AntibioticDB: An Updated and Improved Open-Access Database for the Antibacterial Research and Development CommunityClick to copy article linkArticle link copied!
- Luiza H. GalarionLuiza H. GalarionFaculty of Biological Sciences, School of Molecular and Cellular Biology, University of Leeds, Leeds LS2 9JT, U.K.More by Luiza H. Galarion
- Alan Hennessy*Alan Hennessy*Email: [email protected]The Global Antibiotic Research & Development Partnership (GARDP), Geneva 1202, SwitzerlandMore by Alan Hennessy
- Simon D. HardingSimon D. HardingInstitute for Neuroscience and Cardiovascular Research, University of Edinburgh, Edinburgh EH8 9XD, U.K.More by Simon D. Harding
- Jane F. ArmstrongJane F. ArmstrongInstitute for Neuroscience and Cardiovascular Research, University of Edinburgh, Edinburgh EH8 9XD, U.K.More by Jane F. Armstrong
- Astrid Pentz-MurrAstrid Pentz-MurrThe Global Antibiotic Research & Development Partnership (GARDP), Geneva 1202, SwitzerlandMore by Astrid Pentz-Murr
- Jamie A. DaviesJamie A. DaviesEdinburgh Medical School, University of Edinburgh, Edinburgh EH8 9XD, U.K.More by Jamie A. Davies
- Alex J. O’Neill*Alex J. O’Neill*Email: [email protected]Faculty of Biological Sciences, School of Molecular and Cellular Biology, University of Leeds, Leeds LS2 9JT, U.K.More by Alex J. O’Neill
- Laura J. V. PiddockLaura J. V. PiddockThe Global Antibiotic Research & Development Partnership (GARDP), Geneva 1202, SwitzerlandMore by Laura J. V. Piddock
Abstract
AntibioticDB (https://www.antibioticdb.com/), originally established in 2017 and since 2021 led by the Global Antibiotic Research & Development Partnership (GARDP), is a freely available database of antibacterial agents to facilitate research and development of new antibacterial therapeutics. Here, we describe a new release of AntibioticDB that has been significantly expanded and updated with the aid of user feedback and which offers additional functionality through a redesigned web portal. Improvements include reciprocal integration with the IUPHAR/BPS Guide to Pharmacology (https://www.guidetopharmacology.org), capturing of compound structure information in the form of standard chemical identifiers (canonical and isomeric SMILES, InChI, and InChI Key), chemical 2D structure images, and harmonizing terminology to optimize database searching. Ongoing curation efforts have increased the number of individual entries to >3,500, a process driven mostly by a significant expansion of historical natural product antibiotics that were previously under-represented in the database. The database is continuously updated by mining the published literature and capturing newly discovered antibacterial compounds as they are reported, making AntibioticDB the most complete global resource on antibacterial agents.
This publication is licensed under
License Summary*
You are free to share(copy and redistribute) this article in any medium or format and to adapt(remix, transform, and build upon) the material for any purpose, even commercially within the parameters below:
Creative Commons (CC): This is a Creative Commons license.
Attribution (BY): Credit must be given to the creator.
*Disclaimer
This summary highlights only some of the key features and terms of the actual license. It is not a license and has no legal value. Carefully review the actual license before using these materials.
License Summary*
You are free to share(copy and redistribute) this article in any medium or format and to adapt(remix, transform, and build upon) the material for any purpose, even commercially within the parameters below:
Creative Commons (CC): This is a Creative Commons license.
Attribution (BY): Credit must be given to the creator.
*Disclaimer
This summary highlights only some of the key features and terms of the actual license. It is not a license and has no legal value. Carefully review the actual license before using these materials.
License Summary*
You are free to share(copy and redistribute) this article in any medium or format and to adapt(remix, transform, and build upon) the material for any purpose, even commercially within the parameters below:
Creative Commons (CC): This is a Creative Commons license.
Attribution (BY): Credit must be given to the creator.
*Disclaimer
This summary highlights only some of the key features and terms of the actual license. It is not a license and has no legal value. Carefully review the actual license before using these materials.
Expanding Database Content
| (i) | Agents should demonstrate a degree of selective toxicity against bacteria over mammalian cells. This is a defining feature of an agent with potential for use in systemic antibacterial chemotherapy, since it makes feasible specific targeting of disease-causing bacteria without comparable toxic effects on the patient. It is rarely possible to “introduce” selective toxicity by modification of a compound that lacks selectivity at the outset because such agents typically exert their effects on bacterial and mammalian cells through the same mechanism. Thus, compounds lacking selectivity of action have limited value or prospects for antibacterial chemotherapy and are not the focus of AntibioticDB. An “indication of selectivity” can take many forms in the published literature including a demonstration of safety/efficacy in vivo, a difference in the observed response to the compound between cultured bacterial and mammalian cells, or inhibitory action against a purified target protein that is reduced or absent against the mammalian counterpart. | ||||
| (ii) | Historically, the major focus in antibacterial chemotherapy has been on small molecule drugs that inhibit or kill bacteria (“direct-acting antibacterials”), and such molecules also constitute the bulk of entries in AntibioticDB. Nevertheless, there are agents in clinical use that do not act in this manner (e.g., beta-lactamase inhibitors that primarily act to inhibit an antibiotic resistance mechanism), and there is a growing interest in “nontraditional” agents that act through alternative means. (10) Any modality that has, or might have, potential in the prevention or treatment of bacterial infection is considered appropriate for inclusion in AntibioticDB. Thus, included in AntibioticDB are agents that lack intrinsic antibacterial activity (e.g., those that sensitize bacteria to antibacterial drugs, that target bacterial virulence functions as opposed to viability, or that modulate the host response to infection) and those that are considerably larger than the classical antibacterial drug (e.g., biological materials like enzymes, antibodies, and bacteriophages). Where such agents are intended to be combined with a specific antibacterial drug during treatment, each of the individual components and the combination treatment are curated in AntibioticDB as independent entries. | ||||
| (iii) | Close chemical analogues of antibacterial agents generated during discovery programs are not captured as separate entries in AntibioticDB unless there is a clear rationale for doing so. Often, such “project compounds” are not all individually characterized and/or reported in detail in the published literature, and the assumption that these molecules have the same antibacterial mode of action as the lead compound may not have been verified. Thus, there is typically a lack of robust data available to warrant dedicated entries for such analogues in AntibioticDB. Instead, information is captured for the core or lead scaffold, and the existence of reported analogues is indicated. Exceptions to this approach are made when compelling reported data for an analogue indicate an important difference to other compounds of the series (e.g., distinct antibacterial spectrum or mode of action). | ||||
Improved Information Capture and Harmonization of Terminology in AntibioticDB
| “development status” field term | definition |
|---|---|
| approved | drugs approved by FDA, EMA, or other drug regulatory agencies and currently available in ≥1 country |
| active | agents in clinical trials (whether ongoing or completed, with or without results posted) |
| inactive | agents that have at least reached phase 1 clinical trials and are no longer being taken forward |
| experimental | agents in discovery or preclinical stages |
| withdrawn | drugs previously approved but discontinued/withdrawn from market |
| unknown | agents for which clinical trials have been performed or published but have not been approved for reasons unknown |
| “highest development stage” field term | definition |
|---|---|
| preclinical | agents explicitly reported to be undergoing preclinical evaluation (e.g., in a published article, company press release, or by direct communication from researchers) |
| phase 1–4 | agents undergoing clinical trial (reference number included where available) |
| approved | approved status (with approving agency and year of first approval shown) |
Figure 1
Figure 1. Agents in AntibioticDB as of October 2025 and their development status.
New Functionality in AntibioticDB
An Improved Database and Web Portal
Figure 2
Figure 2. Layout and content of a typical entry page in the new release of AntibioticDB. The entry page is shown in the left-hand panel; linked information is highlighted in red boxes and directs users to relevant information held in other databases.
Concluding Remarks and Future Directions
Acknowledgments
The authors thank the following who participated and provided valuable feedback in beta-testing of the updated AntibioticDB website: Diarmaid Hughes (Uppsala University), Elena Faccenda (Senior curator, Guide to Pharmacology), Paul Hergenrother (University of Illinois), Andrew Tomaras (Senior Vice President─Blacksmith Medicines), Jimmy Nkaiwuatei (Discovery and Innovations─Students Against Superbugs Africa), Olga Genilloud (Scientific Director─Fundación MEDINA), Helen Zgurskaya (University of Oklahoma), Pamela Brown (Director─Pam Brown Consulting Ltd.), Chad Testa (CEO─Curza, Inc.), Martin Everett (CSO─Aurobac Therapeutics), David Davies (Head of Medicinal Chemistry Antabio), Maya Farha (McMaster University), Laura Hollis (University of Birmingham), Samuel Kraus (Universities of Exeter and Queensland), Grant Boyle (H3D, University of Cape Town), and Nikki Cardoso (AMR Investigator─H3D University of Cape Town). We also thank our previous curators and developers: Ursula Theuretzbacher, Elena Faccenda, and Liangcui Chu.
References
This article references 16 other publications.
- 1Murray, C. J. L.; Ikuta, K. S.; Sharara, F.; Swetschinski, L.; Robles Aguilar, G.; Gray, A.; Han, C.; Bisignano, C.; Rao, P.; Wool, E.; Johnson, S. C.; Browne, A. J.; Chipeta, M. G.; Fell, F.; Hackett, S.; Haines-Woodhouse, G.; Kashef Hamadani, B. H.; Kumaran, E. A. P.; McManigal, B.; Achalapong, S.; Agarwal, R.; Akech, S.; Albertson, S.; Amuasi, J.; Andrews, J.; Aravkin, A.; Ashley, E.; Babin, F. X.; Bailey, F.; Baker, S.; Basnyat, B.; Bekker, A.; Bender, R.; Berkley, J. A.; Bethou, A.; Bielicki, J.; Boonkasidecha, S.; Bukosia, J.; Carvalheiro, C.; Castañeda-Orjuela, C.; Chansamouth, V.; Chaurasia, S.; Chiurchiù, S.; Chowdhury, F.; Clotaire Donatien, R.; Cook, A. J.; Cooper, B.; Cressey, T. R.; Criollo-Mora, E.; Cunningham, M.; Darboe, S.; Day, N. P. J.; De Luca, M.; Dokova, K.; Dramowski, A.; Dunachie, S. J.; Duong Bich, T.; Eckmanns, T.; Eibach, D.; Emami, A.; Feasey, N.; Fisher-Pearson, N.; Forrest, K.; Garcia, C.; Garrett, D.; Gastmeier, P.; Giref, A. Z.; Greer, R. C.; Gupta, V.; Haller, S.; Haselbeck, A.; Hay, S. I.; Holm, M.; Hopkins, S.; Hsia, Y.; Iregbu, K. C.; Jacobs, J.; Jarovsky, D.; Javanmardi, F.; Jenney, A. W. J.; Khorana, M.; Khusuwan, S.; Kissoon, N.; Kobeissi, E.; Kostyanev, T.; Krapp, F.; Krumkamp, R.; Kumar, A.; Kyu, H. H.; Lim, C.; Lim, K.; Limmathurotsakul, D.; Loftus, M. J.; Lunn, M.; Ma, J.; Manoharan, A.; Marks, F.; May, J.; Mayxay, M.; Mturi, N.; Munera-Huertas, T.; Musicha, P.; Musila, L. A.; Mussi-Pinhata, M. M.; Naidu, R. N.; Nakamura, T.; Nanavati, R.; Nangia, S.; Newton, P.; Ngoun, C.; Novotney, A.; Nwakanma, D.; Obiero, C. W.; Ochoa, T. J.; Olivas-Martinez, A.; Olliaro, P.; Ooko, E.; Ortiz-Brizuela, E.; Ounchanum, P.; Pak, G. D.; Paredes, J. L.; Peleg, A. Y.; Perrone, C.; Phe, T.; Phommasone, K.; Plakkal, N.; Ponce-de-Leon, A.; Raad, M.; Ramdin, T.; Rattanavong, S.; Riddell, A.; Roberts, T.; Robotham, J. V.; Roca, A.; Rosenthal, V. D.; Rudd, K. E.; Russell, N.; Sader, H. S.; Saengchan, W.; Schnall, J.; Scott, J. A. G.; Seekaew, S.; Sharland, M.; Shivamallappa, M.; Sifuentes-Osornio, J.; Simpson, A. J.; Steenkeste, N.; Stewardson, A. J.; Stoeva, T.; Tasak, N.; Thaiprakong, A.; Thwaites, G.; Tigoi, C.; Turner, C.; Turner, P.; van Doorn, H. R.; Velaphi, S.; Vongpradith, A.; Vongsouvath, M.; Vu, H.; Walsh, T.; Walson, J. L.; Waner, S.; Wangrangsimakul, T.; Wannapinij, P.; Wozniak, T.; Young Sharma, T. E. M. W.; Yu, K. C.; Zheng, P.; Sartorius, B.; Lopez, A. D.; Stergachis, A.; Moore, C.; Dolecek, C.; Naghavi, M.; Antimicrobial Resistance Collaborators Global burden of bacterial antimicrobial resistance in 2019: a systematic analysis. Lancet 2022, 399, 629– 655, DOI: 10.1016/s0140-6736(21)02724-0Google ScholarThere is no corresponding record for this reference.
- 2Naghavi, M.; Vollset, S. E.; Ikuta, K. S.; Swetschinski, L. R.; Gray, A. P.; Wool, E. E.; Robles Aguilar, G.; Mestrovic, T.; Smith, G.; Han, C.; Hsu, R. L.; Chalek, J.; Araki, D. T.; Chung, E.; Raggi, C.; Gershberg Hayoon, A.; Davis Weaver, N.; Lindstedt, P. A.; Smith, A. E.; Altay, U.; Bhattacharjee, N. V.; Giannakis, K.; Fell, F.; McManigal, B.; Ekapirat, N.; Mendes, J. A.; Runghien, T.; Srimokla, O.; Abdelkader, A.; Abd-Elsalam, S.; Aboagye, R. G.; Abolhassani, H.; Abualruz, H.; Abubakar, U.; Abukhadijah, H. J.; Aburuz, S.; Abu-Zaid, A.; Achalapong, S.; Addo, I. Y.; Adekanmbi, V.; Adeyeoluwa, T. E.; Adnani, Q. E. S.; Adzigbli, L. A.; Afzal, M. S.; Afzal, S.; Agodi, A.; Ahlstrom, A. J.; Ahmad, A.; Ahmad, S.; Ahmad, T.; Ahmadi, A.; Ahmed, A.; Ahmed, H.; Ahmed, I.; Ahmed, M.; Ahmed, S.; Ahmed, S. A.; Akkaif, M. A.; Al Awaidy, S.; Al Thaher, Y.; Alalalmeh, S. O.; AlBataineh, M. T.; Aldhaleei, W. A.; Al-Gheethi, A. A. S.; Alhaji, N. B.; Ali, A.; Ali, L.; Ali, S. S.; Ali, W.; Allel, K.; Al-Marwani, S.; Alrawashdeh, A.; Altaf, A.; Al-Tammemi, A. B.; Al-Tawfiq, J. A.; Alzoubi, K. H.; Al-Zyoud, W. A.; Amos, B.; Amuasi, J. H.; Ancuceanu, R.; Andrews, J. R.; Anil, A.; Anuoluwa, I. A.; Anvari, S.; Anyasodor, A. E.; Apostol, G. L. C.; Arabloo, J.; Arafat, M.; Aravkin, A. Y.; Areda, D.; Aremu, A.; Artamonov, A. A.; Ashley, E. A.; Asika, M. O.; Athari, S. S.; Atout, M. M. W.; Awoke, T.; Azadnajafabad, S.; Azam, J. M.; Aziz, S.; Azzam, A. Y.; Babaei, M.; Babin, F. X.; Badar, M.; Baig, A. A.; Bajcetic, M.; Baker, S.; Bardhan, M.; Barqawi, H. J.; Basharat, Z.; Basiru, A.; Bastard, M.; Basu, S.; Bayleyegn, N. S.; Belete, M. A.; Bello, O. O.; Beloukas, A.; Berkley, J. A.; Bhagavathula, A. S.; Bhaskar, S.; Bhuyan, S. S.; Bielicki, J. A.; Briko, N. I.; Brown, C. S.; Browne, A. J.; Buonsenso, D.; Bustanji, Y.; Carvalheiro, C. G.; Castañeda-Orjuela, C. A.; Cenderadewi, M.; Chadwick, J.; Chakraborty, S.; Chandika, R. M.; Chandy, S.; Chansamouth, V.; Chattu, V. K.; Chaudhary, A. A.; Ching, P. R.; Chopra, H.; Chowdhury, F. R.; Chu, D. T.; Chutiyami, M.; Cruz-Martins, N.; da Silva, A. G.; Dadras, O.; Dai, X.; Darcho, S. D.; Das, S.; De la Hoz, F. P.; Dekker, D. M.; Dhama, K.; Diaz, D.; Dickson, B. F. R.; Djorie, S. G.; Dodangeh, M.; Dohare, S.; Dokova, K. G.; Doshi, O. P.; Dowou, R. K.; Dsouza, H. L.; Dunachie, S. J.; Dziedzic, A. M.; Eckmanns, T.; Ed-Dra, A.; Eftekharimehrabad, A.; Ekundayo, T. C.; El Sayed, I.; Elhadi, M.; El-Huneidi, W.; Elias, C.; Ellis, S. J.; Elsheikh, R.; Elsohaby, I.; Eltaha, C.; Eshrati, B.; Eslami, M.; Eyre, D. W.; Fadaka, A. O.; Fagbamigbe, A. F.; Fahim, A.; Fakhri-Demeshghieh, A.; Fasina, F. O.; Fasina, M. M.; Fatehizadeh, A.; Feasey, N. A.; Feizkhah, A.; Fekadu, G.; Fischer, F.; Fitriana, I.; Forrest, K. M.; Fortuna Rodrigues, C.; Fuller, J. E.; Gadanya, M. A.; Gajdács, M.; Gandhi, A. P.; Garcia-Gallo, E. E.; Garrett, D. O.; Gautam, R. K.; Gebregergis, M. W.; Gebrehiwot, M.; Gebremeskel, T. G.; Geffers, C.; Georgalis, L.; Ghazy, R. M.; Golechha, M.; Golinelli, D.; Gordon, M.; Gulati, S.; Gupta, R. D.; Gupta, S.; Gupta, V. K.; Habteyohannes, A. D.; Haller, S.; Harapan, H.; Harrison, M. L.; Hasaballah, A. I.; Hasan, I.; Hasan, R. S.; Hasani, H.; Haselbeck, A. H.; Hasnain, M. S.; Hassan, I. I.; Hassan, S.; Hassan Zadeh Tabatabaei, M. S.; Hayat, K.; He, J.; Hegazi, O. E.; Heidari, M.; Hezam, K.; Holla, R.; Holm, M.; Hopkins, H.; Hossain, M. M.; Hosseinzadeh, M.; Hostiuc, S.; Hussein, N. R.; Huy, L. D.; Ibáñez-Prada, E. D.; Ikiroma, A.; Ilic, I. M.; Islam, S. M. S.; Ismail, F.; Ismail, N. E.; Iwu, C. D.; Iwu-Jaja, C. J.; Jafarzadeh, A.; Jaiteh, F.; Jalilzadeh Yengejeh, R.; Jamora, R. D. G.; Javidnia, J.; Jawaid, T.; Jenney, A. W. J.; Jeon, H. J.; Jokar, M.; Jomehzadeh, N.; Joo, T.; Joseph, N.; Kamal, Z.; Kanmodi, K. K.; Kantar, R. S.; Kapisi, J. A.; Karaye, I. M.; Khader, Y. S.; Khajuria, H.; Khalid, N.; Khamesipour, F.; Khan, A.; Khan, M. J.; Khan, M. T.; Khanal, V.; Khidri, F. F.; Khubchandani, J.; Khusuwan, S.; Kim, M. S.; Kisa, A.; Korshunov, V. A.; Krapp, F.; Krumkamp, R.; Kuddus, M.; Kulimbet, M.; Kumar, D.; Kumaran, E. A. P.; Kuttikkattu, A.; Kyu, H. H.; Landires, I.; Lawal, B. K.; Le, T. T. T.; Lederer, I. M.; Lee, M.; Lee, S. W.; Lepape, A.; Lerango, T. L.; Ligade, V. S.; Lim, C.; Lim, S. S.; Limenh, L. W.; Liu, C.; Liu, X.; Liu, X.; Loftus, M. J.; M Amin, H. I.; Maass, K. L.; Maharaj, S. B.; Mahmoud, M. A.; Maikanti-Charalampous, P.; Makram, O. M.; Malhotra, K.; Malik, A. A.; Mandilara, G. D.; Marks, F.; Martinez-Guerra, B. A.; Martorell, M.; Masoumi-Asl, H.; Mathioudakis, A. G.; May, J.; McHugh, T. A.; Meiring, J.; Meles, H. N.; Melese, A.; Melese, E. B.; Minervini, G.; Mohamed, N. S.; Mohammed, S.; Mohan, S.; Mokdad, A. H.; Monasta, L.; Moodi Ghalibaf, A.; Moore, C. E.; Moradi, Y.; Mossialos, E.; Mougin, V.; Mukoro, G. D.; Mulita, F.; Muller-Pebody, B.; Murillo-Zamora, E.; Musa, S.; Musicha, P.; Musila, L. A.; Muthupandian, S.; Nagarajan, A. J.; Naghavi, P.; Nainu, F.; Nair, T. S.; Najmuldeen, H. H. R.; Natto, Z. S.; Nauman, J.; Nayak, B. P.; Nchanji, G. T.; Ndishimye, P.; Negoi, I.; Negoi, R. I.; Nejadghaderi, S. A.; Nguyen, Q. P.; Noman, E. A.; Nwakanma, D. C.; O’Brien, S.; Ochoa, T. J.; Odetokun, I. A.; Ogundijo, O. A.; Ojo-Akosile, T. R.; Okeke, S. R.; Okonji, O. C.; Olagunju, A. T.; Olivas-Martinez, A.; Olorukooba, A. A.; Olwoch, P.; Onyedibe, K. I.; Ortiz-Brizuela, E.; Osuolale, O.; Ounchanum, P.; Oyeyemi, O. T.; Mahesh Padukudru, P. A.; Paredes, J. L.; Parikh, R. R.; Patel, J.; Patil, S.; Pawar, S.; Peleg, A. Y.; Peprah, P.; Perdigão, J.; Perrone, C.; Petcu, I. R.; Phommasone, K.; Piracha, Z. Z.; Poddighe, D.; Pollard, A. J.; Poluru, R.; Ponce-De-Leon, A.; Puvvula, J.; Qamar, F. N.; Qasim, N. H.; Rafai, C. D.; Raghav, P.; Rahbarnia, L.; Rahim, F.; Rahimi-Movaghar, V.; Rahman, M.; Rahman, M. A.; Ramadan, H.; Ramasamy, S. K.; Ramesh, P. S.; Ramteke, P. W.; Rana, R. K.; Rani, U.; Rashidi, M. M.; Rathish, D.; Rattanavong, S.; Rawaf, S.; Redwan, E. M. M.; Reyes, L. F.; Roberts, T.; Robotham, J. V.; Rosenthal, V. D.; Ross, A. G.; Roy, N.; Rudd, K. E.; Sabet, C. J.; Saddik, B. A.; Saeb, M. R.; Saeed, U.; Saeedi Moghaddam, S.; Saengchan, W.; Safaei, M.; Saghazadeh, A.; Saheb Sharif-Askari, N.; Sahebkar, A.; Sahoo, S. S.; Sahu, M.; Saki, M.; Salam, N.; Saleem, Z.; Saleh, M. A.; Samodra, Y. L.; Samy, A. M.; Saravanan, A.; Satpathy, M.; Schumacher, A. E.; Sedighi, M.; Seekaew, S.; Shafie, M.; Shah, P. A.; Shahid, S.; Shahwan, M. J.; Shakoor, S.; Shalev, N.; Shamim, M. A.; Shamshirgaran, M. A.; Shamsi, A.; Sharifan, A.; Shastry, R. P.; Shetty, M.; Shittu, A.; Shrestha, S.; Siddig, E. E.; Sideroglou, T.; Sifuentes-Osornio, J.; Silva, L. M. L. R.; Simões, E. A. F.; Simpson, A. J. H.; Singh, A.; Singh, S.; Sinto, R.; Soliman, S. S. M.; Soraneh, S.; Stoesser, N.; Stoeva, T. Z.; Swain, C. K.; Szarpak, L.; Sree Sudha, T. Y.; Tabatabai, S.; Tabche, C.; Taha, Z. M. A.; Tan, K. K.; Tasak, N.; Tat, N. Y.; Thaiprakong, A.; Thangaraju, P.; Tigoi, C. C.; Tiwari, K.; Tovani-Palone, M. R.; Tran, T. H.; Tumurkhuu, M.; Turner, P.; Udoakang, A. J.; Udoh, A.; Ullah, N.; Ullah, S.; Vaithinathan, A. G.; Valenti, M.; Vos, T.; Vu, H. T. L.; Waheed, Y.; Walker, A. S.; Walson, J. L.; Wangrangsimakul, T.; Weerakoon, K. G.; Wertheim, H. F. L.; Williams, P. C. M.; Wolde, A. A.; Wozniak, T. M.; Wu, F.; Wu, Z.; Yadav, M. K. K.; Yaghoubi, S.; Yahaya, Z. S.; Yarahmadi, A.; Yezli, S.; Yismaw, Y. E.; Yon, D. K.; Yuan, C. W.; Yusuf, H.; Zakham, F.; Zamagni, G.; Zhang, H.; Zhang, Z. J.; Zielińska, M.; Zumla, A.; Zyoud, S. H. H.; Zyoud, S. H.; Hay, S. I.; Stergachis, A.; Sartorius, B.; Cooper, B. S.; Dolecek, C.; Murray, C. J. L.; GBD 2021 Antimicrobial Resistance Collaborators Global burden of bacterial antimicrobial resistance 1990–2021: a systematic analysis with forecasts to 2050. Lancet 2024, 404, 1199– 1226, DOI: 10.1016/s0140-6736(24)01867-1Google ScholarThere is no corresponding record for this reference.
- 3UK Government; Department of Health and Social Care; Scottish Government; Welsh Government; Department for Environmental Food & Rural Affairs; Department of Health (Northern Ireland); Department of Agriculture, Environment and Rural Affairs (Northern Ireland). Confronting antimicrobial resistance 2024 to 2029, 2024. Available at: https://www.gov.uk/government/publications/uk-5-year-action-plan-for-antimicrobial-resistance-2024-to-2029/confronting-antimicrobial-resistance-2024-to-2029#executive-summary.Google ScholarThere is no corresponding record for this reference.
- 4Piddock, L. J. V.; Alimi, Y.; Anderson, J.; de Felice, D.; Moore, C. E.; Røttingen, J.-A.; Skinner, H.; Beyer, P. Advancing global antibiotic research, development and access. Nat. Med. 2024, 30, 2432– 2443, DOI: 10.1038/s41591-024-03218-wGoogle ScholarThere is no corresponding record for this reference.
- 5Theuretzbacher, U.; Outterson, K.; Engel, A.; Karlén, A. The global preclinical antibacterial pipeline. Nat. Rev. Microbiol. 2020, 18, 275– 285, DOI: 10.1038/s41579-019-0288-0Google ScholarThere is no corresponding record for this reference.
- 6Rybak, M. J. The efficacy and safety of daptomycin: first in a new class of antibiotics for Gram-positive bacteria. Clin. Microbiol. Infect. 2006, 12, 24– 32, DOI: 10.1111/j.1469-0691.2006.01342.xGoogle ScholarThere is no corresponding record for this reference.
- 7Venugopal, A. A.; Johnson, S. Fidaxomicin: A Novel Macrocyclic Antibiotic Approved for Treatment of Clostridium difficile Infection. Clin. Infect. Dis. 2012, 54, 568– 574, DOI: 10.1093/cid/cir830Google ScholarThere is no corresponding record for this reference.
- 8Farrell, L. J.; Lo, R.; Wanford, J. J.; Jenkins, A.; Maxwell, A.; Piddock, L. J. V. Revitalizing the drug pipeline: AntibioticDB, an open access database to aid antibacterial research and development. J. Antimicrob. Chemother. 2018, 73, 2284– 2297, DOI: 10.1093/jac/dky208Google ScholarThere is no corresponding record for this reference.
- 9Glasby, J. S. Encyclopedia of Antibiotics, 3rd ed.; John Wiley & Sons Ltd., 1992.Google ScholarThere is no corresponding record for this reference.
- 10Czaplewski, L.; Bax, R.; Clokie, M.; Dawson, M.; Fairhead, H.; Fischetti, V. A.; Foster, S.; Gilmore, B. F.; Hancock, R. E. W.; Harper, D.; Henderson, I. R.; Hilpert, K.; Jones, B. V.; Kadioglu, A.; Knowles, D.; Ólafsdóttir, S.; Payne, D.; Projan, S.; Shaunak, S.; Silverman, J.; Thomas, C. M.; Trust, T. J.; Warn, P.; Rex, J. H. Alternatives to antibiotics─a pipeline portfolio review. Lancet Infect. Dis. 2016, 16, 239– 251, DOI: 10.1016/S1473-3099(15)00466-1Google ScholarThere is no corresponding record for this reference.
- 11Harding, S. D.; Armstrong, J. F.; Faccenda, E.; Southan, C.; Alexander, S. P. H.; Davenport, A. P.; Spedding, M.; Davies, J. A. The IUPHAR/BPS Guide to PHARMACOLOGY in 2024. Nucleic Acids Res. 2024, 52, D1438– D1449, DOI: 10.1093/nar/gkad944Google ScholarThere is no corresponding record for this reference.
- 12Pawson, A. J.; Sharman, J. L.; Benson, H. E.; Faccenda, E.; Alexander, S. P. H.; Buneman, O. P.; Davenport, A. P.; McGrath, J. C.; Peters, J. A.; Southan, C.; Spedding, M.; Yu, W.; Harmar, A. J. The IUPHAR/BPS Guide to PHARMACOLOGY: an expert-driven knowledgebase of drug targets and their ligands. Nucleic Acids Res. 2014, 42, D1098– D1106, DOI: 10.1093/nar/gkt1143Google ScholarThere is no corresponding record for this reference.
- 13Kim, S.; Chen, J.; Cheng, T.; Gindulyte, A.; He, J.; He, S.; Li, Q.; Shoemaker, B. A.; Thiessen, P. A.; Yu, B.; Zaslavsky, L.; Zhang, J.; Bolton, E. E. PubChem 2023 update. Nucleic Acids Res. 2023, 51, D1373– D1380, DOI: 10.1093/nar/gkac956Google ScholarThere is no corresponding record for this reference.
- 14Zdrazil, B.; Felix, E.; Hunter, F.; Manners, E. J.; Blackshaw, J.; Corbett, S.; de Veij, M.; Ioannidis, H.; Lopez, D. M.; Mosquera, J. F.; Magarinos, M. P.; Bosc, N.; Arcila, R.; Kizilören, T.; Gaulton, A.; Bento, A. P.; Adasme, M. F.; Monecke, P.; Landrum, G. A.; Leach, A. R. The ChEMBL Database in 2023: a drug discovery platform spanning multiple bioactivity data types and time periods. Nucleic Acids Res. 2024, 52, D1180– D1192, DOI: 10.1093/nar/gkad1004Google ScholarThere is no corresponding record for this reference.
- 15Rajan, K.; Zielesny, A.; Steinbeck, C. DECIMER: towards deep learning for chemical image recognition. J. Cheminf. 2020, 12, 65, DOI: 10.1186/s13321-020-00469-wGoogle ScholarThere is no corresponding record for this reference.
- 16Willighagen, E. L.; Mayfield, J. W.; Alvarsson, J.; Berg, A.; Carlsson, L.; Jeliazkova, N.; Kuhn, S.; Pluskal, T.; Rojas-Chertó, M.; Spjuth, O.; Torrance, G.; Evelo, C. T.; Guha, R.; Steinbeck, C. The Chemistry Development Kit (CDK) v2.0: atom typing, depiction, molecular formulas, and substructure searching. J. Cheminf. 2017, 9, 33, DOI: 10.1186/s13321-017-0220-4Google ScholarThere is no corresponding record for this reference.
Cited By
This article has not yet been cited by other publications.
Article Views
Altmetric
Citations
Article Views are the COUNTER-compliant sum of full text article downloads since November 2008 (both PDF and HTML) across all institutions and individuals. These metrics are regularly updated to reflect usage leading up to the last few days.
Citations are the number of other articles citing this article, calculated by Crossref and updated daily. Find more information about Crossref citation counts.
The Altmetric Attention Score is a quantitative measure of the attention that a research article has received online. Clicking on the donut icon will load a page at altmetric.com with additional details about the score and the social media presence for the given article. Find more information on the Altmetric Attention Score and how the score is calculated.
Recommended Articles
Abstract
Figure 1
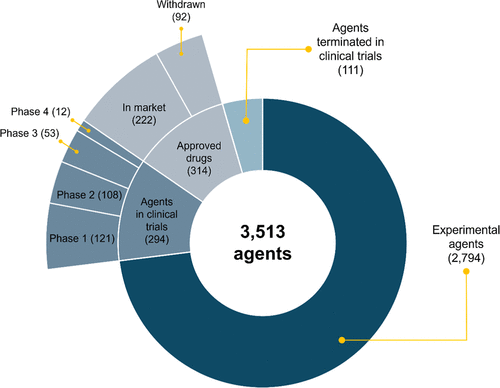
Figure 1. Agents in AntibioticDB as of October 2025 and their development status.
Figure 2

Figure 2. Layout and content of a typical entry page in the new release of AntibioticDB. The entry page is shown in the left-hand panel; linked information is highlighted in red boxes and directs users to relevant information held in other databases.
References
This article references 16 other publications.
- 1Murray, C. J. L.; Ikuta, K. S.; Sharara, F.; Swetschinski, L.; Robles Aguilar, G.; Gray, A.; Han, C.; Bisignano, C.; Rao, P.; Wool, E.; Johnson, S. C.; Browne, A. J.; Chipeta, M. G.; Fell, F.; Hackett, S.; Haines-Woodhouse, G.; Kashef Hamadani, B. H.; Kumaran, E. A. P.; McManigal, B.; Achalapong, S.; Agarwal, R.; Akech, S.; Albertson, S.; Amuasi, J.; Andrews, J.; Aravkin, A.; Ashley, E.; Babin, F. X.; Bailey, F.; Baker, S.; Basnyat, B.; Bekker, A.; Bender, R.; Berkley, J. A.; Bethou, A.; Bielicki, J.; Boonkasidecha, S.; Bukosia, J.; Carvalheiro, C.; Castañeda-Orjuela, C.; Chansamouth, V.; Chaurasia, S.; Chiurchiù, S.; Chowdhury, F.; Clotaire Donatien, R.; Cook, A. J.; Cooper, B.; Cressey, T. R.; Criollo-Mora, E.; Cunningham, M.; Darboe, S.; Day, N. P. J.; De Luca, M.; Dokova, K.; Dramowski, A.; Dunachie, S. J.; Duong Bich, T.; Eckmanns, T.; Eibach, D.; Emami, A.; Feasey, N.; Fisher-Pearson, N.; Forrest, K.; Garcia, C.; Garrett, D.; Gastmeier, P.; Giref, A. Z.; Greer, R. C.; Gupta, V.; Haller, S.; Haselbeck, A.; Hay, S. I.; Holm, M.; Hopkins, S.; Hsia, Y.; Iregbu, K. C.; Jacobs, J.; Jarovsky, D.; Javanmardi, F.; Jenney, A. W. J.; Khorana, M.; Khusuwan, S.; Kissoon, N.; Kobeissi, E.; Kostyanev, T.; Krapp, F.; Krumkamp, R.; Kumar, A.; Kyu, H. H.; Lim, C.; Lim, K.; Limmathurotsakul, D.; Loftus, M. J.; Lunn, M.; Ma, J.; Manoharan, A.; Marks, F.; May, J.; Mayxay, M.; Mturi, N.; Munera-Huertas, T.; Musicha, P.; Musila, L. A.; Mussi-Pinhata, M. M.; Naidu, R. N.; Nakamura, T.; Nanavati, R.; Nangia, S.; Newton, P.; Ngoun, C.; Novotney, A.; Nwakanma, D.; Obiero, C. W.; Ochoa, T. J.; Olivas-Martinez, A.; Olliaro, P.; Ooko, E.; Ortiz-Brizuela, E.; Ounchanum, P.; Pak, G. D.; Paredes, J. L.; Peleg, A. Y.; Perrone, C.; Phe, T.; Phommasone, K.; Plakkal, N.; Ponce-de-Leon, A.; Raad, M.; Ramdin, T.; Rattanavong, S.; Riddell, A.; Roberts, T.; Robotham, J. V.; Roca, A.; Rosenthal, V. D.; Rudd, K. E.; Russell, N.; Sader, H. S.; Saengchan, W.; Schnall, J.; Scott, J. A. G.; Seekaew, S.; Sharland, M.; Shivamallappa, M.; Sifuentes-Osornio, J.; Simpson, A. J.; Steenkeste, N.; Stewardson, A. J.; Stoeva, T.; Tasak, N.; Thaiprakong, A.; Thwaites, G.; Tigoi, C.; Turner, C.; Turner, P.; van Doorn, H. R.; Velaphi, S.; Vongpradith, A.; Vongsouvath, M.; Vu, H.; Walsh, T.; Walson, J. L.; Waner, S.; Wangrangsimakul, T.; Wannapinij, P.; Wozniak, T.; Young Sharma, T. E. M. W.; Yu, K. C.; Zheng, P.; Sartorius, B.; Lopez, A. D.; Stergachis, A.; Moore, C.; Dolecek, C.; Naghavi, M.; Antimicrobial Resistance Collaborators Global burden of bacterial antimicrobial resistance in 2019: a systematic analysis. Lancet 2022, 399, 629– 655, DOI: 10.1016/s0140-6736(21)02724-0There is no corresponding record for this reference.
- 2Naghavi, M.; Vollset, S. E.; Ikuta, K. S.; Swetschinski, L. R.; Gray, A. P.; Wool, E. E.; Robles Aguilar, G.; Mestrovic, T.; Smith, G.; Han, C.; Hsu, R. L.; Chalek, J.; Araki, D. T.; Chung, E.; Raggi, C.; Gershberg Hayoon, A.; Davis Weaver, N.; Lindstedt, P. A.; Smith, A. E.; Altay, U.; Bhattacharjee, N. V.; Giannakis, K.; Fell, F.; McManigal, B.; Ekapirat, N.; Mendes, J. A.; Runghien, T.; Srimokla, O.; Abdelkader, A.; Abd-Elsalam, S.; Aboagye, R. G.; Abolhassani, H.; Abualruz, H.; Abubakar, U.; Abukhadijah, H. J.; Aburuz, S.; Abu-Zaid, A.; Achalapong, S.; Addo, I. Y.; Adekanmbi, V.; Adeyeoluwa, T. E.; Adnani, Q. E. S.; Adzigbli, L. A.; Afzal, M. S.; Afzal, S.; Agodi, A.; Ahlstrom, A. J.; Ahmad, A.; Ahmad, S.; Ahmad, T.; Ahmadi, A.; Ahmed, A.; Ahmed, H.; Ahmed, I.; Ahmed, M.; Ahmed, S.; Ahmed, S. A.; Akkaif, M. A.; Al Awaidy, S.; Al Thaher, Y.; Alalalmeh, S. O.; AlBataineh, M. T.; Aldhaleei, W. A.; Al-Gheethi, A. A. S.; Alhaji, N. B.; Ali, A.; Ali, L.; Ali, S. S.; Ali, W.; Allel, K.; Al-Marwani, S.; Alrawashdeh, A.; Altaf, A.; Al-Tammemi, A. B.; Al-Tawfiq, J. A.; Alzoubi, K. H.; Al-Zyoud, W. A.; Amos, B.; Amuasi, J. H.; Ancuceanu, R.; Andrews, J. R.; Anil, A.; Anuoluwa, I. A.; Anvari, S.; Anyasodor, A. E.; Apostol, G. L. C.; Arabloo, J.; Arafat, M.; Aravkin, A. Y.; Areda, D.; Aremu, A.; Artamonov, A. A.; Ashley, E. A.; Asika, M. O.; Athari, S. S.; Atout, M. M. W.; Awoke, T.; Azadnajafabad, S.; Azam, J. M.; Aziz, S.; Azzam, A. Y.; Babaei, M.; Babin, F. X.; Badar, M.; Baig, A. A.; Bajcetic, M.; Baker, S.; Bardhan, M.; Barqawi, H. J.; Basharat, Z.; Basiru, A.; Bastard, M.; Basu, S.; Bayleyegn, N. S.; Belete, M. A.; Bello, O. O.; Beloukas, A.; Berkley, J. A.; Bhagavathula, A. S.; Bhaskar, S.; Bhuyan, S. S.; Bielicki, J. A.; Briko, N. I.; Brown, C. S.; Browne, A. J.; Buonsenso, D.; Bustanji, Y.; Carvalheiro, C. G.; Castañeda-Orjuela, C. A.; Cenderadewi, M.; Chadwick, J.; Chakraborty, S.; Chandika, R. M.; Chandy, S.; Chansamouth, V.; Chattu, V. K.; Chaudhary, A. A.; Ching, P. R.; Chopra, H.; Chowdhury, F. R.; Chu, D. T.; Chutiyami, M.; Cruz-Martins, N.; da Silva, A. G.; Dadras, O.; Dai, X.; Darcho, S. D.; Das, S.; De la Hoz, F. P.; Dekker, D. M.; Dhama, K.; Diaz, D.; Dickson, B. F. R.; Djorie, S. G.; Dodangeh, M.; Dohare, S.; Dokova, K. G.; Doshi, O. P.; Dowou, R. K.; Dsouza, H. L.; Dunachie, S. J.; Dziedzic, A. M.; Eckmanns, T.; Ed-Dra, A.; Eftekharimehrabad, A.; Ekundayo, T. C.; El Sayed, I.; Elhadi, M.; El-Huneidi, W.; Elias, C.; Ellis, S. J.; Elsheikh, R.; Elsohaby, I.; Eltaha, C.; Eshrati, B.; Eslami, M.; Eyre, D. W.; Fadaka, A. O.; Fagbamigbe, A. F.; Fahim, A.; Fakhri-Demeshghieh, A.; Fasina, F. O.; Fasina, M. M.; Fatehizadeh, A.; Feasey, N. A.; Feizkhah, A.; Fekadu, G.; Fischer, F.; Fitriana, I.; Forrest, K. M.; Fortuna Rodrigues, C.; Fuller, J. E.; Gadanya, M. A.; Gajdács, M.; Gandhi, A. P.; Garcia-Gallo, E. E.; Garrett, D. O.; Gautam, R. K.; Gebregergis, M. W.; Gebrehiwot, M.; Gebremeskel, T. G.; Geffers, C.; Georgalis, L.; Ghazy, R. M.; Golechha, M.; Golinelli, D.; Gordon, M.; Gulati, S.; Gupta, R. D.; Gupta, S.; Gupta, V. K.; Habteyohannes, A. D.; Haller, S.; Harapan, H.; Harrison, M. L.; Hasaballah, A. I.; Hasan, I.; Hasan, R. S.; Hasani, H.; Haselbeck, A. H.; Hasnain, M. S.; Hassan, I. I.; Hassan, S.; Hassan Zadeh Tabatabaei, M. S.; Hayat, K.; He, J.; Hegazi, O. E.; Heidari, M.; Hezam, K.; Holla, R.; Holm, M.; Hopkins, H.; Hossain, M. M.; Hosseinzadeh, M.; Hostiuc, S.; Hussein, N. R.; Huy, L. D.; Ibáñez-Prada, E. D.; Ikiroma, A.; Ilic, I. M.; Islam, S. M. S.; Ismail, F.; Ismail, N. E.; Iwu, C. D.; Iwu-Jaja, C. J.; Jafarzadeh, A.; Jaiteh, F.; Jalilzadeh Yengejeh, R.; Jamora, R. D. G.; Javidnia, J.; Jawaid, T.; Jenney, A. W. J.; Jeon, H. J.; Jokar, M.; Jomehzadeh, N.; Joo, T.; Joseph, N.; Kamal, Z.; Kanmodi, K. K.; Kantar, R. S.; Kapisi, J. A.; Karaye, I. M.; Khader, Y. S.; Khajuria, H.; Khalid, N.; Khamesipour, F.; Khan, A.; Khan, M. J.; Khan, M. T.; Khanal, V.; Khidri, F. F.; Khubchandani, J.; Khusuwan, S.; Kim, M. S.; Kisa, A.; Korshunov, V. A.; Krapp, F.; Krumkamp, R.; Kuddus, M.; Kulimbet, M.; Kumar, D.; Kumaran, E. A. P.; Kuttikkattu, A.; Kyu, H. H.; Landires, I.; Lawal, B. K.; Le, T. T. T.; Lederer, I. M.; Lee, M.; Lee, S. W.; Lepape, A.; Lerango, T. L.; Ligade, V. S.; Lim, C.; Lim, S. S.; Limenh, L. W.; Liu, C.; Liu, X.; Liu, X.; Loftus, M. J.; M Amin, H. I.; Maass, K. L.; Maharaj, S. B.; Mahmoud, M. A.; Maikanti-Charalampous, P.; Makram, O. M.; Malhotra, K.; Malik, A. A.; Mandilara, G. D.; Marks, F.; Martinez-Guerra, B. A.; Martorell, M.; Masoumi-Asl, H.; Mathioudakis, A. G.; May, J.; McHugh, T. A.; Meiring, J.; Meles, H. N.; Melese, A.; Melese, E. B.; Minervini, G.; Mohamed, N. S.; Mohammed, S.; Mohan, S.; Mokdad, A. H.; Monasta, L.; Moodi Ghalibaf, A.; Moore, C. E.; Moradi, Y.; Mossialos, E.; Mougin, V.; Mukoro, G. D.; Mulita, F.; Muller-Pebody, B.; Murillo-Zamora, E.; Musa, S.; Musicha, P.; Musila, L. A.; Muthupandian, S.; Nagarajan, A. J.; Naghavi, P.; Nainu, F.; Nair, T. S.; Najmuldeen, H. H. R.; Natto, Z. S.; Nauman, J.; Nayak, B. P.; Nchanji, G. T.; Ndishimye, P.; Negoi, I.; Negoi, R. I.; Nejadghaderi, S. A.; Nguyen, Q. P.; Noman, E. A.; Nwakanma, D. C.; O’Brien, S.; Ochoa, T. J.; Odetokun, I. A.; Ogundijo, O. A.; Ojo-Akosile, T. R.; Okeke, S. R.; Okonji, O. C.; Olagunju, A. T.; Olivas-Martinez, A.; Olorukooba, A. A.; Olwoch, P.; Onyedibe, K. I.; Ortiz-Brizuela, E.; Osuolale, O.; Ounchanum, P.; Oyeyemi, O. T.; Mahesh Padukudru, P. A.; Paredes, J. L.; Parikh, R. R.; Patel, J.; Patil, S.; Pawar, S.; Peleg, A. Y.; Peprah, P.; Perdigão, J.; Perrone, C.; Petcu, I. R.; Phommasone, K.; Piracha, Z. Z.; Poddighe, D.; Pollard, A. J.; Poluru, R.; Ponce-De-Leon, A.; Puvvula, J.; Qamar, F. N.; Qasim, N. H.; Rafai, C. D.; Raghav, P.; Rahbarnia, L.; Rahim, F.; Rahimi-Movaghar, V.; Rahman, M.; Rahman, M. A.; Ramadan, H.; Ramasamy, S. K.; Ramesh, P. S.; Ramteke, P. W.; Rana, R. K.; Rani, U.; Rashidi, M. M.; Rathish, D.; Rattanavong, S.; Rawaf, S.; Redwan, E. M. M.; Reyes, L. F.; Roberts, T.; Robotham, J. V.; Rosenthal, V. D.; Ross, A. G.; Roy, N.; Rudd, K. E.; Sabet, C. J.; Saddik, B. A.; Saeb, M. R.; Saeed, U.; Saeedi Moghaddam, S.; Saengchan, W.; Safaei, M.; Saghazadeh, A.; Saheb Sharif-Askari, N.; Sahebkar, A.; Sahoo, S. S.; Sahu, M.; Saki, M.; Salam, N.; Saleem, Z.; Saleh, M. A.; Samodra, Y. L.; Samy, A. M.; Saravanan, A.; Satpathy, M.; Schumacher, A. E.; Sedighi, M.; Seekaew, S.; Shafie, M.; Shah, P. A.; Shahid, S.; Shahwan, M. J.; Shakoor, S.; Shalev, N.; Shamim, M. A.; Shamshirgaran, M. A.; Shamsi, A.; Sharifan, A.; Shastry, R. P.; Shetty, M.; Shittu, A.; Shrestha, S.; Siddig, E. E.; Sideroglou, T.; Sifuentes-Osornio, J.; Silva, L. M. L. R.; Simões, E. A. F.; Simpson, A. J. H.; Singh, A.; Singh, S.; Sinto, R.; Soliman, S. S. M.; Soraneh, S.; Stoesser, N.; Stoeva, T. Z.; Swain, C. K.; Szarpak, L.; Sree Sudha, T. Y.; Tabatabai, S.; Tabche, C.; Taha, Z. M. A.; Tan, K. K.; Tasak, N.; Tat, N. Y.; Thaiprakong, A.; Thangaraju, P.; Tigoi, C. C.; Tiwari, K.; Tovani-Palone, M. R.; Tran, T. H.; Tumurkhuu, M.; Turner, P.; Udoakang, A. J.; Udoh, A.; Ullah, N.; Ullah, S.; Vaithinathan, A. G.; Valenti, M.; Vos, T.; Vu, H. T. L.; Waheed, Y.; Walker, A. S.; Walson, J. L.; Wangrangsimakul, T.; Weerakoon, K. G.; Wertheim, H. F. L.; Williams, P. C. M.; Wolde, A. A.; Wozniak, T. M.; Wu, F.; Wu, Z.; Yadav, M. K. K.; Yaghoubi, S.; Yahaya, Z. S.; Yarahmadi, A.; Yezli, S.; Yismaw, Y. E.; Yon, D. K.; Yuan, C. W.; Yusuf, H.; Zakham, F.; Zamagni, G.; Zhang, H.; Zhang, Z. J.; Zielińska, M.; Zumla, A.; Zyoud, S. H. H.; Zyoud, S. H.; Hay, S. I.; Stergachis, A.; Sartorius, B.; Cooper, B. S.; Dolecek, C.; Murray, C. J. L.; GBD 2021 Antimicrobial Resistance Collaborators Global burden of bacterial antimicrobial resistance 1990–2021: a systematic analysis with forecasts to 2050. Lancet 2024, 404, 1199– 1226, DOI: 10.1016/s0140-6736(24)01867-1There is no corresponding record for this reference.
- 3UK Government; Department of Health and Social Care; Scottish Government; Welsh Government; Department for Environmental Food & Rural Affairs; Department of Health (Northern Ireland); Department of Agriculture, Environment and Rural Affairs (Northern Ireland). Confronting antimicrobial resistance 2024 to 2029, 2024. Available at: https://www.gov.uk/government/publications/uk-5-year-action-plan-for-antimicrobial-resistance-2024-to-2029/confronting-antimicrobial-resistance-2024-to-2029#executive-summary.There is no corresponding record for this reference.
- 4Piddock, L. J. V.; Alimi, Y.; Anderson, J.; de Felice, D.; Moore, C. E.; Røttingen, J.-A.; Skinner, H.; Beyer, P. Advancing global antibiotic research, development and access. Nat. Med. 2024, 30, 2432– 2443, DOI: 10.1038/s41591-024-03218-wThere is no corresponding record for this reference.
- 5Theuretzbacher, U.; Outterson, K.; Engel, A.; Karlén, A. The global preclinical antibacterial pipeline. Nat. Rev. Microbiol. 2020, 18, 275– 285, DOI: 10.1038/s41579-019-0288-0There is no corresponding record for this reference.
- 6Rybak, M. J. The efficacy and safety of daptomycin: first in a new class of antibiotics for Gram-positive bacteria. Clin. Microbiol. Infect. 2006, 12, 24– 32, DOI: 10.1111/j.1469-0691.2006.01342.xThere is no corresponding record for this reference.
- 7Venugopal, A. A.; Johnson, S. Fidaxomicin: A Novel Macrocyclic Antibiotic Approved for Treatment of Clostridium difficile Infection. Clin. Infect. Dis. 2012, 54, 568– 574, DOI: 10.1093/cid/cir830There is no corresponding record for this reference.
- 8Farrell, L. J.; Lo, R.; Wanford, J. J.; Jenkins, A.; Maxwell, A.; Piddock, L. J. V. Revitalizing the drug pipeline: AntibioticDB, an open access database to aid antibacterial research and development. J. Antimicrob. Chemother. 2018, 73, 2284– 2297, DOI: 10.1093/jac/dky208There is no corresponding record for this reference.
- 9Glasby, J. S. Encyclopedia of Antibiotics, 3rd ed.; John Wiley & Sons Ltd., 1992.There is no corresponding record for this reference.
- 10Czaplewski, L.; Bax, R.; Clokie, M.; Dawson, M.; Fairhead, H.; Fischetti, V. A.; Foster, S.; Gilmore, B. F.; Hancock, R. E. W.; Harper, D.; Henderson, I. R.; Hilpert, K.; Jones, B. V.; Kadioglu, A.; Knowles, D.; Ólafsdóttir, S.; Payne, D.; Projan, S.; Shaunak, S.; Silverman, J.; Thomas, C. M.; Trust, T. J.; Warn, P.; Rex, J. H. Alternatives to antibiotics─a pipeline portfolio review. Lancet Infect. Dis. 2016, 16, 239– 251, DOI: 10.1016/S1473-3099(15)00466-1There is no corresponding record for this reference.
- 11Harding, S. D.; Armstrong, J. F.; Faccenda, E.; Southan, C.; Alexander, S. P. H.; Davenport, A. P.; Spedding, M.; Davies, J. A. The IUPHAR/BPS Guide to PHARMACOLOGY in 2024. Nucleic Acids Res. 2024, 52, D1438– D1449, DOI: 10.1093/nar/gkad944There is no corresponding record for this reference.
- 12Pawson, A. J.; Sharman, J. L.; Benson, H. E.; Faccenda, E.; Alexander, S. P. H.; Buneman, O. P.; Davenport, A. P.; McGrath, J. C.; Peters, J. A.; Southan, C.; Spedding, M.; Yu, W.; Harmar, A. J. The IUPHAR/BPS Guide to PHARMACOLOGY: an expert-driven knowledgebase of drug targets and their ligands. Nucleic Acids Res. 2014, 42, D1098– D1106, DOI: 10.1093/nar/gkt1143There is no corresponding record for this reference.
- 13Kim, S.; Chen, J.; Cheng, T.; Gindulyte, A.; He, J.; He, S.; Li, Q.; Shoemaker, B. A.; Thiessen, P. A.; Yu, B.; Zaslavsky, L.; Zhang, J.; Bolton, E. E. PubChem 2023 update. Nucleic Acids Res. 2023, 51, D1373– D1380, DOI: 10.1093/nar/gkac956There is no corresponding record for this reference.
- 14Zdrazil, B.; Felix, E.; Hunter, F.; Manners, E. J.; Blackshaw, J.; Corbett, S.; de Veij, M.; Ioannidis, H.; Lopez, D. M.; Mosquera, J. F.; Magarinos, M. P.; Bosc, N.; Arcila, R.; Kizilören, T.; Gaulton, A.; Bento, A. P.; Adasme, M. F.; Monecke, P.; Landrum, G. A.; Leach, A. R. The ChEMBL Database in 2023: a drug discovery platform spanning multiple bioactivity data types and time periods. Nucleic Acids Res. 2024, 52, D1180– D1192, DOI: 10.1093/nar/gkad1004There is no corresponding record for this reference.
- 15Rajan, K.; Zielesny, A.; Steinbeck, C. DECIMER: towards deep learning for chemical image recognition. J. Cheminf. 2020, 12, 65, DOI: 10.1186/s13321-020-00469-wThere is no corresponding record for this reference.
- 16Willighagen, E. L.; Mayfield, J. W.; Alvarsson, J.; Berg, A.; Carlsson, L.; Jeliazkova, N.; Kuhn, S.; Pluskal, T.; Rojas-Chertó, M.; Spjuth, O.; Torrance, G.; Evelo, C. T.; Guha, R.; Steinbeck, C. The Chemistry Development Kit (CDK) v2.0: atom typing, depiction, molecular formulas, and substructure searching. J. Cheminf. 2017, 9, 33, DOI: 10.1186/s13321-017-0220-4There is no corresponding record for this reference.



