phang, I. Y.; Ling, X. Y. Harnessing Cooperative Multivalency in Thioguanine for Surface-Enhanced Raman Scattering (SERS)-Based Differentiation of Polyfunctional Analytes Differing by a Single Functional Group. Angew Chem Int Ed 2024, 63, e202410815 DOI: 10.1002/anie.202410815Google ScholarThere is no corresponding record for this reference.
- 8Simas, M. V.; Davis, G. A., Jr; Hati, S.; Pu, J.; Goodpaster, J. V.; Sardar, R. Anisotropically Shaped Plasmonic WO3–x Nanostructure-Driven Ultrasensitive SERS Detection and Machine Learning-Based Differentiation of Nitro-Explosives. ACS Appl. Mater. Interfaces 2025, 17 (7), 11309– 11324, DOI: 10.1021/acsami.4c19673Google ScholarThere is no corresponding record for this reference.
- 9Chen, J. R. T.; Tan, E. X.; Tang, J.; Leong, S. X.; Hue, S. K. X.; Pun, C. S.; Phang, I. Y.; Ling, X. Y. Machine learning-based SERS chemical space for two-way prediction of structures and spectra of untrained molecules. J. Am. Chem. Soc. 2025, 147 (8), 6654– 6664, DOI: 10.1021/jacs.4c15804Google ScholarThere is no corresponding record for this reference.
- 10Abed, M. M.; Wouters, C. L.; Froehlich, C. E.; Nguyen, T. B.; Caldwell, R.; Riley, K. L.; Roy, P.; Reineke, T. M.; Haynes, C. L. A Machine Learning-Enabled SERS Sensor: Multiplex Detection of Lipopolysaccharides from Foodborne Pathogenic Bacteria. ACS Appl. Mater. Interfaces 2025, 17 (31), 45139– 45149, DOI: 10.1021/acsami.5c08361Google ScholarThere is no corresponding record for this reference.
- 11Du, S.; Su, M.; Wang, C.; Ding, Z.; Jiang, Y.; Liu, H. Pinpointing alkane chain length, saturation, and double bond regio-and stereoisomers by liquid interfacial plasmonic enhanced raman spectroscopy. Anal. Chem. 2022, 94 (6), 2891– 2900, DOI: 10.1021/acs.analchem.1c04774Google ScholarThere is no corresponding record for this reference.
- 12Fahy, E.; Subramaniam, S.; Brown, H. A.; Glass, C. K.; Merrill, A. H., Jr; Murphy, R. C.; Raetz, C. R.; Russell, D. W.; Seyama, Y.; Shaw, W. A comprehensive classification system for lipids. European journal of lipid science and technology 2005, 107 (5), 337– 364, DOI: 10.1002/ejlt.200405001Google ScholarThere is no corresponding record for this reference.
- 13Xia, F.; Wan, J. B. Chemical derivatization strategy for mass spectrometry-based lipidomics. Mass Spectrom. Rev. 2023, 42 (1), 432– 452, DOI: 10.1002/mas.21729Google ScholarThere is no corresponding record for this reference.
- 14Kirkwood, K. I.; Pratt, B. S.; Shulman, N.; Tamura, K.; MacCoss, M. J.; MacLean, B. X.; Baker, E. S. Utilizing Skyline to analyze lipidomics data containing liquid chromatography, ion mobility spectrometry and mass spectrometry dimensions. Nat. Protoc. 2022, 17 (11), 2415– 2430, DOI: 10.1038/s41596-022-00714-6Google ScholarThere is no corresponding record for this reference.
- 15Nagana Gowda, G.; Raftery, D. NMR metabolomics methods for investigating disease. Analytical chemistry 2023, 95 (1), 83– 99, DOI: 10.1021/acs.analchem.2c04606Google ScholarThere is no corresponding record for this reference.
- 16Tan, E. X.; Tang, J.; Leong, Y. X.; Phang, I. Y.; Lee, Y. H.; Pun, C. S.; Ling, X. Y. Creating 3D Nanoparticle Structural Space via Data Augmentation to Bidirectionally Predict Nanoparticle Mixture’s Purity, Size, and Shape from Extinction Spectra. Angew Chem Int Ed 2024, 63 (14), e202317978 DOI: 10.1002/anie.202317978Google ScholarThere is no corresponding record for this reference.
- 17Beattie, J. R.; Bell, S. E.; Moss, B. W. A critical evaluation of Raman spectroscopy for the analysis of lipids: Fatty acid methyl esters. Lipids 2004, 39 (5), 407– 419, DOI: 10.1007/s11745-004-1245-zGoogle ScholarThere is no corresponding record for this reference.
- 18Geerlings, P.; De Proft, F.; Langenaeker, W. Conceptual density functional theory. Chem. Rev. 2003, 103 (5), 1793– 1874, DOI: 10.1021/cr990029pGoogle ScholarThere is no corresponding record for this reference.
- 19Cohen, A. J.; Mori-Sánchez, P.; Yang, W. Challenges for density functional theory. Chem. Rev. 2012, 112 (1), 289– 320, DOI: 10.1021/cr200107zGoogle ScholarThere is no corresponding record for this reference.
- 20Patterson, M. L.; Weaver, M. J. Surface-enhanced Raman spectroscopy as a probe of adsorbate-surface bonding: simple alkenes and alkynes adsorbed at gold electrodes. J. Phys. Chem. 1985, 89 (23), 5046– 5051, DOI: 10.1021/j100269a032Google ScholarThere is no corresponding record for this reference.
- 21Tan, E. X.; Chen, Y.; Lee, Y. H.; Leong, Y. X.; Leong, S. X.; Stanley, C. V.; Pun, C. S.; Ling, X. Y. Incorporating plasmonic featurization with machine learning to achieve accurate and bidirectional prediction of nanoparticle size and size distribution. Nanoscale Horizons 2022, 7 (6), 626– 633, DOI: 10.1039/D2NH00146BGoogle ScholarThere is no corresponding record for this reference.
- 22Breiman, L. Random forests. Machine learning 2001, 45 (1), 5– 32, DOI: 10.1023/A:1010933404324Google ScholarThere is no corresponding record for this reference.
- 23Pisner, D. A.; Schnyer, D. M. Support vector machine. Machine learning 2020, 101– 121, DOI: 10.1016/B978-0-12-815739-8.00006-7Google ScholarThere is no corresponding record for this reference.
- 24Xie, H.; Cheng, H. S.; Ng, J. X. J.; Zhang, S.; Kim, J. H. S.; Tan, S. H.; Tan, C.-H.; Tan, N. S. A gut-liver lipid flux checkpoint mediates FAHFA protection from MASLD. Pharmacol. Res. 2026, 224, 108085 DOI: 10.1016/j.phrs.2025.108085Google ScholarThere is no corresponding record for this reference.
- 25Simopoulos, A. P. Essential fatty acids in health and chronic disease. American journal of clinical nutrition 1999, 70 (3), 560S– 569S, DOI: 10.1093/ajcn/70.3.560sGoogle ScholarThere is no corresponding record for this reference.
- 26Gentile, C. L.; Pagliassotti, M. J. The role of fatty acids in the development and progression of nonalcoholic fatty liver disease. Journal of nutritional biochemistry 2008, 19 (9), 567– 576, DOI: 10.1016/j.jnutbio.2007.10.001Google ScholarThere is no corresponding record for this reference.
- 27Lian, S.; Li, X.; Lv, X. Recent Developments in SERS Microfluidic Chips: From Fundamentals to Biosensing Applications. ACS Appl. Mater. Interfaces 2025, 17 (7), 10193– 10230, DOI: 10.1021/acsami.4c17779Google ScholarThere is no corresponding record for this reference.
- 28Tao, A.; Sinsermsuksakul, P.; Yang, P. Polyhedral silver nanocrystals with distinct scattering signatures. ANGEWANDTE CHEMIE-INTERNATIONAL EDITION IN ENGLISH- 2006, 45 (28), 4597, DOI: 10.1002/anie.200601277Google ScholarThere is no corresponding record for this reference.
- 29Lee, Y. H.; Lee, C. K.; Tan, B.; Rui Tan, J. M.; Phang, I. Y.; Ling, X. Y. Using the Langmuir–Schaefer technique to fabricate large-area dense SERS-active Au nanoprism monolayer films. Nanoscale 2013, 5 (14), 6404– 6412, DOI: 10.1039/c3nr00981eGoogle ScholarThere is no corresponding record for this reference.
- 30You, T.-t.; Yin, P.-g.; Jiang, L.; Lang, X.-f.; Guo, L.; Yang, S.-h. In situ identification of the adsorption of 4, 4′-thiobisbenzenethiol on silver nanoparticles surface: a combined investigation of surface-enhanced Raman scattering and density functional theory study. Phys. Chem. Chem. Phys. 2012, 14 (19), 6817– 6825, DOI: 10.1039/c2cp24147aGoogle ScholarThere is no corresponding record for this reference.
Cited By
This article has not yet been cited by other publications.
Article Views
Altmetric
Citations
Article Views are the COUNTER-compliant sum of full text article downloads since November 2008 (both PDF and HTML) across all institutions and individuals. These metrics are regularly updated to reflect usage leading up to the last few days.
Citations are the number of other articles citing this article, calculated by Crossref and updated daily. Find more information about Crossref citation counts.
The Altmetric Attention Score is a quantitative measure of the attention that a research article has received online. Clicking on the donut icon will load a page at altmetric.com with additional details about the score and the social media presence for the given article. Find more information on the Altmetric Attention Score and how the score is calculated.
Recommended Articles
Abstract
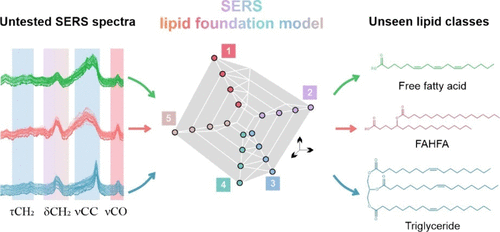
Figure 1
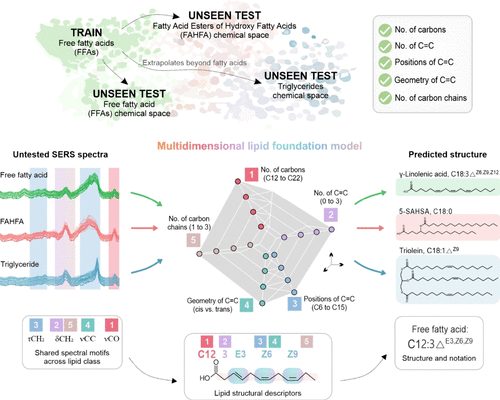
Figure 1. SERS foundation model for fatty-acid-derived lipids enables extrapolative lipid structure elucidation across related chemical spaces. The foundation model is trained on free fatty acids and tested on chemically adjacent but unseen lipid classes, including untested free fatty acids, FAHFAs, and triglycerides. Untested SERS spectra from different lipid classes exhibit conserved vibrational features (τCH2, δCH2, νCC, and νCO) that reflect shared molecular substructures rather than lipid class identity. The SERS lipid foundation model learns quantitative relationships between these spectral features and molecular vectors: (1) number of carbon atoms, (2) number of C═C bonds, positions of C═C bonds, cis/trans geometry of C═C, and total number of carbon chains corresponding directly to standard lipid structural notation. By inferring molecular vector values from untested spectra, the model predicts complete lipid structures outside the training set, exemplified by the fatty acid γ-linolenic acid, the FAHFA 5-SAHSA, and the triglyceride triolein.
Figure 2
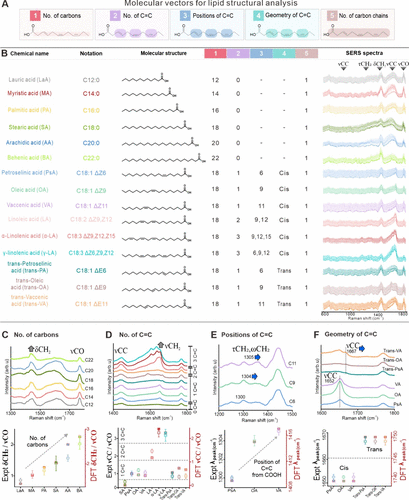
Figure 2. SERS fingerprints encode lipid molecular vectors governing chain length, unsaturation, and isomerism. (A) Lipid structures are encoded using five chemically interpretable molecular vectors: (1) number of carbon atoms, (2) number of C═C bonds, (3) positions of C═C bonds, (4) cis–trans geometry of C═C, and (5) total number of carbon chains. (B) A curated data set of various free fatty acids is shown alongside molecular structures and corresponding SERS spectra. Systematic and reproducible changes in vibrational features accompany variation of individual molecular vectors, indicating that SERS directly encodes lipid structural chemistry. (C) Increasing carbon number (C12–C22) yields a monotonic increase in the δCH2/νCO intensity ratio. (D) Increasing the number of C═C bonds (0–3) systematically modulates the νCC/νCH2 intensity ratio with increasing unsaturation. (E) Positional isomers exhibit correlated shifts in τCH2 and δCH2 modes as a function of C═C bond location, enabling discrimination of double-bond position. (F) Cis–trans geometry of C═C is resolved by a characteristic blue shift of the νCC. Symbols: ν, stretching vibration; δ, deformation (bending) vibration; τ, twisting vibration; ω, wagging vibration (out-of-plane bending).
Figure 3
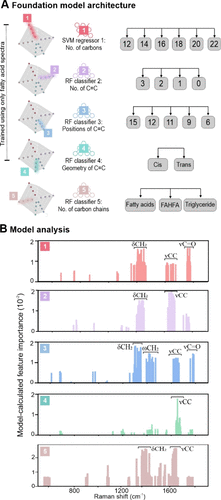
Figure 3. Architecture and interpretability of the SERS lipid foundation model for fatty-acid-derived lipids. (A) The foundation model comprises an ensemble of parallel regressors and classifiers trained exclusively on free fatty acid SERS spectra to predict five lipid molecular vectors independently. (B) Feature-importance analysis for each model reveals distinctive vibrational modes (δCH2, νCC, and νCO) associated with each molecular vector, showing that the models learn physically meaningful spectral–structure relationships from SERS spectra for robust lipid structure elucidation. Symbols: ν, stretching vibration; δ, deformation (bending) vibration; τ, twisting vibration; ω, wagging vibration (out-of-plane bending).
Figure 4

Figure 4. SERS foundation model for vector-based lipid structure elucidation and cross-class generalization. (A) Schematics of the prediction workflow in which feature-matching highlights informative regions in an unknown SERS spectrum, an ensemble infers five molecular vectors, and the vector set is assembled to reconstruct the complete lipid structure. (B) Cross-class spectral comparison showing conserved fatty acid-derived bands alongside added contributions in complex lipids; FAHFAs and triglycerides exhibit broadened CH2 fingerprints, ester features, and glycerol signatures superimposed on fatty-acid modes. (C) Blind-test results across lipid classes demonstrate stable prediction beyond the single-chain FFA training chemical space, achieving overall elucidation accuracies of 91.7% (FFAs), 85.5% (FAHFAs), and 80.0% (triglycerides), with representative reconstructed structures shown for each class. Symbols: ν, stretching vibration; δ, deformation (bending) vibration; τ, twisting vibration; ω, wagging vibration (out-of-plane bending).
Figure 5
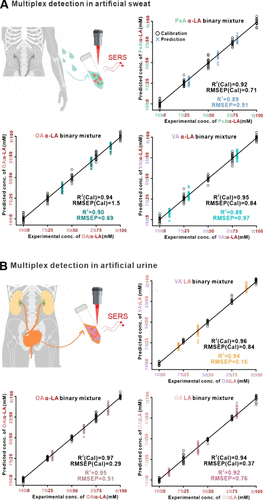
Figure 5. Multiplex quantitative lipid detection in complex biofluids. SERS combined with the support vector machine regressor (SVM-R) enables multiplex quantification of three different binary lipid mixtures in (A) artificial sweat and (B) artificial urine. The blind-test-predicted concentrations closely match experimental values across multiple lipid pairs with high coefficients of determination and low prediction errors, demonstrating matrix-tolerant multiplex analysis in biologically relevant environments and applicability in real-life scenarios.
References
This article references 30 other publications.
- 1Guo, H.; Zhi, S.; Zhao, Z.; Gao, T.; Ma, H.; Luo, S.-H.; Zhang, W.; Guo, P.; Ren, B.; Tian, Z.-Q. Rapid SERS Analysis: From Laboratory to Real Sample. ACS Appl. Mater. Interfaces 2025, 17, 33318, DOI: 10.1021/acsami.5c05699There is no corresponding record for this reference.
- 2Tan, E. X.; Zhong, Q.-Z.; Ting Chen, J. R.; Leong, Y. X.; Leon, G. K.; Tran, C. T.; Phang, I. Y.; Ling, X. Y. Surface-Enhanced Raman Scattering-Based Multimodal Techniques: Advances and Perspectives. ACS Nano 2024, 18 (47), 32315– 32334, DOI: 10.1021/acsnano.4c12996There is no corresponding record for this reference.
- 3Tan, E. X.; Nguyen, L. B. T.; Jin, Y.; Lv, Y.; Phang, I. Y.; Ling, X. Y. SERS Cheminformatics: Opportunities for Data-Driven Discovery and Applications. ACS Central Science 2025, 11 (11), 2034– 2052, DOI: 10.1021/acscentsci.5c00785There is no corresponding record for this reference.
- 4Tan, E. X.; Leong, Y. X.; Lim, S. H.; Chng, M. W. K.; Phang, I. Y.; Ling, X. Y. Chemistry-informed recommender system to predict optimal molecular receptors in SERS nanosensors. Nat. Commun. 2025, 16 (1), 7095, DOI: 10.1038/s41467-025-62519-xThere is no corresponding record for this reference.
- 5Tan, E. X.; Leong, S. X.; Liew, W. A.; Phang, I. Y.; Ng, J. Y.; Tan, N. S.; Lee, Y. H.; Ling, X. Y. Forward-predictive SERS-based chemical taxonomy for untargeted structural elucidation of epimeric cerebrosides. Nat. Commun. 2024, 15 (1), 2582, DOI: 10.1038/s41467-024-46838-zThere is no corresponding record for this reference.
- 6Tan, E. X.; Chen, J. R. T.; Pang, D. W. C.; Tan, N. S.; Phang, I. Y.; Ling, X. Y. Transfer Learning-Assisted SERS: Predicting Molecular Identity and Concentration in Mixtures Using Pure Compound Spectra. Angew Chem Int Ed 2025, 64, e202508717 DOI: 10.1002/anie.202508717There is no corresponding record for this reference.
- 7Nguyen, L. B. T.; Tan, E. X.; Leong, S. X.; Koh, C. S. L.; Madhumita, M.; Phang, I. Y.; Ling, X. Y. Harnessing Cooperative Multivalency in Thioguanine for Surface-Enhanced Raman Scattering (SERS)-Based Differentiation of Polyfunctional Analytes Differing by a Single Functional Group. Angew Chem Int Ed 2024, 63, e202410815 DOI: 10.1002/anie.202410815There is no corresponding record for this reference.
- 8Simas, M. V.; Davis, G. A., Jr; Hati, S.; Pu, J.; Goodpaster, J. V.; Sardar, R. Anisotropically Shaped Plasmonic WO3–x Nanostructure-Driven Ultrasensitive SERS Detection and Machine Learning-Based Differentiation of Nitro-Explosives. ACS Appl. Mater. Interfaces 2025, 17 (7), 11309– 11324, DOI: 10.1021/acsami.4c19673There is no corresponding record for this reference.
- 9Chen, J. R. T.; Tan, E. X.; Tang, J.; Leong, S. X.; Hue, S. K. X.; Pun, C. S.; Phang, I. Y.; Ling, X. Y. Machine learning-based SERS chemical space for two-way prediction of structures and spectra of untrained molecules. J. Am. Chem. Soc. 2025, 147 (8), 6654– 6664, DOI: 10.1021/jacs.4c15804There is no corresponding record for this reference.
- 10Abed, M. M.; Wouters, C. L.; Froehlich, C. E.; Nguyen, T. B.; Caldwell, R.; Riley, K. L.; Roy, P.; Reineke, T. M.; Haynes, C. L. A Machine Learning-Enabled SERS Sensor: Multiplex Detection of Lipopolysaccharides from Foodborne Pathogenic Bacteria. ACS Appl. Mater. Interfaces 2025, 17 (31), 45139– 45149, DOI: 10.1021/acsami.5c08361There is no corresponding record for this reference.
- 11Du, S.; Su, M.; Wang, C.; Ding, Z.; Jiang, Y.; Liu, H. Pinpointing alkane chain length, saturation, and double bond regio-and stereoisomers by liquid interfacial plasmonic enhanced raman spectroscopy. Anal. Chem. 2022, 94 (6), 2891– 2900, DOI: 10.1021/acs.analchem.1c04774There is no corresponding record for this reference.
- 12Fahy, E.; Subramaniam, S.; Brown, H. A.; Glass, C. K.; Merrill, A. H., Jr; Murphy, R. C.; Raetz, C. R.; Russell, D. W.; Seyama, Y.; Shaw, W. A comprehensive classification system for lipids. European journal of lipid science and technology 2005, 107 (5), 337– 364, DOI: 10.1002/ejlt.200405001There is no corresponding record for this reference.
- 13Xia, F.; Wan, J. B. Chemical derivatization strategy for mass spectrometry-based lipidomics. Mass Spectrom. Rev. 2023, 42 (1), 432– 452, DOI: 10.1002/mas.21729There is no corresponding record for this reference.
- 14Kirkwood, K. I.; Pratt, B. S.; Shulman, N.; Tamura, K.; MacCoss, M. J.; MacLean, B. X.; Baker, E. S. Utilizing Skyline to analyze lipidomics data containing liquid chromatography, ion mobility spectrometry and mass spectrometry dimensions. Nat. Protoc. 2022, 17 (11), 2415– 2430, DOI: 10.1038/s41596-022-00714-6There is no corresponding record for this reference.
- 15Nagana Gowda, G.; Raftery, D. NMR metabolomics methods for investigating disease. Analytical chemistry 2023, 95 (1), 83– 99, DOI: 10.1021/acs.analchem.2c04606There is no corresponding record for this reference.
- 16Tan, E. X.; Tang, J.; Leong, Y. X.; Phang, I. Y.; Lee, Y. H.; Pun, C. S.; Ling, X. Y. Creating 3D Nanoparticle Structural Space via Data Augmentation to Bidirectionally Predict Nanoparticle Mixture’s Purity, Size, and Shape from Extinction Spectra. Angew Chem Int Ed 2024, 63 (14), e202317978 DOI: 10.1002/anie.202317978There is no corresponding record for this reference.
- 17Beattie, J. R.; Bell, S. E.; Moss, B. W. A critical evaluation of Raman spectroscopy for the analysis of lipids: Fatty acid methyl esters. Lipids 2004, 39 (5), 407– 419, DOI: 10.1007/s11745-004-1245-zThere is no corresponding record for this reference.
- 18Geerlings, P.; De Proft, F.; Langenaeker, W. Conceptual density functional theory. Chem. Rev. 2003, 103 (5), 1793– 1874, DOI: 10.1021/cr990029pThere is no corresponding record for this reference.
- 19Cohen, A. J.; Mori-Sánchez, P.; Yang, W. Challenges for density functional theory. Chem. Rev. 2012, 112 (1), 289– 320, DOI: 10.1021/cr200107zThere is no corresponding record for this reference.
- 20Patterson, M. L.; Weaver, M. J. Surface-enhanced Raman spectroscopy as a probe of adsorbate-surface bonding: simple alkenes and alkynes adsorbed at gold electrodes. J. Phys. Chem. 1985, 89 (23), 5046– 5051, DOI: 10.1021/j100269a032There is no corresponding record for this reference.
- 21Tan, E. X.; Chen, Y.; Lee, Y. H.; Leong, Y. X.; Leong, S. X.; Stanley, C. V.; Pun, C. S.; Ling, X. Y. Incorporating plasmonic featurization with machine learning to achieve accurate and bidirectional prediction of nanoparticle size and size distribution. Nanoscale Horizons 2022, 7 (6), 626– 633, DOI: 10.1039/D2NH00146BThere is no corresponding record for this reference.
- 22Breiman, L. Random forests. Machine learning 2001, 45 (1), 5– 32, DOI: 10.1023/A:1010933404324There is no corresponding record for this reference.
- 23Pisner, D. A.; Schnyer, D. M. Support vector machine. Machine learning 2020, 101– 121, DOI: 10.1016/B978-0-12-815739-8.00006-7There is no corresponding record for this reference.
- 24Xie, H.; Cheng, H. S.; Ng, J. X. J.; Zhang, S.; Kim, J. H. S.; Tan, S. H.; Tan, C.-H.; Tan, N. S. A gut-liver lipid flux checkpoint mediates FAHFA protection from MASLD. Pharmacol. Res. 2026, 224, 108085 DOI: 10.1016/j.phrs.2025.108085There is no corresponding record for this reference.
- 25Simopoulos, A. P. Essential fatty acids in health and chronic disease. American journal of clinical nutrition 1999, 70 (3), 560S– 569S, DOI: 10.1093/ajcn/70.3.560sThere is no corresponding record for this reference.
- 26Gentile, C. L.; Pagliassotti, M. J. The role of fatty acids in the development and progression of nonalcoholic fatty liver disease. Journal of nutritional biochemistry 2008, 19 (9), 567– 576, DOI: 10.1016/j.jnutbio.2007.10.001There is no corresponding record for this reference.
- 27Lian, S.; Li, X.; Lv, X. Recent Developments in SERS Microfluidic Chips: From Fundamentals to Biosensing Applications. ACS Appl. Mater. Interfaces 2025, 17 (7), 10193– 10230, DOI: 10.1021/acsami.4c17779There is no corresponding record for this reference.
- 28Tao, A.; Sinsermsuksakul, P.; Yang, P. Polyhedral silver nanocrystals with distinct scattering signatures. ANGEWANDTE CHEMIE-INTERNATIONAL EDITION IN ENGLISH- 2006, 45 (28), 4597, DOI: 10.1002/anie.200601277There is no corresponding record for this reference.
- 29Lee, Y. H.; Lee, C. K.; Tan, B.; Rui Tan, J. M.; Phang, I. Y.; Ling, X. Y. Using the Langmuir–Schaefer technique to fabricate large-area dense SERS-active Au nanoprism monolayer films. Nanoscale 2013, 5 (14), 6404– 6412, DOI: 10.1039/c3nr00981eThere is no corresponding record for this reference.
- 30You, T.-t.; Yin, P.-g.; Jiang, L.; Lang, X.-f.; Guo, L.; Yang, S.-h. In situ identification of the adsorption of 4, 4′-thiobisbenzenethiol on silver nanoparticles surface: a combined investigation of surface-enhanced Raman scattering and density functional theory study. Phys. Chem. Chem. Phys. 2012, 14 (19), 6817– 6825, DOI: 10.1039/c2cp24147aThere is no corresponding record for this reference.
Supporting Information
Supporting Information
The Supporting Information is available free of charge at https://pubs.acs.org/doi/10.1021/acsami.6c00759.
Materials and methods, AgNC platform characterization, SERS measurements of lipids, density functional theory calculations, machine learning modeling (PDF)
Terms & Conditions
Most electronic Supporting Information files are available without a subscription to ACS Web Editions. Such files may be downloaded by article for research use (if there is a public use license linked to the relevant article, that license may permit other uses). Permission may be obtained from ACS for other uses through requests via the RightsLink permission system: http://pubs.acs.org/page/copyright/permissions.html.


