Advancements in Technologies Targeting Horizontal Gene Transfer─Routes to Control Drug Resistance EvolutionClick to copy article linkArticle link copied!
- Samuel Chetachukwu AdegokeSamuel Chetachukwu AdegokeDepartment of Nanoscience, Joint School of Nanoscience and Nanoengineering, E Gate City Blvd, Greensboro, North Carolina 27401, United StatesMore by Samuel Chetachukwu Adegoke
- Md Adnan KarimMd Adnan KarimDepartment of Nanoscience, Joint School of Nanoscience and Nanoengineering, E Gate City Blvd, Greensboro, North Carolina 27401, United StatesMore by Md Adnan Karim
- Maurelio Cabo JrMaurelio Cabo JrDepartment of Nanoscience, Joint School of Nanoscience and Nanoengineering, E Gate City Blvd, Greensboro, North Carolina 27401, United StatesMore by Maurelio Cabo Jr
- Ignatius Senyo Yao YawluiIgnatius Senyo Yao YawluiDepartment of Nanoscience, Joint School of Nanoscience and Nanoengineering, E Gate City Blvd, Greensboro, North Carolina 27401, United StatesMore by Ignatius Senyo Yao Yawlui
- Dennis LaJeunesse*Dennis LaJeunesse*Email: [email protected]Department of Nanoscience, Joint School of Nanoscience and Nanoengineering, E Gate City Blvd, Greensboro, North Carolina 27401, United StatesMore by Dennis LaJeunesse
Abstract
The global rise of multidrug-resistant (MDR) bacteria poses a major public health crisis, threatening the effectiveness of modern medicine. Traditional antibiotic development struggles to keep pace with bacterial evolution, largely due to the rapid dissemination of antibiotic resistance genes via horizontal gene transfer (HGT). HGT mechanisms both canonical and noncanonical enable bacteria to acquire resistance traits defining species and even special challenges. In this review, we cover the current understanding of HGT in spreading antibiotic resistance and explore possible strategies to control HGT and slow the spread of antimicrobial resistance. Recent advances highlight the potential of synthetic competence inhibitors, advanced oxidation processes (AOPs), CRISPR-Cas technologies, gene drives, and antiplasmids to disrupt horizontal gene flow and mitigate resistance evolution. Despite promising laboratory results, challenges remain in translating these approaches into clinical and environmental applications. Blocking HGT could complement antimicrobial stewardship programs and traditional antibiotic therapies by curbing the emergence of new resistant strains at their genetic roots. By targeting the foundational mechanisms of resistance acquisition, these strategies offer a proactive pathway to extend the efficacy of existing antibiotics and prevent a “postantibiotic” era. Ongoing research into bacterial pathogenesis, genome defense systems, and innovative gene-editing technologies will be critical to developing effective, scalable solutions for managing MDR infections worldwide.
This publication is licensed under
License Summary*
You are free to share(copy and redistribute) this article in any medium or format and to adapt(remix, transform, and build upon) the material for any purpose, even commercially within the parameters below:
Creative Commons (CC): This is a Creative Commons license.
Attribution (BY): Credit must be given to the creator.
*Disclaimer
This summary highlights only some of the key features and terms of the actual license. It is not a license and has no legal value. Carefully review the actual license before using these materials.
License Summary*
You are free to share(copy and redistribute) this article in any medium or format and to adapt(remix, transform, and build upon) the material for any purpose, even commercially within the parameters below:
Creative Commons (CC): This is a Creative Commons license.
Attribution (BY): Credit must be given to the creator.
*Disclaimer
This summary highlights only some of the key features and terms of the actual license. It is not a license and has no legal value. Carefully review the actual license before using these materials.
License Summary*
You are free to share(copy and redistribute) this article in any medium or format and to adapt(remix, transform, and build upon) the material for any purpose, even commercially within the parameters below:
Creative Commons (CC): This is a Creative Commons license.
Attribution (BY): Credit must be given to the creator.
*Disclaimer
This summary highlights only some of the key features and terms of the actual license. It is not a license and has no legal value. Carefully review the actual license before using these materials.
1. Introduction
1.1. Genetic Elements Associated with Bacterial HGT
| pathway type | mechanisms | examples | key features |
|---|---|---|---|
| MGE-dependent | plasmids (conjugation), ICEs, transposons, bacteriophages (transduction), GTAs | E. coli (plasmid transfer), Vibrio cholerae (ICE), Salmonella (phage-mediated) | requires mobile genetic elements; often encodes transfer machinery; efficient and directional |
| MGE-independent | natural transformation (uptake of free DNA), nanotube-mediated exchange, extracellular vesicles (OMVs, MVs, and O-IMVs) | Acinetobacter baylyi (transformation), Bacillus subtilis (nanotubes), Pseudomonas aeruginosa (OMVs) | does not rely on self-mobilizing elements; opportunistic; influenced by environmental conditions |
1.2. Classes of Genetic Elements in HGT
| Genetic Element | Mobile genetic element (Y/N) | Associated HGT Mechanism | Examples |
|---|---|---|---|
| plasmids | yes | conjugation, nanotube-mediated transfer | pNDM, Inc.F, Inc.A/C, Inc. X, Inc.P (16,20−22,60) |
| bacteriophages | yes | transduction (generalized, specialized, lateral) | ΦCTX, Staphylococcus aureus phages, Enterococcus faecalis phages, Salmonella spp. and E. coli phages, environmental phages (28−30,61) |
| integrative and conjugative elements (ICEs) | yes | conjugation | Tn916 family ICE elements, ICESt1, ICESt, ICEBs1, SXT/R391 family ICEs, ICEclc (33−35,51) |
| transposons | yes | mobilization via plasmids or ICEs | Tn2, Tn3, Tn21, Tn7, Tn1546, Tn125, (39,45,47−49) |
| free environmental DNA | no | natural transformation | Streptococcus pneumoniae, Neisseria gonorrheae |
Figure 1
Figure 1. Canonical and noncanonical routes of HGT. (A) Conjugation involves cell-to-cell contact. (B) Transformation involves the uptake of extracellular DNA from the environment. (C) Transduction involves attachment of phage to bacteria followed by subsequent injection of its DNA into the host. (D) HGT through membrane vesicles, where bacteria release vesicles containing cellular materials that are taken up by recipient cells. (E) HGT through nanotubes, which are membrane extensions forming direct bridges between neighboring bacterial cells to enable exchange of cytoplasmic materials. (F) HGT through autolysis. Autolysis involves the self-lysis of a subpopulation of bacterial cells, releasing extracellular DNA that becomes available for uptake by naturally competent bacterial cells. (G) HGT via gene transfer agents (GTAs), which package host DNA fragments and release them upon lysis for uptake by recipient cells.
Figure 2
Figure 2. Crystal structure of stabilized TEM-1 beta-lactamase variant v.13 carrying G238S mutation. (129) The blue colored region with hydrogen bonding is the mutation region.
Figure 3
Figure 3. Schematic representation of the efflux pump system in the Gram-negative bacterial cell membrane. The outer membrane (OM) at the top, the peptidoglycan layer (PGL) in the middle, and the inner membrane (IM) at the bottom, with the cytoplasm beneath. Porins are water-filled channels embedded in the outer membrane that allow hydrophilic molecules and small metabolites to pass through passively. They are essential for nutrient uptake and waste expulsion, functioning as molecular sieves that exclude larger or hydrophobic molecules.
Figure 4
Figure 4. Structure of class 1 integrons. Integrons function as modular genetic elements that facilitate the acquisition, chromosomal insertion, and expression of gene cassettes, frequently encoding antibiotic resistance determinants, via a site-specific recombination mechanism catalyzed by an integrase enzyme.
1.3. Canonical Horizontal Gene Transfer (HGT): The Big Three
| Type of Transduction | Mechanism | Key Features | Example |
|---|---|---|---|
| generalized | random bacterial DNA fragments are accidentally packaged into phage capsids during the lytic cycle. | transfers any gene; occurs at low frequency. | Salmonella ssp., E. coli |
| specialized | prophage excises incorrectly from host chromosome, taking adjacent bacterial genes with it. | transfers only genes near integration site. | λ phage in E. coli (gal, bio) |
| lateral/progressive | phage replicates while integrated, packaging large contiguous regions of bacterial DNA. | transfers large chromosomal segments; newly discovered. | Salmonella, Staphylococcus aureus |
1.3.1. Transformation
1.3.2. Transduction
1.3.3. Conjugation
1.4. Noncanonical Horizontal Gene Transfer Agents─The New Kids on the Block
1.4.1. Gene Transfer Agents (GTAs)
1.4.2. Bacterial Extracellular Vesicles
| Type | Size | Structure | Origin | Cargo | Role in HGT |
|---|---|---|---|---|---|
| outer membrane vesicles (OMVs) | ∼20–250 nm | single lipid bilayer | Gram-negative outer membrane | DNA, RNA, proteins, toxins | deliver genetic material directly to recipient cells |
| membrane vesicles (MVs) | ∼20–400 nm | single lipid bilayer | Gram-positive cytoplasmic membrane | chromosomal fragments, plasmids | facilitate gene transfer between Gram-positive bacteria |
| outer-inner membrane vesicles (O-IMVs) | ∼100–450 nm (often larger than OMVs because they include cytoplasmic content) | double-layered structures | both inner and outer membranes of Gram-negative bacteria | cytoplasmic content including DNA | enable transfer of larger genetic elements and cytoplasmic molecules |
1.4.3. Nanotube-Mediated DNA Exchange
| Feature | Conjugation | Nanotube exchange |
|---|---|---|
| structure used | sex pilus + type IV secretion system | membrane nanotubes |
| genetic requirement | conjugative plasmid or ICE | nonspecific |
| directionality | unidirectional (donor → recipient) | bidirectional |
| specificity | species-specific or plasmid-specific | broad, even cross-species |
| transfer type | primarily plasmids | plasmids, chromosomal DNA, proteins |
1.4.4. CTCNT-Mediated Plasmid Transfer (CTCNT-P)
2. HGT and the Global Spread of Resistance Evolution
| Mechanism | Description | Examples | Genetic elements driving global ARG spread via HGT |
|---|---|---|---|
| enzymatic inactivation | enzymes degrade or modify antibiotics. | TEM, SHV, mecA, and other β-lactamases, aminoglycoside-modifying enzymes | plasmids (Inc.F, Inc.I, and Inc.P), class 1 integrons, transposons (Tn3 and Tn4401) |
| gain of function mutation | spontaneous changes in chromosomal genes altering antibiotic targets or permeability. | rpoB (rifampin), gyrA (fluoroquinolones) | occasionally mobilized via transposons (e.g., Tn3) |
| efflux pumps | active expulsion of antibiotics from the cell, reducing intracellular concentration. | Tet(A), AcrAB-TolC | plasmid-borne efflux genes, integrons |
| target modification | alteration of antibiotic binding sites to reduce drug affinity. | erm genes (macrolides), mosaic PBPs | plasmid-mediated genes, transposons |
| target protection | proteins shield antibiotic targets without altering their function. | Tet(M), Tet(O), and Qnr proteins | plasmids, integrons, gene cassettes, and examples of innate antibiotic drug resistance |
2.1. Enzymatic Inactivation
2.2. Gain-of-Function Mutation
2.3. Efflux Pumps
2.4. Target Modification
2.4.1. Target Protection
3. Targeting HGT as a Strategy to Control the Spread of Drug Resistance
3.1. HGT in a Biofilm Environment
Figure 5
Figure 5. Proposed routes to control HGT. Nanoparticle disruption of the biofilm architecture, this essentially prevent cell proximity, thus preventing conjugation. In some cases, the nanoparticle may actively compete with the bacteria for the extracellular DNA. Sequestering the extracellular DNA is a route to limit resistance evolution.
3.1.1. Environmental Intervention
3.2. Targeting HGT as a Strategy to Control the Spread of Drug Resistance
3.3. Eliminating ARGs in the Environment
3.3.1. Bulk ARG Ablation Technologies
3.3.2. Membrane Filtration Technologies
3.3.3. Advanced Oxidation Processes
Figure 6
Figure 6. Proposed methods of controlling HGT in bacterial niche. (A) Iron activates the peroxymonosulfate (PMS) resulting to the production of radical species that degrade the cell membrane and disrupt enzyme metabolism, thus preventing conjugative transfer of resistance genes. (B) Silica nanoparticle binds to extracellular DNA by an attractive interaction between DNA phosphate groups and surface silanol groups on silica, thus sequestering the DNA and blocking transformation.
3.3.4. Absorption Technologies
3.3.5. Cellular/Molecular Based Anti-HGT/ARGs Technologies
3.3.6. Chemical/Material Interrogation of Cellular Processes Required for HGT
Figure 7
Figure 7. Strategies to control horizontal gene transfer. (A) Conjugation inhibiting drugs that inactivate components of proteins involved in the plasmid transfer machinery, specifically the Type IV coupling protein that links the relaxosome and the Type IV secretion system (T4SS). (B). Plasmid curing involves a variety of chemicals including detergents, biocides, DNA intercalating agents, antibiotics, psychotropic drugs, and plant-derived compounds that target eDNA for degradation. (C) Competence blockers (COM-blockers) disrupt the proton motive force, which inhibits quorum-sensing signaling via CSP, thereby preventing activation of the DNA uptake machinery.
3.3.7. Competence Blockers: Antievolution Compounds
3.3.8. Activating Bacterial Innate Immunity to Combat HGT
Figure 8
Figure 8. Bacterial innate immune defense against phages/plasmids. (A). Bacteria possess sophisticated innate immune mechanisms capable of mounting effective responses against invading phages and foreign genetic elements. These defenses comprise a diverse array of genetically encoded systems that are frequently colocalized within discrete genomic regions known as “defense islands”. The repertoire of characterized defense systems continues to expand and currently encompass restriction-modification systems, CRISPR-Cas machinery, Argonaute-based defense pathways, cyclic-oligonucleotide-based antiphage signaling systems (CBASS), and abortive-infection modules, each representing a functionally distinct strategy for detecting and neutralizing nonself-nucleic acids. (B). CRISPR-based prokaryotic gene drives targeting plasmid ARGs. Through this mechanism, the DoS plasmid is able to disseminate novel genetic information throughout a target bacterial population while simultaneously outcompeting the conjugative plasmid, displacing it not only as the primary vehicle of horizontal gene transfer but also as a heritable element passed vertically to daughter cells.
3.3.9. Engineered Crispr-Based Anti-HGT Technologies
3.3.10. Gene Drives
3.3.10.1. Antiplasmid Systems to Combat ARGs Spread
4. Conclusion
References
This article references 316 other publications.
- 1Baym, M.; Stone, L. K.; Kishony, R. Multidrug evolutionary strategies to reverse antibiotic resistance. Science. 2016, 351 (6268), aad3292, DOI: 10.1126/science.aad3292Google ScholarThere is no corresponding record for this reference.
- 2Zavaleta-Monestel, E.; Arguedas-Chacón, S.; Rojas-Chinchilla, C.; Díaz-Madriz, J. P. Antimicrobial Resistance: An Emerging Global Threat to Modern Medicine. Cureus. 2025, DOI: 10.7759/cureus.97668Google ScholarThere is no corresponding record for this reference.
- 3Mouzakis, A.; Panagopoulos, P.; Papazoglou, D.; Petrakis, V. A Comprehensive Review of Nanoparticles in the Fight Against Antimicrobial Resistance. Pathogens 2025, 14 (11), 1090, DOI: 10.3390/pathogens14111090Google ScholarThere is no corresponding record for this reference.
- 4Smith, P. A.; Romesberg, F. E. Combating bacteria and drug resistance by inhibiting mechanisms of persistence and adaptation. Nat. Chem. Biol. 2007, 3 (9), 549– 556, DOI: 10.1038/nchembio.2007.27Google ScholarThere is no corresponding record for this reference.
- 5Zhang, S.; Lu, J.; Wang, Y.; Verstraete, W.; Yuan, Z.; Guo, J. Insights of metallic nanoparticles and ions in accelerating the bacterial uptake of antibiotic resistance genes. J. Hazard. Mater. 2022, 421, 126728, DOI: 10.1016/j.jhazmat.2021.126728Google ScholarThere is no corresponding record for this reference.
- 6Basarab, G. S.; Ghorpade, S.; Gibhard, L.; Mueller, R.; Njoroge, M.; Peton, N.; Govender, P.; Massoudi, L. M.; Robertson, G. T.; Lenaerts, A. J. Spiropyrimidinetriones: a Class of DNA Gyrase Inhibitors with Activity against Mycobacterium tuberculosis and without Cross-Resistance to Fluoroquinolones. Antimicrob. Agents Chemother. 2022, 66 (4), e02192-21 DOI: 10.1128/aac.02192-21Google ScholarThere is no corresponding record for this reference.
- 7Dubnau, D.; Blokesch, M. Mechanisms of DNA Uptake by Naturally Competent Bacteria. Annu. Rev. Genet. 2019, 53 (1), 217– 237, DOI: 10.1146/annurev-genet-112618-043641Google ScholarThere is no corresponding record for this reference.
- 8Khedkar, S.; Smyshlyaev, G.; Letunic, I. Landscape of mobile genetic elements and their antibiotic resistance cargo in prokaryotic genomes. Nucleic Acids Res. 2022, 50 (6), 3155– 3168, DOI: 10.1093/nar/gkac163Google ScholarThere is no corresponding record for this reference.
- 9Lang, A. S.; Buchan, A.; Burrus, V. Interactions and evolutionary relationships among bacterial mobile genetic elements. Nat. Rev. Microbiol. 2025, 23 (7), 423– 438, DOI: 10.1038/s41579-025-01157-yGoogle ScholarThere is no corresponding record for this reference.
- 10Haudiquet, M.; De Sousa, J. M.; Touchon, M.; Rocha, E. P. C. Selfish, promiscuous and sometimes useful: how mobile genetic elements drive horizontal gene transfer in microbial populations. Philosophical Transactions of the Royal Society B 2022, 377 (1861), 20210234, DOI: 10.1098/rstb.2021.0234Google ScholarThere is no corresponding record for this reference.
- 11Di Giacomo, S.; Toussaint, F.; Ledesma-García, L. Expanding natural transformation to improve beneficial lactic acid bacteria. FEMS Microbiol. Rev. 2022, 46 (4), fuac014, DOI: 10.1093/femsre/fuac014Google ScholarThere is no corresponding record for this reference.
- 12Marinacci, B.; Krzyżek, P.; Pellegrini, B.; Turacchio, G.; Grande, R. Latest Update on Outer Membrane Vesicles and Their Role in Horizontal Gene Transfer: A Mini-Review. Membranes 2023, 13 (11), 860, DOI: 10.3390/membranes13110860Google ScholarThere is no corresponding record for this reference.
- 13Morawska, L. P.; Kuipers, O. P. Cell-to-cell non-conjugative plasmid transfer between Bacillus subtilis and lactic acid bacteria. Microb. Biotechnol. 2023, 16 (4), 784– 798, DOI: 10.1111/1751-7915.14195Google ScholarThere is no corresponding record for this reference.
- 14Rodríguez-Beltrán, J.; DelaFuente, J.; León-Sampedro, R.; MacLean, R. C.; San Millán, A. ́. Beyond horizontal gene transfer: the role of plasmids in bacterial evolution. Nat. Rev. Microbiol. 2021, 19 (6), 347– 359, DOI: 10.1038/s41579-020-00497-1Google ScholarThere is no corresponding record for this reference.
- 15Hashimoto, Y.; Taniguchi, M.; Uesaka, K. Novel Multidrug-Resistant Enterococcal Mobile Linear Plasmid pELF1 Encoding vanA and vanM Gene Clusters From a Japanese Vancomycin-Resistant Enterococci Isolate. Front. Microbiol. 2019, 10, 2568, DOI: 10.3389/fmicb.2019.02568Google ScholarThere is no corresponding record for this reference.
- 16Rozwandowicz, M.; Brouwer, M. S. M.; Fischer, J. Plasmids carrying antimicrobial resistance genes in Enterobacteriaceae. J. Antimicrob. Chemother. 2018, 73 (5), 1121– 1137, DOI: 10.1093/jac/dkx488Google ScholarThere is no corresponding record for this reference.
- 17Suzuki, H.; Yano, H.; Brown, C. J.; Top, E. M. Predicting Plasmid Promiscuity Based on Genomic Signature. J. Bacteriol. 2010, 192 (22), 6045– 6055, DOI: 10.1128/JB.00277-10Google ScholarThere is no corresponding record for this reference.
- 18Zechner, E. L.; Moncalián, G.; De La Cruz, F. Relaxases and Plasmid Transfer in Gram-Negative Bacteria. Type IV Secretion in Gram-Negative and Gram-Positive Bacteria; Backert, S., Grohmann, E., Eds.; Springer International Publishing, 2017; Vol. 413, pp 93– 113. DOI: 10.1007/978-3-319-75241-9_4 .Google ScholarThere is no corresponding record for this reference.
- 19Smillie, C.; Garcillán-Barcia, M. P.; Francia, M. V.; Rocha, E. P. C.; De La Cruz, F. Mobility of Plasmids. Microbiol. Mol. Biol. Rev. 2010, 74 (3), 434– 452, DOI: 10.1128/MMBR.00020-10Google ScholarThere is no corresponding record for this reference.
- 20Carattoli, A. Resistance Plasmid Families in Enterobacteriaceae. Antimicrob. Agents Chemother. 2009, 53 (6), 2227– 2238, DOI: 10.1128/AAC.01707-08Google ScholarThere is no corresponding record for this reference.
- 21Thomas, C. M.; Nielsen, K. M. Mechanisms of, and Barriers to, Horizontal Gene Transfer between Bacteria. Nat. Rev. Microbiol. 2005, 3 (9), 711– 721, DOI: 10.1038/nrmicro1234Google ScholarThere is no corresponding record for this reference.
- 22Fricke, W. F.; Welch, T. J.; McDermott, P. F. Comparative Genomics of the IncA/C Multidrug Resistance Plasmid Family. J. Bacteriol. 2009, 191 (15), 4750– 4757, DOI: 10.1128/JB.00189-09Google ScholarThere is no corresponding record for this reference.
- 23Guo, X.; Chen, R.; Wang, Q. Global prevalence, characteristics, and future prospects of IncX3 plasmids: A review. Front. Microbiol. 2022, 13, 979558, DOI: 10.3389/fmicb.2022.979558Google ScholarThere is no corresponding record for this reference.
- 24Rasheed, J. K.; Kitchel, B.; Zhu, W. New Delhi Metallo-β-Lactamase-producing Enterobacteriaceae, United States. Emerg Infect Dis. 2013, 19 (6), 870– 878, DOI: 10.3201/eid1906.121515Google ScholarThere is no corresponding record for this reference.
- 25Segundo-Arizmendi, N.; Arellano-Maciel, D.; Rivera-Ramírez, A.; Piña-González, A. M.; López-Leal, G.; Hernández-Baltazar, E. Bacteriophages: A Challenge for Antimicrobial Therapy. Microorganisms 2025, 13 (1), 100, DOI: 10.3390/microorganisms13010100Google ScholarThere is no corresponding record for this reference.
- 26Touchon, M.; Moura De Sousa, J. A.; Rocha, E. P. Embracing the enemy: the diversification of microbial gene repertoires by phage-mediated horizontal gene transfer. Curr. Opin. Microbiol. 2017, 38, 66– 73, DOI: 10.1016/j.mib.2017.04.010Google ScholarThere is no corresponding record for this reference.
- 27Pires, J.; Santos, R.; Monteiro, S. Antibiotic resistance genes in bacteriophages from wastewater treatment plant and hospital wastewaters. Sci. Total Environ. 2023, 892, 164708, DOI: 10.1016/j.scitotenv.2023.164708Google ScholarThere is no corresponding record for this reference.
- 28Waldor, M. K.; Mekalanos, J. J. Lysogenic Conversion by a Filamentous Phage Encoding Cholera Toxin. Science. 1996, 272 (5270), 1910– 1914, DOI: 10.1126/science.272.5270.1910Google ScholarThere is no corresponding record for this reference.
- 29Goerke, C.; Pantucek, R.; Holtfreter, S. Diversity of Prophages in Dominant Staphylococcus aureus Clonal Lineages. J. Bacteriol. 2009, 191 (11), 3462– 3468, DOI: 10.1128/JB.01804-08Google ScholarThere is no corresponding record for this reference.
- 30Colavecchio, A.; Cadieux, B.; Lo, A.; Goodridge, L. D. Bacteriophages Contribute to the Spread of Antibiotic Resistance Genes among Foodborne Pathogens of the Enterobacteriaceae Family - A Review. Front. Microbiol. 2017, 8, 1108, DOI: 10.3389/fmicb.2017.01108Google ScholarThere is no corresponding record for this reference.
- 31Johnson, C. M.; Grossman, A. D. Integrative and Conjugative Elements (ICEs): What They Do and How They Work. Annu. Rev. Genet. 2015, 49 (1), 577– 601, DOI: 10.1146/annurev-genet-112414-055018Google ScholarThere is no corresponding record for this reference.
- 32Benigno, V.; Carraro, N.; Sarton-Lohéac, G.; Romano-Bertrand, S.; Blanc, D. S.; Van, D. M., JR. Diversity and evolution of an abundant ICE clc family of integrative and conjugative elements in Pseudomonas aeruginosa. Gales AC, ed. mSphere 2023, 8 (6), e00517–23 DOI: 10.1128/msphere.00517-23Google ScholarThere is no corresponding record for this reference.
- 33Roberts, A. P.; Mullany, P. Tn 916 -like genetic elements: a diverse group of modular mobile elements conferring antibiotic resistance. FEMS Microbiol. Rev. 2011, 35 (5), 856– 871, DOI: 10.1111/j.1574-6976.2011.00283.xGoogle ScholarThere is no corresponding record for this reference.
- 34Auchtung, J. M.; Aleksanyan, N.; Bulku, A.; Berkmen, M. B. Biology of ICE Bs1, an integrative and conjugative element in Bacillus subtilis. Plasmid 2016, 86, 14– 25, DOI: 10.1016/j.plasmid.2016.07.001Google ScholarThere is no corresponding record for this reference.
- 35Wozniak, R. A. F.; Waldor, M. K. Integrative and conjugative elements: mosaic mobile genetic elements enabling dynamic lateral gene flow. Nat. Rev. Microbiol. 2010, 8 (8), 552– 563, DOI: 10.1038/nrmicro2382Google ScholarThere is no corresponding record for this reference.
- 36Burrus, V.; Pavlovic, G.; Decaris, B.; Guédon, G. The ICESt1 element of Streptococcus thermophilus belongs to a large family of integrative and conjugative elements that exchange modules and change their specificity of integration. Plasmid 2002, 48 (2), 77– 97, DOI: 10.1016/S0147-619X(02)00102-6Google ScholarThere is no corresponding record for this reference.
- 37Blokesch, M. Defence systems encoded by core genomic islands of seventh pandemic Vibrio cholerae. Phil Trans R Soc. B 2025, 380 (1934), 20240083, DOI: 10.1098/rstb.2024.0083Google ScholarThere is no corresponding record for this reference.
- 38Nusrat, S.; Aliyu, M.; Zohora, F. T. Mechanisms of antimicrobial resistance: From genetic evolution to clinical manifestations. AIMSMICRO. 2025, 11 (4), 1007– 1034, DOI: 10.3934/microbiol.2025045Google ScholarThere is no corresponding record for this reference.
- 39Partridge, S. R.; Kwong, S. M.; Firth, N.; Jensen, S. O. Mobile Genetic Elements Associated with Antimicrobial Resistance. Clin. Microbiol. Rev. 2018, 31 (4), e00088-17 DOI: 10.1128/CMR.00088-17Google ScholarThere is no corresponding record for this reference.
- 40Roy, S.; Nandy, S.; Morita, D. Genomic analysis of a novel high-risk ST5217/ExoU+/O11 clone of carbapenem-resistant OXA-181- and VIM-2-producing Pseudomonas aeruginosa in India. J. Global Antimicrob. Resist. 2026, 46, 158– 161, DOI: 10.1016/j.jgar.2025.12.002Google ScholarThere is no corresponding record for this reference.
- 41Calbet, A. Pelagic Shuttles of Antibiotic Resistance Genes: Zooplankton as Overlooked Vectors Across Space and Food Webs. Microb. Ecol. 2026, 89 (1), 12, DOI: 10.1007/s00248-025-02669-zGoogle ScholarThere is no corresponding record for this reference.
- 42Wachino, J. Horizontal Gene Transfer Systems for Spread of Antibiotic Resistance in Gram-Negative Bacteria. Microbiol. Immunol. 2025, 69 (7), 367– 376, DOI: 10.1111/1348-0421.13222Google ScholarThere is no corresponding record for this reference.
- 43Wang, X.; Chen, Z.; Liu, C.; Zhang, Z.; Deng, Y.; Tao, L.; Tiedje, J. M.; Deng, J. Type I-F CRISPR-associated transposons contribute to genomic plasticity in Shewanella and mediate efficient programmable DNA integration. Microb. Genomics 2025, 11(8). DOI: 10.1099/mgen.0.001476 .Google ScholarThere is no corresponding record for this reference.
- 44Deng, Y.; Bao, X.; Ji, L. Resistance integrons: class 1, 2 and 3 integrons. Ann. Clin. Microbiol. Antimicrob. 2015, 14 (1), 45, DOI: 10.1186/s12941-015-0100-6Google ScholarThere is no corresponding record for this reference.
- 45Stokes, H. W.; Gillings, M. R. Gene flow, mobile genetic elements and the recruitment of antibiotic resistance genes into Gram-negative pathogens. FEMS Microbiol. Rev. 2011, 35 (5), 790– 819, DOI: 10.1111/j.1574-6976.2011.00273.xGoogle ScholarThere is no corresponding record for this reference.
- 46Nicolas, E.; Lambin, M.; Dandoy, D.; The Tn 3 -family of Replicative Transposons Mobile DNA III; Craig, N. L., Chandler, M., Gellert, M., Lambowitz, A. M., Rice, P. A., Sandmeyer, S. B., Eds.; ASM Press, 2015; pp 693– 726. DOI: 10.1128/9781555819217.ch32 .Google ScholarThere is no corresponding record for this reference.
- 47Peters, J. E. Targeted transposition with Tn7 elements: safe sites, mobile plasmids, CRISPR/Cas and beyond. Mol. Microbiol. 2019, 112 (6), 1635– 1644, DOI: 10.1111/mmi.14383Google ScholarThere is no corresponding record for this reference.
- 48Peters, J. E. Tn7 Mobile DNA III; Craig, N. L., Chandler, M., Gellert, M., Lambowitz, A. M., Rice, P. A., Sandmeyer, S. B., Eds.; ASM Press, 2015; pp 647– 667. DOI: 10.1128/9781555819217.ch30 .Google ScholarThere is no corresponding record for this reference.
- 49Courvalin, P. Vancomycin Resistance in Gram-Positive Cocci. Clin. Infect. Dis. 2006, 42 (Supplement_1), S25– S34, DOI: 10.1086/491711Google ScholarThere is no corresponding record for this reference.
- 50Bush, K. Evolution of β-Lactamases: Past, Present, and Future. Antibiotic Discovery and Development; Dougherty, T. J., Pucci, M. J., Eds.; Springer US, 2012; pp 427– 453. DOI: 10.1007/978-1-4614-1400-1_12 .Google ScholarThere is no corresponding record for this reference.
- 51Naas, T.; Cuzon, G.; Villegas, M. V.; Lartigue, M. F.; Quinn, J. P.; Nordmann, P. Genetic Structures at the Origin of Acquisition of the β-Lactamase blaKPC Gene. Antimicrob. Agents Chemother. 2008, 52 (4), 1257– 1263, DOI: 10.1128/AAC.01451-07Google ScholarThere is no corresponding record for this reference.
- 52Ameyama, S.; Onodera, S.; Takahata, M. Mosaic-Like Structure of Penicillin-Binding Protein 2 Gene (penA) in Clinical Isolates of Neisseria gonorrhoeae with Reduced Susceptibility to Cefixime. Antimicrob. Agents Chemother. 2002, 46 (12), 3744– 3749, DOI: 10.1128/AAC.46.12.3744-3749.2002Google ScholarThere is no corresponding record for this reference.
- 53Maziero, M.; Juillot, D.; Mortier-Barrière, I. A toxin/antitoxin system targeting the replication sliding-clamp induces competence in Streptococcus pneumoniae. Kjos M, ed. PLoS Genet. 2025, 21 (12), e1011863 DOI: 10.1371/journal.pgen.1011863Google ScholarThere is no corresponding record for this reference.
- 54Hakenbeck, R. Transformation in: mosaic genes and the regulation of competence. Res. Microbiol. 2000, 151 (6), 453– 456, DOI: 10.1016/S0923-2508(00)00170-4Google ScholarThere is no corresponding record for this reference.
- 55Wadsworth, C. B.; Goytia, M.; Shafer, W. M. Commensal Neisseria and Antimicrobial-Resistant Gonorrhea. Annu. Rev. Microbiol. 2025, 79 (1), 215– 240, DOI: 10.1146/annurev-micro-022024-024306Google ScholarThere is no corresponding record for this reference.
- 56Unitt, A.; Maiden, M.; Harrison, O. Characterizing the diversity and commensal origins of penA mosaicism in the genus Neisseria. Microb. Genomics 2024, 10(2). DOI: 10.1099/mgen.0.001209 .Google ScholarThere is no corresponding record for this reference.
- 57Hanao, M.; Aoki, K.; Ishii, Y.; Shimuta, K.; Ohnishi, M.; Tateda, K. Molecular characterization of Neisseria gonorrhoeae isolates collected through a national surveillance programme in Japan, 2013: evidence of the emergence of a ceftriaxone-resistant strain from a ceftriaxone-susceptible lineage. J. Antimicrob. Chemother. 2021, 76 (7), 1769– 1775, DOI: 10.1093/jac/dkab104Google ScholarThere is no corresponding record for this reference.
- 58Liu, E. Y. M.; Chang, J. C.; Lin, J. C.; Chang, F. Y.; Fung, C. P. Important Mutations Contributing to High-Level Penicillin Resistance in Taiwan19F −14, Taiwan23F −15, and Spain23F −1 of Streptococcus pneumoniae Isolated from Taiwan. Microbial Drug Resistance. 2016, 22 (8), 646– 654, DOI: 10.1089/mdr.2015.0261Google ScholarThere is no corresponding record for this reference.
- 59Panickar, A.; Manoharan, A.; Ramaiah, S. Machine learning-based virtual screening and density functional theory characterisation of natural inhibitors targeting mutant PBP2x in Streptococcus pneumoniae. Sci. Rep. 2025, 15 (1), 39164, DOI: 10.1038/s41598-025-24222-1Google ScholarThere is no corresponding record for this reference.
- 60Fang, L.; Chen, R.; Li, C. The association between the genetic structures of commonly incompatible plasmids in Gram-negative bacteria, their distribution and the resistance genes. Front. Cell. Infect. Microbiol. 2024, 14, 1472876, DOI: 10.3389/fcimb.2024.1472876Google ScholarThere is no corresponding record for this reference.
- 61Mazaheri Nezhad Fard, R.; Barton, M. D.; Heuzenroeder, M. W. Bacteriophage-mediated transduction of antibiotic resistance in enterococci: Transduction in Enterococcus spp. Lett. Appl. Microbiol. 2011, 52 (6), 559– 564, DOI: 10.1111/j.1472-765X.2011.03043.xGoogle ScholarThere is no corresponding record for this reference.
- 62Huang, M.; Liu, M.; Huang, L. The activation and limitation of the bacterial natural transformation system: The function in genome evolution and stability. Microbiol. Res. 2021, 252, 126856, DOI: 10.1016/j.micres.2021.126856Google ScholarThere is no corresponding record for this reference.
- 63Soucy, S. M.; Huang, J.; Gogarten, J. P. Horizontal gene transfer: building the web of life. Nat. Rev. Genet. 2015, 16 (8), 472– 482, DOI: 10.1038/nrg3962Google ScholarThere is no corresponding record for this reference.
- 64Arnold, B. J.; Huang, I. T.; Hanage, W. P. Horizontal gene transfer and adaptive evolution in bacteria. Nat. Rev. Microbiol. 2022, 20 (4), 206– 218, DOI: 10.1038/s41579-021-00650-4Google ScholarThere is no corresponding record for this reference.
- 65McCallum, M., Burrows, L. L., Howell, P. L. The Dynamic Structures of the Type IV Pilus. Microbiol Spectr , 2019, 7 (2) DOI: 10.1128/microbiolspec.PSIB-0006-2018 .Google ScholarThere is no corresponding record for this reference.
- 66Luna Rico, A.; Zheng, W.; Petiot, N.; Egelman, E. H.; Francetic, O. Functional reconstitution of the type IVa pilus assembly system from enterohaemorrhagic Escherichia coli. Mol. Microbiol. 2019, 111 (3), 732– 749, DOI: 10.1111/mmi.14188Google ScholarThere is no corresponding record for this reference.
- 67Ellison, C. K.; Whitfield, G. B.; Brun, Y. V. Type IV Pili: dynamic bacterial nanomachines. FEMS Microbiol. Rev. 2022, 46 (2), fuab053, DOI: 10.1093/femsre/fuab053Google ScholarThere is no corresponding record for this reference.
- 68Blokesch, M. Natural competence for transformation. Curr. Biol. 2016, 26 (21), R1126– R1130, DOI: 10.1016/j.cub.2016.08.058Google ScholarThere is no corresponding record for this reference.
- 69Yu, Z.; Wang, Y.; Henderson, I. R.; Guo, J. Artificial sweeteners stimulate horizontal transfer of extracellular antibiotic resistance genes through natural transformation. ISME J. 2022, 16 (2), 543– 554, DOI: 10.1038/s41396-021-01095-6Google ScholarThere is no corresponding record for this reference.
- 70Johnsborg, O.; Eldholm, V.; Håvarstein, L. S. Natural genetic transformation: prevalence, mechanisms and function. Res. Microbiol. 2007, 158 (10), 767– 778, DOI: 10.1016/j.resmic.2007.09.004Google ScholarThere is no corresponding record for this reference.
- 71Johnston, C.; Martin, B.; Fichant, G.; Polard, P.; Claverys, J. P. Bacterial transformation: distribution, shared mechanisms and divergent control. Nat. Rev. Microbiol. 2014, 12 (3), 181– 196, DOI: 10.1038/nrmicro3199Google ScholarThere is no corresponding record for this reference.
- 72Chen, J.; Quiles-Puchalt, N.; Chiang, Y. N. Genome hypermobility by lateral transduction. Science. 2018, 362 (6411), 207– 212, DOI: 10.1126/science.aat5867Google ScholarThere is no corresponding record for this reference.
- 73Bhattacharya, T.; Chatterjee, S.; Maiti, D. Molecular analysis of the rstR and orfU genes of the CTX prophages integrated in the small chromosomes of environmental Vibrio cholerae non-O1, non-O139 strains. Environ. Microbiol. 2006, 8 (3), 526– 634, DOI: 10.1111/j.1462-2920.2005.00932.xGoogle ScholarThere is no corresponding record for this reference.
- 74Zhao, Y.; Ma, Y.; Vasileiou, C.; Farr, A. D.; Rogers, D. W.; Rainey, P. B. Jumbo phage-mediated transduction of genomic islands. Proc. Natl. Acad. Sci. U. S. A. 2025, 122 (44), e2512465122 DOI: 10.1073/pnas.2512465122Google ScholarThere is no corresponding record for this reference.
- 75Waldor, M. K.; Friedman, D. I. Phage regulatory circuits and virulence gene expression. Curr. Opin. Microbiol. 2005, 8 (4), 459– 465, DOI: 10.1016/j.mib.2005.06.001Google ScholarThere is no corresponding record for this reference.
- 76Chiang, Y. N.; Penadés, J. R.; Chen, J. Genetic transduction by phages and chromosomal islands: The new and noncanonical. Kline KA, ed. PLoS Pathog. 2019, 15 (8), e1007878 DOI: 10.1371/journal.ppat.1007878Google ScholarThere is no corresponding record for this reference.
- 77Leclerc, Q. J.; Wildfire, J.; Gupta, A.; Lindsay, J. A.; Knight, G. M. Growth-Dependent Predation and Generalized Transduction of Antimicrobial Resistance by Bacteriophage. Gilbert JA, ed. mSystems 2022, 7 (2), e00135–22 DOI: 10.1128/msystems.00135-22Google ScholarThere is no corresponding record for this reference.
- 78Waksman, G. From conjugation to T4S systems in Gram-negative bacteria: a mechanistic biology perspective. EMBO Rep. 2019, 20 (2), e47012 DOI: 10.15252/embr.201847012Google ScholarThere is no corresponding record for this reference.
- 79Humphrey, S.; Fillol-Salom, A.; Quiles-Puchalt, N. Bacterial chromosomal mobility via lateral transduction exceeds that of classical mobile genetic elements. Nat. Commun. 2021, 12 (1), 6509, DOI: 10.1038/s41467-021-26004-5Google ScholarThere is no corresponding record for this reference.
- 80Fillol-Salom, A.; Bacigalupe, R.; Humphrey, S.; Chiang, Y. N.; Chen, J.; Penadés, JR. Lateral transduction is inherent to the life cycle of the archetypical Salmonella phage P22. Nat. Commun. 2021, 12 (1), 6510, DOI: 10.1038/s41467-021-26520-4Google ScholarThere is no corresponding record for this reference.
- 81Cabezón, E.; De La Cruz, F.; Arechaga, I. Conjugation Inhibitors and Their Potential Use to Prevent Dissemination of Antibiotic Resistance Genes in Bacteria. Front. Microbiol. 2017, 8, 2329, DOI: 10.3389/fmicb.2017.02329Google ScholarThere is no corresponding record for this reference.
- 82Sher, A. A.; VanAllen, M. E.; Ahmed, H. Conjugative RP4 Plasmid-Mediated Transfer of Antibiotic Resistance Genes to Commensal and Multidrug-Resistant Enteric Bacteria In Vitro. Microorganisms 2023, 11 (1), 193, DOI: 10.3390/microorganisms11010193Google ScholarThere is no corresponding record for this reference.
- 83Schröder, G.; Lanka, E. The mating pair formation system of conjugative plasmids–A versatile secretion machinery for transfer of proteins and DNA. Plasmid 2005, 54 (1), 1– 25, DOI: 10.1016/j.plasmid.2005.02.001Google ScholarThere is no corresponding record for this reference.
- 84Bañuelos-Vazquez, L. A.; Torres Tejerizo, G.; Brom, S. Regulation of conjugative transfer of plasmids and integrative conjugative elements. Plasmid 2017, 91, 82– 89, DOI: 10.1016/j.plasmid.2017.04.002Google ScholarThere is no corresponding record for this reference.
- 85Rutherford, S. T.; Bassler, B. L. Bacterial Quorum Sensing: Its Role in Virulence and Possibilities for Its Control. Cold Spring Harbor Perspect. Med. 2012, 2 (11), a012427, DOI: 10.1101/cshperspect.a012427Google ScholarThere is no corresponding record for this reference.
- 86Chen, G.; Swem, L. R.; Swem, D. L. A Strategy for Antagonizing Quorum Sensing. Mol. Cell 2011, 42 (2), 199– 209, DOI: 10.1016/j.molcel.2011.04.003Google ScholarThere is no corresponding record for this reference.
- 87Lu, Y.; Zeng, J.; Wu, B.; E, S.; Wang, L.; Cai, R.; Zhang, N.; Li, Y.; Huang, X.; Huang, B.; Quorum Sensing N-acyl Homoserine Lactones-SdiA Suppresses Escherichia coli-Pseudomonas aeruginosa Conjugation through Inhibiting traI Expression. Front. Cell. Infect. Microbiol. 2017, 7. DOI: 10.3389/fcimb.2017.00007 .Google ScholarThere is no corresponding record for this reference.
- 88García-Aljaro, C.; Ballesté, E.; Muniesa, M. Beyond the canonical strategies of horizontal gene transfer in prokaryotes. Curr. Opin. Microbiol. 2017, 38, 95– 105, DOI: 10.1016/j.mib.2017.04.011Google ScholarThere is no corresponding record for this reference.
- 89Vos, M.; Buckling, A.; Kuijper, B. Why do mobile genetic elements transfer DNA of their hosts?. Trends Genet. 2024, 40 (11), 927– 938, DOI: 10.1016/j.tig.2024.07.008Google ScholarThere is no corresponding record for this reference.
- 90Bárdy, P.; Füzik, T.; Hrebík, D.; Pantůček, R.; Thomas Beatty, J.; Plevka, P. Structure and mechanism of DNA delivery of a gene transfer agent. Nat. Commun. 2020, 11 (1), 3034, DOI: 10.1038/s41467-020-16669-9Google ScholarThere is no corresponding record for this reference.
- 91Savory, E. A.; Fuller, S. L.; Weisberg, A. J. Evolutionary transitions between beneficial and phytopathogenic Rhodococcus challenge disease management. eLife 2017, 6, e30925 DOI: 10.7554/eLife.30925Google ScholarThere is no corresponding record for this reference.
- 92Rivard, N., Colwell, R. R., Burrus, V. Antibiotic Resistance in Vibrio cholerae: Mechanistic Insights from IncC Plasmid-Mediated Dissemination of a Novel Family of Genomic Islands Inserted at trmE. mSphere , 2020; 5(4) DOI: 10.1128/msphere.00748-20 . doi: 10.1128/msphere.00748-20.Google ScholarThere is no corresponding record for this reference.
- 93Fogg, P. C. M. Identification and characterization of a direct activator of a gene transfer agent. Nat. Commun. 2019, 10 (1), 595, DOI: 10.1038/s41467-019-08526-1Google ScholarThere is no corresponding record for this reference.
- 94Tran, N. T.; Le, T. B. K. Control of a gene transfer agent cluster in Caulobacter crescentus by transcriptional activation and anti-termination. Nat. Commun. 2024, 15 (1), 4749, DOI: 10.1038/s41467-024-49114-2Google ScholarThere is no corresponding record for this reference.
- 95Devati, M. S.; Jnana, A.; Kidd, S. P. Decoding bacterial extracellular vesicles: A review on isolation and characterization techniques. Arch. Microbiol. 2026, 208 (1), 63, DOI: 10.1007/s00203-025-04628-1Google ScholarThere is no corresponding record for this reference.
- 96Xu, Y.; Xie, C.; Liu, Y.; Qin, X.; Liu, J. An update on our understanding of Gram-positive bacterial membrane vesicles: discovery, functions, and applications. Front. Cell. Infect. Microbiol. 2023, 13, 1273813, DOI: 10.3389/fcimb.2023.1273813Google ScholarThere is no corresponding record for this reference.
- 97Pérez-Cruz, C.; Delgado, L.; López-Iglesias, C.; Mercade, E. Outer-Inner Membrane Vesicles Naturally Secreted by Gram-Negative Pathogenic Bacteria. Rudel T, ed. PLoS One 2015, 10 (1), e0116896 DOI: 10.1371/journal.pone.0116896Google ScholarThere is no corresponding record for this reference.
- 98Aktar, S.; Okamoto, Y.; Ueno, S. Incorporation of Plasmid DNA Into Bacterial Membrane Vesicles by Peptidoglycan Defects in Escherichia coli. Front. Microbiol. 2021, 12, 747606, DOI: 10.3389/fmicb.2021.747606Google ScholarThere is no corresponding record for this reference.
- 99Zhao, X.; Wei, Y.; Bu, Y.; Ren, X.; Dong, Z. Review on bacterial outer membrane vesicles: structure, vesicle formation, separation and biotechnological applications. Microb. Cell Fact. 2025, 24 (1), 27, DOI: 10.1186/s12934-025-02653-9Google ScholarThere is no corresponding record for this reference.
- 100Nagakubo, T.; Nomura, N.; Toyofuku, M. Cracking Open Bacterial Membrane Vesicles. Front. Microbiol. 2020, 10, 3026, DOI: 10.3389/fmicb.2019.03026Google ScholarThere is no corresponding record for this reference.
- 101Toyofuku, M.; Schild, S.; Kaparakis-Liaskos, M.; Eberl, L. Composition and functions of bacterial membrane vesicles. Nat. Rev. Microbiol. 2023, 21 (7), 415– 430, DOI: 10.1038/s41579-023-00875-5Google ScholarThere is no corresponding record for this reference.
- 102Juodeikis, R.; Carding, S. R. Outer Membrane Vesicles: Biogenesis, Functions, and Issues. Microbiol. Mol. Biol. Rev. 2022, 86 (4), e00032-22 DOI: 10.1128/mmbr.00032-22Google ScholarThere is no corresponding record for this reference.
- 103Furuyama, N.; Sircili, M. P. Outer Membrane Vesicles (OMVs) Produced by Gram-Negative Bacteria: Structure, Functions, Biogenesis, and Vaccine Application. Gebre AK, ed. BioMed Res. Int. 2021, 2021 (1), 1490732, DOI: 10.1155/2021/1490732Google ScholarThere is no corresponding record for this reference.
- 104Molina-Santiago, C.; Bernal, P. Nanotube-mediated plasmid transfer as a natural alternative for the improvement of industrially relevant bacteria. Microb. Biotechnol. 2023, 16 (4), 706– 708, DOI: 10.1111/1751-7915.14225Google ScholarThere is no corresponding record for this reference.
- 105Dubey, G. P.; Malli Mohan, G. B.; Dubrovsky, A. Architecture and Characteristics of Bacterial Nanotubes. Dev. Cell 2016, 36 (4), 453– 461, DOI: 10.1016/j.devcel.2016.01.013Google ScholarThere is no corresponding record for this reference.
- 106Dubey, G. P.; Ben-Yehuda, S. Intercellular Nanotubes Mediate Bacterial Communication. Cell. 2011, 144 (4), 590– 600, DOI: 10.1016/j.cell.2011.01.015Google ScholarThere is no corresponding record for this reference.
- 107Baidya, A. K.; Rosenshine, I.; Ben-Yehuda, S. Donor-delivered cell wall hydrolases facilitate nanotube penetration into recipient bacteria. Nat. Commun. 2020, 11 (1), 1938, DOI: 10.1038/s41467-020-15605-1Google ScholarThere is no corresponding record for this reference.
- 108Pospíšil, J.; Vítovská, D.; Kofroňová, O. Bacterial nanotubes as a manifestation of cell death. Nat. Commun. 2020, 11 (1), 4963, DOI: 10.1038/s41467-020-18800-2Google ScholarThere is no corresponding record for this reference.
- 109Wang, C.; Zhao, R.; Yang, W. Cell-to-Cell Natural Transformation Mediated Efficient Plasmid Transfer Between Bacillus Species. IJMS. 2025, 26 (2), 621, DOI: 10.3390/ijms26020621Google ScholarThere is no corresponding record for this reference.
- 110McInerney, J. O.; McNally, A.; O’Connell, M. J. Why prokaryotes have pangenomes. Nat. Microbiol. 2017, 2 (4), 17040, DOI: 10.1038/nmicrobiol.2017.40Google ScholarThere is no corresponding record for this reference.
- 111Lee, I. P. A.; Eldakar, O. T.; Gogarten, J. P.; Andam, C. P. Bacterial cooperation through horizontal gene transfer. Trends Ecol. Evol. 2022, 37 (3), 223– 232, DOI: 10.1016/j.tree.2021.11.006Google ScholarThere is no corresponding record for this reference.
- 112Husnik, F.; McCutcheon, J. P. Functional horizontal gene transfer from bacteria to eukaryotes. Nat. Rev. Microbiol. 2018, 16 (2), 67– 79, DOI: 10.1038/nrmicro.2017.137Google ScholarThere is no corresponding record for this reference.
- 113Niero, G.; Bortolaia, V.; Vanni, M.; Intorre, L.; Guardabassi, L.; Piccirillo, A. High diversity of genes and plasmids encoding resistance to third-generation cephalosporins and quinolones in clinical Escherichia coli from commercial poultry flocks in Italy. Vet. Microbiol. 2018, 216, 93– 98, DOI: 10.1016/j.vetmic.2018.02.012Google ScholarThere is no corresponding record for this reference.
- 114López, L.; Jumbo, M.; Mosquera, P.; Donoso, G.; Graham, J.; Trueba, G. Oral and parenteral treatment with a third-generation cephalosporin promotes the proliferation of diverse ESBL-producing Escherichia coli in the chicken intestinal tract. Rao K, ed. mSphere 2025, 10 (7), e00227–25 DOI: 10.1128/msphere.00227-25Google ScholarThere is no corresponding record for this reference.
- 115Akhtar, A.; Fatima, N.; Khan, H. M. Beta-Lactamases and Their Classification: An Overview. Beta-Lactam Resistance in Gram-Negative Bacteria; Shahid, M., Singh, A., Sami, H., Eds.; Springer Nature Singapore, 2022; pp 25– 33. DOI: 10.1007/978-981-16-9097-6_3 .Google ScholarThere is no corresponding record for this reference.
- 116Philippon, A.; Jacquier, H.; Ruppé, E.; Labia, R. Structure-based classification of class A beta-lactamases, an update. Curr. Res. Transl. Med. 2019, 67 (4), 115– 122, DOI: 10.1016/j.retram.2019.05.003Google ScholarThere is no corresponding record for this reference.
- 117Page, M. I.; Badarau, A. The Mechanisms of Catalysis by Metallo β -Lactamases. Mugesh G, ed. Bioinorg. Chem. Appl. 2008, 2008 (1), 576297, DOI: 10.1155/2008/576297Google ScholarThere is no corresponding record for this reference.
- 118Fernandes, R.; Amador, P.; Prudêncio, C. β-Lactams: chemical structure, mode of action and mechanisms of resistance. Rev. Med. Microbiol. 2013, 24 (1), 7– 17, DOI: 10.1097/MRM.0b013e3283587727Google ScholarThere is no corresponding record for this reference.
- 119Carcione, D.; Siracusa, C.; Sulejmani, A.; Leoni, V.; Intra, J. Old and New Beta-Lactamase Inhibitors: Molecular Structure, Mechanism of Action, and Clinical Use. Antibiotics 2021, 10 (8), 995, DOI: 10.3390/antibiotics10080995Google ScholarThere is no corresponding record for this reference.
- 120Chaves, J.; Ladona, M. G.; Segura, C.; Coira, A.; Reig, R.; Ampurdanés, C. SHV-1 β-Lactamase Is Mainly a Chromosomally Encoded Species-Specific Enzyme in Klebsiella pneumoniae. Antimicrob. Agents Chemother. 2001, 45 (10), 2856– 2861, DOI: 10.1128/AAC.45.10.2856-2861.2001Google ScholarThere is no corresponding record for this reference.
- 121Acman, M.; Wang, R.; Van Dorp, L. Role of mobile genetic elements in the global dissemination of the carbapenem resistance gene blaNDM. Nat. Commun. 2022, 13 (1), 1131, DOI: 10.1038/s41467-022-28819-2Google ScholarThere is no corresponding record for this reference.
- 122Roca, I.; Mosqueda, N.; Altun, B.; Espinal, P.; Akova, M.; Vila, J. Molecular characterization of NDM-1-producing Acinetobacter pittii isolated from Turkey in 2006. J. Antimicrob. Chemother. 2014, 69 (12), 3437– 3438, DOI: 10.1093/jac/dku306Google ScholarThere is no corresponding record for this reference.
- 123Zhou, M.; Cai, Q.; Zhang, C.; Ouyang, P.; Yu, L.; Xu, Y. Antibiotic resistance bacteria and antibiotic resistance genes survived from the extremely acidity posing a risk on intestinal bacteria in an in vitro digestion model by horizontal gene transfer. Ecotoxicol. Environ. Saf. 2022, 247, 114247, DOI: 10.1016/j.ecoenv.2022.114247Google ScholarThere is no corresponding record for this reference.
- 124Haverkate, M. R.; Dautzenberg, M. J. D.; Ossewaarde, T. J. M. Within-Host and Population Transmission of blaOXA-48 in K. pneumoniae and E. coli. Friedrich A, ed. PLoS One 2015, 10 (10), e0140960 DOI: 10.1371/journal.pone.0140960Google ScholarThere is no corresponding record for this reference.
- 125Tofteland, S.; Naseer, U.; Lislevand, J. H.; Sundsfjord, A.; Samuelsen, Ø. A Long-Term Low-Frequency Hospital Outbreak of KPC-Producing Klebsiella pneumoniae Involving Intergenus Plasmid Diffusion and a Persisting Environmental Reservoir. Kluytmans J, ed. PLoS One 2013, 8 (3), e59015 DOI: 10.1371/journal.pone.0059015Google ScholarThere is no corresponding record for this reference.
- 126Liu, Y.; Gao, J.; Zhao, M.; Fu, X.; Zhang, Y.; Zhang, H. Removal of antibiotic resistant bacteria, genes and inhibition of plasmid-mediated horizontal transfer by peroxymonosulfate: Efficiency and mechanisms. Chem. Eng. J. 2023, 453, 139728, DOI: 10.1016/j.cej.2022.139728Google ScholarThere is no corresponding record for this reference.
- 127Jones, L. S.; Toleman, M. A.; Weeks, J. L.; Howe, R. A.; Walsh, T. R.; Kumarasamy, K. K. Plasmid Carriage of blaNDM-1 in Clinical Acinetobacter baumannii Isolates from India. Antimicrob. Agents Chemother. 2014, 58 (7), 4211– 4213, DOI: 10.1128/AAC.02500-14Google ScholarThere is no corresponding record for this reference.
- 128Poirel, L.; Bonnin, R. A.; Nordmann, P. Analysis of the Resistome of a Multidrug-Resistant NDM-1-Producing Escherichia coli Strain by High-Throughput Genome Sequencing. Antimicrob. Agents Chemother. 2011, 55 (9), 4224– 4229, DOI: 10.1128/AAC.00165-11Google ScholarThere is no corresponding record for this reference.
- 129Dellus-Gur, E.; Elias, M.; Caselli, E. Negative Epistasis and Evolvability in TEM-1 β-Lactamase–The Thin Line between an Enzyme’s Conformational Freedom and Disorder. J. Mol. Biol. 2015, 427 (14), 2396– 2409, DOI: 10.1016/j.jmb.2015.05.011Google ScholarThere is no corresponding record for this reference.
- 130Prescott, J. F. Beta-lactam Antibiotics: Cephalosporins. Antimicrobial Therapy in Veterinary Medicine, 1 ed.; Giguère, S., Prescott, J. F., Dowling, P. M., Eds.; Wiley, 2013; pp 153– 173 DOI: 10.1002/9781118675014.ch9 .Google ScholarThere is no corresponding record for this reference.
- 131Galleni, M.; Lamotte-Brasseur, J.; Raquet, X. The enigmatic catalytic mechanism of active-site serine β-lactamases. Biochem. Pharmacol. 1995, 49 (9), 1171– 1178, DOI: 10.1016/0006-2952(94)00502-DGoogle ScholarThere is no corresponding record for this reference.
- 132Dubus, A.; Wilkin, J. M.; Raquet, X.; Normark, S.; Frère, J. M. Catalytic mechanism of active-site serine β -lactamases: role of the conserved hydroxy group of the Lys-Thr(Ser)-Gly triad. Biochem. J. 1994, 301 (2), 485– 494, DOI: 10.1042/bj3010485Google ScholarThere is no corresponding record for this reference.
- 133Lamotte-Brasseur, J.; Knox, J.; Kelly, J. A. The Structures and Catalytic Mechanisms of Active-Site Serine β-Lactamases. Biotechnol. Genet. Eng. Rev. 1994, 12 (1), 189– 230, DOI: 10.1080/02648725.1994.10647912Google ScholarThere is no corresponding record for this reference.
- 134Tooke, C. L.; Hinchliffe, P.; Bragginton, E. C. β-Lactamases and β-Lactamase Inhibitors in the 21st Century. J. Mol. Biol. 2019, 431 (18), 3472– 3500, DOI: 10.1016/j.jmb.2019.04.002Google ScholarThere is no corresponding record for this reference.
- 135Gniadkowski, M. Evolution of extended-spectrum β-lactamases by mutation. Clin. Microbiol. Infect. 2008, 14, 11– 32, DOI: 10.1111/j.1469-0691.2007.01854.xGoogle ScholarThere is no corresponding record for this reference.
- 136Castañeda-Barba, S.; Top, E. M.; Stalder, T. Plasmids, a molecular cornerstone of antimicrobial resistance in the One Health era. Nat. Rev. Microbiol. 2024, 22 (1), 18– 32, DOI: 10.1038/s41579-023-00926-xGoogle ScholarThere is no corresponding record for this reference.
- 137Razavi, M.; Kristiansson, E.; Flach, C. F.; Larsson, D. G. J. The Association between Insertion Sequences and Antibiotic Resistance Genes. mSphere 2020, 5 (5), e00418–20 DOI: 10.1128/mSphere.00418-20Google ScholarThere is no corresponding record for this reference.
- 138Lipszyc, A.; Szuplewska, M.; Bartosik, D. How Do Transposable Elements Activate Expression of Transcriptionally Silent Antibiotic Resistance Genes?. IJMS. 2022, 23 (15), 8063, DOI: 10.3390/ijms23158063Google ScholarThere is no corresponding record for this reference.
- 139Bhat, B. A.; Mir, R. A.; Qadri, H. Integrons in the development of antimicrobial resistance: critical review and perspectives. Front. Microbiol. 2023, 14, 1231938, DOI: 10.3389/fmicb.2023.1231938Google ScholarThere is no corresponding record for this reference.
- 140Kotloff, K. L.; Riddle, M. S.; Platts-Mills, J. A.; Pavlinac, P.; Zaidi, A. K. M. Shigellosis. Lancet 2018, 391 (10122), 801– 812, DOI: 10.1016/S0140-6736(17)33296-8Google ScholarThere is no corresponding record for this reference.
- 141Khalil, I.; Troeger, C. E.; Blacker, B. F.; Reiner, R. C. Capturing the true burden of Shigella and ETEC: The way forward. Vaccine 2019, 37 (34), 4784– 4786, DOI: 10.1016/j.vaccine.2019.01.031Google ScholarThere is no corresponding record for this reference.
- 142Mason, L. C. E.; Greig, D. R.; Cowley, L. A. The evolution and international spread of extensively drug resistant Shigella sonnei. Nat. Commun. 2023, 14 (1), 1983, DOI: 10.1038/s41467-023-37672-wGoogle ScholarThere is no corresponding record for this reference.
- 143Davies, J. R.; Farrant, W. N.; Tomlinson, A. J. H. Further studies on the antibiotic resistance of Shigella sonnei: II. The acquisition of transferable antibiotic resistance in vivo. J. Hyg. 1968, 66 (3), 479– 487, DOI: 10.1017/S0022172400041346Google ScholarThere is no corresponding record for this reference.
- 144Thanh Duy, P.; Thi Nguyen, T. N.; Vu Thuy, D. Commensal Escherichia coli are a reservoir for the transfer of XDR plasmids into epidemic fluoroquinolone-resistant Shigella sonnei. Nat. Microbiol. 2020, 5 (2), 256– 264, DOI: 10.1038/s41564-019-0645-9Google ScholarThere is no corresponding record for this reference.
- 145Baker, K. S.; Dallman, T. J.; Field, N. Horizontal antimicrobial resistance transfer drives epidemics of multiple Shigella species. Nat. Commun. 2018, 9 (1), 1462, DOI: 10.1038/s41467-018-03949-8Google ScholarThere is no corresponding record for this reference.
- 146Goodman, R. N.; Tansirichaiya, S.; Brouwer, M. S. M.; Roberts, A. P. Intracellular Transposition of Mobile Genetic Elements Associated with the Colistin Resistance Gene mcr-1. Microbiol Spectr. 2023, 11 (1), e03278–22 DOI: 10.1128/spectrum.03278-22Google ScholarThere is no corresponding record for this reference.
- 147De La Cadena, E.; Mahecha, M.; Velandia, A. M. Identification of mcr-1 Genes and Characterization of Resistance Mechanisms to Colistin in Escherichia coli Isolates from Colombian Hospitals. Antibiotics 2023, 12 (3), 488, DOI: 10.3390/antibiotics12030488Google ScholarThere is no corresponding record for this reference.
- 148Liu, Y. Y.; Wang, Y.; Walsh, T. R. Emergence of plasmid-mediated colistin resistance mechanism MCR-1 in animals and human beings in China: a microbiological and molecular biological study. Lancet Infect. Dis. 2016, 16 (2), 161– 168, DOI: 10.1016/S1473-3099(15)00424-7Google ScholarThere is no corresponding record for this reference.
- 149Falgenhauer, L.; Waezsada, S. E.; Yao, Y. Colistin resistance gene mcr-1 in extended-spectrum β-lactamase-producing and carbapenemase-producing Gram-negative bacteria in Germany. Lancet Infect. Dis. 2016, 16 (3), 282– 283, DOI: 10.1016/S1473-3099(16)00009-8Google ScholarThere is no corresponding record for this reference.
- 150Haenni, M.; Poirel, L.; Kieffer, N. Co-occurrence of extended spectrum β lactamase and MCR-1 encoding genes on plasmids. Lancet Infect. Dis. 2016, 16 (3), 281– 282, DOI: 10.1016/S1473-3099(16)00007-4Google ScholarThere is no corresponding record for this reference.
- 151Zurfuh, K.; Poirel, L.; Nordmann, P.; Nüesch-Inderbinen, M.; Hächler, H.; Stephan, R. Occurrence of the Plasmid-Borne mcr-1 Colistin Resistance Gene in Extended-Spectrum-β-Lactamase-Producing Enterobacteriaceae in River Water and Imported Vegetable Samples in Switzerland. Antimicrob. Agents Chemother. 2016, 60 (4), 2594– 2595, DOI: 10.1128/AAC.00066-16Google ScholarThere is no corresponding record for this reference.
- 152Du, H.; Chen, L.; Tang, Y. W.; Kreiswirth, B. N. Emergence of the mcr-1 colistin resistance gene in carbapenem-resistant Enterobacteriaceae. Lancet Infect. Dis. 2016, 16 (3), 287– 288, DOI: 10.1016/S1473-3099(16)00056-6Google ScholarThere is no corresponding record for this reference.
- 153Nang, S. C.; Li, J.; Velkov, T. The rise and spread of mcr plasmid-mediated polymyxin resistance. Crit. Rev. Microbiol. 2019, 45 (2), 131– 161, DOI: 10.1080/1040841X.2018.1492902Google ScholarThere is no corresponding record for this reference.
- 154Blair, J. M. A.; Webber, M. A.; Baylay, A. J.; Ogbolu, D. O.; Piddock, L. J. V. Molecular mechanisms of antibiotic resistance. Nat. Rev. Microbiol. 2015, 13 (1), 42– 51, DOI: 10.1038/nrmicro3380Google ScholarThere is no corresponding record for this reference.
- 155Cox, G.; Wright, G. D. Intrinsic antibiotic resistance: Mechanisms, origins, challenges and solutions. Int. J. Med. Microbiol. 2013, 303 (6–7), 287– 292, DOI: 10.1016/j.ijmm.2013.02.009Google ScholarThere is no corresponding record for this reference.
- 156Palzkill, T. Structural and Mechanistic Basis for Extended-Spectrum Drug-Resistance Mutations in Altering the Specificity of TEM, CTX-M, and KPC β-lactamases. Front. Mol. Biosci. 2018, 5, 16, DOI: 10.3389/fmolb.2018.00016Google ScholarThere is no corresponding record for this reference.
- 157Zhang, H.; Seward, C. H.; Wu, Z.; Ye, H.; Feng, Y. Genomic insights into the ESBL and MCR-1-producing ST648 Escherichia coli with multi-drug resistance. Sci. Bull. 2016, 61 (11), 875– 878, DOI: 10.1007/s11434-016-1086-yGoogle ScholarThere is no corresponding record for this reference.
- 158Kluytmans-van Den Bergh, M. F.; Huizinga, P.; Bonten, M. J.; Presence of mcr-1-positive Enterobacteriaceae in retail chicken meat but not in humans in the Netherlands since 2009. Eurosurveillance 2016, 21(9). DOI: 10.2807/1560-7917.ES.2016.21.9.30149 .Google ScholarThere is no corresponding record for this reference.
- 159Von Wintersdorff, C. J. H.; Penders, J.; Van Niekerk, J. M.; Mills, N. D.; Majumder, S.; van Alphen, L. B.; Savelkoul, P. H. M.; Wolffs, P. F. G. Dissemination of Antimicrobial Resistance in Microbial Ecosystems through Horizontal Gene Transfer. Front. Microbiol. 2016, 7. DOI: 10.3389/fmicb.2016.00173 .Google ScholarThere is no corresponding record for this reference.
- 160Zhang, G.; Wang, C.; Sui, Z.; Feng, J. Insights into the evolutionary trajectories of fluoroquinolone resistance in Streptococcus pneumoniae. J. Antimicrob. Chemother. 2015, 70 (9), 2499– 2506, DOI: 10.1093/jac/dkv134Google ScholarThere is no corresponding record for this reference.
- 161Cuypers, W. L.; Meysman, P.; Weill, F. X. A global genomic analysis of Salmonella Concord reveals lineages with high antimicrobial resistance in Ethiopia. Nat. Commun. 2023, 14 (1), 3517, DOI: 10.1038/s41467-023-38902-xGoogle ScholarThere is no corresponding record for this reference.
- 162Hassan, R.; Tantawy, M.; Gouda, N. A. Genotypic characterization of multiple drug resistant Escherichia coli isolates from a pediatric cancer hospital in Egypt. Sci. Rep. 2020, 10 (1), 4165, DOI: 10.1038/s41598-020-61159-zGoogle ScholarThere is no corresponding record for this reference.
- 163Thiolas, A.; Bornet, C.; Davin-Régli, A.; Pagès, J. M.; Bollet, C. Resistance to imipenem, cefepime, and cefpirome associated with mutation in Omp36 osmoporin of Enterobacter aerogenes. Biochem. Biophys. Res. Commun. 2004, 317 (3), 851– 856, DOI: 10.1016/j.bbrc.2004.03.130Google ScholarThere is no corresponding record for this reference.
- 164Dé, E.; Baslé, A.; Jaquinod, M. A new mechanism of antibiotic resistance in Enterobacteriaceae induced by a structural modification of the major porin. Mol. Microbiol. 2001, 41 (1), 189– 198, DOI: 10.1046/j.1365-2958.2001.02501.xGoogle ScholarThere is no corresponding record for this reference.
- 165Sharma, A.; Gupta, V. K.; Pathania, R. Efflux pump inhibitors for bacterial pathogens: From bench to bedside. Indian J. Med. Res. 2019, 149 (2), 129– 145, DOI: 10.4103/ijmr.IJMR_2079_17Google ScholarThere is no corresponding record for this reference.
- 166Wilson, D. N.; Hauryliuk, V.; Atkinson, G. C.; O’Neill, A. J. Target protection as a key antibiotic resistance mechanism. Nat. Rev. Microbiol. 2020, 18 (11), 637– 648, DOI: 10.1038/s41579-020-0386-zGoogle ScholarThere is no corresponding record for this reference.
- 167Stephan, J.; Mailaender, C.; Etienne, G.; Daffé, M.; Niederweis, M. Multidrug Resistance of a Porin Deletion Mutant of Mycobacterium smegmatis. Antimicrob. Agents Chemother. 2004, 48 (11), 4163– 4170, DOI: 10.1128/AAC.48.11.4163-4170.2004Google ScholarThere is no corresponding record for this reference.
- 168Stahl, C.; Kubetzko, S.; Kaps, I.; Seeber, S.; Engelhardt, H.; Niederweis, M. MspA provides the main hydrophilic pathway through the cell wall of Mycobacterium smegmatis. Mol. Microbiol. 2001, 40 (2), 451– 464, DOI: 10.1046/j.1365-2958.2001.02394.xGoogle ScholarThere is no corresponding record for this reference.
- 169Masi, M.; Réfregiers, M.; Pos, K. M.; Pagès, J. M. Mechanisms of envelope permeability and antibiotic influx and efflux in Gram-negative bacteria. Nat. Microbiol. 2017, 2 (3), 17001, DOI: 10.1038/nmicrobiol.2017.1Google ScholarThere is no corresponding record for this reference.
- 170Vergalli, J.; Bodrenko, I. V.; Masi, M. Porins and small-molecule translocation across the outer membrane of Gram-negative bacteria. Nat. Rev. Microbiol. 2020, 18 (3), 164– 176, DOI: 10.1038/s41579-019-0294-2Google ScholarThere is no corresponding record for this reference.
- 171Chen, S.; Fu, J.; Zhao, K. Class 1 integron carrying qacEΔ1 gene confers resistance to disinfectant and antibiotics in Salmonella. Int. J. Food Microbiol. 2023, 404, 110319, DOI: 10.1016/j.ijfoodmicro.2023.110319Google ScholarThere is no corresponding record for this reference.
- 172Ovung, A.; Bhattacharyya, J. Sulfonamide drugs: structure, antibacterial property, toxicity, and biophysical interactions. Biophys Rev. 2021, 13 (2), 259– 272, DOI: 10.1007/s12551-021-00795-9Google ScholarThere is no corresponding record for this reference.
- 173Roberts, M. C. Update on macrolide-lincosamide-streptogramin, ketolide, and oxazolidinone resistance genes: MLSKO genes. FEMS Microbiol. Lett. 2008, 282 (2), 147– 159, DOI: 10.1111/j.1574-6968.2008.01145.xGoogle ScholarThere is no corresponding record for this reference.
- 174Chen, L.; Huang, J.; Huang, X. Horizontal Transfer of Different erm(B)-Carrying Mobile Elements Among Streptococcus suis Strains With Different Serotypes. Front. Microbiol. 2021, 12, 628740, DOI: 10.3389/fmicb.2021.628740Google ScholarThere is no corresponding record for this reference.
- 175Sun, S. Emerging antibiotic resistance by various novel proteins/enzymes. Eur. J. Clin. Microbiol. Infect. Dis. 2025, 44 (7), 1551– 1566, DOI: 10.1007/s10096-025-05126-4Google ScholarThere is no corresponding record for this reference.
- 176Unemo, M.; Lahra, M. M.; Cole, M. J. WHO global gonococcal antimicrobial surveillance programmes, 2019–22: a retrospective observational study. Lancet Microbe 2025, 6 (10), 101181, DOI: 10.1016/j.lanmic.2025.101181Google ScholarThere is no corresponding record for this reference.
- 177Unemo, M.; Lahra, M. M.; Cole, M. World Health Organization Global Gonococcal Antimicrobial Surveillance Program (WHO GASP): review of new data and evidence to inform international collaborative actions and research efforts. Sex. Health 2019, 16 (5), 412– 425, DOI: 10.1071/SH19023Google ScholarThere is no corresponding record for this reference.
- 178Mo, Y.; Tan, W. C.; Cooper, B. S. Antibiotic duration for common bacterial infections–a systematic review. JAC-Antimicrobial Resistance. 2024, 7 (1), dlae215, DOI: 10.1093/jacamr/dlae215Google ScholarThere is no corresponding record for this reference.
- 179Mollah, F.; Khatun, M. M.; Chowdhury, R. Therapeutic Promises of Bioactive Linarin, a Glycosylated Flavonoid: A Comprehensive Review With Mechanistic Insight. Buch L, ed. J. Trop. Med. 2025, 2025 (1), 9989759, DOI: 10.1155/jotm/9989759Google ScholarThere is no corresponding record for this reference.
- 180Vassilopoulos, S.; Mylonakis, E. Advances in methicillin-resistant staphylococcus aureus drug discovery: developments and challenges. Expert Opin. Drug Discovery 2026, 21, 231– 244, DOI: 10.1080/17460441.2026.2618787Google ScholarThere is no corresponding record for this reference.
- 181Ghazaei, C. The role of bacteriophages and CRISPR-Cas in combating multidrug-resistant bacteria. Nat. Prod. Bioprospect. 2026, 16 (1), 14, DOI: 10.1007/s13659-025-00567-yGoogle ScholarThere is no corresponding record for this reference.
- 182Khan, M. F.; Javed, M.; Kaur, J.; Badwal, A. K.; Singh, S. CRISPR-Cas mediated targeting of resistance genes for combating ESKAPE pathogen infections: A Review. Int. J. Biol. Macromol. 2025, 334, 149180, DOI: 10.1016/j.ijbiomac.2025.149180Google ScholarThere is no corresponding record for this reference.
- 183Antelo-Riveiro, P.; Garcia-Fandino, R.; Piñeiro, A. ́. Antimicrobial peptides at (lipid) interfaces: Insights from monolayer models. Adv. Colloid Interface Sci. 2026, 350, 103775, DOI: 10.1016/j.cis.2025.103775Google ScholarThere is no corresponding record for this reference.
- 184Galhano, J.; Capelo-Martinez, J. L.; Lorenzo, J.; Lodeiro, C.; Oliveira, E. Advances in Antimicrobial Applications of Ag, Cu, and AgCu Nanoparticle-Doped Polymeric Composite Materials: A Comprehensive Review. ACS Nano 2025, 19 (35), 31301– 31330, DOI: 10.1021/acsnano.5c08822Google ScholarThere is no corresponding record for this reference.
- 185Ravikumar, S.; Velappan, K.; Munusamy, S.; M, V.; S, N.; Shanmugam, R. Emerging nanotechnologies in wound care: The role of metal and polymeric nanocomposites in enhancing healing and combating infections. Int. J. Pharm. 2025, 684, 126143, DOI: 10.1016/j.ijpharm.2025.126143Google ScholarThere is no corresponding record for this reference.
- 186Wijethunge, D.; Mathew, A.; Yarlagadda, P. K. D. V. Comprehensive review of bacterial death mechanism on nanopillared nanostructured surfaces. Biophys Rev. 2025, 17 (3), 893– 908, DOI: 10.1007/s12551-025-01319-5Google ScholarThere is no corresponding record for this reference.
- 187Linklater, D. P.; Baulin, V. A.; Juodkazis, S.; Crawford, R. J.; Stoodley, P.; Ivanova, E. P. Mechano-bactericidal actions of nanostructured surfaces. Nat. Rev. Microbiol. 2021, 19 (1), 8– 22, DOI: 10.1038/s41579-020-0414-zGoogle ScholarThere is no corresponding record for this reference.
- 188Cao, H.; De La Fuente-Nunez, C. Microbial Primer: Artificial intelligence for microbiologists: This article is part of the < span style = ″color: rgb(33, 51, 104);″>Microbial Primers</span> collection.Š. Microbiology. 2025, 171(11). DOI: 10.1099/mic.0.001629 .Google ScholarThere is no corresponding record for this reference.
- 189Adams, J. M. E.; El-Halfawy, O. M. Chemical-mediated alteration of antibiotic susceptibility: mechanisms and potential new targets for antibiotic adjuvant discovery. J. Bacteriol. 2026, e00351–25 DOI: 10.1128/jb.00351-25Google ScholarThere is no corresponding record for this reference.
- 190Michaelis, C.; Grohmann, E. Horizontal Gene Transfer of Antibiotic Resistance Genes in Biofilms. Antibiotics 2023, 12 (2), 328, DOI: 10.3390/antibiotics12020328Google ScholarThere is no corresponding record for this reference.
- 191Abe, K.; Nomura, N.; Suzuki, S. Biofilms: hot spots of horizontal gene transfer (HGT) in aquatic environments, with a focus on a new HGT mechanism. FEMS Microbiol. Ecol. 2020, 96 (5), fiaa031, DOI: 10.1093/femsec/fiaa031Google ScholarThere is no corresponding record for this reference.
- 192Djermoun, S.; Rode, D. K. H.; Jiménez-Siebert, E. Biofilm architecture determines the dissemination of conjugative plasmids. Proc. Natl. Acad. Sci. U. S. A. 2025, 122 (17), e2417452122 DOI: 10.1073/pnas.2417452122Google ScholarThere is no corresponding record for this reference.
- 193Tian, S.; Van Der Mei, H. C.; Ren, Y.; Busscher, H. J.; Shi, L. Recent advances and future challenges in the use of nanoparticles for the dispersal of infectious biofilms. J. Mater. Sci. Technol. 2021, 84, 208– 218, DOI: 10.1016/j.jmst.2021.02.007Google ScholarThere is no corresponding record for this reference.
- 194Ma, L.; Konkel, M. E.; Lu, X. Antimicrobial Resistance Gene Transfer from Campylobacter jejuni in Mono- and Dual-Species Biofilms. Elkins CA, ed. Appl. Environ. Microbiol. 2021, 87 (15), e00659–21 DOI: 10.1128/AEM.00659-21Google ScholarThere is no corresponding record for this reference.
- 195Lerminiaux, N. A.; Cameron, A. D. S. Horizontal transfer of antibiotic resistance genes in clinical environments. Can. J. Microbiol. 2019, 65 (1), 34– 44, DOI: 10.1139/cjm-2018-0275Google ScholarThere is no corresponding record for this reference.
- 196Hutinel, M.; Fick, J.; Larsson, D. G. J.; Flach, C. F. Investigating the effects of municipal and hospital wastewaters on horizontal gene transfer. Environ. Pollut. 2021, 276, 116733, DOI: 10.1016/j.envpol.2021.116733Google ScholarThere is no corresponding record for this reference.
- 197Sridhar, S.; Worby, C. J.; Bronson, R. A.; Turbett, S. E.; Oliver, E.; Shea, T.; Rao, S. R.; Sanchez, V.; Becker, M. V.; Holliday, L. K. Insights Into Global Antimicrobial Resistance Dynamics Through the Sequencing of Enteric Bacteria From US International Travelers. J. Infect. Dis. 2026, 233 (1), e164– e173, DOI: 10.1093/infdis/jiaf469Google ScholarThere is no corresponding record for this reference.
- 198Sassi, A.; Basher, N. S.; Kirat, H. The Role of the Environment (Water, Air, Soil) in the Emergence and Dissemination of Antimicrobial Resistance: A One Health Perspective. Antibiotics 2025, 14 (8), 764, DOI: 10.3390/antibiotics14080764Google ScholarThere is no corresponding record for this reference.
- 199Karim, M. A.; KianvashRad, N.; Cabo, M., Jr Cell Adhesion and Biofilm Development via Force-Sensitive Mechanisms: A Perspective. ACS Biomater. Sci. Eng. 2026, 12 (1), 3– 12, DOI: 10.1021/acsbiomaterials.5c01755Google ScholarThere is no corresponding record for this reference.
- 200Liu, N.; Du, J.; Ge, J.; Liu, S. B. DNA damage-inducing endogenous and exogenous factors and research progress. Nucleosides Nucleotides Nucleic Acids 2025, 44 (12), 969– 1001, DOI: 10.1080/15257770.2024.2428436Google ScholarThere is no corresponding record for this reference.
- 201Adamou, P.; Entwistle, J.; Graham, D. W.; Neumann, A. Mineral-Based Advanced Oxidation Processes for Enhancing the Removal of Antibiotic Resistance Genes from Domestic Wastewater. ACS EST Water. 2025, 5 (5), 2310– 2321, DOI: 10.1021/acsestwater.4c01213Google ScholarThere is no corresponding record for this reference.
- 202Feng, M.; Liu, Y.; Yang, L.; Li, Z. Antibiotics and antibiotic resistance gene dynamics in the composting of antibiotic fermentation waste - A review. Bioresour. Technol. 2023, 390, 129861, DOI: 10.1016/j.biortech.2023.129861Google ScholarThere is no corresponding record for this reference.
- 203Liao, H.; Bai, Y.; Liu, C. Airborne and indigenous microbiomes co-drive the rebound of antibiotic resistome during compost storage. Environ. Microbiol. 2021, 23 (12), 7483– 7496, DOI: 10.1111/1462-2920.15672Google ScholarThere is no corresponding record for this reference.
- 204Xia, R.; Zhang, L.; Li, G.; Luo, W.; Xu, Z. A small technology for big health: Blocking the potential spread of antibiotic resistomes from home composting of food waste by mature compost. Waste Manage. 2026, 211, 115312, DOI: 10.1016/j.wasman.2025.115312Google ScholarThere is no corresponding record for this reference.
- 205Yin, D.; Wang, K.; Wu, C. Refluxing mature compost to replace bulking agents: A low-cost solution for suppressing antibiotic resistance genes rebound in sewage sludge composting. Environ. Res. 2025, 269, 120811, DOI: 10.1016/j.envres.2025.120811Google ScholarThere is no corresponding record for this reference.
- 206Yoon, Y.; Chung, H. J.; Wen Di, D. Y.; Dodd, M. C.; Hur, H. G.; Lee, Y. Inactivation efficiency of plasmid-encoded antibiotic resistance genes during water treatment with chlorine, UV, and UV/H2O2. Water Res. 2017, 123, 783– 793, DOI: 10.1016/j.watres.2017.06.056Google ScholarThere is no corresponding record for this reference.
- 207He, H.; Choi, Y.; Wu, S. J. Application of Nucleotide-Based Kinetic Modeling Approaches to Predict Antibiotic Resistance Gene Degradation during UV- and Chlorine-Based Wastewater Disinfection Processes: From Bench- to Full-Scale. Environ. Sci. Technol. 2022, 56 (21), 15141– 15155, DOI: 10.1021/acs.est.2c00567Google ScholarThere is no corresponding record for this reference.
- 208Yao, M. C.; Zhang, X.; Huang, Q.; Huang, J.; Sheng, G. P. Chlorine oxide radical (ClO) enables the enhanced degradation of antibiotic resistance genes during UV/chlorine treatment by selectively inducing base damage. Environ. Int. 2023, 178, 108121, DOI: 10.1016/j.envint.2023.108121Google ScholarThere is no corresponding record for this reference.
- 209Guerra-Rodríguez, S.; Abeledo-Lameiro, M. J.; Polo-López, M. I. Pilot-scale sulfate radical-based advanced oxidation for wastewater reuse: simultaneous disinfection, removal of contaminants of emerging concern, and antibiotic resistance genes. Chem. Eng. J. 2023, 477, 146916, DOI: 10.1016/j.cej.2023.146916Google ScholarThere is no corresponding record for this reference.
- 210Liang, C.; Wei, D.; Zhang, S.; Ren, Q.; Shi, J.; Liu, L. Removal of antibiotic resistance genes from swine wastewater by membrane filtration treatment. Ecotoxicol. Environ. Saf. 2021, 210, 111885, DOI: 10.1016/j.ecoenv.2020.111885Google ScholarThere is no corresponding record for this reference.
- 211Li, R.; Wei, Z.; Li, P. Novel visible-light activated photocatalytic ultrafiltration membrane for simultaneous separation and degradation of emerging contaminants. J. Hazard. Mater. 2024, 478, 135634, DOI: 10.1016/j.jhazmat.2024.135634Google ScholarThere is no corresponding record for this reference.
- 212Sun, L.; Shi, P.; He, N.; Zhang, Q.; Duan, X. Antibiotic resistance genes removal and membrane fouling in secondary effluents by combined processes of PAC/BPAC-UF. J. Water Health 2019, 17 (6), 910– 920, DOI: 10.2166/wh.2019.160Google ScholarThere is no corresponding record for this reference.
- 213Berruti, I.; Nahim-Granados, S.; Abeledo-Lameiro, M. J.; Oller, I.; Polo-López, M. I. Peroxymonosulfate/Solar process for urban wastewater purification at a pilot plant scale: A techno-economic assessment. Sci. Total Environ. 2023, 881, 163407, DOI: 10.1016/j.scitotenv.2023.163407Google ScholarThere is no corresponding record for this reference.
- 214Rutten, S. B.; Ojobe, B. L.; Hernández Leal, L. Evaluation of Membrane Integrity Monitoring Methods for Hollow Fiber Nanofiltration Membranes: Applicability in Gray Water Reclamation Systems. ACS EST Water. 2023, 3 (12), 3884– 3892, DOI: 10.1021/acsestwater.3c00307Google ScholarThere is no corresponding record for this reference.
- 215Slipko, K.; Reif, D.; Wögerbauer, M.; Hufnagl, P.; Krampe, J.; Kreuzinger, N. Removal of extracellular free DNA and antibiotic resistance genes from water and wastewater by membranes ranging from microfiltration to reverse osmosis. Water Res. 2019, 164, 114916, DOI: 10.1016/j.watres.2019.114916Google ScholarThere is no corresponding record for this reference.
- 216Li, M.; Wang, P.; Zhang, K. Single cobalt atoms anchored on Ti3 C2 Tx with dual reaction sites for efficient adsorption-degradation of antibiotic resistance genes. Proc. Natl. Acad. Sci. U. S. A. 2023, 120 (29), e2305705120 DOI: 10.1073/pnas.2305705120Google ScholarThere is no corresponding record for this reference.
- 217Liu, Y.; Gao, J.; Wang, Y. The removal of antibiotic resistant bacteria and genes and inhibition of the horizontal gene transfer by contrastive research on sulfidated nanoscale zerovalent iron activating peroxymonosulfate or peroxydisulfate. J. Hazard. Mater. 2022, 423, 126866, DOI: 10.1016/j.jhazmat.2021.126866Google ScholarThere is no corresponding record for this reference.
- 218Liu, Y.; Cai, Y.; Li, G.; Wang, W.; Wong, P. K.; An, T. Response mechanisms of different antibiotic-resistant bacteria with different resistance action targets to the stress from photocatalytic oxidation. Water Res. 2022, 218, 118407, DOI: 10.1016/j.watres.2022.118407Google ScholarThere is no corresponding record for this reference.
- 219Poyatos, J. M.; Muñio, M. M.; Almecija, M. C.; Torres, J. C.; Hontoria, E.; Osorio, F. Advanced Oxidation Processes for Wastewater Treatment: State of the Art. Water, Air, Soil Pollut. 2010, 205 (1–4), 187– 204, DOI: 10.1007/s11270-009-0065-1Google ScholarThere is no corresponding record for this reference.
- 220Li, F.; Liu, K.; Bao, Y. Molecular level removal of antibiotic resistant bacteria and genes: A review of interfacial chemical in advanced oxidation processes. Water Res. 2024, 254, 121373, DOI: 10.1016/j.watres.2024.121373Google ScholarThere is no corresponding record for this reference.
- 221Manoharan, R. K.; Ishaque, F.; Ahn, Y. H. Fate of antibiotic resistant genes in wastewater environments and treatment strategies - A review. Chemosphere 2022, 298, 134671, DOI: 10.1016/j.chemosphere.2022.134671Google ScholarThere is no corresponding record for this reference.
- 222Stefaniak, K.; Harnisz, M.; Męcik, M.; Korzeniewska, E. ARB inactivation, ARGs and antibiotics degradation in hospital wastewater. J. Hazard. Mater. 2025, 495, 138833, DOI: 10.1016/j.jhazmat.2025.138833Google ScholarThere is no corresponding record for this reference.
- 223Emamalipour, M.; Seidi, K.; Zununi Vahed, S. Horizontal Gene Transfer: From Evolutionary Flexibility to Disease Progression. Front Cell Dev Biol. 2020, 8, 229, DOI: 10.3389/fcell.2020.00229Google ScholarThere is no corresponding record for this reference.
- 224Brito, I. L. Examining horizontal gene transfer in microbial communities. Nat. Rev. Microbiol. 2021, 19 (7), 442– 453, DOI: 10.1038/s41579-021-00534-7Google ScholarThere is no corresponding record for this reference.
- 225Abdulhussien, A. A.; Farhan, Z. M.; Yaseen, L. J.; Mohsein, O. A. The Clinico-Pathological Role of Horizontal Gene Transfer in the Ability of Pseudomonas Aeruginosa to Cause Disease in Lung Tissue. ejmhr. 2025, 3 (1), 27– 34, DOI: 10.59324/ejmhr.2025.3(1).06Google ScholarThere is no corresponding record for this reference.
- 226Evans, D. R.; Griffith, M. P.; Sundermann, A. J. Systematic detection of horizontal gene transfer across genera among multidrug-resistant bacteria in a single hospital. eLife 2020, 9, e53886 DOI: 10.7554/eLife.53886Google ScholarThere is no corresponding record for this reference.
- 227McCallum, G. E.; Hall, J. P. J. The hospital sink drain microbiome as a melting pot for AMR transmission to nosocomial pathogens. npj Antimicrob Resist. 2025, 3 (1), 68, DOI: 10.1038/s44259-025-00137-9Google ScholarThere is no corresponding record for this reference.
- 228McInnes, R. S.; McCallum, G. E.; Lamberte, L. E.; Van Schaik, W. Horizontal transfer of antibiotic resistance genes in the human gut microbiome. Curr. Opin. Microbiol. 2020, 53, 35– 43, DOI: 10.1016/j.mib.2020.02.002Google ScholarThere is no corresponding record for this reference.
- 229Woods, L. C.; Gorrell, R. J.; Taylor, F.; Connallon, T.; Kwok, T.; McDonald, M. J. Horizontal gene transfer potentiates adaptation by reducing selective constraints on the spread of genetic variation. Proc. Natl. Acad. Sci. U. S. A. 2020, 117 (43), 26868– 26875, DOI: 10.1073/pnas.2005331117Google ScholarThere is no corresponding record for this reference.
- 230Wang, J.; Huo, L.; Bian, K. Efficacy and Mechanism of Antibiotic Resistance Gene Degradation and Cell Membrane Damage during Ultraviolet Advanced Oxidation Processes. ACS EST Water. 2024, 4 (6), 2746– 2755, DOI: 10.1021/acsestwater.4c00350Google ScholarThere is no corresponding record for this reference.
- 231Liu, Y.; Hull, N. M. Emerging investigator series: Inactivation of antibiotic resistant bacteria and inhibition of horizontal resistance gene transfer is more effective by 222 than 254 nm UV. Environ. Sci: Water Res. Technol. 2025, 11 (2), 306– 316, DOI: 10.1039/D4EW00530AGoogle ScholarThere is no corresponding record for this reference.
- 232Han, J.; Li, W.; Yang, Y. UV-Based Advanced Oxidation Processes for Antibiotic Resistance Control: Efficiency, Influencing Factors, and Energy Consumption. Engineering. 2024, 37, 27– 39, DOI: 10.1016/j.eng.2023.09.021Google ScholarThere is no corresponding record for this reference.
- 233Zheng, T. H.; Zhang, Z. Z.; Liu, Y.; Zou, L. H. Recent Progress in Catalytically Driven Advanced Oxidation Processes for Wastewater Treatment. Catalysts 2025, 15 (8), 761, DOI: 10.3390/catal15080761Google ScholarThere is no corresponding record for this reference.
- 234Satyam, S.; Patra, S. The Evolving Landscape of Advanced Oxidation Processes in Wastewater Treatment: Challenges and Recent Innovations. Processes 2025, 13 (4), 987, DOI: 10.3390/pr13040987Google ScholarThere is no corresponding record for this reference.
- 235Ahmed, B.; Ameen, F.; Rizvi, A. Destruction of Cell Topography, Morphology, Membrane, Inhibition of Respiration, Biofilm Formation, and Bioactive Molecule Production by Nanoparticles of Ag, ZnO, CuO, TiO2, and Al2 O3 toward Beneficial Soil Bacteria. ACS Omega 2020, 5 (14), 7861– 7876, DOI: 10.1021/acsomega.9b04084Google ScholarThere is no corresponding record for this reference.
- 236Moreira, N. F. F.; Narciso-da-Rocha, C.; Polo-López, M. I. Solar treatment (H2O2, TiO2-P25 and GO-TiO2 photocatalysis, photo-Fenton) of organic micropollutants, human pathogen indicators, antibiotic resistant bacteria and related genes in urban wastewater. Water Res. 2018, 135, 195– 206, DOI: 10.1016/j.watres.2018.01.064Google ScholarThere is no corresponding record for this reference.
- 237Fu, X.; Gao, J.; Wang, Q. Mechanisms on the removal of gram-negative/positive antibiotic resistant bacteria and inhibition of horizontal gene transfer by ferrate coupled with peroxydisulfate or peroxymonosulfate. J. Hazard. Mater. 2024, 470, 134254, DOI: 10.1016/j.jhazmat.2024.134254Google ScholarThere is no corresponding record for this reference.
- 238Mahmoodi, M.; Pishbin, E. Ozone-based advanced oxidation processes in water treatment: recent advances, challenges, and perspective. Environ. Sci. Pollut. Res. 2025, 32 (7), 3531– 3570, DOI: 10.1007/s11356-024-35835-wGoogle ScholarThere is no corresponding record for this reference.
- 239Macedo, G.; Olesen, A. K.; Maccario, L. Horizontal Gene Transfer of an IncP1 Plasmid to Soil Bacterial Community Introduced by Escherichia coli through Manure Amendment in Soil Microcosms. Environ. Sci. Technol. 2022, 56 (16), 11398– 11408, DOI: 10.1021/acs.est.2c02686Google ScholarThere is no corresponding record for this reference.
- 240Mei, Z.; He, C.; Balcazar, J. L. Antibiotic-degrading bacteria shape resistome dynamics and horizontal gene transfer potential in soils with contrasting properties. ISME Communications. 2026, 6 (1), ycaf246, DOI: 10.1093/ismeco/ycaf246Google ScholarThere is no corresponding record for this reference.
- 241Shrestha, P.; Bajracharya, B.; Shrestha, D.; Karna, A. K.; Shah, P.; Gurung, K.; Ghimire, S.; Shrestha, A. Molecular detection of antibiotic resistance genes in Staphylococcus aureus isolated from poultry farms of Kathmandu Valley, Nepal. BMC Microbiol. 2026, 26, 291, DOI: 10.1186/s12866-026-04713-5Google ScholarThere is no corresponding record for this reference.
- 242Lima, T.; Domingues, S.; Da Silva, G. J. Manure as a Potential Hotspot for Antibiotic Resistance Dissemination by Horizontal Gene Transfer Events. Vet. Sci. 2020, 7 (3), 110, DOI: 10.3390/vetsci7030110Google ScholarThere is no corresponding record for this reference.
- 243Goh, Y. X.; Anupoju, S. M. B.; Nguyen, A. Evidence of horizontal gene transfer and environmental selection impacting antibiotic resistance evolution in soil-dwelling Listeria. Nat. Commun. 2024, 15 (1), 10034, DOI: 10.1038/s41467-024-54459-9Google ScholarThere is no corresponding record for this reference.
- 244Rocha, K. O.; Brandão, F.; Mendes, C. á.; Carvalho, M. G.; Mazierski, P.; Zaleska-Medynska, A.; Gomes, J.; Martins, R. C.; Domingues, E. Olive mill waste bio-based catalyst application in advanced oxidation processes for wastewater treatment. Catal. Today 2024, 432, 114618, DOI: 10.1016/j.cattod.2024.114618Google ScholarThere is no corresponding record for this reference.
- 245Martins, R. B.; Jorge, N.; Lucas, M. S.; Raymundo, A.; Barros, A. I. R. N. A.; Peres, J. A. Food By-Product Valorization by Using Plant-Based Coagulants Combined with AOPs for Agro-Industrial Wastewater Treatment. IJERPH. 2022, 19 (7), 4134, DOI: 10.3390/ijerph19074134Google ScholarThere is no corresponding record for this reference.
- 246Feng, Y.; Xie, T.; Li, F. New challenge: Mitigation and control of antibiotic resistant genes in aquatic environments by biochar. Sci. Total Environ. 2024, 946, 174385, DOI: 10.1016/j.scitotenv.2024.174385Google ScholarThere is no corresponding record for this reference.
- 247Fu, Y.; Wang, F.; Sheng, H. Removal of extracellular antibiotic resistance genes using magnetic biochar/quaternary phosphonium salt in aquatic environments: A mechanistic study. J. Hazard. Mater. 2021, 411, 125048, DOI: 10.1016/j.jhazmat.2021.125048Google ScholarThere is no corresponding record for this reference.
- 248Wu, C.; Fu, L.; Li, H.; Liu, X.; Wan, C. Using biochar to strengthen the removal of antibiotic resistance genes: Performance and mechanism. Sci. Total Environ. 2022, 816, 151554, DOI: 10.1016/j.scitotenv.2021.151554Google ScholarThere is no corresponding record for this reference.
- 249Wijitkosum, S. Biochar derived from agricultural wastes and wood residues for sustainable agricultural and environmental applications. Int. Soil Water Conserv. Res. 2022, 10 (2), 335– 341, DOI: 10.1016/j.iswcr.2021.09.006Google ScholarThere is no corresponding record for this reference.
- 250Sun, X.; Shi, L.; Zhang, H.; Li, F.; Long, Y.; Zhang, D. Chain-length-dependent adsorption of extracellular DNA on biochar: Behaviors, mechanisms, and structural Stability. Water Res.:X 2026, 30, 100496, DOI: 10.1016/j.wroa.2026.100496Google ScholarThere is no corresponding record for this reference.
- 251Sun, X.; Shi, L.; He, Z.; Zhang, H.; Li, F.; Zhang, D. Divalent metal ions facilitate environmental DNA adsorption on biochar by inducing new hydrogen bonds. Int. J. Biol. Macromol. 2025, 319, 145619, DOI: 10.1016/j.ijbiomac.2025.145619Google ScholarThere is no corresponding record for this reference.
- 252Wang, C.; Wang, Y.; Yan, S. Biochar-amended composting of lincomycin fermentation dregs promoted microbial metabolism and reduced antibiotic resistance genes. Bioresour. Technol. 2023, 367, 128253, DOI: 10.1016/j.biortech.2022.128253Google ScholarThere is no corresponding record for this reference.
- 253Kui, H.; Jingyang, C.; Mengxin, G.; Hui, X.; Li, L. Effects of biochars on the fate of antibiotics and their resistance genes during vermicomposting of dewatered sludge. J. Hazard. Mater. 2020, 397, 122767, DOI: 10.1016/j.jhazmat.2020.122767Google ScholarThere is no corresponding record for this reference.
- 254Li, J.; Lu, H.; Yang, H.; Wen, X.; Huang, Y.; Li, Q. Performances of antibiotic resistance genes profile upon the action of biochar-activated peroxydisulfate in composting. J. Environ. Manage. 2023, 334, 117509, DOI: 10.1016/j.jenvman.2023.117509Google ScholarThere is no corresponding record for this reference.
- 255Shi, L.; Zhang, D.; Yang, M. New discovery of extremely high adsorption of environmental DNA on cuttlefish bone pyrolysis derivative via large pore structure and carbon film. Waste Manage. 2024, 175, 286– 293, DOI: 10.1016/j.wasman.2024.01.016Google ScholarThere is no corresponding record for this reference.
- 256Saraei, F.; Amini, K.; Goodarzi, R.; Khanifar, A.; Asghari, B. A magnetically recoverable Chitosan-Based nanocomposite for simultaneous removal of tetracycline and antibiotic resistance genes from wastewater. Sci. Rep. 2026, 16 (1), 817, DOI: 10.1038/s41598-025-30583-4Google ScholarThere is no corresponding record for this reference.
- 257Xie, T.; Lin, D.; Cai, X. D. Nano-biochar regulates phage-host interactions, reducing antibiotic resistance genes in vermicomposting systems. Proc. Natl. Acad. Sci. U. S. A. 2025, 122 (34), e2511986122 DOI: 10.1073/pnas.2511986122Google ScholarThere is no corresponding record for this reference.
- 258Majaneva, M.; Diserud, O. H.; Eagle, S. H. C.; Boström, E.; Hajibabaei, M.; Ekrem, T. Environmental DNA filtration techniques affect recovered biodiversity. Sci. Rep. 2018, 8 (1), 4682, DOI: 10.1038/s41598-018-23052-8Google ScholarThere is no corresponding record for this reference.
- 259Schefer, R. B.; Paolucci, C.; Mitrano, D. M. Adsorption and Protection of Environmental DNA (eDNA) on Polymer and Silica Surfaces. Environ. Sci. Technol. 2025, 59 (36), 19479– 19489, DOI: 10.1021/acs.est.5c05439Google ScholarThere is no corresponding record for this reference.
- 260Boom, R.; Sol, C. J.; Salimans, M. M.; Jansen, C. L.; Wertheim-van Dillen, P. M.; Van Der Noordaa, J. Rapid and simple method for purification of nucleic acids. J. Clin. Microbiol. 1990, 28 (3), 495– 503, DOI: 10.1128/jcm.28.3.495-503.1990Google ScholarThere is no corresponding record for this reference.
- 261Lorenz, M. G.; Wackernagel, W. Adsorption of DNA to sand and variable degradation rates of adsorbed DNA. Appl. Environ. Microbiol. 1987, 53 (12), 2948– 2952, DOI: 10.1128/aem.53.12.2948-2952.1987Google ScholarThere is no corresponding record for this reference.
- 262Bag, S.; Rauwolf, S.; Schwaminger, S. P.; Wenzel, W.; Berensmeier, S. DNA Binding to the Silica: Cooperative Adsorption in Action. Langmuir 2021, 37 (19), 5902– 5908, DOI: 10.1021/acs.langmuir.1c00381Google ScholarThere is no corresponding record for this reference.
- 263Adegoke, S. C.; Yawlui, I. S. Y.; LaJeunesse, D. Silica Nanoparticles Block Natural Genetic Transformation in Acinetobacter baylyi ADP1. ACS Omega 2025, 10, 62609– 62620, DOI: 10.1021/acsomega.5c06566Google ScholarThere is no corresponding record for this reference.
- 264Cai, P.; Huang, Q.; Zhang, X.; Chen, H. Adsorption of DNA on clay minerals and various colloidal particles from an Alfisol. Soil Biol. Biochem. 2006, 38 (3), 471– 476, DOI: 10.1016/j.soilbio.2005.05.019Google ScholarThere is no corresponding record for this reference.
- 265Khanna, M.; Stotzky, G. Transformation of Bacillus subtilis by DNA bound on montmorillonite and effect of DNase on the transforming ability of bound DNA. Appl. Environ. Microbiol. 1992, 58 (6), 1930– 1939, DOI: 10.1128/aem.58.6.1930-1939.1992Google ScholarThere is no corresponding record for this reference.
- 266Buckner, M. M. C.; Ciusa, M. L.; Piddock, L. J. V. Strategies to combat antimicrobial resistance: anti-plasmid and plasmid curing. FEMS Microbiol. Rev. 2018, 42 (6), 781– 804, DOI: 10.1093/femsre/fuy031Google ScholarThere is no corresponding record for this reference.
- 267Zulauf, K. E.; Kirby, J. E. Discovery of small-molecule inhibitors of multidrug-resistance plasmid maintenance using a high-throughput screening approach. Proc. Natl. Acad. Sci. U. S. A. 2020, 117 (47), 29839– 29850, DOI: 10.1073/pnas.2005948117Google ScholarThere is no corresponding record for this reference.
- 268Beg, A. Z.; Ahmad, I. Effect of Plumbago zeylanica extract and certain curing agents on multidrug resistant bacteria of clinical origin. World J. Microbiol. Biotechnol. 2000, 16 (8–9), 841– 844, DOI: 10.1023/A:1008991724288Google ScholarThere is no corresponding record for this reference.
- 269Bharathi, A.; Polasa, H. Elimination of broad-host range plasmid vectors in Escherichia coli by curring agents. FEMS Microbiol. Lett. 1991, 84 (1), 37– 40, DOI: 10.1111/j.1574-6968.1991.tb04565.xGoogle ScholarThere is no corresponding record for this reference.
- 270Gómez-Rubio, E.; Arana, L.; Vicario-Martín, R. Exploring Inhibition of Bacterial Conjugation Coupling Protein TrwB: Novel Ligands to Fight Antimicrobial Resistance Spread. ACS Omega 2025, 10 (31), 34645– 34658, DOI: 10.1021/acsomega.5c03425Google ScholarThere is no corresponding record for this reference.
- 271Fernandez-Lopez, R.; Machón, C.; Longshaw, C. M. Unsaturated fatty acids are inhibitors of bacterial conjugation. Microbiology. 2005, 151 (11), 3517– 3526, DOI: 10.1099/mic.0.28216-0Google ScholarThere is no corresponding record for this reference.
- 272Álvarez-Rodríguez, I.; Arana, L.; Ugarte-Uribe, B. Type IV Coupling Proteins as Potential Targets to Control the Dissemination of Antibiotic Resistance. Front. Mol. Biosci. 2020, 7, 201, DOI: 10.3389/fmolb.2020.00201Google ScholarThere is no corresponding record for this reference.
- 273Wei, F.; Cui, X.; Wang, Z.; Dong, C.; Li, J.; Han, X. Recoverable peroxidase-like Fe3O4@MoS2-Ag nanozyme with enhanced antibacterial ability. Chem. Eng. J. 2021, 408, 127240, DOI: 10.1016/j.cej.2020.127240Google ScholarThere is no corresponding record for this reference.
- 274Doolan, J. A.; Williams, G. T.; Hilton, K. L. F. Advancements in antimicrobial nanoscale materials and self-assembling systems. Chem. Soc. Rev. 2022, 51 (20), 8696– 8755, DOI: 10.1039/D1CS00915JGoogle ScholarThere is no corresponding record for this reference.
- 275Bharathan, S.; Sundaramoorthy, N. S.; Chandrasekaran, H. Sub lethal levels of platinum nanoparticle cures plasmid and in combination with carbapenem, curtails carbapenem resistant Escherichia coli. Sci. Rep. 2019, 9 (1), 5305, DOI: 10.1038/s41598-019-41489-3Google ScholarThere is no corresponding record for this reference.
- 276Markowicz, A.; Borymski, S.; Adamek, A.; Sułowicz, S. The influence of ZnO nanoparticles on horizontal transfer of resistance genes in lab and soil conditions. Environ. Res. 2023, 223, 115420, DOI: 10.1016/j.envres.2023.115420Google ScholarThere is no corresponding record for this reference.
- 277Otinov, G. D.; Lokteva, A. V.; Petrova, A. D. Positive and Negative Effects of Metal Oxide Nanoparticles on Antibiotic Resistance Genes Transfer. Antibiotics 2020, 9 (11), 742, DOI: 10.3390/antibiotics9110742Google ScholarThere is no corresponding record for this reference.
- 278Okkeh, M.; Bloise, N.; Restivo, E.; De Vita, L.; Pallavicini, P.; Visai, L. Gold Nanoparticles: Can They Be the Next Magic Bullet for Multidrug-Resistant Bacteria?. Nanomaterials 2021, 11 (2), 312, DOI: 10.3390/nano11020312Google ScholarThere is no corresponding record for this reference.
- 279Wang, H.; Qi, H.; Gong, S. Fe3O4 composited with MoS2 blocks horizontal gene transfer. Colloids Surf., B 2020, 185, 110569, DOI: 10.1016/j.colsurfb.2019.110569Google ScholarThere is no corresponding record for this reference.
- 280Chen, X.; Zhan, M.; Shao, H. Mechanism of reducing antibiotic resistance genes by nano-selenium during composting: insight into host microorganisms and a two-component system. J. Environ. Manage. 2025, 386, 125704, DOI: 10.1016/j.jenvman.2025.125704Google ScholarThere is no corresponding record for this reference.
- 281Das, A.; Biswas, R.; Najiar, R.; Thatharaju, P.; Biswas, S.; Mohan, U. Quorum sensing in bacteria: insights into communication and inhibition strategies─a review. Arch. Microbiol. 2026, 208 (4), 157, DOI: 10.1007/s00203-025-04610-xGoogle ScholarThere is no corresponding record for this reference.
- 282Zhu, L.; Lau, G. W. Inhibition of Competence Development, Horizontal Gene Transfer and Virulence in Streptococcus pneumoniae by a Modified Competence Stimulating Peptide. Bessen DE, ed. PLoS Pathog. 2011, 7 (9), e1002241 DOI: 10.1371/journal.ppat.1002241Google ScholarThere is no corresponding record for this reference.
- 283Malshikare, H.; Priyakumar, U. D.; Chatterjee, P.; Sengupta, D. Mechanistic principles of antimicrobial peptides uncovered by charge density-based machine learning. Chem. Commun. 2026, 62, 4067, DOI: 10.1039/D5CC06374DGoogle ScholarThere is no corresponding record for this reference.
- 284Domenech, A.; Brochado, A. R.; Sender, V. Proton Motive Force Disruptors Block Bacterial Competence and Horizontal Gene Transfer. Cell Host Microbe 2020, 27 (4), 544– 555, DOI: 10.1016/j.chom.2020.02.002Google ScholarThere is no corresponding record for this reference.
- 285Jia, Y.; Zheng, Z.; Yang, B.; Zhang, H.; Wang, Z.; Liu, Y. A Broad-Spectrum Horizontal Transfer Inhibitor Prevents Transmission of Plasmids Carrying Multiple Antibiotic Resistance Genes. Khurshid M, ed. Transboundary Emerging Dis. 2024, 2024 (1), 7063673, DOI: 10.1155/2024/7063673Google ScholarThere is no corresponding record for this reference.
- 286Kogay, R.; Wolf, Y. I.; Koonin, E. V. Defence systems and horizontal gene transfer in bacteria. Environ. Microbiol. 2024, 26 (4), e16630 DOI: 10.1111/1462-2920.16630Google ScholarThere is no corresponding record for this reference.
- 287Hwang, S.; Maxwell, K. L. Meet the Anti-CRISPRs: Widespread Protein Inhibitors of CRISPR-Cas Systems. CRISPR J. 2019, 2 (1), 23– 30, DOI: 10.1089/crispr.2018.0052Google ScholarThere is no corresponding record for this reference.
- 288Isaev, A. B.; Musharova, O. S.; Severinov, K. V. Microbial Arsenal of Antiviral Defenses - Part I. Biochemistry (Moscow) 2021, 86 (3), 319– 337, DOI: 10.1134/S0006297921030081Google ScholarThere is no corresponding record for this reference.
- 289Isaev, A. B.; Musharova, O. S.; Severinov, K. V. Microbial Arsenal of Antiviral Defenses. Part II. Biochemistry (Moscow) 2021, 86 (4), 449– 470, DOI: 10.1134/S0006297921040064Google ScholarThere is no corresponding record for this reference.
- 290Price, V. J.; McBride, S. W.; Hullahalli, K.; Chatterjee, A.; Duerkop, B. A.; Palmer, K. L. Enterococcus faecalis CRISPR-Cas Is a Robust Barrier to Conjugative Antibiotic Resistance Dissemination in the Murine Intestine. Ellermeier CD, ed. mSphere 2019, 4 (4), e00464–19 DOI: 10.1128/mSphere.00464-19Google ScholarThere is no corresponding record for this reference.
- 291Rafiq, M. S.; Shabbir, M. A.; Raza, A. CRISPR-Cas System: A New Dawn to Combat Antibiotic Resistance. BioDrugs 2024, 38 (3), 387– 404, DOI: 10.1007/s40259-024-00656-3Google ScholarThere is no corresponding record for this reference.
- 292O’Meara, D.; Nunney, L. A phylogenetic test of the role of CRISPR-Cas in limiting plasmid acquisition and prophage integration in bacteria. Plasmid 2019, 104, 102418, DOI: 10.1016/j.plasmid.2019.102418Google ScholarThere is no corresponding record for this reference.
- 293Wu, Y.; Garushyants, S. K.; Van Den Hurk, A. Bacterial defense systems exhibit synergistic anti-phage activity. Cell Host Microbe 2024, 32 (4), 557– 572, DOI: 10.1016/j.chom.2024.01.015Google ScholarThere is no corresponding record for this reference.
- 294Wheatley, R. M.; MacLean, R. C. CRISPR-Cas systems restrict horizontal gene transfer in Pseudomonas aeruginosa. ISME J. 2021, 15 (5), 1420– 1433, DOI: 10.1038/s41396-020-00860-3Google ScholarThere is no corresponding record for this reference.
- 295Mahendra, C.; Christie, K. A.; Osuna, B. A.; Pinilla-Redondo, R.; Kleinstiver, B. P.; Bondy-Denomy, J. Broad-spectrum anti-CRISPR proteins facilitate horizontal gene transfer. Nat. Microbiol. 2020, 5 (4), 620– 629, DOI: 10.1038/s41564-020-0692-2Google ScholarThere is no corresponding record for this reference.
- 296Park, S. J.; Lee, G. E.; Cho, S. M.; Choi, E. H. Recent applications, future perspectives, and limitations of the CRISPR-Cas system. Mol. Ther. Nucleic Acids 2025, 36 (3), 102634, DOI: 10.1016/j.omtn.2025.102634Google ScholarThere is no corresponding record for this reference.
- 297Palacios Araya, D.; Palmer, K. L.; Duerkop, B. A. CRISPR-based antimicrobials to obstruct antibiotic-resistant and pathogenic bacteria. Kline KA, ed. PLoS Pathog. 2021, 17 (7), e1009672 DOI: 10.1371/journal.ppat.1009672Google ScholarThere is no corresponding record for this reference.
- 298Wu, Y.; Battalapalli, D.; Hakeem, M. J. Engineered CRISPR-Cas systems for the detection and control of antibiotic-resistant infections. J. Nanobiotechnol. 2021, 19 (1), 401, DOI: 10.1186/s12951-021-01132-8Google ScholarThere is no corresponding record for this reference.
- 299Gomaa, A. A.; Klumpe, H. E.; Luo, M. L.; Selle, K.; Barrangou, R.; Beisel, C. L. Programmable Removal of Bacterial Strains by Use of Genome-Targeting CRISPR-Cas Systems. mBio 2014, 5 (1), e00928–13 DOI: 10.1128/mBio.00928-13Google ScholarThere is no corresponding record for this reference.
- 300Kippnich, J.; Benz, F.; Uecker, H.; Baumdicker, F. Effectiveness of CRISPR-Cas in sensitizing bacterial populations with plasmid-encoded antimicrobial resistance. Lenormand T, ed. GENETICS. 2025, 231 (3), iyaf192, DOI: 10.1093/genetics/iyaf192Google ScholarThere is no corresponding record for this reference.
- 301Saffari Natanzi, A.; Poudineh, M.; Karimi, E.; Khaledi, A.; Haddad Kashani, H. Innovative approaches to combat antibiotic resistance: integrating CRISPR/Cas9 and nanoparticles against biofilm-driven infections. BMC Med. 2025, 23 (1), 486, DOI: 10.1186/s12916-025-04323-4Google ScholarThere is no corresponding record for this reference.
- 302Zuberi, A.; Ahmad, N.; Ahmad, H.; Saeed, M.; Ahmad, I. Beyond antibiotics: CRISPR/Cas9 triumph over biofilm-associated antibiotic resistance infections. Front. Cell. Infect. Microbiol. 2024, 14, 1408569, DOI: 10.3389/fcimb.2024.1408569Google ScholarThere is no corresponding record for this reference.
- 303Joseph, J.; Abraham, J. S.; Abraham, R.; Nagella, P.; Anand, A. Filamentous Fungi: Heterologous Protein Expression. Prospects of Fungal Biotechnologies for Livestock Vol. 2; Gupta, A., Ed.; Springer Nature Switzerland, 2025; pp 197– 231. DOI: 10.1007/978-3-032-06478-3_8 .Google ScholarThere is no corresponding record for this reference.
- 304Pandey, P.; Vavilala, S. L. From Gene Editing to Biofilm Busting: CRISPR-CAS9 Against Antibiotic Resistance–A Review. Cell Biochem. Biophys. 2024, 82 (2), 549– 560, DOI: 10.1007/s12013-024-01276-yGoogle ScholarThere is no corresponding record for this reference.
- 305Rodrigues, M.; McBride, S. W.; Hullahalli, K.; Palmer, K. L.; Duerkop, B. A. Conjugative Delivery of CRISPR-Cas9 for the Selective Depletion of Antibiotic-Resistant Enterococci. Antimicrob. Agents Chemother. 2019, 63 (11), e01454-19 DOI: 10.1128/AAC.01454-19Google ScholarThere is no corresponding record for this reference.
- 306Peng, H.; Chen, I. A. Rapid Colorimetric Detection of Bacterial Species through the Capture of Gold Nanoparticles by Chimeric Phages. ACS Nano 2019, acsnano.8b06395, DOI: 10.1021/acsnano.8b06395Google ScholarThere is no corresponding record for this reference.
- 307Bier, E.; Nizet, V. Driving to Safety: CRISPR-Based Genetic Approaches to Reducing Antibiotic Resistance. Trends Genet. 2021, 37 (8), 745– 757, DOI: 10.1016/j.tig.2021.02.007Google ScholarThere is no corresponding record for this reference.
- 308Kaduwal, S.; Stuart, E. C.; Auradkar, A.; Washabaugh, S.; Meyer, J. R.; Bier, E. A conjugal gene drive-like system efficiently suppresses antibiotic resistance in a bacterial population. npj Antimicrob Resist. 2026, 4 (1), 8, DOI: 10.1038/s44259-026-00181-zGoogle ScholarThere is no corresponding record for this reference.
- 309Tsoi, R.; Son, H. I.; Hamrick, G. S. A predatory gene drive for targeted control of self-transmissible plasmids. Sci. Adv. 2025, 11 (14), eads4735 DOI: 10.1126/sciadv.ads4735Google ScholarThere is no corresponding record for this reference.
- 310Walter, M.; Verdin, E. Viral gene drive in herpesviruses. Nat. Commun. 2020, 11 (1), 4884, DOI: 10.1038/s41467-020-18678-0Google ScholarThere is no corresponding record for this reference.
- 311Gantz, V. M.; Jasinskiene, N.; Tatarenkova, O.; Fazekas, A.; Macias, V. M.; Bier, E.; James, A. A. Highly efficient Cas9-mediated gene drive for population modification of the malaria vector mosquito Anopheles stephensi. Proc. Natl. Acad. Sci. U. S. A. 2015, 112(49). DOI: 10.1073/pnas.1521077112 .Google ScholarThere is no corresponding record for this reference.
- 312Valderrama, J. A.; Kulkarni, S. S.; Nizet, V.; Bier, E. A bacterial gene-drive system efficiently edits and inactivates a high copy number antibiotic resistance locus. Nat. Commun. 2019, 10 (1), 5726, DOI: 10.1038/s41467-019-13649-6Google ScholarThere is no corresponding record for this reference.
- 313Asim, M.; Rizvi, S. A.; Haq, Q. M. R. Antiplasmid systems: a novel strategy to combat antibiotic resistance. J. Antimicrob. Chemother. 2026, 81 (1), dkaf472, DOI: 10.1093/jac/dkaf472Google ScholarThere is no corresponding record for this reference.
- 314Bravo, J. P. K. Anti-plasmid immunity: a key to pathogen success?. Future Microbiol. 2024, 19 (15), 1269– 1272, DOI: 10.1080/17460913.2024.2389720Google ScholarThere is no corresponding record for this reference.
- 315Karaiskos, I.; Galani, I.; Papoutsaki, V.; Galani, L.; Giamarellou, H. Carbapenemase producing Klebsiella pneumoniae: implication on future therapeutic strategies. Expert Rev. Anti-Infect. Ther. 2022, 20 (1), 53– 69, DOI: 10.1080/14787210.2021.1935237Google ScholarThere is no corresponding record for this reference.
- 316Zongo, P. D.; Cabanel, N.; Royer, G. An antiplasmid system drives antibiotic resistance gene integration in carbapenemase-producing Escherichia coli lineages. Nat. Commun. 2024, 15 (1), 4093, DOI: 10.1038/s41467-024-48219-yGoogle ScholarThere is no corresponding record for this reference.
Cited By
This article has not yet been cited by other publications.
Article Views
Altmetric
Citations
Article Views are the COUNTER-compliant sum of full text article downloads since November 2008 (both PDF and HTML) across all institutions and individuals. These metrics are regularly updated to reflect usage leading up to the last few days.
Citations are the number of other articles citing this article, calculated by Crossref and updated daily. Find more information about Crossref citation counts.
The Altmetric Attention Score is a quantitative measure of the attention that a research article has received online. Clicking on the donut icon will load a page at altmetric.com with additional details about the score and the social media presence for the given article. Find more information on the Altmetric Attention Score and how the score is calculated.
Recommended Articles
Abstract
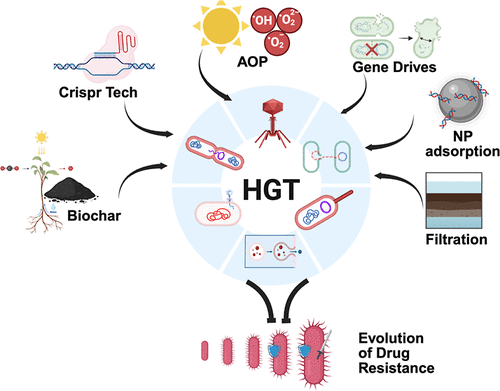
Figure 1
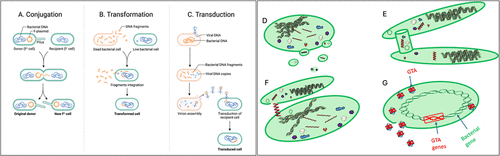
Figure 1. Canonical and noncanonical routes of HGT. (A) Conjugation involves cell-to-cell contact. (B) Transformation involves the uptake of extracellular DNA from the environment. (C) Transduction involves attachment of phage to bacteria followed by subsequent injection of its DNA into the host. (D) HGT through membrane vesicles, where bacteria release vesicles containing cellular materials that are taken up by recipient cells. (E) HGT through nanotubes, which are membrane extensions forming direct bridges between neighboring bacterial cells to enable exchange of cytoplasmic materials. (F) HGT through autolysis. Autolysis involves the self-lysis of a subpopulation of bacterial cells, releasing extracellular DNA that becomes available for uptake by naturally competent bacterial cells. (G) HGT via gene transfer agents (GTAs), which package host DNA fragments and release them upon lysis for uptake by recipient cells.
Figure 2
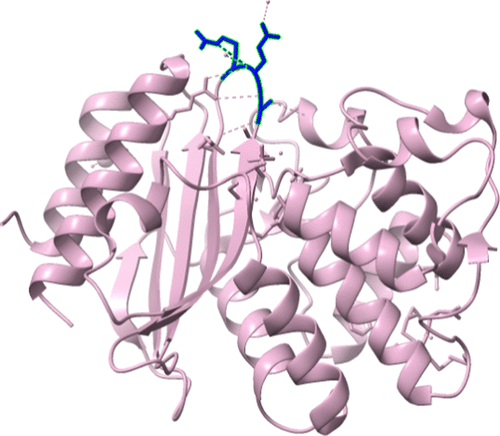
Figure 2. Crystal structure of stabilized TEM-1 beta-lactamase variant v.13 carrying G238S mutation. (129) The blue colored region with hydrogen bonding is the mutation region.
Figure 3
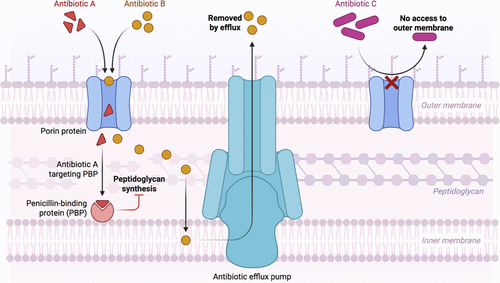
Figure 3. Schematic representation of the efflux pump system in the Gram-negative bacterial cell membrane. The outer membrane (OM) at the top, the peptidoglycan layer (PGL) in the middle, and the inner membrane (IM) at the bottom, with the cytoplasm beneath. Porins are water-filled channels embedded in the outer membrane that allow hydrophilic molecules and small metabolites to pass through passively. They are essential for nutrient uptake and waste expulsion, functioning as molecular sieves that exclude larger or hydrophobic molecules.
Figure 4
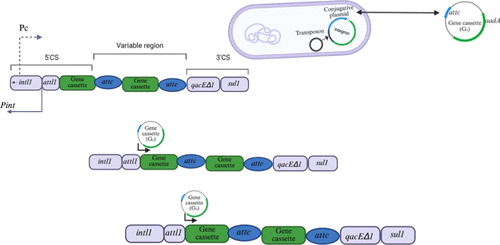
Figure 4. Structure of class 1 integrons. Integrons function as modular genetic elements that facilitate the acquisition, chromosomal insertion, and expression of gene cassettes, frequently encoding antibiotic resistance determinants, via a site-specific recombination mechanism catalyzed by an integrase enzyme.
Figure 5
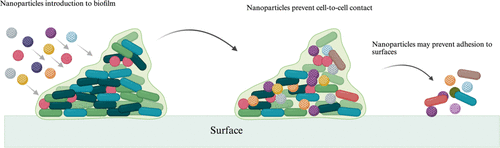
Figure 5. Proposed routes to control HGT. Nanoparticle disruption of the biofilm architecture, this essentially prevent cell proximity, thus preventing conjugation. In some cases, the nanoparticle may actively compete with the bacteria for the extracellular DNA. Sequestering the extracellular DNA is a route to limit resistance evolution.
Figure 6
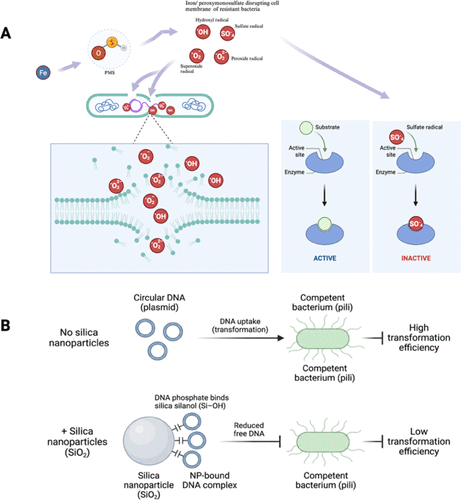
Figure 6. Proposed methods of controlling HGT in bacterial niche. (A) Iron activates the peroxymonosulfate (PMS) resulting to the production of radical species that degrade the cell membrane and disrupt enzyme metabolism, thus preventing conjugative transfer of resistance genes. (B) Silica nanoparticle binds to extracellular DNA by an attractive interaction between DNA phosphate groups and surface silanol groups on silica, thus sequestering the DNA and blocking transformation.
Figure 7
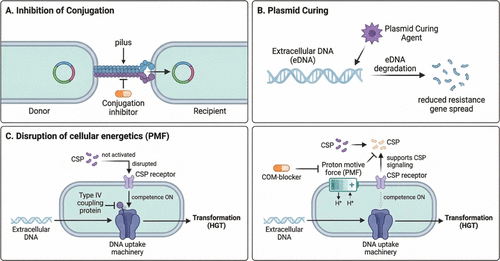
Figure 7. Strategies to control horizontal gene transfer. (A) Conjugation inhibiting drugs that inactivate components of proteins involved in the plasmid transfer machinery, specifically the Type IV coupling protein that links the relaxosome and the Type IV secretion system (T4SS). (B). Plasmid curing involves a variety of chemicals including detergents, biocides, DNA intercalating agents, antibiotics, psychotropic drugs, and plant-derived compounds that target eDNA for degradation. (C) Competence blockers (COM-blockers) disrupt the proton motive force, which inhibits quorum-sensing signaling via CSP, thereby preventing activation of the DNA uptake machinery.
Figure 8
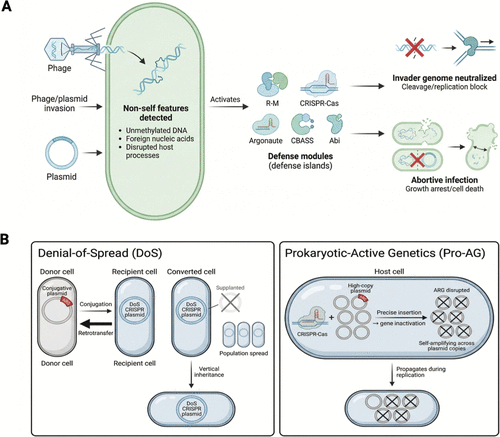
Figure 8. Bacterial innate immune defense against phages/plasmids. (A). Bacteria possess sophisticated innate immune mechanisms capable of mounting effective responses against invading phages and foreign genetic elements. These defenses comprise a diverse array of genetically encoded systems that are frequently colocalized within discrete genomic regions known as “defense islands”. The repertoire of characterized defense systems continues to expand and currently encompass restriction-modification systems, CRISPR-Cas machinery, Argonaute-based defense pathways, cyclic-oligonucleotide-based antiphage signaling systems (CBASS), and abortive-infection modules, each representing a functionally distinct strategy for detecting and neutralizing nonself-nucleic acids. (B). CRISPR-based prokaryotic gene drives targeting plasmid ARGs. Through this mechanism, the DoS plasmid is able to disseminate novel genetic information throughout a target bacterial population while simultaneously outcompeting the conjugative plasmid, displacing it not only as the primary vehicle of horizontal gene transfer but also as a heritable element passed vertically to daughter cells.
References
This article references 316 other publications.
- 1Baym, M.; Stone, L. K.; Kishony, R. Multidrug evolutionary strategies to reverse antibiotic resistance. Science. 2016, 351 (6268), aad3292, DOI: 10.1126/science.aad3292There is no corresponding record for this reference.
- 2Zavaleta-Monestel, E.; Arguedas-Chacón, S.; Rojas-Chinchilla, C.; Díaz-Madriz, J. P. Antimicrobial Resistance: An Emerging Global Threat to Modern Medicine. Cureus. 2025, DOI: 10.7759/cureus.97668There is no corresponding record for this reference.
- 3Mouzakis, A.; Panagopoulos, P.; Papazoglou, D.; Petrakis, V. A Comprehensive Review of Nanoparticles in the Fight Against Antimicrobial Resistance. Pathogens 2025, 14 (11), 1090, DOI: 10.3390/pathogens14111090There is no corresponding record for this reference.
- 4Smith, P. A.; Romesberg, F. E. Combating bacteria and drug resistance by inhibiting mechanisms of persistence and adaptation. Nat. Chem. Biol. 2007, 3 (9), 549– 556, DOI: 10.1038/nchembio.2007.27There is no corresponding record for this reference.
- 5Zhang, S.; Lu, J.; Wang, Y.; Verstraete, W.; Yuan, Z.; Guo, J. Insights of metallic nanoparticles and ions in accelerating the bacterial uptake of antibiotic resistance genes. J. Hazard. Mater. 2022, 421, 126728, DOI: 10.1016/j.jhazmat.2021.126728There is no corresponding record for this reference.
- 6Basarab, G. S.; Ghorpade, S.; Gibhard, L.; Mueller, R.; Njoroge, M.; Peton, N.; Govender, P.; Massoudi, L. M.; Robertson, G. T.; Lenaerts, A. J. Spiropyrimidinetriones: a Class of DNA Gyrase Inhibitors with Activity against Mycobacterium tuberculosis and without Cross-Resistance to Fluoroquinolones. Antimicrob. Agents Chemother. 2022, 66 (4), e02192-21 DOI: 10.1128/aac.02192-21There is no corresponding record for this reference.
- 7Dubnau, D.; Blokesch, M. Mechanisms of DNA Uptake by Naturally Competent Bacteria. Annu. Rev. Genet. 2019, 53 (1), 217– 237, DOI: 10.1146/annurev-genet-112618-043641There is no corresponding record for this reference.
- 8Khedkar, S.; Smyshlyaev, G.; Letunic, I. Landscape of mobile genetic elements and their antibiotic resistance cargo in prokaryotic genomes. Nucleic Acids Res. 2022, 50 (6), 3155– 3168, DOI: 10.1093/nar/gkac163There is no corresponding record for this reference.
- 9Lang, A. S.; Buchan, A.; Burrus, V. Interactions and evolutionary relationships among bacterial mobile genetic elements. Nat. Rev. Microbiol. 2025, 23 (7), 423– 438, DOI: 10.1038/s41579-025-01157-yThere is no corresponding record for this reference.
- 10Haudiquet, M.; De Sousa, J. M.; Touchon, M.; Rocha, E. P. C. Selfish, promiscuous and sometimes useful: how mobile genetic elements drive horizontal gene transfer in microbial populations. Philosophical Transactions of the Royal Society B 2022, 377 (1861), 20210234, DOI: 10.1098/rstb.2021.0234There is no corresponding record for this reference.
- 11Di Giacomo, S.; Toussaint, F.; Ledesma-García, L. Expanding natural transformation to improve beneficial lactic acid bacteria. FEMS Microbiol. Rev. 2022, 46 (4), fuac014, DOI: 10.1093/femsre/fuac014There is no corresponding record for this reference.
- 12Marinacci, B.; Krzyżek, P.; Pellegrini, B.; Turacchio, G.; Grande, R. Latest Update on Outer Membrane Vesicles and Their Role in Horizontal Gene Transfer: A Mini-Review. Membranes 2023, 13 (11), 860, DOI: 10.3390/membranes13110860There is no corresponding record for this reference.
- 13Morawska, L. P.; Kuipers, O. P. Cell-to-cell non-conjugative plasmid transfer between Bacillus subtilis and lactic acid bacteria. Microb. Biotechnol. 2023, 16 (4), 784– 798, DOI: 10.1111/1751-7915.14195There is no corresponding record for this reference.
- 14Rodríguez-Beltrán, J.; DelaFuente, J.; León-Sampedro, R.; MacLean, R. C.; San Millán, A. ́. Beyond horizontal gene transfer: the role of plasmids in bacterial evolution. Nat. Rev. Microbiol. 2021, 19 (6), 347– 359, DOI: 10.1038/s41579-020-00497-1There is no corresponding record for this reference.
- 15Hashimoto, Y.; Taniguchi, M.; Uesaka, K. Novel Multidrug-Resistant Enterococcal Mobile Linear Plasmid pELF1 Encoding vanA and vanM Gene Clusters From a Japanese Vancomycin-Resistant Enterococci Isolate. Front. Microbiol. 2019, 10, 2568, DOI: 10.3389/fmicb.2019.02568There is no corresponding record for this reference.
- 16Rozwandowicz, M.; Brouwer, M. S. M.; Fischer, J. Plasmids carrying antimicrobial resistance genes in Enterobacteriaceae. J. Antimicrob. Chemother. 2018, 73 (5), 1121– 1137, DOI: 10.1093/jac/dkx488There is no corresponding record for this reference.
- 17Suzuki, H.; Yano, H.; Brown, C. J.; Top, E. M. Predicting Plasmid Promiscuity Based on Genomic Signature. J. Bacteriol. 2010, 192 (22), 6045– 6055, DOI: 10.1128/JB.00277-10There is no corresponding record for this reference.
- 18Zechner, E. L.; Moncalián, G.; De La Cruz, F. Relaxases and Plasmid Transfer in Gram-Negative Bacteria. Type IV Secretion in Gram-Negative and Gram-Positive Bacteria; Backert, S., Grohmann, E., Eds.; Springer International Publishing, 2017; Vol. 413, pp 93– 113. DOI: 10.1007/978-3-319-75241-9_4 .There is no corresponding record for this reference.
- 19Smillie, C.; Garcillán-Barcia, M. P.; Francia, M. V.; Rocha, E. P. C.; De La Cruz, F. Mobility of Plasmids. Microbiol. Mol. Biol. Rev. 2010, 74 (3), 434– 452, DOI: 10.1128/MMBR.00020-10There is no corresponding record for this reference.
- 20Carattoli, A. Resistance Plasmid Families in Enterobacteriaceae. Antimicrob. Agents Chemother. 2009, 53 (6), 2227– 2238, DOI: 10.1128/AAC.01707-08There is no corresponding record for this reference.
- 21Thomas, C. M.; Nielsen, K. M. Mechanisms of, and Barriers to, Horizontal Gene Transfer between Bacteria. Nat. Rev. Microbiol. 2005, 3 (9), 711– 721, DOI: 10.1038/nrmicro1234There is no corresponding record for this reference.
- 22Fricke, W. F.; Welch, T. J.; McDermott, P. F. Comparative Genomics of the IncA/C Multidrug Resistance Plasmid Family. J. Bacteriol. 2009, 191 (15), 4750– 4757, DOI: 10.1128/JB.00189-09There is no corresponding record for this reference.
- 23Guo, X.; Chen, R.; Wang, Q. Global prevalence, characteristics, and future prospects of IncX3 plasmids: A review. Front. Microbiol. 2022, 13, 979558, DOI: 10.3389/fmicb.2022.979558There is no corresponding record for this reference.
- 24Rasheed, J. K.; Kitchel, B.; Zhu, W. New Delhi Metallo-β-Lactamase-producing Enterobacteriaceae, United States. Emerg Infect Dis. 2013, 19 (6), 870– 878, DOI: 10.3201/eid1906.121515There is no corresponding record for this reference.
- 25Segundo-Arizmendi, N.; Arellano-Maciel, D.; Rivera-Ramírez, A.; Piña-González, A. M.; López-Leal, G.; Hernández-Baltazar, E. Bacteriophages: A Challenge for Antimicrobial Therapy. Microorganisms 2025, 13 (1), 100, DOI: 10.3390/microorganisms13010100There is no corresponding record for this reference.
- 26Touchon, M.; Moura De Sousa, J. A.; Rocha, E. P. Embracing the enemy: the diversification of microbial gene repertoires by phage-mediated horizontal gene transfer. Curr. Opin. Microbiol. 2017, 38, 66– 73, DOI: 10.1016/j.mib.2017.04.010There is no corresponding record for this reference.
- 27Pires, J.; Santos, R.; Monteiro, S. Antibiotic resistance genes in bacteriophages from wastewater treatment plant and hospital wastewaters. Sci. Total Environ. 2023, 892, 164708, DOI: 10.1016/j.scitotenv.2023.164708There is no corresponding record for this reference.
- 28Waldor, M. K.; Mekalanos, J. J. Lysogenic Conversion by a Filamentous Phage Encoding Cholera Toxin. Science. 1996, 272 (5270), 1910– 1914, DOI: 10.1126/science.272.5270.1910There is no corresponding record for this reference.
- 29Goerke, C.; Pantucek, R.; Holtfreter, S. Diversity of Prophages in Dominant Staphylococcus aureus Clonal Lineages. J. Bacteriol. 2009, 191 (11), 3462– 3468, DOI: 10.1128/JB.01804-08There is no corresponding record for this reference.
- 30Colavecchio, A.; Cadieux, B.; Lo, A.; Goodridge, L. D. Bacteriophages Contribute to the Spread of Antibiotic Resistance Genes among Foodborne Pathogens of the Enterobacteriaceae Family - A Review. Front. Microbiol. 2017, 8, 1108, DOI: 10.3389/fmicb.2017.01108There is no corresponding record for this reference.
- 31Johnson, C. M.; Grossman, A. D. Integrative and Conjugative Elements (ICEs): What They Do and How They Work. Annu. Rev. Genet. 2015, 49 (1), 577– 601, DOI: 10.1146/annurev-genet-112414-055018There is no corresponding record for this reference.
- 32Benigno, V.; Carraro, N.; Sarton-Lohéac, G.; Romano-Bertrand, S.; Blanc, D. S.; Van, D. M., JR. Diversity and evolution of an abundant ICE clc family of integrative and conjugative elements in Pseudomonas aeruginosa. Gales AC, ed. mSphere 2023, 8 (6), e00517–23 DOI: 10.1128/msphere.00517-23There is no corresponding record for this reference.
- 33Roberts, A. P.; Mullany, P. Tn 916 -like genetic elements: a diverse group of modular mobile elements conferring antibiotic resistance. FEMS Microbiol. Rev. 2011, 35 (5), 856– 871, DOI: 10.1111/j.1574-6976.2011.00283.xThere is no corresponding record for this reference.
- 34Auchtung, J. M.; Aleksanyan, N.; Bulku, A.; Berkmen, M. B. Biology of ICE Bs1, an integrative and conjugative element in Bacillus subtilis. Plasmid 2016, 86, 14– 25, DOI: 10.1016/j.plasmid.2016.07.001There is no corresponding record for this reference.
- 35Wozniak, R. A. F.; Waldor, M. K. Integrative and conjugative elements: mosaic mobile genetic elements enabling dynamic lateral gene flow. Nat. Rev. Microbiol. 2010, 8 (8), 552– 563, DOI: 10.1038/nrmicro2382There is no corresponding record for this reference.
- 36Burrus, V.; Pavlovic, G.; Decaris, B.; Guédon, G. The ICESt1 element of Streptococcus thermophilus belongs to a large family of integrative and conjugative elements that exchange modules and change their specificity of integration. Plasmid 2002, 48 (2), 77– 97, DOI: 10.1016/S0147-619X(02)00102-6There is no corresponding record for this reference.
- 37Blokesch, M. Defence systems encoded by core genomic islands of seventh pandemic Vibrio cholerae. Phil Trans R Soc. B 2025, 380 (1934), 20240083, DOI: 10.1098/rstb.2024.0083There is no corresponding record for this reference.
- 38Nusrat, S.; Aliyu, M.; Zohora, F. T. Mechanisms of antimicrobial resistance: From genetic evolution to clinical manifestations. AIMSMICRO. 2025, 11 (4), 1007– 1034, DOI: 10.3934/microbiol.2025045There is no corresponding record for this reference.
- 39Partridge, S. R.; Kwong, S. M.; Firth, N.; Jensen, S. O. Mobile Genetic Elements Associated with Antimicrobial Resistance. Clin. Microbiol. Rev. 2018, 31 (4), e00088-17 DOI: 10.1128/CMR.00088-17There is no corresponding record for this reference.
- 40Roy, S.; Nandy, S.; Morita, D. Genomic analysis of a novel high-risk ST5217/ExoU+/O11 clone of carbapenem-resistant OXA-181- and VIM-2-producing Pseudomonas aeruginosa in India. J. Global Antimicrob. Resist. 2026, 46, 158– 161, DOI: 10.1016/j.jgar.2025.12.002There is no corresponding record for this reference.
- 41Calbet, A. Pelagic Shuttles of Antibiotic Resistance Genes: Zooplankton as Overlooked Vectors Across Space and Food Webs. Microb. Ecol. 2026, 89 (1), 12, DOI: 10.1007/s00248-025-02669-zThere is no corresponding record for this reference.
- 42Wachino, J. Horizontal Gene Transfer Systems for Spread of Antibiotic Resistance in Gram-Negative Bacteria. Microbiol. Immunol. 2025, 69 (7), 367– 376, DOI: 10.1111/1348-0421.13222There is no corresponding record for this reference.
- 43Wang, X.; Chen, Z.; Liu, C.; Zhang, Z.; Deng, Y.; Tao, L.; Tiedje, J. M.; Deng, J. Type I-F CRISPR-associated transposons contribute to genomic plasticity in Shewanella and mediate efficient programmable DNA integration. Microb. Genomics 2025, 11(8). DOI: 10.1099/mgen.0.001476 .There is no corresponding record for this reference.
- 44Deng, Y.; Bao, X.; Ji, L. Resistance integrons: class 1, 2 and 3 integrons. Ann. Clin. Microbiol. Antimicrob. 2015, 14 (1), 45, DOI: 10.1186/s12941-015-0100-6There is no corresponding record for this reference.
- 45Stokes, H. W.; Gillings, M. R. Gene flow, mobile genetic elements and the recruitment of antibiotic resistance genes into Gram-negative pathogens. FEMS Microbiol. Rev. 2011, 35 (5), 790– 819, DOI: 10.1111/j.1574-6976.2011.00273.xThere is no corresponding record for this reference.
- 46Nicolas, E.; Lambin, M.; Dandoy, D.; The Tn 3 -family of Replicative Transposons Mobile DNA III; Craig, N. L., Chandler, M., Gellert, M., Lambowitz, A. M., Rice, P. A., Sandmeyer, S. B., Eds.; ASM Press, 2015; pp 693– 726. DOI: 10.1128/9781555819217.ch32 .There is no corresponding record for this reference.
- 47Peters, J. E. Targeted transposition with Tn7 elements: safe sites, mobile plasmids, CRISPR/Cas and beyond. Mol. Microbiol. 2019, 112 (6), 1635– 1644, DOI: 10.1111/mmi.14383There is no corresponding record for this reference.
- 48Peters, J. E. Tn7 Mobile DNA III; Craig, N. L., Chandler, M., Gellert, M., Lambowitz, A. M., Rice, P. A., Sandmeyer, S. B., Eds.; ASM Press, 2015; pp 647– 667. DOI: 10.1128/9781555819217.ch30 .There is no corresponding record for this reference.
- 49Courvalin, P. Vancomycin Resistance in Gram-Positive Cocci. Clin. Infect. Dis. 2006, 42 (Supplement_1), S25– S34, DOI: 10.1086/491711There is no corresponding record for this reference.
- 50Bush, K. Evolution of β-Lactamases: Past, Present, and Future. Antibiotic Discovery and Development; Dougherty, T. J., Pucci, M. J., Eds.; Springer US, 2012; pp 427– 453. DOI: 10.1007/978-1-4614-1400-1_12 .There is no corresponding record for this reference.
- 51Naas, T.; Cuzon, G.; Villegas, M. V.; Lartigue, M. F.; Quinn, J. P.; Nordmann, P. Genetic Structures at the Origin of Acquisition of the β-Lactamase blaKPC Gene. Antimicrob. Agents Chemother. 2008, 52 (4), 1257– 1263, DOI: 10.1128/AAC.01451-07There is no corresponding record for this reference.
- 52Ameyama, S.; Onodera, S.; Takahata, M. Mosaic-Like Structure of Penicillin-Binding Protein 2 Gene (penA) in Clinical Isolates of Neisseria gonorrhoeae with Reduced Susceptibility to Cefixime. Antimicrob. Agents Chemother. 2002, 46 (12), 3744– 3749, DOI: 10.1128/AAC.46.12.3744-3749.2002There is no corresponding record for this reference.
- 53Maziero, M.; Juillot, D.; Mortier-Barrière, I. A toxin/antitoxin system targeting the replication sliding-clamp induces competence in Streptococcus pneumoniae. Kjos M, ed. PLoS Genet. 2025, 21 (12), e1011863 DOI: 10.1371/journal.pgen.1011863There is no corresponding record for this reference.
- 54Hakenbeck, R. Transformation in: mosaic genes and the regulation of competence. Res. Microbiol. 2000, 151 (6), 453– 456, DOI: 10.1016/S0923-2508(00)00170-4There is no corresponding record for this reference.
- 55Wadsworth, C. B.; Goytia, M.; Shafer, W. M. Commensal Neisseria and Antimicrobial-Resistant Gonorrhea. Annu. Rev. Microbiol. 2025, 79 (1), 215– 240, DOI: 10.1146/annurev-micro-022024-024306There is no corresponding record for this reference.
- 56Unitt, A.; Maiden, M.; Harrison, O. Characterizing the diversity and commensal origins of penA mosaicism in the genus Neisseria. Microb. Genomics 2024, 10(2). DOI: 10.1099/mgen.0.001209 .There is no corresponding record for this reference.
- 57Hanao, M.; Aoki, K.; Ishii, Y.; Shimuta, K.; Ohnishi, M.; Tateda, K. Molecular characterization of Neisseria gonorrhoeae isolates collected through a national surveillance programme in Japan, 2013: evidence of the emergence of a ceftriaxone-resistant strain from a ceftriaxone-susceptible lineage. J. Antimicrob. Chemother. 2021, 76 (7), 1769– 1775, DOI: 10.1093/jac/dkab104There is no corresponding record for this reference.
- 58Liu, E. Y. M.; Chang, J. C.; Lin, J. C.; Chang, F. Y.; Fung, C. P. Important Mutations Contributing to High-Level Penicillin Resistance in Taiwan19F −14, Taiwan23F −15, and Spain23F −1 of Streptococcus pneumoniae Isolated from Taiwan. Microbial Drug Resistance. 2016, 22 (8), 646– 654, DOI: 10.1089/mdr.2015.0261There is no corresponding record for this reference.
- 59Panickar, A.; Manoharan, A.; Ramaiah, S. Machine learning-based virtual screening and density functional theory characterisation of natural inhibitors targeting mutant PBP2x in Streptococcus pneumoniae. Sci. Rep. 2025, 15 (1), 39164, DOI: 10.1038/s41598-025-24222-1There is no corresponding record for this reference.
- 60Fang, L.; Chen, R.; Li, C. The association between the genetic structures of commonly incompatible plasmids in Gram-negative bacteria, their distribution and the resistance genes. Front. Cell. Infect. Microbiol. 2024, 14, 1472876, DOI: 10.3389/fcimb.2024.1472876There is no corresponding record for this reference.
- 61Mazaheri Nezhad Fard, R.; Barton, M. D.; Heuzenroeder, M. W. Bacteriophage-mediated transduction of antibiotic resistance in enterococci: Transduction in Enterococcus spp. Lett. Appl. Microbiol. 2011, 52 (6), 559– 564, DOI: 10.1111/j.1472-765X.2011.03043.xThere is no corresponding record for this reference.
- 62Huang, M.; Liu, M.; Huang, L. The activation and limitation of the bacterial natural transformation system: The function in genome evolution and stability. Microbiol. Res. 2021, 252, 126856, DOI: 10.1016/j.micres.2021.126856There is no corresponding record for this reference.
- 63Soucy, S. M.; Huang, J.; Gogarten, J. P. Horizontal gene transfer: building the web of life. Nat. Rev. Genet. 2015, 16 (8), 472– 482, DOI: 10.1038/nrg3962There is no corresponding record for this reference.
- 64Arnold, B. J.; Huang, I. T.; Hanage, W. P. Horizontal gene transfer and adaptive evolution in bacteria. Nat. Rev. Microbiol. 2022, 20 (4), 206– 218, DOI: 10.1038/s41579-021-00650-4There is no corresponding record for this reference.
- 65McCallum, M., Burrows, L. L., Howell, P. L. The Dynamic Structures of the Type IV Pilus. Microbiol Spectr , 2019, 7 (2) DOI: 10.1128/microbiolspec.PSIB-0006-2018 .There is no corresponding record for this reference.
- 66Luna Rico, A.; Zheng, W.; Petiot, N.; Egelman, E. H.; Francetic, O. Functional reconstitution of the type IVa pilus assembly system from enterohaemorrhagic Escherichia coli. Mol. Microbiol. 2019, 111 (3), 732– 749, DOI: 10.1111/mmi.14188There is no corresponding record for this reference.
- 67Ellison, C. K.; Whitfield, G. B.; Brun, Y. V. Type IV Pili: dynamic bacterial nanomachines. FEMS Microbiol. Rev. 2022, 46 (2), fuab053, DOI: 10.1093/femsre/fuab053There is no corresponding record for this reference.
- 68Blokesch, M. Natural competence for transformation. Curr. Biol. 2016, 26 (21), R1126– R1130, DOI: 10.1016/j.cub.2016.08.058There is no corresponding record for this reference.
- 69Yu, Z.; Wang, Y.; Henderson, I. R.; Guo, J. Artificial sweeteners stimulate horizontal transfer of extracellular antibiotic resistance genes through natural transformation. ISME J. 2022, 16 (2), 543– 554, DOI: 10.1038/s41396-021-01095-6There is no corresponding record for this reference.
- 70Johnsborg, O.; Eldholm, V.; Håvarstein, L. S. Natural genetic transformation: prevalence, mechanisms and function. Res. Microbiol. 2007, 158 (10), 767– 778, DOI: 10.1016/j.resmic.2007.09.004There is no corresponding record for this reference.
- 71Johnston, C.; Martin, B.; Fichant, G.; Polard, P.; Claverys, J. P. Bacterial transformation: distribution, shared mechanisms and divergent control. Nat. Rev. Microbiol. 2014, 12 (3), 181– 196, DOI: 10.1038/nrmicro3199There is no corresponding record for this reference.
- 72Chen, J.; Quiles-Puchalt, N.; Chiang, Y. N. Genome hypermobility by lateral transduction. Science. 2018, 362 (6411), 207– 212, DOI: 10.1126/science.aat5867There is no corresponding record for this reference.
- 73Bhattacharya, T.; Chatterjee, S.; Maiti, D. Molecular analysis of the rstR and orfU genes of the CTX prophages integrated in the small chromosomes of environmental Vibrio cholerae non-O1, non-O139 strains. Environ. Microbiol. 2006, 8 (3), 526– 634, DOI: 10.1111/j.1462-2920.2005.00932.xThere is no corresponding record for this reference.
- 74Zhao, Y.; Ma, Y.; Vasileiou, C.; Farr, A. D.; Rogers, D. W.; Rainey, P. B. Jumbo phage-mediated transduction of genomic islands. Proc. Natl. Acad. Sci. U. S. A. 2025, 122 (44), e2512465122 DOI: 10.1073/pnas.2512465122There is no corresponding record for this reference.
- 75Waldor, M. K.; Friedman, D. I. Phage regulatory circuits and virulence gene expression. Curr. Opin. Microbiol. 2005, 8 (4), 459– 465, DOI: 10.1016/j.mib.2005.06.001There is no corresponding record for this reference.
- 76Chiang, Y. N.; Penadés, J. R.; Chen, J. Genetic transduction by phages and chromosomal islands: The new and noncanonical. Kline KA, ed. PLoS Pathog. 2019, 15 (8), e1007878 DOI: 10.1371/journal.ppat.1007878There is no corresponding record for this reference.
- 77Leclerc, Q. J.; Wildfire, J.; Gupta, A.; Lindsay, J. A.; Knight, G. M. Growth-Dependent Predation and Generalized Transduction of Antimicrobial Resistance by Bacteriophage. Gilbert JA, ed. mSystems 2022, 7 (2), e00135–22 DOI: 10.1128/msystems.00135-22There is no corresponding record for this reference.
- 78Waksman, G. From conjugation to T4S systems in Gram-negative bacteria: a mechanistic biology perspective. EMBO Rep. 2019, 20 (2), e47012 DOI: 10.15252/embr.201847012There is no corresponding record for this reference.
- 79Humphrey, S.; Fillol-Salom, A.; Quiles-Puchalt, N. Bacterial chromosomal mobility via lateral transduction exceeds that of classical mobile genetic elements. Nat. Commun. 2021, 12 (1), 6509, DOI: 10.1038/s41467-021-26004-5There is no corresponding record for this reference.
- 80Fillol-Salom, A.; Bacigalupe, R.; Humphrey, S.; Chiang, Y. N.; Chen, J.; Penadés, JR. Lateral transduction is inherent to the life cycle of the archetypical Salmonella phage P22. Nat. Commun. 2021, 12 (1), 6510, DOI: 10.1038/s41467-021-26520-4There is no corresponding record for this reference.
- 81Cabezón, E.; De La Cruz, F.; Arechaga, I. Conjugation Inhibitors and Their Potential Use to Prevent Dissemination of Antibiotic Resistance Genes in Bacteria. Front. Microbiol. 2017, 8, 2329, DOI: 10.3389/fmicb.2017.02329There is no corresponding record for this reference.
- 82Sher, A. A.; VanAllen, M. E.; Ahmed, H. Conjugative RP4 Plasmid-Mediated Transfer of Antibiotic Resistance Genes to Commensal and Multidrug-Resistant Enteric Bacteria In Vitro. Microorganisms 2023, 11 (1), 193, DOI: 10.3390/microorganisms11010193There is no corresponding record for this reference.
- 83Schröder, G.; Lanka, E. The mating pair formation system of conjugative plasmids–A versatile secretion machinery for transfer of proteins and DNA. Plasmid 2005, 54 (1), 1– 25, DOI: 10.1016/j.plasmid.2005.02.001There is no corresponding record for this reference.
- 84Bañuelos-Vazquez, L. A.; Torres Tejerizo, G.; Brom, S. Regulation of conjugative transfer of plasmids and integrative conjugative elements. Plasmid 2017, 91, 82– 89, DOI: 10.1016/j.plasmid.2017.04.002There is no corresponding record for this reference.
- 85Rutherford, S. T.; Bassler, B. L. Bacterial Quorum Sensing: Its Role in Virulence and Possibilities for Its Control. Cold Spring Harbor Perspect. Med. 2012, 2 (11), a012427, DOI: 10.1101/cshperspect.a012427There is no corresponding record for this reference.
- 86Chen, G.; Swem, L. R.; Swem, D. L. A Strategy for Antagonizing Quorum Sensing. Mol. Cell 2011, 42 (2), 199– 209, DOI: 10.1016/j.molcel.2011.04.003There is no corresponding record for this reference.
- 87Lu, Y.; Zeng, J.; Wu, B.; E, S.; Wang, L.; Cai, R.; Zhang, N.; Li, Y.; Huang, X.; Huang, B.; Quorum Sensing N-acyl Homoserine Lactones-SdiA Suppresses Escherichia coli-Pseudomonas aeruginosa Conjugation through Inhibiting traI Expression. Front. Cell. Infect. Microbiol. 2017, 7. DOI: 10.3389/fcimb.2017.00007 .There is no corresponding record for this reference.
- 88García-Aljaro, C.; Ballesté, E.; Muniesa, M. Beyond the canonical strategies of horizontal gene transfer in prokaryotes. Curr. Opin. Microbiol. 2017, 38, 95– 105, DOI: 10.1016/j.mib.2017.04.011There is no corresponding record for this reference.
- 89Vos, M.; Buckling, A.; Kuijper, B. Why do mobile genetic elements transfer DNA of their hosts?. Trends Genet. 2024, 40 (11), 927– 938, DOI: 10.1016/j.tig.2024.07.008There is no corresponding record for this reference.
- 90Bárdy, P.; Füzik, T.; Hrebík, D.; Pantůček, R.; Thomas Beatty, J.; Plevka, P. Structure and mechanism of DNA delivery of a gene transfer agent. Nat. Commun. 2020, 11 (1), 3034, DOI: 10.1038/s41467-020-16669-9There is no corresponding record for this reference.
- 91Savory, E. A.; Fuller, S. L.; Weisberg, A. J. Evolutionary transitions between beneficial and phytopathogenic Rhodococcus challenge disease management. eLife 2017, 6, e30925 DOI: 10.7554/eLife.30925There is no corresponding record for this reference.
- 92Rivard, N., Colwell, R. R., Burrus, V. Antibiotic Resistance in Vibrio cholerae: Mechanistic Insights from IncC Plasmid-Mediated Dissemination of a Novel Family of Genomic Islands Inserted at trmE. mSphere , 2020; 5(4) DOI: 10.1128/msphere.00748-20 . doi: 10.1128/msphere.00748-20.There is no corresponding record for this reference.
- 93Fogg, P. C. M. Identification and characterization of a direct activator of a gene transfer agent. Nat. Commun. 2019, 10 (1), 595, DOI: 10.1038/s41467-019-08526-1There is no corresponding record for this reference.
- 94Tran, N. T.; Le, T. B. K. Control of a gene transfer agent cluster in Caulobacter crescentus by transcriptional activation and anti-termination. Nat. Commun. 2024, 15 (1), 4749, DOI: 10.1038/s41467-024-49114-2There is no corresponding record for this reference.
- 95Devati, M. S.; Jnana, A.; Kidd, S. P. Decoding bacterial extracellular vesicles: A review on isolation and characterization techniques. Arch. Microbiol. 2026, 208 (1), 63, DOI: 10.1007/s00203-025-04628-1There is no corresponding record for this reference.
- 96Xu, Y.; Xie, C.; Liu, Y.; Qin, X.; Liu, J. An update on our understanding of Gram-positive bacterial membrane vesicles: discovery, functions, and applications. Front. Cell. Infect. Microbiol. 2023, 13, 1273813, DOI: 10.3389/fcimb.2023.1273813There is no corresponding record for this reference.
- 97Pérez-Cruz, C.; Delgado, L.; López-Iglesias, C.; Mercade, E. Outer-Inner Membrane Vesicles Naturally Secreted by Gram-Negative Pathogenic Bacteria. Rudel T, ed. PLoS One 2015, 10 (1), e0116896 DOI: 10.1371/journal.pone.0116896There is no corresponding record for this reference.
- 98Aktar, S.; Okamoto, Y.; Ueno, S. Incorporation of Plasmid DNA Into Bacterial Membrane Vesicles by Peptidoglycan Defects in Escherichia coli. Front. Microbiol. 2021, 12, 747606, DOI: 10.3389/fmicb.2021.747606There is no corresponding record for this reference.
- 99Zhao, X.; Wei, Y.; Bu, Y.; Ren, X.; Dong, Z. Review on bacterial outer membrane vesicles: structure, vesicle formation, separation and biotechnological applications. Microb. Cell Fact. 2025, 24 (1), 27, DOI: 10.1186/s12934-025-02653-9There is no corresponding record for this reference.
- 100Nagakubo, T.; Nomura, N.; Toyofuku, M. Cracking Open Bacterial Membrane Vesicles. Front. Microbiol. 2020, 10, 3026, DOI: 10.3389/fmicb.2019.03026There is no corresponding record for this reference.
- 101Toyofuku, M.; Schild, S.; Kaparakis-Liaskos, M.; Eberl, L. Composition and functions of bacterial membrane vesicles. Nat. Rev. Microbiol. 2023, 21 (7), 415– 430, DOI: 10.1038/s41579-023-00875-5There is no corresponding record for this reference.
- 102Juodeikis, R.; Carding, S. R. Outer Membrane Vesicles: Biogenesis, Functions, and Issues. Microbiol. Mol. Biol. Rev. 2022, 86 (4), e00032-22 DOI: 10.1128/mmbr.00032-22There is no corresponding record for this reference.
- 103Furuyama, N.; Sircili, M. P. Outer Membrane Vesicles (OMVs) Produced by Gram-Negative Bacteria: Structure, Functions, Biogenesis, and Vaccine Application. Gebre AK, ed. BioMed Res. Int. 2021, 2021 (1), 1490732, DOI: 10.1155/2021/1490732There is no corresponding record for this reference.
- 104Molina-Santiago, C.; Bernal, P. Nanotube-mediated plasmid transfer as a natural alternative for the improvement of industrially relevant bacteria. Microb. Biotechnol. 2023, 16 (4), 706– 708, DOI: 10.1111/1751-7915.14225There is no corresponding record for this reference.
- 105Dubey, G. P.; Malli Mohan, G. B.; Dubrovsky, A. Architecture and Characteristics of Bacterial Nanotubes. Dev. Cell 2016, 36 (4), 453– 461, DOI: 10.1016/j.devcel.2016.01.013There is no corresponding record for this reference.
- 106Dubey, G. P.; Ben-Yehuda, S. Intercellular Nanotubes Mediate Bacterial Communication. Cell. 2011, 144 (4), 590– 600, DOI: 10.1016/j.cell.2011.01.015There is no corresponding record for this reference.
- 107Baidya, A. K.; Rosenshine, I.; Ben-Yehuda, S. Donor-delivered cell wall hydrolases facilitate nanotube penetration into recipient bacteria. Nat. Commun. 2020, 11 (1), 1938, DOI: 10.1038/s41467-020-15605-1There is no corresponding record for this reference.
- 108Pospíšil, J.; Vítovská, D.; Kofroňová, O. Bacterial nanotubes as a manifestation of cell death. Nat. Commun. 2020, 11 (1), 4963, DOI: 10.1038/s41467-020-18800-2There is no corresponding record for this reference.
- 109Wang, C.; Zhao, R.; Yang, W. Cell-to-Cell Natural Transformation Mediated Efficient Plasmid Transfer Between Bacillus Species. IJMS. 2025, 26 (2), 621, DOI: 10.3390/ijms26020621There is no corresponding record for this reference.
- 110McInerney, J. O.; McNally, A.; O’Connell, M. J. Why prokaryotes have pangenomes. Nat. Microbiol. 2017, 2 (4), 17040, DOI: 10.1038/nmicrobiol.2017.40There is no corresponding record for this reference.
- 111Lee, I. P. A.; Eldakar, O. T.; Gogarten, J. P.; Andam, C. P. Bacterial cooperation through horizontal gene transfer. Trends Ecol. Evol. 2022, 37 (3), 223– 232, DOI: 10.1016/j.tree.2021.11.006There is no corresponding record for this reference.
- 112Husnik, F.; McCutcheon, J. P. Functional horizontal gene transfer from bacteria to eukaryotes. Nat. Rev. Microbiol. 2018, 16 (2), 67– 79, DOI: 10.1038/nrmicro.2017.137There is no corresponding record for this reference.
- 113Niero, G.; Bortolaia, V.; Vanni, M.; Intorre, L.; Guardabassi, L.; Piccirillo, A. High diversity of genes and plasmids encoding resistance to third-generation cephalosporins and quinolones in clinical Escherichia coli from commercial poultry flocks in Italy. Vet. Microbiol. 2018, 216, 93– 98, DOI: 10.1016/j.vetmic.2018.02.012There is no corresponding record for this reference.
- 114López, L.; Jumbo, M.; Mosquera, P.; Donoso, G.; Graham, J.; Trueba, G. Oral and parenteral treatment with a third-generation cephalosporin promotes the proliferation of diverse ESBL-producing Escherichia coli in the chicken intestinal tract. Rao K, ed. mSphere 2025, 10 (7), e00227–25 DOI: 10.1128/msphere.00227-25There is no corresponding record for this reference.
- 115Akhtar, A.; Fatima, N.; Khan, H. M. Beta-Lactamases and Their Classification: An Overview. Beta-Lactam Resistance in Gram-Negative Bacteria; Shahid, M., Singh, A., Sami, H., Eds.; Springer Nature Singapore, 2022; pp 25– 33. DOI: 10.1007/978-981-16-9097-6_3 .There is no corresponding record for this reference.
- 116Philippon, A.; Jacquier, H.; Ruppé, E.; Labia, R. Structure-based classification of class A beta-lactamases, an update. Curr. Res. Transl. Med. 2019, 67 (4), 115– 122, DOI: 10.1016/j.retram.2019.05.003There is no corresponding record for this reference.
- 117Page, M. I.; Badarau, A. The Mechanisms of Catalysis by Metallo β -Lactamases. Mugesh G, ed. Bioinorg. Chem. Appl. 2008, 2008 (1), 576297, DOI: 10.1155/2008/576297There is no corresponding record for this reference.
- 118Fernandes, R.; Amador, P.; Prudêncio, C. β-Lactams: chemical structure, mode of action and mechanisms of resistance. Rev. Med. Microbiol. 2013, 24 (1), 7– 17, DOI: 10.1097/MRM.0b013e3283587727There is no corresponding record for this reference.
- 119Carcione, D.; Siracusa, C.; Sulejmani, A.; Leoni, V.; Intra, J. Old and New Beta-Lactamase Inhibitors: Molecular Structure, Mechanism of Action, and Clinical Use. Antibiotics 2021, 10 (8), 995, DOI: 10.3390/antibiotics10080995There is no corresponding record for this reference.
- 120Chaves, J.; Ladona, M. G.; Segura, C.; Coira, A.; Reig, R.; Ampurdanés, C. SHV-1 β-Lactamase Is Mainly a Chromosomally Encoded Species-Specific Enzyme in Klebsiella pneumoniae. Antimicrob. Agents Chemother. 2001, 45 (10), 2856– 2861, DOI: 10.1128/AAC.45.10.2856-2861.2001There is no corresponding record for this reference.
- 121Acman, M.; Wang, R.; Van Dorp, L. Role of mobile genetic elements in the global dissemination of the carbapenem resistance gene blaNDM. Nat. Commun. 2022, 13 (1), 1131, DOI: 10.1038/s41467-022-28819-2There is no corresponding record for this reference.
- 122Roca, I.; Mosqueda, N.; Altun, B.; Espinal, P.; Akova, M.; Vila, J. Molecular characterization of NDM-1-producing Acinetobacter pittii isolated from Turkey in 2006. J. Antimicrob. Chemother. 2014, 69 (12), 3437– 3438, DOI: 10.1093/jac/dku306There is no corresponding record for this reference.
- 123Zhou, M.; Cai, Q.; Zhang, C.; Ouyang, P.; Yu, L.; Xu, Y. Antibiotic resistance bacteria and antibiotic resistance genes survived from the extremely acidity posing a risk on intestinal bacteria in an in vitro digestion model by horizontal gene transfer. Ecotoxicol. Environ. Saf. 2022, 247, 114247, DOI: 10.1016/j.ecoenv.2022.114247There is no corresponding record for this reference.
- 124Haverkate, M. R.; Dautzenberg, M. J. D.; Ossewaarde, T. J. M. Within-Host and Population Transmission of blaOXA-48 in K. pneumoniae and E. coli. Friedrich A, ed. PLoS One 2015, 10 (10), e0140960 DOI: 10.1371/journal.pone.0140960There is no corresponding record for this reference.
- 125Tofteland, S.; Naseer, U.; Lislevand, J. H.; Sundsfjord, A.; Samuelsen, Ø. A Long-Term Low-Frequency Hospital Outbreak of KPC-Producing Klebsiella pneumoniae Involving Intergenus Plasmid Diffusion and a Persisting Environmental Reservoir. Kluytmans J, ed. PLoS One 2013, 8 (3), e59015 DOI: 10.1371/journal.pone.0059015There is no corresponding record for this reference.
- 126Liu, Y.; Gao, J.; Zhao, M.; Fu, X.; Zhang, Y.; Zhang, H. Removal of antibiotic resistant bacteria, genes and inhibition of plasmid-mediated horizontal transfer by peroxymonosulfate: Efficiency and mechanisms. Chem. Eng. J. 2023, 453, 139728, DOI: 10.1016/j.cej.2022.139728There is no corresponding record for this reference.
- 127Jones, L. S.; Toleman, M. A.; Weeks, J. L.; Howe, R. A.; Walsh, T. R.; Kumarasamy, K. K. Plasmid Carriage of blaNDM-1 in Clinical Acinetobacter baumannii Isolates from India. Antimicrob. Agents Chemother. 2014, 58 (7), 4211– 4213, DOI: 10.1128/AAC.02500-14There is no corresponding record for this reference.
- 128Poirel, L.; Bonnin, R. A.; Nordmann, P. Analysis of the Resistome of a Multidrug-Resistant NDM-1-Producing Escherichia coli Strain by High-Throughput Genome Sequencing. Antimicrob. Agents Chemother. 2011, 55 (9), 4224– 4229, DOI: 10.1128/AAC.00165-11There is no corresponding record for this reference.
- 129Dellus-Gur, E.; Elias, M.; Caselli, E. Negative Epistasis and Evolvability in TEM-1 β-Lactamase–The Thin Line between an Enzyme’s Conformational Freedom and Disorder. J. Mol. Biol. 2015, 427 (14), 2396– 2409, DOI: 10.1016/j.jmb.2015.05.011There is no corresponding record for this reference.
- 130Prescott, J. F. Beta-lactam Antibiotics: Cephalosporins. Antimicrobial Therapy in Veterinary Medicine, 1 ed.; Giguère, S., Prescott, J. F., Dowling, P. M., Eds.; Wiley, 2013; pp 153– 173 DOI: 10.1002/9781118675014.ch9 .There is no corresponding record for this reference.
- 131Galleni, M.; Lamotte-Brasseur, J.; Raquet, X. The enigmatic catalytic mechanism of active-site serine β-lactamases. Biochem. Pharmacol. 1995, 49 (9), 1171– 1178, DOI: 10.1016/0006-2952(94)00502-DThere is no corresponding record for this reference.
- 132Dubus, A.; Wilkin, J. M.; Raquet, X.; Normark, S.; Frère, J. M. Catalytic mechanism of active-site serine β -lactamases: role of the conserved hydroxy group of the Lys-Thr(Ser)-Gly triad. Biochem. J. 1994, 301 (2), 485– 494, DOI: 10.1042/bj3010485There is no corresponding record for this reference.
- 133Lamotte-Brasseur, J.; Knox, J.; Kelly, J. A. The Structures and Catalytic Mechanisms of Active-Site Serine β-Lactamases. Biotechnol. Genet. Eng. Rev. 1994, 12 (1), 189– 230, DOI: 10.1080/02648725.1994.10647912There is no corresponding record for this reference.
- 134Tooke, C. L.; Hinchliffe, P.; Bragginton, E. C. β-Lactamases and β-Lactamase Inhibitors in the 21st Century. J. Mol. Biol. 2019, 431 (18), 3472– 3500, DOI: 10.1016/j.jmb.2019.04.002There is no corresponding record for this reference.
- 135Gniadkowski, M. Evolution of extended-spectrum β-lactamases by mutation. Clin. Microbiol. Infect. 2008, 14, 11– 32, DOI: 10.1111/j.1469-0691.2007.01854.xThere is no corresponding record for this reference.
- 136Castañeda-Barba, S.; Top, E. M.; Stalder, T. Plasmids, a molecular cornerstone of antimicrobial resistance in the One Health era. Nat. Rev. Microbiol. 2024, 22 (1), 18– 32, DOI: 10.1038/s41579-023-00926-xThere is no corresponding record for this reference.
- 137Razavi, M.; Kristiansson, E.; Flach, C. F.; Larsson, D. G. J. The Association between Insertion Sequences and Antibiotic Resistance Genes. mSphere 2020, 5 (5), e00418–20 DOI: 10.1128/mSphere.00418-20There is no corresponding record for this reference.
- 138Lipszyc, A.; Szuplewska, M.; Bartosik, D. How Do Transposable Elements Activate Expression of Transcriptionally Silent Antibiotic Resistance Genes?. IJMS. 2022, 23 (15), 8063, DOI: 10.3390/ijms23158063There is no corresponding record for this reference.
- 139Bhat, B. A.; Mir, R. A.; Qadri, H. Integrons in the development of antimicrobial resistance: critical review and perspectives. Front. Microbiol. 2023, 14, 1231938, DOI: 10.3389/fmicb.2023.1231938There is no corresponding record for this reference.
- 140Kotloff, K. L.; Riddle, M. S.; Platts-Mills, J. A.; Pavlinac, P.; Zaidi, A. K. M. Shigellosis. Lancet 2018, 391 (10122), 801– 812, DOI: 10.1016/S0140-6736(17)33296-8There is no corresponding record for this reference.
- 141Khalil, I.; Troeger, C. E.; Blacker, B. F.; Reiner, R. C. Capturing the true burden of Shigella and ETEC: The way forward. Vaccine 2019, 37 (34), 4784– 4786, DOI: 10.1016/j.vaccine.2019.01.031There is no corresponding record for this reference.
- 142Mason, L. C. E.; Greig, D. R.; Cowley, L. A. The evolution and international spread of extensively drug resistant Shigella sonnei. Nat. Commun. 2023, 14 (1), 1983, DOI: 10.1038/s41467-023-37672-wThere is no corresponding record for this reference.
- 143Davies, J. R.; Farrant, W. N.; Tomlinson, A. J. H. Further studies on the antibiotic resistance of Shigella sonnei: II. The acquisition of transferable antibiotic resistance in vivo. J. Hyg. 1968, 66 (3), 479– 487, DOI: 10.1017/S0022172400041346There is no corresponding record for this reference.
- 144Thanh Duy, P.; Thi Nguyen, T. N.; Vu Thuy, D. Commensal Escherichia coli are a reservoir for the transfer of XDR plasmids into epidemic fluoroquinolone-resistant Shigella sonnei. Nat. Microbiol. 2020, 5 (2), 256– 264, DOI: 10.1038/s41564-019-0645-9There is no corresponding record for this reference.
- 145Baker, K. S.; Dallman, T. J.; Field, N. Horizontal antimicrobial resistance transfer drives epidemics of multiple Shigella species. Nat. Commun. 2018, 9 (1), 1462, DOI: 10.1038/s41467-018-03949-8There is no corresponding record for this reference.
- 146Goodman, R. N.; Tansirichaiya, S.; Brouwer, M. S. M.; Roberts, A. P. Intracellular Transposition of Mobile Genetic Elements Associated with the Colistin Resistance Gene mcr-1. Microbiol Spectr. 2023, 11 (1), e03278–22 DOI: 10.1128/spectrum.03278-22There is no corresponding record for this reference.
- 147De La Cadena, E.; Mahecha, M.; Velandia, A. M. Identification of mcr-1 Genes and Characterization of Resistance Mechanisms to Colistin in Escherichia coli Isolates from Colombian Hospitals. Antibiotics 2023, 12 (3), 488, DOI: 10.3390/antibiotics12030488There is no corresponding record for this reference.
- 148Liu, Y. Y.; Wang, Y.; Walsh, T. R. Emergence of plasmid-mediated colistin resistance mechanism MCR-1 in animals and human beings in China: a microbiological and molecular biological study. Lancet Infect. Dis. 2016, 16 (2), 161– 168, DOI: 10.1016/S1473-3099(15)00424-7There is no corresponding record for this reference.
- 149Falgenhauer, L.; Waezsada, S. E.; Yao, Y. Colistin resistance gene mcr-1 in extended-spectrum β-lactamase-producing and carbapenemase-producing Gram-negative bacteria in Germany. Lancet Infect. Dis. 2016, 16 (3), 282– 283, DOI: 10.1016/S1473-3099(16)00009-8There is no corresponding record for this reference.
- 150Haenni, M.; Poirel, L.; Kieffer, N. Co-occurrence of extended spectrum β lactamase and MCR-1 encoding genes on plasmids. Lancet Infect. Dis. 2016, 16 (3), 281– 282, DOI: 10.1016/S1473-3099(16)00007-4There is no corresponding record for this reference.
- 151Zurfuh, K.; Poirel, L.; Nordmann, P.; Nüesch-Inderbinen, M.; Hächler, H.; Stephan, R. Occurrence of the Plasmid-Borne mcr-1 Colistin Resistance Gene in Extended-Spectrum-β-Lactamase-Producing Enterobacteriaceae in River Water and Imported Vegetable Samples in Switzerland. Antimicrob. Agents Chemother. 2016, 60 (4), 2594– 2595, DOI: 10.1128/AAC.00066-16There is no corresponding record for this reference.
- 152Du, H.; Chen, L.; Tang, Y. W.; Kreiswirth, B. N. Emergence of the mcr-1 colistin resistance gene in carbapenem-resistant Enterobacteriaceae. Lancet Infect. Dis. 2016, 16 (3), 287– 288, DOI: 10.1016/S1473-3099(16)00056-6There is no corresponding record for this reference.
- 153Nang, S. C.; Li, J.; Velkov, T. The rise and spread of mcr plasmid-mediated polymyxin resistance. Crit. Rev. Microbiol. 2019, 45 (2), 131– 161, DOI: 10.1080/1040841X.2018.1492902There is no corresponding record for this reference.
- 154Blair, J. M. A.; Webber, M. A.; Baylay, A. J.; Ogbolu, D. O.; Piddock, L. J. V. Molecular mechanisms of antibiotic resistance. Nat. Rev. Microbiol. 2015, 13 (1), 42– 51, DOI: 10.1038/nrmicro3380There is no corresponding record for this reference.
- 155Cox, G.; Wright, G. D. Intrinsic antibiotic resistance: Mechanisms, origins, challenges and solutions. Int. J. Med. Microbiol. 2013, 303 (6–7), 287– 292, DOI: 10.1016/j.ijmm.2013.02.009There is no corresponding record for this reference.
- 156Palzkill, T. Structural and Mechanistic Basis for Extended-Spectrum Drug-Resistance Mutations in Altering the Specificity of TEM, CTX-M, and KPC β-lactamases. Front. Mol. Biosci. 2018, 5, 16, DOI: 10.3389/fmolb.2018.00016There is no corresponding record for this reference.
- 157Zhang, H.; Seward, C. H.; Wu, Z.; Ye, H.; Feng, Y. Genomic insights into the ESBL and MCR-1-producing ST648 Escherichia coli with multi-drug resistance. Sci. Bull. 2016, 61 (11), 875– 878, DOI: 10.1007/s11434-016-1086-yThere is no corresponding record for this reference.
- 158Kluytmans-van Den Bergh, M. F.; Huizinga, P.; Bonten, M. J.; Presence of mcr-1-positive Enterobacteriaceae in retail chicken meat but not in humans in the Netherlands since 2009. Eurosurveillance 2016, 21(9). DOI: 10.2807/1560-7917.ES.2016.21.9.30149 .There is no corresponding record for this reference.
- 159Von Wintersdorff, C. J. H.; Penders, J.; Van Niekerk, J. M.; Mills, N. D.; Majumder, S.; van Alphen, L. B.; Savelkoul, P. H. M.; Wolffs, P. F. G. Dissemination of Antimicrobial Resistance in Microbial Ecosystems through Horizontal Gene Transfer. Front. Microbiol. 2016, 7. DOI: 10.3389/fmicb.2016.00173 .There is no corresponding record for this reference.
- 160Zhang, G.; Wang, C.; Sui, Z.; Feng, J. Insights into the evolutionary trajectories of fluoroquinolone resistance in Streptococcus pneumoniae. J. Antimicrob. Chemother. 2015, 70 (9), 2499– 2506, DOI: 10.1093/jac/dkv134There is no corresponding record for this reference.
- 161Cuypers, W. L.; Meysman, P.; Weill, F. X. A global genomic analysis of Salmonella Concord reveals lineages with high antimicrobial resistance in Ethiopia. Nat. Commun. 2023, 14 (1), 3517, DOI: 10.1038/s41467-023-38902-xThere is no corresponding record for this reference.
- 162Hassan, R.; Tantawy, M.; Gouda, N. A. Genotypic characterization of multiple drug resistant Escherichia coli isolates from a pediatric cancer hospital in Egypt. Sci. Rep. 2020, 10 (1), 4165, DOI: 10.1038/s41598-020-61159-zThere is no corresponding record for this reference.
- 163Thiolas, A.; Bornet, C.; Davin-Régli, A.; Pagès, J. M.; Bollet, C. Resistance to imipenem, cefepime, and cefpirome associated with mutation in Omp36 osmoporin of Enterobacter aerogenes. Biochem. Biophys. Res. Commun. 2004, 317 (3), 851– 856, DOI: 10.1016/j.bbrc.2004.03.130There is no corresponding record for this reference.
- 164Dé, E.; Baslé, A.; Jaquinod, M. A new mechanism of antibiotic resistance in Enterobacteriaceae induced by a structural modification of the major porin. Mol. Microbiol. 2001, 41 (1), 189– 198, DOI: 10.1046/j.1365-2958.2001.02501.xThere is no corresponding record for this reference.
- 165Sharma, A.; Gupta, V. K.; Pathania, R. Efflux pump inhibitors for bacterial pathogens: From bench to bedside. Indian J. Med. Res. 2019, 149 (2), 129– 145, DOI: 10.4103/ijmr.IJMR_2079_17There is no corresponding record for this reference.
- 166Wilson, D. N.; Hauryliuk, V.; Atkinson, G. C.; O’Neill, A. J. Target protection as a key antibiotic resistance mechanism. Nat. Rev. Microbiol. 2020, 18 (11), 637– 648, DOI: 10.1038/s41579-020-0386-zThere is no corresponding record for this reference.
- 167Stephan, J.; Mailaender, C.; Etienne, G.; Daffé, M.; Niederweis, M. Multidrug Resistance of a Porin Deletion Mutant of Mycobacterium smegmatis. Antimicrob. Agents Chemother. 2004, 48 (11), 4163– 4170, DOI: 10.1128/AAC.48.11.4163-4170.2004There is no corresponding record for this reference.
- 168Stahl, C.; Kubetzko, S.; Kaps, I.; Seeber, S.; Engelhardt, H.; Niederweis, M. MspA provides the main hydrophilic pathway through the cell wall of Mycobacterium smegmatis. Mol. Microbiol. 2001, 40 (2), 451– 464, DOI: 10.1046/j.1365-2958.2001.02394.xThere is no corresponding record for this reference.
- 169Masi, M.; Réfregiers, M.; Pos, K. M.; Pagès, J. M. Mechanisms of envelope permeability and antibiotic influx and efflux in Gram-negative bacteria. Nat. Microbiol. 2017, 2 (3), 17001, DOI: 10.1038/nmicrobiol.2017.1There is no corresponding record for this reference.
- 170Vergalli, J.; Bodrenko, I. V.; Masi, M. Porins and small-molecule translocation across the outer membrane of Gram-negative bacteria. Nat. Rev. Microbiol. 2020, 18 (3), 164– 176, DOI: 10.1038/s41579-019-0294-2There is no corresponding record for this reference.
- 171Chen, S.; Fu, J.; Zhao, K. Class 1 integron carrying qacEΔ1 gene confers resistance to disinfectant and antibiotics in Salmonella. Int. J. Food Microbiol. 2023, 404, 110319, DOI: 10.1016/j.ijfoodmicro.2023.110319There is no corresponding record for this reference.
- 172Ovung, A.; Bhattacharyya, J. Sulfonamide drugs: structure, antibacterial property, toxicity, and biophysical interactions. Biophys Rev. 2021, 13 (2), 259– 272, DOI: 10.1007/s12551-021-00795-9There is no corresponding record for this reference.
- 173Roberts, M. C. Update on macrolide-lincosamide-streptogramin, ketolide, and oxazolidinone resistance genes: MLSKO genes. FEMS Microbiol. Lett. 2008, 282 (2), 147– 159, DOI: 10.1111/j.1574-6968.2008.01145.xThere is no corresponding record for this reference.
- 174Chen, L.; Huang, J.; Huang, X. Horizontal Transfer of Different erm(B)-Carrying Mobile Elements Among Streptococcus suis Strains With Different Serotypes. Front. Microbiol. 2021, 12, 628740, DOI: 10.3389/fmicb.2021.628740There is no corresponding record for this reference.
- 175Sun, S. Emerging antibiotic resistance by various novel proteins/enzymes. Eur. J. Clin. Microbiol. Infect. Dis. 2025, 44 (7), 1551– 1566, DOI: 10.1007/s10096-025-05126-4There is no corresponding record for this reference.
- 176Unemo, M.; Lahra, M. M.; Cole, M. J. WHO global gonococcal antimicrobial surveillance programmes, 2019–22: a retrospective observational study. Lancet Microbe 2025, 6 (10), 101181, DOI: 10.1016/j.lanmic.2025.101181There is no corresponding record for this reference.
- 177Unemo, M.; Lahra, M. M.; Cole, M. World Health Organization Global Gonococcal Antimicrobial Surveillance Program (WHO GASP): review of new data and evidence to inform international collaborative actions and research efforts. Sex. Health 2019, 16 (5), 412– 425, DOI: 10.1071/SH19023There is no corresponding record for this reference.
- 178Mo, Y.; Tan, W. C.; Cooper, B. S. Antibiotic duration for common bacterial infections–a systematic review. JAC-Antimicrobial Resistance. 2024, 7 (1), dlae215, DOI: 10.1093/jacamr/dlae215There is no corresponding record for this reference.
- 179Mollah, F.; Khatun, M. M.; Chowdhury, R. Therapeutic Promises of Bioactive Linarin, a Glycosylated Flavonoid: A Comprehensive Review With Mechanistic Insight. Buch L, ed. J. Trop. Med. 2025, 2025 (1), 9989759, DOI: 10.1155/jotm/9989759There is no corresponding record for this reference.
- 180Vassilopoulos, S.; Mylonakis, E. Advances in methicillin-resistant staphylococcus aureus drug discovery: developments and challenges. Expert Opin. Drug Discovery 2026, 21, 231– 244, DOI: 10.1080/17460441.2026.2618787There is no corresponding record for this reference.
- 181Ghazaei, C. The role of bacteriophages and CRISPR-Cas in combating multidrug-resistant bacteria. Nat. Prod. Bioprospect. 2026, 16 (1), 14, DOI: 10.1007/s13659-025-00567-yThere is no corresponding record for this reference.
- 182Khan, M. F.; Javed, M.; Kaur, J.; Badwal, A. K.; Singh, S. CRISPR-Cas mediated targeting of resistance genes for combating ESKAPE pathogen infections: A Review. Int. J. Biol. Macromol. 2025, 334, 149180, DOI: 10.1016/j.ijbiomac.2025.149180There is no corresponding record for this reference.
- 183Antelo-Riveiro, P.; Garcia-Fandino, R.; Piñeiro, A. ́. Antimicrobial peptides at (lipid) interfaces: Insights from monolayer models. Adv. Colloid Interface Sci. 2026, 350, 103775, DOI: 10.1016/j.cis.2025.103775There is no corresponding record for this reference.
- 184Galhano, J.; Capelo-Martinez, J. L.; Lorenzo, J.; Lodeiro, C.; Oliveira, E. Advances in Antimicrobial Applications of Ag, Cu, and AgCu Nanoparticle-Doped Polymeric Composite Materials: A Comprehensive Review. ACS Nano 2025, 19 (35), 31301– 31330, DOI: 10.1021/acsnano.5c08822There is no corresponding record for this reference.
- 185Ravikumar, S.; Velappan, K.; Munusamy, S.; M, V.; S, N.; Shanmugam, R. Emerging nanotechnologies in wound care: The role of metal and polymeric nanocomposites in enhancing healing and combating infections. Int. J. Pharm. 2025, 684, 126143, DOI: 10.1016/j.ijpharm.2025.126143There is no corresponding record for this reference.
- 186Wijethunge, D.; Mathew, A.; Yarlagadda, P. K. D. V. Comprehensive review of bacterial death mechanism on nanopillared nanostructured surfaces. Biophys Rev. 2025, 17 (3), 893– 908, DOI: 10.1007/s12551-025-01319-5There is no corresponding record for this reference.
- 187Linklater, D. P.; Baulin, V. A.; Juodkazis, S.; Crawford, R. J.; Stoodley, P.; Ivanova, E. P. Mechano-bactericidal actions of nanostructured surfaces. Nat. Rev. Microbiol. 2021, 19 (1), 8– 22, DOI: 10.1038/s41579-020-0414-zThere is no corresponding record for this reference.
- 188Cao, H.; De La Fuente-Nunez, C. Microbial Primer: Artificial intelligence for microbiologists: This article is part of the < span style = ″color: rgb(33, 51, 104);″>Microbial Primers</span> collection.Š. Microbiology. 2025, 171(11). DOI: 10.1099/mic.0.001629 .There is no corresponding record for this reference.
- 189Adams, J. M. E.; El-Halfawy, O. M. Chemical-mediated alteration of antibiotic susceptibility: mechanisms and potential new targets for antibiotic adjuvant discovery. J. Bacteriol. 2026, e00351–25 DOI: 10.1128/jb.00351-25There is no corresponding record for this reference.
- 190Michaelis, C.; Grohmann, E. Horizontal Gene Transfer of Antibiotic Resistance Genes in Biofilms. Antibiotics 2023, 12 (2), 328, DOI: 10.3390/antibiotics12020328There is no corresponding record for this reference.
- 191Abe, K.; Nomura, N.; Suzuki, S. Biofilms: hot spots of horizontal gene transfer (HGT) in aquatic environments, with a focus on a new HGT mechanism. FEMS Microbiol. Ecol. 2020, 96 (5), fiaa031, DOI: 10.1093/femsec/fiaa031There is no corresponding record for this reference.
- 192Djermoun, S.; Rode, D. K. H.; Jiménez-Siebert, E. Biofilm architecture determines the dissemination of conjugative plasmids. Proc. Natl. Acad. Sci. U. S. A. 2025, 122 (17), e2417452122 DOI: 10.1073/pnas.2417452122There is no corresponding record for this reference.
- 193Tian, S.; Van Der Mei, H. C.; Ren, Y.; Busscher, H. J.; Shi, L. Recent advances and future challenges in the use of nanoparticles for the dispersal of infectious biofilms. J. Mater. Sci. Technol. 2021, 84, 208– 218, DOI: 10.1016/j.jmst.2021.02.007There is no corresponding record for this reference.
- 194Ma, L.; Konkel, M. E.; Lu, X. Antimicrobial Resistance Gene Transfer from Campylobacter jejuni in Mono- and Dual-Species Biofilms. Elkins CA, ed. Appl. Environ. Microbiol. 2021, 87 (15), e00659–21 DOI: 10.1128/AEM.00659-21There is no corresponding record for this reference.
- 195Lerminiaux, N. A.; Cameron, A. D. S. Horizontal transfer of antibiotic resistance genes in clinical environments. Can. J. Microbiol. 2019, 65 (1), 34– 44, DOI: 10.1139/cjm-2018-0275There is no corresponding record for this reference.
- 196Hutinel, M.; Fick, J.; Larsson, D. G. J.; Flach, C. F. Investigating the effects of municipal and hospital wastewaters on horizontal gene transfer. Environ. Pollut. 2021, 276, 116733, DOI: 10.1016/j.envpol.2021.116733There is no corresponding record for this reference.
- 197Sridhar, S.; Worby, C. J.; Bronson, R. A.; Turbett, S. E.; Oliver, E.; Shea, T.; Rao, S. R.; Sanchez, V.; Becker, M. V.; Holliday, L. K. Insights Into Global Antimicrobial Resistance Dynamics Through the Sequencing of Enteric Bacteria From US International Travelers. J. Infect. Dis. 2026, 233 (1), e164– e173, DOI: 10.1093/infdis/jiaf469There is no corresponding record for this reference.
- 198Sassi, A.; Basher, N. S.; Kirat, H. The Role of the Environment (Water, Air, Soil) in the Emergence and Dissemination of Antimicrobial Resistance: A One Health Perspective. Antibiotics 2025, 14 (8), 764, DOI: 10.3390/antibiotics14080764There is no corresponding record for this reference.
- 199Karim, M. A.; KianvashRad, N.; Cabo, M., Jr Cell Adhesion and Biofilm Development via Force-Sensitive Mechanisms: A Perspective. ACS Biomater. Sci. Eng. 2026, 12 (1), 3– 12, DOI: 10.1021/acsbiomaterials.5c01755There is no corresponding record for this reference.
- 200Liu, N.; Du, J.; Ge, J.; Liu, S. B. DNA damage-inducing endogenous and exogenous factors and research progress. Nucleosides Nucleotides Nucleic Acids 2025, 44 (12), 969– 1001, DOI: 10.1080/15257770.2024.2428436There is no corresponding record for this reference.
- 201Adamou, P.; Entwistle, J.; Graham, D. W.; Neumann, A. Mineral-Based Advanced Oxidation Processes for Enhancing the Removal of Antibiotic Resistance Genes from Domestic Wastewater. ACS EST Water. 2025, 5 (5), 2310– 2321, DOI: 10.1021/acsestwater.4c01213There is no corresponding record for this reference.
- 202Feng, M.; Liu, Y.; Yang, L.; Li, Z. Antibiotics and antibiotic resistance gene dynamics in the composting of antibiotic fermentation waste - A review. Bioresour. Technol. 2023, 390, 129861, DOI: 10.1016/j.biortech.2023.129861There is no corresponding record for this reference.
- 203Liao, H.; Bai, Y.; Liu, C. Airborne and indigenous microbiomes co-drive the rebound of antibiotic resistome during compost storage. Environ. Microbiol. 2021, 23 (12), 7483– 7496, DOI: 10.1111/1462-2920.15672There is no corresponding record for this reference.
- 204Xia, R.; Zhang, L.; Li, G.; Luo, W.; Xu, Z. A small technology for big health: Blocking the potential spread of antibiotic resistomes from home composting of food waste by mature compost. Waste Manage. 2026, 211, 115312, DOI: 10.1016/j.wasman.2025.115312There is no corresponding record for this reference.
- 205Yin, D.; Wang, K.; Wu, C. Refluxing mature compost to replace bulking agents: A low-cost solution for suppressing antibiotic resistance genes rebound in sewage sludge composting. Environ. Res. 2025, 269, 120811, DOI: 10.1016/j.envres.2025.120811There is no corresponding record for this reference.
- 206Yoon, Y.; Chung, H. J.; Wen Di, D. Y.; Dodd, M. C.; Hur, H. G.; Lee, Y. Inactivation efficiency of plasmid-encoded antibiotic resistance genes during water treatment with chlorine, UV, and UV/H2O2. Water Res. 2017, 123, 783– 793, DOI: 10.1016/j.watres.2017.06.056There is no corresponding record for this reference.
- 207He, H.; Choi, Y.; Wu, S. J. Application of Nucleotide-Based Kinetic Modeling Approaches to Predict Antibiotic Resistance Gene Degradation during UV- and Chlorine-Based Wastewater Disinfection Processes: From Bench- to Full-Scale. Environ. Sci. Technol. 2022, 56 (21), 15141– 15155, DOI: 10.1021/acs.est.2c00567There is no corresponding record for this reference.
- 208Yao, M. C.; Zhang, X.; Huang, Q.; Huang, J.; Sheng, G. P. Chlorine oxide radical (ClO) enables the enhanced degradation of antibiotic resistance genes during UV/chlorine treatment by selectively inducing base damage. Environ. Int. 2023, 178, 108121, DOI: 10.1016/j.envint.2023.108121There is no corresponding record for this reference.
- 209Guerra-Rodríguez, S.; Abeledo-Lameiro, M. J.; Polo-López, M. I. Pilot-scale sulfate radical-based advanced oxidation for wastewater reuse: simultaneous disinfection, removal of contaminants of emerging concern, and antibiotic resistance genes. Chem. Eng. J. 2023, 477, 146916, DOI: 10.1016/j.cej.2023.146916There is no corresponding record for this reference.
- 210Liang, C.; Wei, D.; Zhang, S.; Ren, Q.; Shi, J.; Liu, L. Removal of antibiotic resistance genes from swine wastewater by membrane filtration treatment. Ecotoxicol. Environ. Saf. 2021, 210, 111885, DOI: 10.1016/j.ecoenv.2020.111885There is no corresponding record for this reference.
- 211Li, R.; Wei, Z.; Li, P. Novel visible-light activated photocatalytic ultrafiltration membrane for simultaneous separation and degradation of emerging contaminants. J. Hazard. Mater. 2024, 478, 135634, DOI: 10.1016/j.jhazmat.2024.135634There is no corresponding record for this reference.
- 212Sun, L.; Shi, P.; He, N.; Zhang, Q.; Duan, X. Antibiotic resistance genes removal and membrane fouling in secondary effluents by combined processes of PAC/BPAC-UF. J. Water Health 2019, 17 (6), 910– 920, DOI: 10.2166/wh.2019.160There is no corresponding record for this reference.
- 213Berruti, I.; Nahim-Granados, S.; Abeledo-Lameiro, M. J.; Oller, I.; Polo-López, M. I. Peroxymonosulfate/Solar process for urban wastewater purification at a pilot plant scale: A techno-economic assessment. Sci. Total Environ. 2023, 881, 163407, DOI: 10.1016/j.scitotenv.2023.163407There is no corresponding record for this reference.
- 214Rutten, S. B.; Ojobe, B. L.; Hernández Leal, L. Evaluation of Membrane Integrity Monitoring Methods for Hollow Fiber Nanofiltration Membranes: Applicability in Gray Water Reclamation Systems. ACS EST Water. 2023, 3 (12), 3884– 3892, DOI: 10.1021/acsestwater.3c00307There is no corresponding record for this reference.
- 215Slipko, K.; Reif, D.; Wögerbauer, M.; Hufnagl, P.; Krampe, J.; Kreuzinger, N. Removal of extracellular free DNA and antibiotic resistance genes from water and wastewater by membranes ranging from microfiltration to reverse osmosis. Water Res. 2019, 164, 114916, DOI: 10.1016/j.watres.2019.114916There is no corresponding record for this reference.
- 216Li, M.; Wang, P.; Zhang, K. Single cobalt atoms anchored on Ti3 C2 Tx with dual reaction sites for efficient adsorption-degradation of antibiotic resistance genes. Proc. Natl. Acad. Sci. U. S. A. 2023, 120 (29), e2305705120 DOI: 10.1073/pnas.2305705120There is no corresponding record for this reference.
- 217Liu, Y.; Gao, J.; Wang, Y. The removal of antibiotic resistant bacteria and genes and inhibition of the horizontal gene transfer by contrastive research on sulfidated nanoscale zerovalent iron activating peroxymonosulfate or peroxydisulfate. J. Hazard. Mater. 2022, 423, 126866, DOI: 10.1016/j.jhazmat.2021.126866There is no corresponding record for this reference.
- 218Liu, Y.; Cai, Y.; Li, G.; Wang, W.; Wong, P. K.; An, T. Response mechanisms of different antibiotic-resistant bacteria with different resistance action targets to the stress from photocatalytic oxidation. Water Res. 2022, 218, 118407, DOI: 10.1016/j.watres.2022.118407There is no corresponding record for this reference.
- 219Poyatos, J. M.; Muñio, M. M.; Almecija, M. C.; Torres, J. C.; Hontoria, E.; Osorio, F. Advanced Oxidation Processes for Wastewater Treatment: State of the Art. Water, Air, Soil Pollut. 2010, 205 (1–4), 187– 204, DOI: 10.1007/s11270-009-0065-1There is no corresponding record for this reference.
- 220Li, F.; Liu, K.; Bao, Y. Molecular level removal of antibiotic resistant bacteria and genes: A review of interfacial chemical in advanced oxidation processes. Water Res. 2024, 254, 121373, DOI: 10.1016/j.watres.2024.121373There is no corresponding record for this reference.
- 221Manoharan, R. K.; Ishaque, F.; Ahn, Y. H. Fate of antibiotic resistant genes in wastewater environments and treatment strategies - A review. Chemosphere 2022, 298, 134671, DOI: 10.1016/j.chemosphere.2022.134671There is no corresponding record for this reference.
- 222Stefaniak, K.; Harnisz, M.; Męcik, M.; Korzeniewska, E. ARB inactivation, ARGs and antibiotics degradation in hospital wastewater. J. Hazard. Mater. 2025, 495, 138833, DOI: 10.1016/j.jhazmat.2025.138833There is no corresponding record for this reference.
- 223Emamalipour, M.; Seidi, K.; Zununi Vahed, S. Horizontal Gene Transfer: From Evolutionary Flexibility to Disease Progression. Front Cell Dev Biol. 2020, 8, 229, DOI: 10.3389/fcell.2020.00229There is no corresponding record for this reference.
- 224Brito, I. L. Examining horizontal gene transfer in microbial communities. Nat. Rev. Microbiol. 2021, 19 (7), 442– 453, DOI: 10.1038/s41579-021-00534-7There is no corresponding record for this reference.
- 225Abdulhussien, A. A.; Farhan, Z. M.; Yaseen, L. J.; Mohsein, O. A. The Clinico-Pathological Role of Horizontal Gene Transfer in the Ability of Pseudomonas Aeruginosa to Cause Disease in Lung Tissue. ejmhr. 2025, 3 (1), 27– 34, DOI: 10.59324/ejmhr.2025.3(1).06There is no corresponding record for this reference.
- 226Evans, D. R.; Griffith, M. P.; Sundermann, A. J. Systematic detection of horizontal gene transfer across genera among multidrug-resistant bacteria in a single hospital. eLife 2020, 9, e53886 DOI: 10.7554/eLife.53886There is no corresponding record for this reference.
- 227McCallum, G. E.; Hall, J. P. J. The hospital sink drain microbiome as a melting pot for AMR transmission to nosocomial pathogens. npj Antimicrob Resist. 2025, 3 (1), 68, DOI: 10.1038/s44259-025-00137-9There is no corresponding record for this reference.
- 228McInnes, R. S.; McCallum, G. E.; Lamberte, L. E.; Van Schaik, W. Horizontal transfer of antibiotic resistance genes in the human gut microbiome. Curr. Opin. Microbiol. 2020, 53, 35– 43, DOI: 10.1016/j.mib.2020.02.002There is no corresponding record for this reference.
- 229Woods, L. C.; Gorrell, R. J.; Taylor, F.; Connallon, T.; Kwok, T.; McDonald, M. J. Horizontal gene transfer potentiates adaptation by reducing selective constraints on the spread of genetic variation. Proc. Natl. Acad. Sci. U. S. A. 2020, 117 (43), 26868– 26875, DOI: 10.1073/pnas.2005331117There is no corresponding record for this reference.
- 230Wang, J.; Huo, L.; Bian, K. Efficacy and Mechanism of Antibiotic Resistance Gene Degradation and Cell Membrane Damage during Ultraviolet Advanced Oxidation Processes. ACS EST Water. 2024, 4 (6), 2746– 2755, DOI: 10.1021/acsestwater.4c00350There is no corresponding record for this reference.
- 231Liu, Y.; Hull, N. M. Emerging investigator series: Inactivation of antibiotic resistant bacteria and inhibition of horizontal resistance gene transfer is more effective by 222 than 254 nm UV. Environ. Sci: Water Res. Technol. 2025, 11 (2), 306– 316, DOI: 10.1039/D4EW00530AThere is no corresponding record for this reference.
- 232Han, J.; Li, W.; Yang, Y. UV-Based Advanced Oxidation Processes for Antibiotic Resistance Control: Efficiency, Influencing Factors, and Energy Consumption. Engineering. 2024, 37, 27– 39, DOI: 10.1016/j.eng.2023.09.021There is no corresponding record for this reference.
- 233Zheng, T. H.; Zhang, Z. Z.; Liu, Y.; Zou, L. H. Recent Progress in Catalytically Driven Advanced Oxidation Processes for Wastewater Treatment. Catalysts 2025, 15 (8), 761, DOI: 10.3390/catal15080761There is no corresponding record for this reference.
- 234Satyam, S.; Patra, S. The Evolving Landscape of Advanced Oxidation Processes in Wastewater Treatment: Challenges and Recent Innovations. Processes 2025, 13 (4), 987, DOI: 10.3390/pr13040987There is no corresponding record for this reference.
- 235Ahmed, B.; Ameen, F.; Rizvi, A. Destruction of Cell Topography, Morphology, Membrane, Inhibition of Respiration, Biofilm Formation, and Bioactive Molecule Production by Nanoparticles of Ag, ZnO, CuO, TiO2, and Al2 O3 toward Beneficial Soil Bacteria. ACS Omega 2020, 5 (14), 7861– 7876, DOI: 10.1021/acsomega.9b04084There is no corresponding record for this reference.
- 236Moreira, N. F. F.; Narciso-da-Rocha, C.; Polo-López, M. I. Solar treatment (H2O2, TiO2-P25 and GO-TiO2 photocatalysis, photo-Fenton) of organic micropollutants, human pathogen indicators, antibiotic resistant bacteria and related genes in urban wastewater. Water Res. 2018, 135, 195– 206, DOI: 10.1016/j.watres.2018.01.064There is no corresponding record for this reference.
- 237Fu, X.; Gao, J.; Wang, Q. Mechanisms on the removal of gram-negative/positive antibiotic resistant bacteria and inhibition of horizontal gene transfer by ferrate coupled with peroxydisulfate or peroxymonosulfate. J. Hazard. Mater. 2024, 470, 134254, DOI: 10.1016/j.jhazmat.2024.134254There is no corresponding record for this reference.
- 238Mahmoodi, M.; Pishbin, E. Ozone-based advanced oxidation processes in water treatment: recent advances, challenges, and perspective. Environ. Sci. Pollut. Res. 2025, 32 (7), 3531– 3570, DOI: 10.1007/s11356-024-35835-wThere is no corresponding record for this reference.
- 239Macedo, G.; Olesen, A. K.; Maccario, L. Horizontal Gene Transfer of an IncP1 Plasmid to Soil Bacterial Community Introduced by Escherichia coli through Manure Amendment in Soil Microcosms. Environ. Sci. Technol. 2022, 56 (16), 11398– 11408, DOI: 10.1021/acs.est.2c02686There is no corresponding record for this reference.
- 240Mei, Z.; He, C.; Balcazar, J. L. Antibiotic-degrading bacteria shape resistome dynamics and horizontal gene transfer potential in soils with contrasting properties. ISME Communications. 2026, 6 (1), ycaf246, DOI: 10.1093/ismeco/ycaf246There is no corresponding record for this reference.
- 241Shrestha, P.; Bajracharya, B.; Shrestha, D.; Karna, A. K.; Shah, P.; Gurung, K.; Ghimire, S.; Shrestha, A. Molecular detection of antibiotic resistance genes in Staphylococcus aureus isolated from poultry farms of Kathmandu Valley, Nepal. BMC Microbiol. 2026, 26, 291, DOI: 10.1186/s12866-026-04713-5There is no corresponding record for this reference.
- 242Lima, T.; Domingues, S.; Da Silva, G. J. Manure as a Potential Hotspot for Antibiotic Resistance Dissemination by Horizontal Gene Transfer Events. Vet. Sci. 2020, 7 (3), 110, DOI: 10.3390/vetsci7030110There is no corresponding record for this reference.
- 243Goh, Y. X.; Anupoju, S. M. B.; Nguyen, A. Evidence of horizontal gene transfer and environmental selection impacting antibiotic resistance evolution in soil-dwelling Listeria. Nat. Commun. 2024, 15 (1), 10034, DOI: 10.1038/s41467-024-54459-9There is no corresponding record for this reference.
- 244Rocha, K. O.; Brandão, F.; Mendes, C. á.; Carvalho, M. G.; Mazierski, P.; Zaleska-Medynska, A.; Gomes, J.; Martins, R. C.; Domingues, E. Olive mill waste bio-based catalyst application in advanced oxidation processes for wastewater treatment. Catal. Today 2024, 432, 114618, DOI: 10.1016/j.cattod.2024.114618There is no corresponding record for this reference.
- 245Martins, R. B.; Jorge, N.; Lucas, M. S.; Raymundo, A.; Barros, A. I. R. N. A.; Peres, J. A. Food By-Product Valorization by Using Plant-Based Coagulants Combined with AOPs for Agro-Industrial Wastewater Treatment. IJERPH. 2022, 19 (7), 4134, DOI: 10.3390/ijerph19074134There is no corresponding record for this reference.
- 246Feng, Y.; Xie, T.; Li, F. New challenge: Mitigation and control of antibiotic resistant genes in aquatic environments by biochar. Sci. Total Environ. 2024, 946, 174385, DOI: 10.1016/j.scitotenv.2024.174385There is no corresponding record for this reference.
- 247Fu, Y.; Wang, F.; Sheng, H. Removal of extracellular antibiotic resistance genes using magnetic biochar/quaternary phosphonium salt in aquatic environments: A mechanistic study. J. Hazard. Mater. 2021, 411, 125048, DOI: 10.1016/j.jhazmat.2021.125048There is no corresponding record for this reference.
- 248Wu, C.; Fu, L.; Li, H.; Liu, X.; Wan, C. Using biochar to strengthen the removal of antibiotic resistance genes: Performance and mechanism. Sci. Total Environ. 2022, 816, 151554, DOI: 10.1016/j.scitotenv.2021.151554There is no corresponding record for this reference.
- 249Wijitkosum, S. Biochar derived from agricultural wastes and wood residues for sustainable agricultural and environmental applications. Int. Soil Water Conserv. Res. 2022, 10 (2), 335– 341, DOI: 10.1016/j.iswcr.2021.09.006There is no corresponding record for this reference.
- 250Sun, X.; Shi, L.; Zhang, H.; Li, F.; Long, Y.; Zhang, D. Chain-length-dependent adsorption of extracellular DNA on biochar: Behaviors, mechanisms, and structural Stability. Water Res.:X 2026, 30, 100496, DOI: 10.1016/j.wroa.2026.100496There is no corresponding record for this reference.
- 251Sun, X.; Shi, L.; He, Z.; Zhang, H.; Li, F.; Zhang, D. Divalent metal ions facilitate environmental DNA adsorption on biochar by inducing new hydrogen bonds. Int. J. Biol. Macromol. 2025, 319, 145619, DOI: 10.1016/j.ijbiomac.2025.145619There is no corresponding record for this reference.
- 252Wang, C.; Wang, Y.; Yan, S. Biochar-amended composting of lincomycin fermentation dregs promoted microbial metabolism and reduced antibiotic resistance genes. Bioresour. Technol. 2023, 367, 128253, DOI: 10.1016/j.biortech.2022.128253There is no corresponding record for this reference.
- 253Kui, H.; Jingyang, C.; Mengxin, G.; Hui, X.; Li, L. Effects of biochars on the fate of antibiotics and their resistance genes during vermicomposting of dewatered sludge. J. Hazard. Mater. 2020, 397, 122767, DOI: 10.1016/j.jhazmat.2020.122767There is no corresponding record for this reference.
- 254Li, J.; Lu, H.; Yang, H.; Wen, X.; Huang, Y.; Li, Q. Performances of antibiotic resistance genes profile upon the action of biochar-activated peroxydisulfate in composting. J. Environ. Manage. 2023, 334, 117509, DOI: 10.1016/j.jenvman.2023.117509There is no corresponding record for this reference.
- 255Shi, L.; Zhang, D.; Yang, M. New discovery of extremely high adsorption of environmental DNA on cuttlefish bone pyrolysis derivative via large pore structure and carbon film. Waste Manage. 2024, 175, 286– 293, DOI: 10.1016/j.wasman.2024.01.016There is no corresponding record for this reference.
- 256Saraei, F.; Amini, K.; Goodarzi, R.; Khanifar, A.; Asghari, B. A magnetically recoverable Chitosan-Based nanocomposite for simultaneous removal of tetracycline and antibiotic resistance genes from wastewater. Sci. Rep. 2026, 16 (1), 817, DOI: 10.1038/s41598-025-30583-4There is no corresponding record for this reference.
- 257Xie, T.; Lin, D.; Cai, X. D. Nano-biochar regulates phage-host interactions, reducing antibiotic resistance genes in vermicomposting systems. Proc. Natl. Acad. Sci. U. S. A. 2025, 122 (34), e2511986122 DOI: 10.1073/pnas.2511986122There is no corresponding record for this reference.
- 258Majaneva, M.; Diserud, O. H.; Eagle, S. H. C.; Boström, E.; Hajibabaei, M.; Ekrem, T. Environmental DNA filtration techniques affect recovered biodiversity. Sci. Rep. 2018, 8 (1), 4682, DOI: 10.1038/s41598-018-23052-8There is no corresponding record for this reference.
- 259Schefer, R. B.; Paolucci, C.; Mitrano, D. M. Adsorption and Protection of Environmental DNA (eDNA) on Polymer and Silica Surfaces. Environ. Sci. Technol. 2025, 59 (36), 19479– 19489, DOI: 10.1021/acs.est.5c05439There is no corresponding record for this reference.
- 260Boom, R.; Sol, C. J.; Salimans, M. M.; Jansen, C. L.; Wertheim-van Dillen, P. M.; Van Der Noordaa, J. Rapid and simple method for purification of nucleic acids. J. Clin. Microbiol. 1990, 28 (3), 495– 503, DOI: 10.1128/jcm.28.3.495-503.1990There is no corresponding record for this reference.
- 261Lorenz, M. G.; Wackernagel, W. Adsorption of DNA to sand and variable degradation rates of adsorbed DNA. Appl. Environ. Microbiol. 1987, 53 (12), 2948– 2952, DOI: 10.1128/aem.53.12.2948-2952.1987There is no corresponding record for this reference.
- 262Bag, S.; Rauwolf, S.; Schwaminger, S. P.; Wenzel, W.; Berensmeier, S. DNA Binding to the Silica: Cooperative Adsorption in Action. Langmuir 2021, 37 (19), 5902– 5908, DOI: 10.1021/acs.langmuir.1c00381There is no corresponding record for this reference.
- 263Adegoke, S. C.; Yawlui, I. S. Y.; LaJeunesse, D. Silica Nanoparticles Block Natural Genetic Transformation in Acinetobacter baylyi ADP1. ACS Omega 2025, 10, 62609– 62620, DOI: 10.1021/acsomega.5c06566There is no corresponding record for this reference.
- 264Cai, P.; Huang, Q.; Zhang, X.; Chen, H. Adsorption of DNA on clay minerals and various colloidal particles from an Alfisol. Soil Biol. Biochem. 2006, 38 (3), 471– 476, DOI: 10.1016/j.soilbio.2005.05.019There is no corresponding record for this reference.
- 265Khanna, M.; Stotzky, G. Transformation of Bacillus subtilis by DNA bound on montmorillonite and effect of DNase on the transforming ability of bound DNA. Appl. Environ. Microbiol. 1992, 58 (6), 1930– 1939, DOI: 10.1128/aem.58.6.1930-1939.1992There is no corresponding record for this reference.
- 266Buckner, M. M. C.; Ciusa, M. L.; Piddock, L. J. V. Strategies to combat antimicrobial resistance: anti-plasmid and plasmid curing. FEMS Microbiol. Rev. 2018, 42 (6), 781– 804, DOI: 10.1093/femsre/fuy031There is no corresponding record for this reference.
- 267Zulauf, K. E.; Kirby, J. E. Discovery of small-molecule inhibitors of multidrug-resistance plasmid maintenance using a high-throughput screening approach. Proc. Natl. Acad. Sci. U. S. A. 2020, 117 (47), 29839– 29850, DOI: 10.1073/pnas.2005948117There is no corresponding record for this reference.
- 268Beg, A. Z.; Ahmad, I. Effect of Plumbago zeylanica extract and certain curing agents on multidrug resistant bacteria of clinical origin. World J. Microbiol. Biotechnol. 2000, 16 (8–9), 841– 844, DOI: 10.1023/A:1008991724288There is no corresponding record for this reference.
- 269Bharathi, A.; Polasa, H. Elimination of broad-host range plasmid vectors in Escherichia coli by curring agents. FEMS Microbiol. Lett. 1991, 84 (1), 37– 40, DOI: 10.1111/j.1574-6968.1991.tb04565.xThere is no corresponding record for this reference.
- 270Gómez-Rubio, E.; Arana, L.; Vicario-Martín, R. Exploring Inhibition of Bacterial Conjugation Coupling Protein TrwB: Novel Ligands to Fight Antimicrobial Resistance Spread. ACS Omega 2025, 10 (31), 34645– 34658, DOI: 10.1021/acsomega.5c03425There is no corresponding record for this reference.
- 271Fernandez-Lopez, R.; Machón, C.; Longshaw, C. M. Unsaturated fatty acids are inhibitors of bacterial conjugation. Microbiology. 2005, 151 (11), 3517– 3526, DOI: 10.1099/mic.0.28216-0There is no corresponding record for this reference.
- 272Álvarez-Rodríguez, I.; Arana, L.; Ugarte-Uribe, B. Type IV Coupling Proteins as Potential Targets to Control the Dissemination of Antibiotic Resistance. Front. Mol. Biosci. 2020, 7, 201, DOI: 10.3389/fmolb.2020.00201There is no corresponding record for this reference.
- 273Wei, F.; Cui, X.; Wang, Z.; Dong, C.; Li, J.; Han, X. Recoverable peroxidase-like Fe3O4@MoS2-Ag nanozyme with enhanced antibacterial ability. Chem. Eng. J. 2021, 408, 127240, DOI: 10.1016/j.cej.2020.127240There is no corresponding record for this reference.
- 274Doolan, J. A.; Williams, G. T.; Hilton, K. L. F. Advancements in antimicrobial nanoscale materials and self-assembling systems. Chem. Soc. Rev. 2022, 51 (20), 8696– 8755, DOI: 10.1039/D1CS00915JThere is no corresponding record for this reference.
- 275Bharathan, S.; Sundaramoorthy, N. S.; Chandrasekaran, H. Sub lethal levels of platinum nanoparticle cures plasmid and in combination with carbapenem, curtails carbapenem resistant Escherichia coli. Sci. Rep. 2019, 9 (1), 5305, DOI: 10.1038/s41598-019-41489-3There is no corresponding record for this reference.
- 276Markowicz, A.; Borymski, S.; Adamek, A.; Sułowicz, S. The influence of ZnO nanoparticles on horizontal transfer of resistance genes in lab and soil conditions. Environ. Res. 2023, 223, 115420, DOI: 10.1016/j.envres.2023.115420There is no corresponding record for this reference.
- 277Otinov, G. D.; Lokteva, A. V.; Petrova, A. D. Positive and Negative Effects of Metal Oxide Nanoparticles on Antibiotic Resistance Genes Transfer. Antibiotics 2020, 9 (11), 742, DOI: 10.3390/antibiotics9110742There is no corresponding record for this reference.
- 278Okkeh, M.; Bloise, N.; Restivo, E.; De Vita, L.; Pallavicini, P.; Visai, L. Gold Nanoparticles: Can They Be the Next Magic Bullet for Multidrug-Resistant Bacteria?. Nanomaterials 2021, 11 (2), 312, DOI: 10.3390/nano11020312There is no corresponding record for this reference.
- 279Wang, H.; Qi, H.; Gong, S. Fe3O4 composited with MoS2 blocks horizontal gene transfer. Colloids Surf., B 2020, 185, 110569, DOI: 10.1016/j.colsurfb.2019.110569There is no corresponding record for this reference.
- 280Chen, X.; Zhan, M.; Shao, H. Mechanism of reducing antibiotic resistance genes by nano-selenium during composting: insight into host microorganisms and a two-component system. J. Environ. Manage. 2025, 386, 125704, DOI: 10.1016/j.jenvman.2025.125704There is no corresponding record for this reference.
- 281Das, A.; Biswas, R.; Najiar, R.; Thatharaju, P.; Biswas, S.; Mohan, U. Quorum sensing in bacteria: insights into communication and inhibition strategies─a review. Arch. Microbiol. 2026, 208 (4), 157, DOI: 10.1007/s00203-025-04610-xThere is no corresponding record for this reference.
- 282Zhu, L.; Lau, G. W. Inhibition of Competence Development, Horizontal Gene Transfer and Virulence in Streptococcus pneumoniae by a Modified Competence Stimulating Peptide. Bessen DE, ed. PLoS Pathog. 2011, 7 (9), e1002241 DOI: 10.1371/journal.ppat.1002241There is no corresponding record for this reference.
- 283Malshikare, H.; Priyakumar, U. D.; Chatterjee, P.; Sengupta, D. Mechanistic principles of antimicrobial peptides uncovered by charge density-based machine learning. Chem. Commun. 2026, 62, 4067, DOI: 10.1039/D5CC06374DThere is no corresponding record for this reference.
- 284Domenech, A.; Brochado, A. R.; Sender, V. Proton Motive Force Disruptors Block Bacterial Competence and Horizontal Gene Transfer. Cell Host Microbe 2020, 27 (4), 544– 555, DOI: 10.1016/j.chom.2020.02.002There is no corresponding record for this reference.
- 285Jia, Y.; Zheng, Z.; Yang, B.; Zhang, H.; Wang, Z.; Liu, Y. A Broad-Spectrum Horizontal Transfer Inhibitor Prevents Transmission of Plasmids Carrying Multiple Antibiotic Resistance Genes. Khurshid M, ed. Transboundary Emerging Dis. 2024, 2024 (1), 7063673, DOI: 10.1155/2024/7063673There is no corresponding record for this reference.
- 286Kogay, R.; Wolf, Y. I.; Koonin, E. V. Defence systems and horizontal gene transfer in bacteria. Environ. Microbiol. 2024, 26 (4), e16630 DOI: 10.1111/1462-2920.16630There is no corresponding record for this reference.
- 287Hwang, S.; Maxwell, K. L. Meet the Anti-CRISPRs: Widespread Protein Inhibitors of CRISPR-Cas Systems. CRISPR J. 2019, 2 (1), 23– 30, DOI: 10.1089/crispr.2018.0052There is no corresponding record for this reference.
- 288Isaev, A. B.; Musharova, O. S.; Severinov, K. V. Microbial Arsenal of Antiviral Defenses - Part I. Biochemistry (Moscow) 2021, 86 (3), 319– 337, DOI: 10.1134/S0006297921030081There is no corresponding record for this reference.
- 289Isaev, A. B.; Musharova, O. S.; Severinov, K. V. Microbial Arsenal of Antiviral Defenses. Part II. Biochemistry (Moscow) 2021, 86 (4), 449– 470, DOI: 10.1134/S0006297921040064There is no corresponding record for this reference.
- 290Price, V. J.; McBride, S. W.; Hullahalli, K.; Chatterjee, A.; Duerkop, B. A.; Palmer, K. L. Enterococcus faecalis CRISPR-Cas Is a Robust Barrier to Conjugative Antibiotic Resistance Dissemination in the Murine Intestine. Ellermeier CD, ed. mSphere 2019, 4 (4), e00464–19 DOI: 10.1128/mSphere.00464-19There is no corresponding record for this reference.
- 291Rafiq, M. S.; Shabbir, M. A.; Raza, A. CRISPR-Cas System: A New Dawn to Combat Antibiotic Resistance. BioDrugs 2024, 38 (3), 387– 404, DOI: 10.1007/s40259-024-00656-3There is no corresponding record for this reference.
- 292O’Meara, D.; Nunney, L. A phylogenetic test of the role of CRISPR-Cas in limiting plasmid acquisition and prophage integration in bacteria. Plasmid 2019, 104, 102418, DOI: 10.1016/j.plasmid.2019.102418There is no corresponding record for this reference.
- 293Wu, Y.; Garushyants, S. K.; Van Den Hurk, A. Bacterial defense systems exhibit synergistic anti-phage activity. Cell Host Microbe 2024, 32 (4), 557– 572, DOI: 10.1016/j.chom.2024.01.015There is no corresponding record for this reference.
- 294Wheatley, R. M.; MacLean, R. C. CRISPR-Cas systems restrict horizontal gene transfer in Pseudomonas aeruginosa. ISME J. 2021, 15 (5), 1420– 1433, DOI: 10.1038/s41396-020-00860-3There is no corresponding record for this reference.
- 295Mahendra, C.; Christie, K. A.; Osuna, B. A.; Pinilla-Redondo, R.; Kleinstiver, B. P.; Bondy-Denomy, J. Broad-spectrum anti-CRISPR proteins facilitate horizontal gene transfer. Nat. Microbiol. 2020, 5 (4), 620– 629, DOI: 10.1038/s41564-020-0692-2There is no corresponding record for this reference.
- 296Park, S. J.; Lee, G. E.; Cho, S. M.; Choi, E. H. Recent applications, future perspectives, and limitations of the CRISPR-Cas system. Mol. Ther. Nucleic Acids 2025, 36 (3), 102634, DOI: 10.1016/j.omtn.2025.102634There is no corresponding record for this reference.
- 297Palacios Araya, D.; Palmer, K. L.; Duerkop, B. A. CRISPR-based antimicrobials to obstruct antibiotic-resistant and pathogenic bacteria. Kline KA, ed. PLoS Pathog. 2021, 17 (7), e1009672 DOI: 10.1371/journal.ppat.1009672There is no corresponding record for this reference.
- 298Wu, Y.; Battalapalli, D.; Hakeem, M. J. Engineered CRISPR-Cas systems for the detection and control of antibiotic-resistant infections. J. Nanobiotechnol. 2021, 19 (1), 401, DOI: 10.1186/s12951-021-01132-8There is no corresponding record for this reference.
- 299Gomaa, A. A.; Klumpe, H. E.; Luo, M. L.; Selle, K.; Barrangou, R.; Beisel, C. L. Programmable Removal of Bacterial Strains by Use of Genome-Targeting CRISPR-Cas Systems. mBio 2014, 5 (1), e00928–13 DOI: 10.1128/mBio.00928-13There is no corresponding record for this reference.
- 300Kippnich, J.; Benz, F.; Uecker, H.; Baumdicker, F. Effectiveness of CRISPR-Cas in sensitizing bacterial populations with plasmid-encoded antimicrobial resistance. Lenormand T, ed. GENETICS. 2025, 231 (3), iyaf192, DOI: 10.1093/genetics/iyaf192There is no corresponding record for this reference.
- 301Saffari Natanzi, A.; Poudineh, M.; Karimi, E.; Khaledi, A.; Haddad Kashani, H. Innovative approaches to combat antibiotic resistance: integrating CRISPR/Cas9 and nanoparticles against biofilm-driven infections. BMC Med. 2025, 23 (1), 486, DOI: 10.1186/s12916-025-04323-4There is no corresponding record for this reference.
- 302Zuberi, A.; Ahmad, N.; Ahmad, H.; Saeed, M.; Ahmad, I. Beyond antibiotics: CRISPR/Cas9 triumph over biofilm-associated antibiotic resistance infections. Front. Cell. Infect. Microbiol. 2024, 14, 1408569, DOI: 10.3389/fcimb.2024.1408569There is no corresponding record for this reference.
- 303Joseph, J.; Abraham, J. S.; Abraham, R.; Nagella, P.; Anand, A. Filamentous Fungi: Heterologous Protein Expression. Prospects of Fungal Biotechnologies for Livestock Vol. 2; Gupta, A., Ed.; Springer Nature Switzerland, 2025; pp 197– 231. DOI: 10.1007/978-3-032-06478-3_8 .There is no corresponding record for this reference.
- 304Pandey, P.; Vavilala, S. L. From Gene Editing to Biofilm Busting: CRISPR-CAS9 Against Antibiotic Resistance–A Review. Cell Biochem. Biophys. 2024, 82 (2), 549– 560, DOI: 10.1007/s12013-024-01276-yThere is no corresponding record for this reference.
- 305Rodrigues, M.; McBride, S. W.; Hullahalli, K.; Palmer, K. L.; Duerkop, B. A. Conjugative Delivery of CRISPR-Cas9 for the Selective Depletion of Antibiotic-Resistant Enterococci. Antimicrob. Agents Chemother. 2019, 63 (11), e01454-19 DOI: 10.1128/AAC.01454-19There is no corresponding record for this reference.
- 306Peng, H.; Chen, I. A. Rapid Colorimetric Detection of Bacterial Species through the Capture of Gold Nanoparticles by Chimeric Phages. ACS Nano 2019, acsnano.8b06395, DOI: 10.1021/acsnano.8b06395There is no corresponding record for this reference.
- 307Bier, E.; Nizet, V. Driving to Safety: CRISPR-Based Genetic Approaches to Reducing Antibiotic Resistance. Trends Genet. 2021, 37 (8), 745– 757, DOI: 10.1016/j.tig.2021.02.007There is no corresponding record for this reference.
- 308Kaduwal, S.; Stuart, E. C.; Auradkar, A.; Washabaugh, S.; Meyer, J. R.; Bier, E. A conjugal gene drive-like system efficiently suppresses antibiotic resistance in a bacterial population. npj Antimicrob Resist. 2026, 4 (1), 8, DOI: 10.1038/s44259-026-00181-zThere is no corresponding record for this reference.
- 309Tsoi, R.; Son, H. I.; Hamrick, G. S. A predatory gene drive for targeted control of self-transmissible plasmids. Sci. Adv. 2025, 11 (14), eads4735 DOI: 10.1126/sciadv.ads4735There is no corresponding record for this reference.
- 310Walter, M.; Verdin, E. Viral gene drive in herpesviruses. Nat. Commun. 2020, 11 (1), 4884, DOI: 10.1038/s41467-020-18678-0There is no corresponding record for this reference.
- 311Gantz, V. M.; Jasinskiene, N.; Tatarenkova, O.; Fazekas, A.; Macias, V. M.; Bier, E.; James, A. A. Highly efficient Cas9-mediated gene drive for population modification of the malaria vector mosquito Anopheles stephensi. Proc. Natl. Acad. Sci. U. S. A. 2015, 112(49). DOI: 10.1073/pnas.1521077112 .There is no corresponding record for this reference.
- 312Valderrama, J. A.; Kulkarni, S. S.; Nizet, V.; Bier, E. A bacterial gene-drive system efficiently edits and inactivates a high copy number antibiotic resistance locus. Nat. Commun. 2019, 10 (1), 5726, DOI: 10.1038/s41467-019-13649-6There is no corresponding record for this reference.
- 313Asim, M.; Rizvi, S. A.; Haq, Q. M. R. Antiplasmid systems: a novel strategy to combat antibiotic resistance. J. Antimicrob. Chemother. 2026, 81 (1), dkaf472, DOI: 10.1093/jac/dkaf472There is no corresponding record for this reference.
- 314Bravo, J. P. K. Anti-plasmid immunity: a key to pathogen success?. Future Microbiol. 2024, 19 (15), 1269– 1272, DOI: 10.1080/17460913.2024.2389720There is no corresponding record for this reference.
- 315Karaiskos, I.; Galani, I.; Papoutsaki, V.; Galani, L.; Giamarellou, H. Carbapenemase producing Klebsiella pneumoniae: implication on future therapeutic strategies. Expert Rev. Anti-Infect. Ther. 2022, 20 (1), 53– 69, DOI: 10.1080/14787210.2021.1935237There is no corresponding record for this reference.
- 316Zongo, P. D.; Cabanel, N.; Royer, G. An antiplasmid system drives antibiotic resistance gene integration in carbapenemase-producing Escherichia coli lineages. Nat. Commun. 2024, 15 (1), 4093, DOI: 10.1038/s41467-024-48219-yThere is no corresponding record for this reference.