
Direct Observation of Metastable Fragment Ions in Ultraviolet Photodissociation of UbiquitinClick to copy article linkArticle link copied!
- Fanny C. LiuFanny C. LiuDepartment of Chemistry and Biochemistry, Florida State University, Tallahassee, Florida 32304, United StatesMore by Fanny C. Liu
- Jusung LeeJusung LeeDepartment of Chemistry and Biochemistry, Florida State University, Tallahassee, Florida 32304, United StatesMore by Jusung Lee
- Kaira A. MayberryKaira A. MayberryDepartment of Chemistry and Biochemistry, Florida State University, Tallahassee, Florida 32304, United StatesMore by Kaira A. Mayberry
- Mark E. RidgewayMark E. RidgewayBruker Daltonics, Billerica, Massachusetts 01821, United StatesMore by Mark E. Ridgeway
- Christopher A. Wootton
- Alina Theisen
- Erin M. PanczykErin M. PanczykBruker Daltonics, Billerica, Massachusetts 01821, United StatesMore by Erin M. Panczyk
- Benjamin J. JonesBenjamin J. JonesBruker Daltonics, Billerica, Massachusetts 01821, United StatesMore by Benjamin J. Jones
- Lea NienhausLea NienhausDepartment of Chemistry and Biochemistry, Florida State University, Tallahassee, Florida 32304, United StatesMore by Lea Nienhaus
- Melvin A. ParkMelvin A. ParkBruker Switzerland AG, Industriestrasse 26, Fällanden 8117, SwitzerlandMore by Melvin A. Park
- Christian Bleiholder*Christian Bleiholder*Email: [email protected]Department of Chemistry and Biochemistry, Florida State University, Tallahassee, Florida 32304, United StatesInstitute of Molecular Biophysics, Florida State University, Tallahassee, Florida 32304, United StatesMore by Christian Bleiholder
Abstract
Ultraviolet photodissociation (UVPD) of proteins is known to exhibit conformation-dependent fragmentation patterns, but direct structural evidence linking precursor protein and fragment ions has been limited. Here, we apply tandem trapped-ion mobility spectrometry/tandem-mass spectrometry to compare collision cross sections of UVPD fragment ions generated from distinct conformers of ubiquitin. Under the high-pressure (∼4 mbar) and low-photon density (∼10 μJ laser pulse energies) conditions employed here, UVPD produces predominantly [b + 2] and [y – 2] ions at proline residues, consistent with direct bond cleavage from the electronically excited state. Our data show that these ions can retain a clear structural relationship to the precursor conformation: UVPD of compact, native-like ubiquitin yields fragments with collision cross sections ∼20% smaller than the corresponding ions produced from extended precursors or by collision-induced dissociation. Further, these compact UVPD fragments are kinetically trapped in metastable conformations, with substantial barriers preventing relaxation toward energetically favored gas-phase structures. We attribute this behavior to limited vibrational energy deposition per absorbed 213 nm photon combined with rapid collisional cooling, which suppress cumulative thermal activation and disfavor statistical fragmentation pathways, leaving direct excited-state dissociation as the dominant observable process. Together with prior UVPD studies on holo-myoglobin, our results suggest that UVPD fragments can retain aspects of their precursor tertiary structure.
This publication is licensed for personal use by The American Chemical Society.
Special Issue
Published as part of Journal of the American Society for Mass Spectrometryspecial issue “Fenn: Photoactivation and Ion Activation”.
Introduction
Figure 1
Figure 1. (A) NMR solution structure of the model protein ubiquitin used in this study. (B) Schematic of the tandem-trapped ion mobility spectrometer/tandem-mass spectrometer coupling two trapped ion mobility spectrometers (TIMS-1, blue; TIMS-2, red), an ion trap for ion storage, and a UV laser for photodissociation of the stored ions (green). This platform allows characterizing the precursor protein conformations, performing UVPD of a selected precursor conformation, and subsequently characterizing the structures of the produced fragment ions.
Materials and Methods
Materials and Sample Preparation
Tandem-TIMS Measurements
Computational Details
Results and Discussion
Direct Cleavage from the Photoexcited State Predominates UVPD in Tandem-TIMS
Figure 2
Figure 2. Conformationally selective UVPD of ubiquitin in Tandem-TIMS. (A-i) Native ion mobility/mass spectrometry of ubiquitin showing charge states 7+ and 8+. (ii) Charge state 7+ exhibits mainly a native-like conformer (blue), whereas (iii) charge state 8+ shows a dominant native-like conformation (1283 Å2, blue) and a minor gas-phase structure (1947 Å2, orange). (B-i) Conformationally selective top-down mass spectrometry of ubiquitin charge state 8+. (ii) CID produces typical a-, b-, and y-type fragments, whereas UVPD of the (iii) native-like and (iv) gas-phase ubiquitin conformations shows formation of [a + 2], [b + 2], and [y – 2] fragment ions.
Figure 3
Figure 3. y585+and [y58 – 2]5+ fragment ion isotope patterns under various measurement conditions. (A) CID in the TIMS-1/TIMS-2 interface produces y585+ ions exclusively. (B, C) UVPD of the compact and extended conformations produces [y58 – 2]5+ fragment ions but no y585+ ions. (D, E) The [y58 – 2]5+ fragment ions are not observed if the ions (D) are not irradiated by UV photons or (E) are irradiated without storage in the ion trap.
UVPD in Tandem-TIMS Preserves a Structural Relationship between Precursor and Fragment Ions
Figure 4
Figure 4. (A) Ion mobility spectra of CID fragment ions produced from the compact (blue) and extended (orange) ubiquitin conformations agree closely. Hence, the produced fragment ions adopt the same gas-phase favored structures, irrespective of the precursor conformation. (B) Ion mobility spectra for the equivalent fragment ions produced by UVPD from the native-like conformation (blue) differ strongly from those produced from the extended, gas-phase conformation (orange). Here, a compact precursor produces fragments with compact conformations, whereas an extended conformation yields fragments with extended structures. This shows that UVPD can generate fragment ions that exist in a nonannealed, metastable conformation.
UVPD of Native-like Ubiquitin Produces Metastable Fragment Ions
Figure 5
Figure 5. Compact UVPD fragment ions produced from the native-like ubiquitin conformation are metastable. The compact (A) [y40 – 2]4+ and (B) [y58 – 2]5+ ions produced by UVPD from the native-like ubiquitin conformation require substantial activation to overcome an energy barrier and unfold into gas-phase favored conformations. The cross sections of the unfolded, gas-phase conformations are consistent with those of the extended fragment ions produced by UVPD from the extended gas-phase ubiquitin conformation and by CID.
Increased Photon Exposure Increases Fragment Yield but Does Not Change Fragment Conformation
Figure 6
Figure 6. Ion mobility spectra of the [y58 – 2]5+ fragment ion generated by UV photodissociation from mobility-selected ubiquitin 8+ precursors stored for 50–500 ms (≈50–500 laser pulses, colored traces). Control spectra after 140 V collisional activation in TIMS-2 (black trace) and without trapping (black dashed trace) are included for comparison. The data show that increasing irradiation time increases fragment yield but does not alter their collision cross sections.
Energy Redistribution, Collisional Energy Transfer, and Fragment Ion Conformational Dynamics
Figure 7
Figure 7. Langevin dynamics simulations of vibrational energy redistribution, collisional cooling, and structural dynamics of an energetically activated fragment ion. (A) Simulation setup for the y404+ fragment (cyan) with 8.5 eV of excess vibrational energy initially localized at Pro1. The peptide is immersed in a N2 heat bath (red, ∼4 mbar) in a 120 nm × 120 nm × 120 nm cubic box under periodic boundary conditions. (B) Time evolution of the polypeptide vibrational temperature, T(t), showing that intramolecular vibrational energy redistribution completes within ∼100 ps and raises the temperature to ∼350 K. This is followed by collisional cooling that is well described by a single-exponential decay with τ ≈ 680 ns and equilibration with the 300 K bath within a few microseconds. (C) Comparison of the initial (black) and final (blue, 2.75 μs) polypeptide structures shows that >80% of heavy-atom contacts are maintained, indicating some rearrangement in the loop region but no major global restructuring during gas-phase relaxation.
Conclusions
| 1) | UVPD in Tandem-TIMS carried out with low-energy UV laser pulse energies (10 μJ) at elevated pressures (∼4 mbar) produces primarily [b+2] and [y-2] type ions at proline residues. Prior reports (16,48) have established these fragment types as markers of direct bond cleavage from the photoexcited electronic state. | ||||
| 2) | These ‘direct-cleavage’ ions produced by UVPD in Tandem-TIMS preserve a structural relationship between precursor and fragment ions: UVPD of the compact, native-like ubiquitin precursor yields fragment ions that maintain a compact structure, with collision cross sections roughly 20% smaller than the equivalent fragments generated from the extended, gas-phase precursor or ions produced by CID. | ||||
| 3) | The compact “direct-cleavage” ions produced by UVPD from the native-like ubiquitin are kinetically trapped in a metastable conformation, with a substantial energy barrier preventing their annealing into energetically favored gas-phase conformations. | ||||
Supporting Information
The Supporting Information is available free of charge at https://pubs.acs.org/doi/10.1021/jasms.5c00432.
Seven figures with details on tandem-TIMS measurements (PDF)
Terms & Conditions
Most electronic Supporting Information files are available without a subscription to ACS Web Editions. Such files may be downloaded by article for research use (if there is a public use license linked to the relevant article, that license may permit other uses). Permission may be obtained from ACS for other uses through requests via the RightsLink permission system: http://pubs.acs.org/page/copyright/permissions.html.
Acknowledgments
This research was supported by the National Institutes of Health under award R01GM135682 (C.B.) and the National Science Foundation under grant CHE-2305173 (C.B. and F.C.L.). L.N. acknowledges support from a Camille Dreyfus Teacher-Scholar Award (TC-23-050). K.A.M. acknowledges support from the American Chemical Society Bridge Program.
References
This article references 55 other publications.
- 1Brodbelt, J. S. Photodissociation Mass Spectrometry: New Tools for Characterization of Biological Molecules. Chem. Soc. Rev. 2014, 43 (8), 2757– 2783, DOI: 10.1039/C3CS60444FGoogle ScholarThere is no corresponding record for this reference.
- 2Christofi, E.; Barran, P. Ion Mobility Mass Spectrometry (IM-MS) for Structural Biology: Insights Gained by Measuring Mass, Charge, and Collision Cross Section. Chem. Rev. 2023, 123 (6), 2902– 2949, DOI: 10.1021/acs.chemrev.2c00600Google ScholarThere is no corresponding record for this reference.
- 3Bernstein, S. L.; Dupuis, N. F.; Lazo, N. D.; Wyttenbach, T.; Condron, M. M.; Bitan, G.; Teplow, D. B.; Shea, J.-E.; Ruotolo, B. T.; Robinson, C. V.; Bowers, M. T. Amyloid-β Protein Oligomerization and the Importance of Tetramers and Dodecamers in the Aetiology of Alzheimer’s Disease. Nat. Chem. 2009, 1 (4), 326– 331, DOI: 10.1038/nchem.247Google ScholarThere is no corresponding record for this reference.
- 4Bleiholder, C.; Dupuis, N. F.; Wyttenbach, T.; Bowers, M. T. Ion Mobility–Mass Spectrometry Reveals a Conformational Conversion from Random Assembly to β-Sheet in Amyloid Fibril Formation. Nat. Chem. 2011, 3 (2), 172– 177, DOI: 10.1038/nchem.945Google ScholarThere is no corresponding record for this reference.
- 5Zhou, M.; Politis, A.; Davies, R. B.; Liko, I.; Wu, K.-J.; Stewart, A. G.; Stock, D.; Robinson, C. V. Ion Mobility–Mass Spectrometry of a Rotary ATPase Reveals ATP-Induced Reduction in Conformational Flexibility. Nat. Chem. 2014, 6 (3), 208– 215, DOI: 10.1038/nchem.1868Google ScholarThere is no corresponding record for this reference.
- 6Beach, H.; Cole, R.; Gill, M. L.; Loria, J. P. Conservation of Μs–ms Enzyme Motions in the Apo- and Substrate-Mimicked State. J. Am. Chem. Soc. 2005, 127 (25), 9167– 9176, DOI: 10.1021/ja0514949Google ScholarThere is no corresponding record for this reference.
- 7Henzler-Wildman, K. A.; Lei, M.; Thai, V.; Kerns, S. J.; Karplus, M.; Kern, D. A Hierarchy of Timescales in Protein Dynamics Is Linked to Enzyme Catalysis. Nature 2007, 450 (7171), 913– 916, DOI: 10.1038/nature06407Google ScholarThere is no corresponding record for this reference.
- 8Tzeng, S.-R.; Kalodimos, C. G. Dynamic Activation of an Allosteric Regulatory Protein. Nature 2009, 462 (7271), 368– 372, DOI: 10.1038/nature08560Google ScholarThere is no corresponding record for this reference.
- 9Marchand, A.; Czar, M. F.; Eggel, E. N.; Kaeslin, J.; Zenobi, R. Studying Biomolecular Folding and Binding Using Temperature-Jump Mass Spectrometry. Nat. Commun. 2020, 11 (1), 566, DOI: 10.1038/s41467-019-14179-xGoogle ScholarThere is no corresponding record for this reference.
- 10McLuckey, S. A.; Goeringer, D. E. Slow Heating Methods in Tandem Mass Spectrometry. J. Mass Spectrom. 1997, 32, 461– 474, DOI: 10.1002/(SICI)1096-9888(199705)32:5<461::AID-JMS515>3.0.CO;2-HGoogle ScholarThere is no corresponding record for this reference.
- 11Breuker, K.; Oh, H.; Horn, D. M.; Cerda, B. A.; McLafferty, F. W. Detailed Unfolding and Folding of Gaseous Ubiquitin Ions Characterized by Electron Capture Dissociation. J. Am. Chem. Soc. 2002, 124 (22), 6407– 6420, DOI: 10.1021/ja012267jGoogle ScholarThere is no corresponding record for this reference.
- 12Stiving, A. Q.; VanAernum, Z. L.; Busch, F.; Harvey, S. R.; Sarni, S. H.; Wysocki, V. H. Surface-Induced Dissociation: An Effective Method for Characterization of Protein Quaternary Structure. Anal. Chem. 2019, 91 (1), 190– 209, DOI: 10.1021/acs.analchem.8b05071Google ScholarThere is no corresponding record for this reference.
- 13Shaw, J. B.; Li, W.; Holden, D. D.; Zhang, Y.; Griep-Raming, J.; Fellers, R. T.; Early, B. P.; Thomas, P. M.; Kelleher, N. L.; Brodbelt, J. S. Complete Protein Characterization Using Top-Down Mass Spectrometry and Ultraviolet Photodissociation. J. Am. Chem. Soc. 2013, 135 (34), 12646– 12651, DOI: 10.1021/ja4029654Google ScholarThere is no corresponding record for this reference.
- 14Cammarata, M. B.; Thyer, R.; Rosenberg, J.; Ellington, A.; Brodbelt, J. S. Structural Characterization of Dihydrofolate Reductase Complexes by Top-Down Ultraviolet Photodissociation Mass Spectrometry. J. Am. Chem. Soc. 2015, 137 (28), 9128– 9135, DOI: 10.1021/jacs.5b04628Google ScholarThere is no corresponding record for this reference.
- 15Morrison, L. J.; Brodbelt, J. S. 193 Nm Ultraviolet Photodissociation Mass Spectrometry of Tetrameric Protein Complexes Provides Insight into Quaternary and Secondary Protein Topology. J. Am. Chem. Soc. 2016, 138 (34), 10849– 10859, DOI: 10.1021/jacs.6b03905Google ScholarThere is no corresponding record for this reference.
- 16Girod, M.; Sanader, Z.; Vojkovic, M.; Antoine, R.; MacAleese, L.; Lemoine, J.; Bonacic-Koutecky, V.; Dugourd, P. UV Photodissociation of Proline-Containing Peptide Ions: Insights from Molecular Dynamics. J. Am. Soc. Mass Spectrom. 2015, 26 (3), 432– 443, DOI: 10.1007/s13361-014-1038-1Google ScholarThere is no corresponding record for this reference.
- 17Zhang, L.; Reilly, J. P. Peptide Photodissociation with 157 Nm Light in a Commercial Tandem Time-of-Flight Mass Spectrometer. Anal. Chem. 2009, 81 (18), 7829– 7838, DOI: 10.1021/ac9012557Google ScholarThere is no corresponding record for this reference.
- 18Cammarata, M. B.; Brodbelt, J. S. Structural Characterization of Holo- and Apo-Myoglobin in the Gas Phase by Ultraviolet Photodissociation Mass Spectrometry. Chem. Sci. 2015, 6 (2), 1324– 1333, DOI: 10.1039/C4SC03200DGoogle ScholarThere is no corresponding record for this reference.
- 19Cammarata, M.; Thyer, R.; Lombardo, M.; Anderson, A.; Wright, D.; Ellington, A.; Brodbelt, J. S. Characterization of Trimethoprim Resistant E. Coli Dihydrofolate Reductase Mutants by Mass Spectrometry and Inhibition by Propargyl-Linked Antifolates. Chem. Sci. 2017, 8 (5), 4062– 4072, DOI: 10.1039/C6SC05235EGoogle ScholarThere is no corresponding record for this reference.
- 20Butalewicz, J. P.; Escobar, E. E.; Wootton, C. A.; Theisen, A.; Park, M. A.; Seeley, E. H.; Brodbelt, J. S. Conformational Characterization of Peptides and Proteins by 193 Nm Ultraviolet Photodissociation in the Collision Cell of a Trapped Ion Mobility Spectrometry-Time-of-Flight Mass Spectrometer. Anal. Chem. 2024, 96 (41), 16154– 16161, DOI: 10.1021/acs.analchem.4c02686Google ScholarThere is no corresponding record for this reference.
- 21Butalewicz, J. P.; Sanders, J. D.; Juetten, K. J.; Buzitis, N. W.; Clowers, B. H.; Brodbelt, J. S. Advancing Protein Analysis: A Low-Pressure Drift Tube Orbitrap Mass Spectrometer for Ultraviolet Photodissociation-Based Structural Characterization. Anal. Chem. 2024, 96 (39), 15674– 15681, DOI: 10.1021/acs.analchem.4c03119Google ScholarThere is no corresponding record for this reference.
- 22Theisen, A.; Black, R.; Corinti, D.; Brown, J. M.; Bellina, B.; Barran, P. E. Initial Protein Unfolding Events in Ubiquitin, Cytochrome c and Myoglobin Are Revealed with the Use of 213 Nm UVPD Coupled to IM-MS. J. Am. Soc. Mass Spectrom. 2019, 30 (1), 24– 33, DOI: 10.1007/s13361-018-1992-0Google ScholarThere is no corresponding record for this reference.
- 23Liu, F. C.; Ridgeway, M. E.; Winfred, J. S. R. V.; Polfer, N. C.; Lee, J.; Theisen, A.; Wootton, C. A.; Park, M. A.; Bleiholder, C. Tandem-Trapped Ion Mobility Spectrometry/Mass Spectrometry Coupled with Ultraviolet Photodissociation. Rapid Commun. Mass Spectrom. 2021, 35, e9192 DOI: 10.1002/rcm.9192Google ScholarThere is no corresponding record for this reference.
- 24Liu, F. C.; Ridgeway, M. E.; Park, M. A.; Bleiholder, C. Tandem Trapped Ion Mobility Spectrometry. Analyst 2018, 143 (10), 2249– 2258, DOI: 10.1039/C7AN02054FGoogle ScholarThere is no corresponding record for this reference.
- 25Vijay-Kumar, S.; Bugg, C. E.; Wilkinson, K. D.; Cook, W. J. Three-Dimensional Structure of Ubiquitin at 2.8 A Resolution. Proc. Natl. Acad. Sci. U. S. A. 1985, 82 (11), 3582– 3585, DOI: 10.1073/pnas.82.11.3582Google ScholarThere is no corresponding record for this reference.
- 26Lange, O. F.; Lakomek, N.-A.; Farès, C.; Schröder, G. F.; Walter, K. F. A.; Becker, S.; Meiler, J.; Grubmüller, H.; Griesinger, C.; de Groot, B. L. Recognition Dynamics Up to Microseconds Revealed from an RDC-Derived Ubiquitin Ensemble in Solution. Science 2008, 320 (5882), 1471– 1475, DOI: 10.1126/science.1157092Google ScholarThere is no corresponding record for this reference.
- 27Jourdan, M.; Searle, M. S. Insights into the Stability of Native and Partially Folded States of Ubiquitin: Effects of Cosolvents and Denaturants on the Thermodynamics of Protein Folding. Biochemistry 2001, 40 (34), 10317– 10325, DOI: 10.1021/bi010767jGoogle ScholarThere is no corresponding record for this reference.
- 28Chai, M.; Young, M. N.; Liu, F. C.; Bleiholder, C. A Transferable, Sample-Independent Calibration Procedure for Trapped Ion Mobility Spectrometry (TIMS). Anal. Chem. 2018, 90 (15), 9040– 9047, DOI: 10.1021/acs.analchem.8b01326Google ScholarThere is no corresponding record for this reference.
- 29Hernandez, D. R.; DeBord, J. D.; Ridgeway, M. E.; Kaplan, D. A.; Park, M. A.; Fernandez-Lima, F. Ion Dynamics in a Trapped Ion Mobility Spectrometer. Analyst 2014, 139 (8), 1913– 1921, DOI: 10.1039/C3AN02174BGoogle ScholarThere is no corresponding record for this reference.
- 30Liu, F. C.; Ridgeway, M. E.; Wootton, C. A.; Theisen, A.; Panczyk, E. M.; Meier, F.; Park, M. A.; Bleiholder, C. Top-Down Protein Analysis by Tandem-Trapped Ion Mobility Spectrometry/Mass Spectrometry (Tandem-TIMS/MS) Coupled with Ultraviolet Photodissociation (UVPD) and Parallel Accumulation/Serial Fragmentation (PASEF) MS/MS Analysis. J. Am. Soc. Mass Spectrom. 2023, 34 (10), 2232– 2246, DOI: 10.1021/jasms.3c00187Google ScholarThere is no corresponding record for this reference.
- 31Fernandez-Lima, F. A.; Kaplan, D. A.; Park, M. A. Note: Integration of Trapped Ion Mobility Spectrometry with Mass Spectrometry. Rev. Sci. Instrum. 2011, 82 (12), 126106, DOI: 10.1063/1.3665933Google ScholarThere is no corresponding record for this reference.
- 32Liu, F. C.; Kirk, S. R.; Bleiholder, C. On the Structural Denaturation of Biological Analytes in Trapped Ion Mobility Spectrometry – Mass Spectrometry. Analyst 2016, 141 (12), 3722– 3730, DOI: 10.1039/C5AN02399HGoogle ScholarThere is no corresponding record for this reference.
- 33Liu, F. C.; Cropley, T. C.; Ridgeway, M. E.; Park, M. A.; Bleiholder, C. Structural Analysis of the Glycoprotein Complex Avidin by Tandem-Trapped Ion Mobility Spectrometry–Mass Spectrometry (Tandem-TIMS/MS). Anal. Chem. 2020, 92 (6), 4459– 4467, DOI: 10.1021/acs.analchem.9b05481Google ScholarThere is no corresponding record for this reference.
- 34Cropley, T. C.; Liu, F. C.; Chai, M.; Bush, M. F.; Bleiholder, C. Metastability of Protein Solution Structures in the Absence of a Solvent: Rugged Energy Landscape and Glass-like Behavior. J. Am. Chem. Soc. 2024, 146, 11115– 11125, DOI: 10.1021/jacs.3c12892Google ScholarThere is no corresponding record for this reference.
- 35Borotto, N. B.; Graham, K. A. Fragmentation and Mobility Separation of Peptide and Protein Ions in a Trapped-Ion Mobility Device. Anal. Chem. 2021, 93 (29), 9959– 9964, DOI: 10.1021/acs.analchem.1c01188Google ScholarThere is no corresponding record for this reference.
- 36Silveira, J. A.; Ridgeway, M. E.; Park, M. A. High Resolution Trapped Ion Mobility Spectrometery of Peptides. Anal. Chem. 2014, 86 (12), 5624– 5627, DOI: 10.1021/ac501261hGoogle ScholarThere is no corresponding record for this reference.
- 37Stow, S. M.; Causon, T. J.; Zheng, X.; Kurulugama, R. T.; Mairinger, T.; May, J. C.; Rennie, E. E.; Baker, E. S.; Smith, R. D.; McLean, J. A.; Hann, S.; Fjeldsted, J. C. An Interlaboratory Evaluation of Drift Tube Ion Mobility–Mass Spectrometry Collision Cross Section Measurements. Anal. Chem. 2017, 89 (17), 9048– 9055, DOI: 10.1021/acs.analchem.7b01729Google ScholarThere is no corresponding record for this reference.
- 38Loos, M.; Gerber, C.; Corona, F.; Hollender, J.; Singer, H. Accelerated Isotope Fine Structure Calculation Using Pruned Transition Trees. Anal. Chem. 2015, 87 (11), 5738– 5744, DOI: 10.1021/acs.analchem.5b00941Google ScholarThere is no corresponding record for this reference.
- 39Berendsen, H. J. C.; van der Spoel, D.; van Drunen, R. GROMACS: A Message-Passing Parallel Molecular Dynamics Implementation. Comput. Phys. Commun. 1995, 91 (1–3), 43– 56, DOI: 10.1016/0010-4655(95)00042-EGoogle ScholarThere is no corresponding record for this reference.
- 40Kaminski, G. A.; Friesner, R. A.; Tirado-Rives, J.; Jorgensen, W. L. Evaluation and Reparametrization of the OPLS-AA Force Field for Proteins via Comparison with Accurate Quantum Chemical Calculations on Peptides. J. Phys. Chem. B 2001, 105 (28), 6474– 6487, DOI: 10.1021/jp003919dGoogle ScholarThere is no corresponding record for this reference.
- 41Wang, S.; Hou, K.; Heinz, H. Accurate and Compatible Force Fields for Molecular Oxygen, Nitrogen, and Hydrogen to Simulate Gases, Electrolytes, and Heterogeneous Interfaces. J. Chem. Theory Comput. 2021, 17 (8), 5198– 5213, DOI: 10.1021/acs.jctc.0c01132Google ScholarThere is no corresponding record for this reference.
- 42Wyttenbach, T.; Bowers, M. T. Structural Stability from Solution to the Gas Phase: Native Solution Structure of Ubiquitin Survives Analysis in a Solvent-Free Ion Mobility–Mass Spectrometry Environment. J. Phys. Chem. B 2011, 115 (42), 12266– 12275, DOI: 10.1021/jp206867aGoogle ScholarThere is no corresponding record for this reference.
- 43Laszlo, K. J.; Munger, E. B.; Bush, M. F. Folding of Protein Ions in the Gas Phase after Cation-to-Anion Proton-Transfer Reactions. J. Am. Chem. Soc. 2016, 138 (30), 9581– 9588, DOI: 10.1021/jacs.6b04282Google ScholarThere is no corresponding record for this reference.
- 44Warnke, S.; Von Helden, G.; Pagel, K. Protein Structure in the Gas Phase: The Influence of Side-Chain Microsolvation. J. Am. Chem. Soc. 2013, 135 (4), 1177– 1180, DOI: 10.1021/ja308528dGoogle ScholarThere is no corresponding record for this reference.
- 45Liu, F. C.; Kirk, S. R.; Caldwell, K. A.; Pedrete, T.; Meier, F.; Bleiholder, C. Tandem Trapped Ion Mobility Spectrometry/Mass Spectrometry (tTIMS/MS) Reveals Sequence-Specific Determinants of Top-Down Protein Fragment Ion Cross Sections. Anal. Chem. 2022, 94 (23), 8146– 8155, DOI: 10.1021/acs.analchem.1c05171Google ScholarThere is no corresponding record for this reference.
- 46Warnke, S.; Baldauf, C.; Bowers, M. T.; Pagel, K.; von Helden, G. Photodissociation of Conformer-Selected Ubiquitin Ions Reveals Site-Specific Cis/Trans Isomerization of Proline Peptide Bonds. J. Am. Chem. Soc. 2014, 136 (29), 10308– 10314, DOI: 10.1021/ja502994bGoogle ScholarThere is no corresponding record for this reference.
- 47Fornelli, L.; Srzentić, K.; Toby, T. K.; Doubleday, P. F.; Huguet, R.; Mullen, C.; Melani, R. D.; dos Santos Seckler, H.; DeHart, C. J.; Weisbrod, C. R.; Durbin, K. R.; Greer, J. B.; Early, B. P.; Fellers, R. T.; Zabrouskov, V.; Thomas, P. M.; Compton, P. D.; Kelleher, N. L. Thorough Performance Evaluation of 213 Nm Ultraviolet Photodissociation for Top-down Proteomics. Mol. Cell. Proteomics 2020, 19 (2), 405– 420, DOI: 10.1074/mcp.TIR119.001638Google ScholarThere is no corresponding record for this reference.
- 48Devakumar, A.; Thompson, M. S.; Reilly, J. P. Fragmentation of Oligosaccharide Ions with 157 Nm Vacuum Ultraviolet Light. Rapid Commun. Mass Spectrom. 2005, 19 (16), 2313– 2320, DOI: 10.1002/rcm.2058Google ScholarThere is no corresponding record for this reference.
- 49Shelimov, K. B.; Jarrold, M. F. Conformations, Unfolding, and Refolding of Apomyoglobin in Vacuum: An Activation Barrier for Gas-Phase Protein Folding. J. Am. Chem. Soc. 1997, 119 (13), 2987– 2994, DOI: 10.1021/ja962914kGoogle ScholarThere is no corresponding record for this reference.
- 50Julian, R. R. The Mechanism Behind Top-Down UVPD Experiments: Making Sense of Apparent Contradictions. J. Am. Soc. Mass Spectrom. 2017, 28 (9), 1823– 1826, DOI: 10.1007/s13361-017-1721-0Google ScholarThere is no corresponding record for this reference.
- 51Brodbelt, J. S.; Morrison, L. J.; Santos, I. Ultraviolet Photodissociation Mass Spectrometry for Analysis of Biological Molecules. Chem. Rev. 2020, 120 (7), 3328– 3380, DOI: 10.1021/acs.chemrev.9b00440Google ScholarThere is no corresponding record for this reference.
- 52Kuipers, B. J. H.; Gruppen, H. Prediction of Molar Extinction Coefficients of Proteins and Peptides Using UV Absorption of the Constituent Amino Acids at 214 Nm To Enable Quantitative Reverse Phase High-Performance Liquid Chromatography–Mass Spectrometry Analysis. J. Agric. Food Chem. 2007, 55 (14), 5445– 5451, DOI: 10.1021/jf070337lGoogle ScholarThere is no corresponding record for this reference.
- 53Grégoire, G.; Jouvet, C.; Dedonder, C.; Sobolewski, A. L. Ab Initio Study of the Excited-State Deactivation Pathways of Protonated Tryptophan and Tyrosine. J. Am. Chem. Soc. 2007, 129 (19), 6223– 6231, DOI: 10.1021/ja069050fGoogle ScholarThere is no corresponding record for this reference.
- 54Paizs, B.; Suhai, S. Fragmentation Pathways of Protonated Peptides. Mass Spectrom. Rev. 2005, 24 (4), 508– 548, DOI: 10.1002/mas.20024Google ScholarThere is no corresponding record for this reference.
- 55Bleiholder, C.; Osburn, S.; Williams, T. D.; Suhai, S.; Van Stipdonk, M.; Harrison, A. G.; Paizs, B. Sequence-Scrambling Fragmentation Pathways of Protonated Peptides. J. Am. Chem. Soc. 2008, 130 (52), 17774– 17789, DOI: 10.1021/ja805074dGoogle ScholarThere is no corresponding record for this reference.
Cited By
This article has not yet been cited by other publications.
Article Views
Altmetric
Citations
Article Views are the COUNTER-compliant sum of full text article downloads since November 2008 (both PDF and HTML) across all institutions and individuals. These metrics are regularly updated to reflect usage leading up to the last few days.
Citations are the number of other articles citing this article, calculated by Crossref and updated daily. Find more information about Crossref citation counts.
The Altmetric Attention Score is a quantitative measure of the attention that a research article has received online. Clicking on the donut icon will load a page at altmetric.com with additional details about the score and the social media presence for the given article. Find more information on the Altmetric Attention Score and how the score is calculated.
Recommended Articles
Abstract
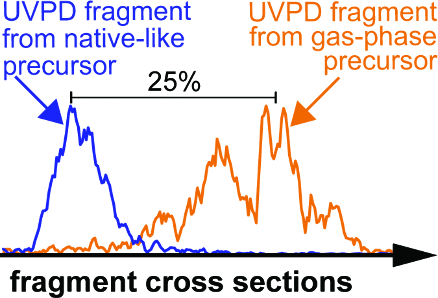
Figure 1
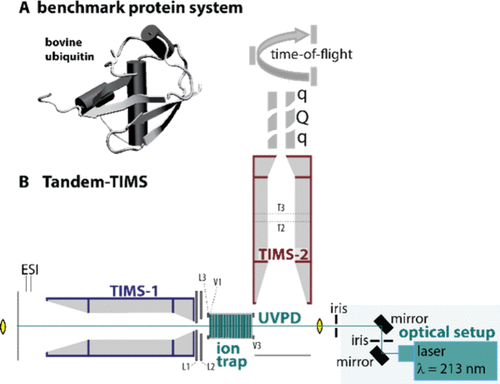
Figure 1. (A) NMR solution structure of the model protein ubiquitin used in this study. (B) Schematic of the tandem-trapped ion mobility spectrometer/tandem-mass spectrometer coupling two trapped ion mobility spectrometers (TIMS-1, blue; TIMS-2, red), an ion trap for ion storage, and a UV laser for photodissociation of the stored ions (green). This platform allows characterizing the precursor protein conformations, performing UVPD of a selected precursor conformation, and subsequently characterizing the structures of the produced fragment ions.
Figure 2
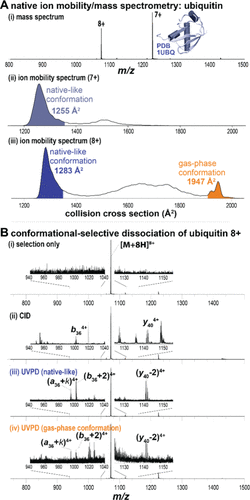
Figure 2. Conformationally selective UVPD of ubiquitin in Tandem-TIMS. (A-i) Native ion mobility/mass spectrometry of ubiquitin showing charge states 7+ and 8+. (ii) Charge state 7+ exhibits mainly a native-like conformer (blue), whereas (iii) charge state 8+ shows a dominant native-like conformation (1283 Å2, blue) and a minor gas-phase structure (1947 Å2, orange). (B-i) Conformationally selective top-down mass spectrometry of ubiquitin charge state 8+. (ii) CID produces typical a-, b-, and y-type fragments, whereas UVPD of the (iii) native-like and (iv) gas-phase ubiquitin conformations shows formation of [a + 2], [b + 2], and [y – 2] fragment ions.
Figure 3
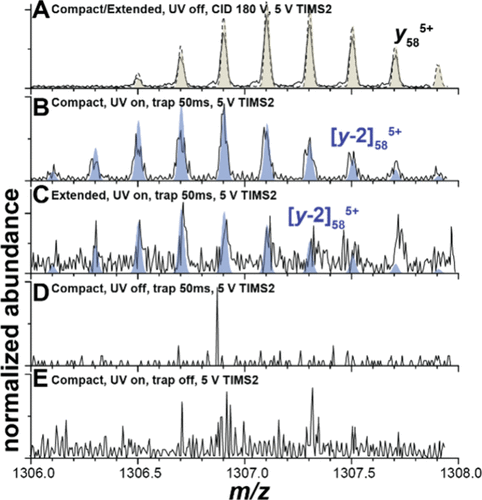
Figure 3. y585+and [y58 – 2]5+ fragment ion isotope patterns under various measurement conditions. (A) CID in the TIMS-1/TIMS-2 interface produces y585+ ions exclusively. (B, C) UVPD of the compact and extended conformations produces [y58 – 2]5+ fragment ions but no y585+ ions. (D, E) The [y58 – 2]5+ fragment ions are not observed if the ions (D) are not irradiated by UV photons or (E) are irradiated without storage in the ion trap.
Figure 4
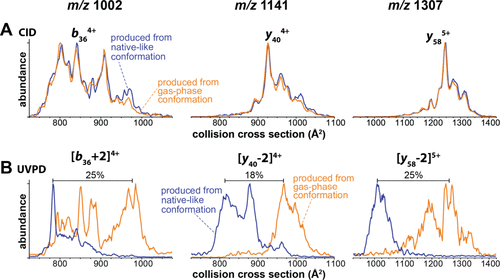
Figure 4. (A) Ion mobility spectra of CID fragment ions produced from the compact (blue) and extended (orange) ubiquitin conformations agree closely. Hence, the produced fragment ions adopt the same gas-phase favored structures, irrespective of the precursor conformation. (B) Ion mobility spectra for the equivalent fragment ions produced by UVPD from the native-like conformation (blue) differ strongly from those produced from the extended, gas-phase conformation (orange). Here, a compact precursor produces fragments with compact conformations, whereas an extended conformation yields fragments with extended structures. This shows that UVPD can generate fragment ions that exist in a nonannealed, metastable conformation.
Figure 5
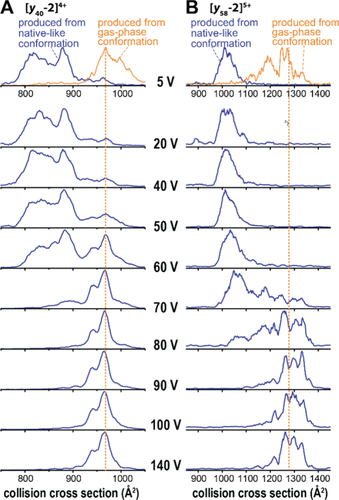
Figure 5. Compact UVPD fragment ions produced from the native-like ubiquitin conformation are metastable. The compact (A) [y40 – 2]4+ and (B) [y58 – 2]5+ ions produced by UVPD from the native-like ubiquitin conformation require substantial activation to overcome an energy barrier and unfold into gas-phase favored conformations. The cross sections of the unfolded, gas-phase conformations are consistent with those of the extended fragment ions produced by UVPD from the extended gas-phase ubiquitin conformation and by CID.
Figure 6
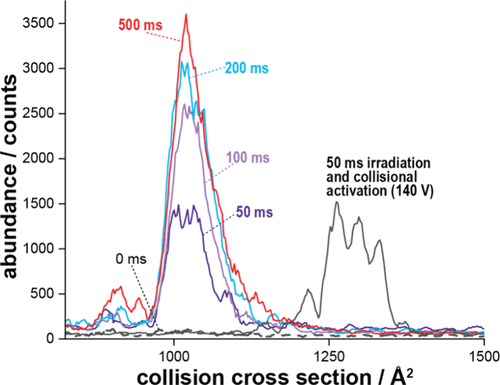
Figure 6. Ion mobility spectra of the [y58 – 2]5+ fragment ion generated by UV photodissociation from mobility-selected ubiquitin 8+ precursors stored for 50–500 ms (≈50–500 laser pulses, colored traces). Control spectra after 140 V collisional activation in TIMS-2 (black trace) and without trapping (black dashed trace) are included for comparison. The data show that increasing irradiation time increases fragment yield but does not alter their collision cross sections.
Figure 7
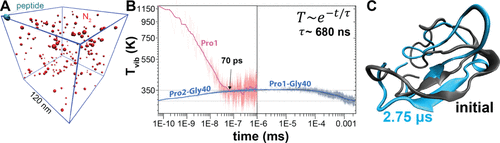
Figure 7. Langevin dynamics simulations of vibrational energy redistribution, collisional cooling, and structural dynamics of an energetically activated fragment ion. (A) Simulation setup for the y404+ fragment (cyan) with 8.5 eV of excess vibrational energy initially localized at Pro1. The peptide is immersed in a N2 heat bath (red, ∼4 mbar) in a 120 nm × 120 nm × 120 nm cubic box under periodic boundary conditions. (B) Time evolution of the polypeptide vibrational temperature, T(t), showing that intramolecular vibrational energy redistribution completes within ∼100 ps and raises the temperature to ∼350 K. This is followed by collisional cooling that is well described by a single-exponential decay with τ ≈ 680 ns and equilibration with the 300 K bath within a few microseconds. (C) Comparison of the initial (black) and final (blue, 2.75 μs) polypeptide structures shows that >80% of heavy-atom contacts are maintained, indicating some rearrangement in the loop region but no major global restructuring during gas-phase relaxation.
References
This article references 55 other publications.
- 1Brodbelt, J. S. Photodissociation Mass Spectrometry: New Tools for Characterization of Biological Molecules. Chem. Soc. Rev. 2014, 43 (8), 2757– 2783, DOI: 10.1039/C3CS60444FThere is no corresponding record for this reference.
- 2Christofi, E.; Barran, P. Ion Mobility Mass Spectrometry (IM-MS) for Structural Biology: Insights Gained by Measuring Mass, Charge, and Collision Cross Section. Chem. Rev. 2023, 123 (6), 2902– 2949, DOI: 10.1021/acs.chemrev.2c00600There is no corresponding record for this reference.
- 3Bernstein, S. L.; Dupuis, N. F.; Lazo, N. D.; Wyttenbach, T.; Condron, M. M.; Bitan, G.; Teplow, D. B.; Shea, J.-E.; Ruotolo, B. T.; Robinson, C. V.; Bowers, M. T. Amyloid-β Protein Oligomerization and the Importance of Tetramers and Dodecamers in the Aetiology of Alzheimer’s Disease. Nat. Chem. 2009, 1 (4), 326– 331, DOI: 10.1038/nchem.247There is no corresponding record for this reference.
- 4Bleiholder, C.; Dupuis, N. F.; Wyttenbach, T.; Bowers, M. T. Ion Mobility–Mass Spectrometry Reveals a Conformational Conversion from Random Assembly to β-Sheet in Amyloid Fibril Formation. Nat. Chem. 2011, 3 (2), 172– 177, DOI: 10.1038/nchem.945There is no corresponding record for this reference.
- 5Zhou, M.; Politis, A.; Davies, R. B.; Liko, I.; Wu, K.-J.; Stewart, A. G.; Stock, D.; Robinson, C. V. Ion Mobility–Mass Spectrometry of a Rotary ATPase Reveals ATP-Induced Reduction in Conformational Flexibility. Nat. Chem. 2014, 6 (3), 208– 215, DOI: 10.1038/nchem.1868There is no corresponding record for this reference.
- 6Beach, H.; Cole, R.; Gill, M. L.; Loria, J. P. Conservation of Μs–ms Enzyme Motions in the Apo- and Substrate-Mimicked State. J. Am. Chem. Soc. 2005, 127 (25), 9167– 9176, DOI: 10.1021/ja0514949There is no corresponding record for this reference.
- 7Henzler-Wildman, K. A.; Lei, M.; Thai, V.; Kerns, S. J.; Karplus, M.; Kern, D. A Hierarchy of Timescales in Protein Dynamics Is Linked to Enzyme Catalysis. Nature 2007, 450 (7171), 913– 916, DOI: 10.1038/nature06407There is no corresponding record for this reference.
- 8Tzeng, S.-R.; Kalodimos, C. G. Dynamic Activation of an Allosteric Regulatory Protein. Nature 2009, 462 (7271), 368– 372, DOI: 10.1038/nature08560There is no corresponding record for this reference.
- 9Marchand, A.; Czar, M. F.; Eggel, E. N.; Kaeslin, J.; Zenobi, R. Studying Biomolecular Folding and Binding Using Temperature-Jump Mass Spectrometry. Nat. Commun. 2020, 11 (1), 566, DOI: 10.1038/s41467-019-14179-xThere is no corresponding record for this reference.
- 10McLuckey, S. A.; Goeringer, D. E. Slow Heating Methods in Tandem Mass Spectrometry. J. Mass Spectrom. 1997, 32, 461– 474, DOI: 10.1002/(SICI)1096-9888(199705)32:5<461::AID-JMS515>3.0.CO;2-HThere is no corresponding record for this reference.
- 11Breuker, K.; Oh, H.; Horn, D. M.; Cerda, B. A.; McLafferty, F. W. Detailed Unfolding and Folding of Gaseous Ubiquitin Ions Characterized by Electron Capture Dissociation. J. Am. Chem. Soc. 2002, 124 (22), 6407– 6420, DOI: 10.1021/ja012267jThere is no corresponding record for this reference.
- 12Stiving, A. Q.; VanAernum, Z. L.; Busch, F.; Harvey, S. R.; Sarni, S. H.; Wysocki, V. H. Surface-Induced Dissociation: An Effective Method for Characterization of Protein Quaternary Structure. Anal. Chem. 2019, 91 (1), 190– 209, DOI: 10.1021/acs.analchem.8b05071There is no corresponding record for this reference.
- 13Shaw, J. B.; Li, W.; Holden, D. D.; Zhang, Y.; Griep-Raming, J.; Fellers, R. T.; Early, B. P.; Thomas, P. M.; Kelleher, N. L.; Brodbelt, J. S. Complete Protein Characterization Using Top-Down Mass Spectrometry and Ultraviolet Photodissociation. J. Am. Chem. Soc. 2013, 135 (34), 12646– 12651, DOI: 10.1021/ja4029654There is no corresponding record for this reference.
- 14Cammarata, M. B.; Thyer, R.; Rosenberg, J.; Ellington, A.; Brodbelt, J. S. Structural Characterization of Dihydrofolate Reductase Complexes by Top-Down Ultraviolet Photodissociation Mass Spectrometry. J. Am. Chem. Soc. 2015, 137 (28), 9128– 9135, DOI: 10.1021/jacs.5b04628There is no corresponding record for this reference.
- 15Morrison, L. J.; Brodbelt, J. S. 193 Nm Ultraviolet Photodissociation Mass Spectrometry of Tetrameric Protein Complexes Provides Insight into Quaternary and Secondary Protein Topology. J. Am. Chem. Soc. 2016, 138 (34), 10849– 10859, DOI: 10.1021/jacs.6b03905There is no corresponding record for this reference.
- 16Girod, M.; Sanader, Z.; Vojkovic, M.; Antoine, R.; MacAleese, L.; Lemoine, J.; Bonacic-Koutecky, V.; Dugourd, P. UV Photodissociation of Proline-Containing Peptide Ions: Insights from Molecular Dynamics. J. Am. Soc. Mass Spectrom. 2015, 26 (3), 432– 443, DOI: 10.1007/s13361-014-1038-1There is no corresponding record for this reference.
- 17Zhang, L.; Reilly, J. P. Peptide Photodissociation with 157 Nm Light in a Commercial Tandem Time-of-Flight Mass Spectrometer. Anal. Chem. 2009, 81 (18), 7829– 7838, DOI: 10.1021/ac9012557There is no corresponding record for this reference.
- 18Cammarata, M. B.; Brodbelt, J. S. Structural Characterization of Holo- and Apo-Myoglobin in the Gas Phase by Ultraviolet Photodissociation Mass Spectrometry. Chem. Sci. 2015, 6 (2), 1324– 1333, DOI: 10.1039/C4SC03200DThere is no corresponding record for this reference.
- 19Cammarata, M.; Thyer, R.; Lombardo, M.; Anderson, A.; Wright, D.; Ellington, A.; Brodbelt, J. S. Characterization of Trimethoprim Resistant E. Coli Dihydrofolate Reductase Mutants by Mass Spectrometry and Inhibition by Propargyl-Linked Antifolates. Chem. Sci. 2017, 8 (5), 4062– 4072, DOI: 10.1039/C6SC05235EThere is no corresponding record for this reference.
- 20Butalewicz, J. P.; Escobar, E. E.; Wootton, C. A.; Theisen, A.; Park, M. A.; Seeley, E. H.; Brodbelt, J. S. Conformational Characterization of Peptides and Proteins by 193 Nm Ultraviolet Photodissociation in the Collision Cell of a Trapped Ion Mobility Spectrometry-Time-of-Flight Mass Spectrometer. Anal. Chem. 2024, 96 (41), 16154– 16161, DOI: 10.1021/acs.analchem.4c02686There is no corresponding record for this reference.
- 21Butalewicz, J. P.; Sanders, J. D.; Juetten, K. J.; Buzitis, N. W.; Clowers, B. H.; Brodbelt, J. S. Advancing Protein Analysis: A Low-Pressure Drift Tube Orbitrap Mass Spectrometer for Ultraviolet Photodissociation-Based Structural Characterization. Anal. Chem. 2024, 96 (39), 15674– 15681, DOI: 10.1021/acs.analchem.4c03119There is no corresponding record for this reference.
- 22Theisen, A.; Black, R.; Corinti, D.; Brown, J. M.; Bellina, B.; Barran, P. E. Initial Protein Unfolding Events in Ubiquitin, Cytochrome c and Myoglobin Are Revealed with the Use of 213 Nm UVPD Coupled to IM-MS. J. Am. Soc. Mass Spectrom. 2019, 30 (1), 24– 33, DOI: 10.1007/s13361-018-1992-0There is no corresponding record for this reference.
- 23Liu, F. C.; Ridgeway, M. E.; Winfred, J. S. R. V.; Polfer, N. C.; Lee, J.; Theisen, A.; Wootton, C. A.; Park, M. A.; Bleiholder, C. Tandem-Trapped Ion Mobility Spectrometry/Mass Spectrometry Coupled with Ultraviolet Photodissociation. Rapid Commun. Mass Spectrom. 2021, 35, e9192 DOI: 10.1002/rcm.9192There is no corresponding record for this reference.
- 24Liu, F. C.; Ridgeway, M. E.; Park, M. A.; Bleiholder, C. Tandem Trapped Ion Mobility Spectrometry. Analyst 2018, 143 (10), 2249– 2258, DOI: 10.1039/C7AN02054FThere is no corresponding record for this reference.
- 25Vijay-Kumar, S.; Bugg, C. E.; Wilkinson, K. D.; Cook, W. J. Three-Dimensional Structure of Ubiquitin at 2.8 A Resolution. Proc. Natl. Acad. Sci. U. S. A. 1985, 82 (11), 3582– 3585, DOI: 10.1073/pnas.82.11.3582There is no corresponding record for this reference.
- 26Lange, O. F.; Lakomek, N.-A.; Farès, C.; Schröder, G. F.; Walter, K. F. A.; Becker, S.; Meiler, J.; Grubmüller, H.; Griesinger, C.; de Groot, B. L. Recognition Dynamics Up to Microseconds Revealed from an RDC-Derived Ubiquitin Ensemble in Solution. Science 2008, 320 (5882), 1471– 1475, DOI: 10.1126/science.1157092There is no corresponding record for this reference.
- 27Jourdan, M.; Searle, M. S. Insights into the Stability of Native and Partially Folded States of Ubiquitin: Effects of Cosolvents and Denaturants on the Thermodynamics of Protein Folding. Biochemistry 2001, 40 (34), 10317– 10325, DOI: 10.1021/bi010767jThere is no corresponding record for this reference.
- 28Chai, M.; Young, M. N.; Liu, F. C.; Bleiholder, C. A Transferable, Sample-Independent Calibration Procedure for Trapped Ion Mobility Spectrometry (TIMS). Anal. Chem. 2018, 90 (15), 9040– 9047, DOI: 10.1021/acs.analchem.8b01326There is no corresponding record for this reference.
- 29Hernandez, D. R.; DeBord, J. D.; Ridgeway, M. E.; Kaplan, D. A.; Park, M. A.; Fernandez-Lima, F. Ion Dynamics in a Trapped Ion Mobility Spectrometer. Analyst 2014, 139 (8), 1913– 1921, DOI: 10.1039/C3AN02174BThere is no corresponding record for this reference.
- 30Liu, F. C.; Ridgeway, M. E.; Wootton, C. A.; Theisen, A.; Panczyk, E. M.; Meier, F.; Park, M. A.; Bleiholder, C. Top-Down Protein Analysis by Tandem-Trapped Ion Mobility Spectrometry/Mass Spectrometry (Tandem-TIMS/MS) Coupled with Ultraviolet Photodissociation (UVPD) and Parallel Accumulation/Serial Fragmentation (PASEF) MS/MS Analysis. J. Am. Soc. Mass Spectrom. 2023, 34 (10), 2232– 2246, DOI: 10.1021/jasms.3c00187There is no corresponding record for this reference.
- 31Fernandez-Lima, F. A.; Kaplan, D. A.; Park, M. A. Note: Integration of Trapped Ion Mobility Spectrometry with Mass Spectrometry. Rev. Sci. Instrum. 2011, 82 (12), 126106, DOI: 10.1063/1.3665933There is no corresponding record for this reference.
- 32Liu, F. C.; Kirk, S. R.; Bleiholder, C. On the Structural Denaturation of Biological Analytes in Trapped Ion Mobility Spectrometry – Mass Spectrometry. Analyst 2016, 141 (12), 3722– 3730, DOI: 10.1039/C5AN02399HThere is no corresponding record for this reference.
- 33Liu, F. C.; Cropley, T. C.; Ridgeway, M. E.; Park, M. A.; Bleiholder, C. Structural Analysis of the Glycoprotein Complex Avidin by Tandem-Trapped Ion Mobility Spectrometry–Mass Spectrometry (Tandem-TIMS/MS). Anal. Chem. 2020, 92 (6), 4459– 4467, DOI: 10.1021/acs.analchem.9b05481There is no corresponding record for this reference.
- 34Cropley, T. C.; Liu, F. C.; Chai, M.; Bush, M. F.; Bleiholder, C. Metastability of Protein Solution Structures in the Absence of a Solvent: Rugged Energy Landscape and Glass-like Behavior. J. Am. Chem. Soc. 2024, 146, 11115– 11125, DOI: 10.1021/jacs.3c12892There is no corresponding record for this reference.
- 35Borotto, N. B.; Graham, K. A. Fragmentation and Mobility Separation of Peptide and Protein Ions in a Trapped-Ion Mobility Device. Anal. Chem. 2021, 93 (29), 9959– 9964, DOI: 10.1021/acs.analchem.1c01188There is no corresponding record for this reference.
- 36Silveira, J. A.; Ridgeway, M. E.; Park, M. A. High Resolution Trapped Ion Mobility Spectrometery of Peptides. Anal. Chem. 2014, 86 (12), 5624– 5627, DOI: 10.1021/ac501261hThere is no corresponding record for this reference.
- 37Stow, S. M.; Causon, T. J.; Zheng, X.; Kurulugama, R. T.; Mairinger, T.; May, J. C.; Rennie, E. E.; Baker, E. S.; Smith, R. D.; McLean, J. A.; Hann, S.; Fjeldsted, J. C. An Interlaboratory Evaluation of Drift Tube Ion Mobility–Mass Spectrometry Collision Cross Section Measurements. Anal. Chem. 2017, 89 (17), 9048– 9055, DOI: 10.1021/acs.analchem.7b01729There is no corresponding record for this reference.
- 38Loos, M.; Gerber, C.; Corona, F.; Hollender, J.; Singer, H. Accelerated Isotope Fine Structure Calculation Using Pruned Transition Trees. Anal. Chem. 2015, 87 (11), 5738– 5744, DOI: 10.1021/acs.analchem.5b00941There is no corresponding record for this reference.
- 39Berendsen, H. J. C.; van der Spoel, D.; van Drunen, R. GROMACS: A Message-Passing Parallel Molecular Dynamics Implementation. Comput. Phys. Commun. 1995, 91 (1–3), 43– 56, DOI: 10.1016/0010-4655(95)00042-EThere is no corresponding record for this reference.
- 40Kaminski, G. A.; Friesner, R. A.; Tirado-Rives, J.; Jorgensen, W. L. Evaluation and Reparametrization of the OPLS-AA Force Field for Proteins via Comparison with Accurate Quantum Chemical Calculations on Peptides. J. Phys. Chem. B 2001, 105 (28), 6474– 6487, DOI: 10.1021/jp003919dThere is no corresponding record for this reference.
- 41Wang, S.; Hou, K.; Heinz, H. Accurate and Compatible Force Fields for Molecular Oxygen, Nitrogen, and Hydrogen to Simulate Gases, Electrolytes, and Heterogeneous Interfaces. J. Chem. Theory Comput. 2021, 17 (8), 5198– 5213, DOI: 10.1021/acs.jctc.0c01132There is no corresponding record for this reference.
- 42Wyttenbach, T.; Bowers, M. T. Structural Stability from Solution to the Gas Phase: Native Solution Structure of Ubiquitin Survives Analysis in a Solvent-Free Ion Mobility–Mass Spectrometry Environment. J. Phys. Chem. B 2011, 115 (42), 12266– 12275, DOI: 10.1021/jp206867aThere is no corresponding record for this reference.
- 43Laszlo, K. J.; Munger, E. B.; Bush, M. F. Folding of Protein Ions in the Gas Phase after Cation-to-Anion Proton-Transfer Reactions. J. Am. Chem. Soc. 2016, 138 (30), 9581– 9588, DOI: 10.1021/jacs.6b04282There is no corresponding record for this reference.
- 44Warnke, S.; Von Helden, G.; Pagel, K. Protein Structure in the Gas Phase: The Influence of Side-Chain Microsolvation. J. Am. Chem. Soc. 2013, 135 (4), 1177– 1180, DOI: 10.1021/ja308528dThere is no corresponding record for this reference.
- 45Liu, F. C.; Kirk, S. R.; Caldwell, K. A.; Pedrete, T.; Meier, F.; Bleiholder, C. Tandem Trapped Ion Mobility Spectrometry/Mass Spectrometry (tTIMS/MS) Reveals Sequence-Specific Determinants of Top-Down Protein Fragment Ion Cross Sections. Anal. Chem. 2022, 94 (23), 8146– 8155, DOI: 10.1021/acs.analchem.1c05171There is no corresponding record for this reference.
- 46Warnke, S.; Baldauf, C.; Bowers, M. T.; Pagel, K.; von Helden, G. Photodissociation of Conformer-Selected Ubiquitin Ions Reveals Site-Specific Cis/Trans Isomerization of Proline Peptide Bonds. J. Am. Chem. Soc. 2014, 136 (29), 10308– 10314, DOI: 10.1021/ja502994bThere is no corresponding record for this reference.
- 47Fornelli, L.; Srzentić, K.; Toby, T. K.; Doubleday, P. F.; Huguet, R.; Mullen, C.; Melani, R. D.; dos Santos Seckler, H.; DeHart, C. J.; Weisbrod, C. R.; Durbin, K. R.; Greer, J. B.; Early, B. P.; Fellers, R. T.; Zabrouskov, V.; Thomas, P. M.; Compton, P. D.; Kelleher, N. L. Thorough Performance Evaluation of 213 Nm Ultraviolet Photodissociation for Top-down Proteomics. Mol. Cell. Proteomics 2020, 19 (2), 405– 420, DOI: 10.1074/mcp.TIR119.001638There is no corresponding record for this reference.
- 48Devakumar, A.; Thompson, M. S.; Reilly, J. P. Fragmentation of Oligosaccharide Ions with 157 Nm Vacuum Ultraviolet Light. Rapid Commun. Mass Spectrom. 2005, 19 (16), 2313– 2320, DOI: 10.1002/rcm.2058There is no corresponding record for this reference.
- 49Shelimov, K. B.; Jarrold, M. F. Conformations, Unfolding, and Refolding of Apomyoglobin in Vacuum: An Activation Barrier for Gas-Phase Protein Folding. J. Am. Chem. Soc. 1997, 119 (13), 2987– 2994, DOI: 10.1021/ja962914kThere is no corresponding record for this reference.
- 50Julian, R. R. The Mechanism Behind Top-Down UVPD Experiments: Making Sense of Apparent Contradictions. J. Am. Soc. Mass Spectrom. 2017, 28 (9), 1823– 1826, DOI: 10.1007/s13361-017-1721-0There is no corresponding record for this reference.
- 51Brodbelt, J. S.; Morrison, L. J.; Santos, I. Ultraviolet Photodissociation Mass Spectrometry for Analysis of Biological Molecules. Chem. Rev. 2020, 120 (7), 3328– 3380, DOI: 10.1021/acs.chemrev.9b00440There is no corresponding record for this reference.
- 52Kuipers, B. J. H.; Gruppen, H. Prediction of Molar Extinction Coefficients of Proteins and Peptides Using UV Absorption of the Constituent Amino Acids at 214 Nm To Enable Quantitative Reverse Phase High-Performance Liquid Chromatography–Mass Spectrometry Analysis. J. Agric. Food Chem. 2007, 55 (14), 5445– 5451, DOI: 10.1021/jf070337lThere is no corresponding record for this reference.
- 53Grégoire, G.; Jouvet, C.; Dedonder, C.; Sobolewski, A. L. Ab Initio Study of the Excited-State Deactivation Pathways of Protonated Tryptophan and Tyrosine. J. Am. Chem. Soc. 2007, 129 (19), 6223– 6231, DOI: 10.1021/ja069050fThere is no corresponding record for this reference.
- 54Paizs, B.; Suhai, S. Fragmentation Pathways of Protonated Peptides. Mass Spectrom. Rev. 2005, 24 (4), 508– 548, DOI: 10.1002/mas.20024There is no corresponding record for this reference.
- 55Bleiholder, C.; Osburn, S.; Williams, T. D.; Suhai, S.; Van Stipdonk, M.; Harrison, A. G.; Paizs, B. Sequence-Scrambling Fragmentation Pathways of Protonated Peptides. J. Am. Chem. Soc. 2008, 130 (52), 17774– 17789, DOI: 10.1021/ja805074dThere is no corresponding record for this reference.
Supporting Information
Supporting Information
The Supporting Information is available free of charge at https://pubs.acs.org/doi/10.1021/jasms.5c00432.
Seven figures with details on tandem-TIMS measurements (PDF)
Terms & Conditions
Most electronic Supporting Information files are available without a subscription to ACS Web Editions. Such files may be downloaded by article for research use (if there is a public use license linked to the relevant article, that license may permit other uses). Permission may be obtained from ACS for other uses through requests via the RightsLink permission system: http://pubs.acs.org/page/copyright/permissions.html.


