
Profiling of Low-Abundance Branched-Chain Fatty Acids via Radical Directed Dissociation Tandem Mass SpectrometryClick to copy article linkArticle link copied!
- Ruijun JianRuijun JianMOE Key Laboratory of Bioorganic Phosphorus Chemistry & Chemical Biological, Department of Chemistry, Tsinghua University, Beijing 100084, ChinaMore by Ruijun Jian
- Shengzhuo WangShengzhuo WangMOE Key Laboratory of Bioorganic Phosphorus Chemistry & Chemical Biological, Department of Chemistry, Tsinghua University, Beijing 100084, ChinaMore by Shengzhuo Wang
- Lipeng QiaoLipeng QiaoMOE Key Laboratory of Bioorganic Phosphorus Chemistry & Chemical Biological, Department of Chemistry, Tsinghua University, Beijing 100084, ChinaMore by Lipeng Qiao
- Xue Zhao*Xue Zhao*Email: [email protected]College of Chemistry and Chemical Engineering, Inner Mongolia University, Hohhot, 010070 Inner Mongolia ChinaMore by Xue Zhao
- Yu Xia*Yu Xia*Email: [email protected]MOE Key Laboratory of Bioorganic Phosphorus Chemistry & Chemical Biological, Department of Chemistry, Tsinghua University, Beijing 100084, ChinaMore by Yu Xia
Abstract
Branched-chain fatty acids (BCFAs) are key components of the bacterial lipidome, playing a role in regulating membrane fluidity and permeability. In mammals, BCFAs occur at much lower concentrations, and their functions remain largely unexplored. Conventional lipid analysis methods, employing collision-induced dissociation (CID)-tandem mass spectrometry (MS/MS), often fail to locate methyl branching, as fragmentation rarely occurs around the branching site. Here, we introduce a bifunctional derivatization reagent, 1-(8-methoxy-5-quinolinyl) methanamine (MeO-QN), for pinpointing methyl branching in BCFAs with high sensitivity. MeO-QN enhances ionization efficiency of derivatized BCFAs in positive ion mode due to its quinoline moiety and serves as a precursor for radical-directed dissociation (RDD). Upon CID, the quinoline-O radical (QN-O•) is generated, which subsequently induces RDD along the fatty acyl chain and forms a characteristic 28 Da spacing indicative of the branching point. By integrating this MS/MS method with reversed-phase liquid chromatography, we have developed a sensitive analytical workflow, detecting BCFAs at sub-nM levels in mammalian samples. We detected the rarely reported n-5 methyl branched fatty acid (FA 16:0;12Me) in pooled human plasma. We also observed significantly reduced even-chain isobranched fatty acids in breast cancer cells (MDA-MB-468) versus normal breast cells (MCF-10A), suggesting its potential in cancer biomarker discovery.
This publication is licensed for personal use by The American Chemical Society.
Introduction
Experimental Section
Lipid Nomenclature
Materials
Free Fatty Acid (FFA) Extraction and Total Fatty Acid (TFA) Extraction
Derivatization by MeO-QN
RPLC-MS analysis
Quantum Chemical Computation
Results and Discussion
CID Triggered Quinoline-O· RDD
Figure 1
Figure 1. (a) BDE of the C–O bond in MeO-QN before and after protonation. (b) MS2 CID spectrum of [MeO-QN+H]+. (c) Derivatization of FA by MeO-QN and proposed fragmentation pathways of derivatized FA ([FA# + H]+) under CID.
Locating Methyl Branching via QN-O• RDD
Figure 2
Figure 2. Ion-trap CID spectra of MeO-QN derivatized FAs and observed fragmentation sites of protonated FA-QN-O•. (a) [n-17:0#+H]+, (b) [i-17:0#+H]+, (c) [a-17:0#+H]+, and (d) [FA 16:0;3Me,7Me,11Me,15Me#+H]+.
Analysis of Other Types of Chain Modifications
An RPLC-MS/MS Workflow for Profiling of BCFAs from Biological Samples
Figure 3
Figure 3. (a) XIC from PIS of m/z 158 for equimolar mixtures of FA 15:0 and FA 17:0 isomers. (b) Calibration curves and limit of identification (LOI) for various fatty acids employing PIS of m/z 158 (IS: FA d4–18:0).
Scheme 1
Profiling of FFAs from Human Plasma
Figure 4
Figure 4. (a) Profiling of FFAs in human plasma (IS: FA 18:0-D4, 50 μ M). (b) XIC of [FA 17:0#+H]+ (m/z 441) in human plasma. (c) Ion trap CID spectrum of [FA 17:0#+H]+ eluted at 21.0 min in panel b.
Profiling of TFAs in Human Breast Cancerous and Normal Cells
Figure 5
Figure 5. (a) Volcano plot of TFA at the sum composition level in breast cancer cells and (b) comparisons of BCFA isomer compositions in breast cancer cells (MDA-MB-468) to normal breast epithelial cells (MCF-10A).
Conclusions
Supporting Information
The Supporting Information is available free of charge at https://pubs.acs.org/doi/10.1021/jasms.6c00001.
Experimental details; figures of optimized MeO-QN structure through DFT calculations, mass spectra of MeO-QN derivatized FAs, and background contaminations, tables of QTRAP 4500 parameters and gas basicity of functional groups (PDF)
Terms & Conditions
Most electronic Supporting Information files are available without a subscription to ACS Web Editions. Such files may be downloaded by article for research use (if there is a public use license linked to the relevant article, that license may permit other uses). Permission may be obtained from ACS for other uses through requests via the RightsLink permission system: http://pubs.acs.org/page/copyright/permissions.html.
Acknowledgments
Financial support from the National Natural Science Foundation of China (no. 22225404, 22074075) and the National Key R&D Program of China (2023YFA0913902) is greatly appreciated.
References
This article references 33 other publications.
- 1Rustan, A. C.; Drevon, C. A., Fatty Acids: Structures and Properties. In Encycl. Life Sci. , 2005. DOI: 10.1038/npg.els.0003894 .Google ScholarThere is no corresponding record for this reference.
- 2Currie, E.; Schulze, A.; Zechner, R.; Walther, T. C.; Farese, R. V. Cellular Fatty Acid Metabolism and Cancer. Cell Metab 2013, 18, 153– 161, DOI: 10.1016/j.cmet.2013.05.017Google ScholarThere is no corresponding record for this reference.
- 3Taormina, V. M.; Unger, A. L.; Schiksnis, M. R.; Torres-Gonzalez, M.; Kraft, J. Branched-Chain Fatty Acids─An Underexplored Class of Dairy-Derived Fatty Acids. Nutrients 2020, 12, 2875– 2890, DOI: 10.3390/nu12092875Google ScholarThere is no corresponding record for this reference.
- 4Martínez-Montoro, J. I.; Núñez-Sánchez, M. Á.; Martinez-Sanchez, M. A.; Balaguer-Román, A.; Fernández-Ruiz, V. E.; Ferrer-Gómez, M.; Sledzinski, T.; Frutos, M. D.; Fernández-García, J. C.; Mika, A.; Ramos-Molina, B. Hepatic and serum branched-chain fatty acid profile in patients with nonalcoholic fatty liver disease: A case–control study. Obesity 2023, 31, 1064– 1074, DOI: 10.1002/oby.23711Google ScholarThere is no corresponding record for this reference.
- 5Mika, A.; Stepnowski, P.; Kaska, L.; Proczko, M.; Wisniewski, P.; Sledzinski, M.; Sledzinski, T. A comprehensive study of serum odd- and branched-chain fatty acids in patients with excess weight. Obesity 2016, 24, 1669– 1676, DOI: 10.1002/oby.21560Google ScholarThere is no corresponding record for this reference.
- 6Wang, Z.; Yang, T.; Brenna, J. T.; Wang, D. H. Fatty acid isomerism: analysis and selected biological functions. Food Funct 2024, 15, 1071– 1088, DOI: 10.1039/D3FO03716AGoogle ScholarThere is no corresponding record for this reference.
- 7Ran-Ressler, R. R.; Lawrence, P.; Brenna, J. T. Structural characterization of saturated branched chain fatty acid methyl esters by collisional dissociation of molecular ions generated by electron ionization. J. Lipid Res. 2012, 53, 195– 203, DOI: 10.1194/jlr.D020651Google ScholarThere is no corresponding record for this reference.
- 8Wang, D. H.; Wang, Z.; Brenna, J. T. Gas Chromatography Chemical Ionization Mass Spectrometry and Tandem Mass Spectrometry for Identification and Straightforward Quantification of Branched Chain Fatty Acids in Foods. J. Agric. Food. Chem. 2020, 68, 4973– 4980, DOI: 10.1021/acs.jafc.0c01075Google ScholarThere is no corresponding record for this reference.
- 9Wang, D. H.; Wang, Z.; Chen, R.; Kothapalli, K. S. D.; Brenna, J. T. Very Long-Chain Branched-Chain Fatty Acids in Chia Seeds: Implications for Human Use. J. Agric. Food. Chem. 2020, 68, 13871– 13878, DOI: 10.1021/acs.jafc.0c05612Google ScholarThere is no corresponding record for this reference.
- 10Kerwin, J. L.; Wiens, A. M.; Ericsson, L. H. Identification of fatty acids by electrospray mass spectrometry and tandem mass spectrometry. J. Mass Spectrom. 1996, 31, 184– 192, DOI: 10.1002/(SICI)1096-9888(199602)31:2<184::AID-JMS283>3.0.CO;2-2Google ScholarThere is no corresponding record for this reference.
- 11Randolph, C. E.; Beveridge, C. H.; Iyer, S.; Blanksby, S. J.; McLuckey, S. A.; Chopra, G. Identification of Monomethyl Branched-Chain Lipids by a Combination of Liquid Chromatography Tandem Mass Spectrometry and Charge-Switching Chemistries. J. Am. Soc. Mass Spectrom. 2022, 33, 2156– 2164, DOI: 10.1021/jasms.2c00225Google ScholarThere is no corresponding record for this reference.
- 12Wang, M.; Han, R. H.; Han, X. Fatty Acidomics: Global Analysis of Lipid Species Containing a Carboxyl Group with a Charge-Remote Fragmentation-Assisted Approach. Anal. Chem. 2013, 85, 9312– 9320, DOI: 10.1021/ac402078pGoogle ScholarThere is no corresponding record for this reference.
- 13Randolph, C. E.; Foreman, D. J.; Betancourt, S. K.; Blanksby, S. J.; McLuckey, S. A. Gas-Phase Ion/Ion Reactions Involving Tris-Phenanthroline Alkaline Earth Metal Complexes as Charge Inversion Reagents for the Identification of Fatty Acids. Anal. Chem. 2018, 90, 12861– 12869, DOI: 10.1021/acs.analchem.8b03441Google ScholarThere is no corresponding record for this reference.
- 14Shenault, D. S. M.; Fabijanczuk, K. C.; Murtada, R.; Finn, S.; Gonzalez, L. E.; Gao, J.; McLuckey, S. A. Gas-Phase Ion/Ion Reactions to Enable Radical-Directed Dissociation of Fatty Acid Ions: Application to Localization of Methyl Branching. Anal. Chem. 2024, 96, 3389– 3401, DOI: 10.1021/acs.analchem.3c04510Google ScholarThere is no corresponding record for this reference.
- 15Pham, H. T.; Ly, T.; Trevitt, A. J.; Mitchell, T. W.; Blanksby, S. J. Differentiation of complex lipid isomers by radical-directed dissociation mass spectrometry. Anal. Chem. 2012, 84, 7525– 7532, DOI: 10.1021/ac301652aGoogle ScholarThere is no corresponding record for this reference.
- 16Narreddula, V. R.; Boase, N. R.; Ailuri, R.; Marshall, D. L.; Poad, B. L. J.; Kelso, M. J.; Trevitt, A. J.; Mitchell, T. W.; Blanksby, S. J. Introduction of a Fixed-Charge, Photolabile Derivative for Enhanced Structural Elucidation of Fatty Acids. Anal. Chem. 2019, 91, 9901– 9909, DOI: 10.1021/acs.analchem.9b01566Google ScholarThere is no corresponding record for this reference.
- 17Nsiah, S. T.; Fabijanczuk, K. C.; McLuckey, S. A. Structural characterization of fatty acid anions via gas-phase charge inversion using Mg(tri-butyl-terpyridine)22+ reagent ions. Rapid Commun. Mass Spectrom. 2024, 38, e9741 DOI: 10.1002/rcm.9741Google ScholarThere is no corresponding record for this reference.
- 18Jian, R.; Zhao, X.; Lin, Q.; Xia, Y. Profiling of branched-chain fatty acids via nitroxide radical-directed dissociation integrated on an LC-MS/MS workflow. Analyst 2022, 147, 2115– 2123, DOI: 10.1039/D2AN00266CGoogle ScholarThere is no corresponding record for this reference.
- 19Liebisch, G.; Fahy, E.; Aoki, J.; Dennis, E. A.; Durand, T.; Ejsing, C. S.; Fedorova, M.; Feussner, I.; Griffiths, W. J.; Kofeler, H.; Merrill, A. H., Jr.; Murphy, R. C.; O’Donnell, V. B.; Oskolkova, O.; Subramaniam, S.; Wakelam, M. J. O.; Spener, F. Update on LIPID MAPS classification, nomenclature, and shorthand notation for MS-derived lipid structures. J. Lipid Res. 2020, 61, 1539– 1555, DOI: 10.1194/jlr.S120001025Google ScholarThere is no corresponding record for this reference.
- 20Quehenberger, O.; Armando, A. M.; Dennis, E. A. High sensitivity quantitative lipidomics analysis of fatty acids in biological samples by gas chromatography–mass spectrometry. BBA-Mol. Cell Biol. L. 2011, 1811, 648– 656, DOI: 10.1016/j.bbalip.2011.07.006Google ScholarThere is no corresponding record for this reference.
- 21Matyash, V.; Liebisch, G.; Kurzchalia, T. V.; Shevchenko, A.; Schwudke, D. Lipid extraction by methyl-tert-butyl ether for high-throughput lipidomics. J. Lipid Res. 2008, 49, 1137– 1146, DOI: 10.1194/jlr.D700041-JLR200Google ScholarThere is no corresponding record for this reference.
- 22Carpino, L. A.; El-Faham, A.; Minor, C. A.; Albericio, F. Advantageous Applications of Azabenzotriazole (Triazo1opyridine)-based Coupling Reagents to Solid-phase Peptide Synthesis. J. Chem. Soc., Chem. Commun. 1994, 2, 201– 203, DOI: 10.1039/c39940000201Google ScholarThere is no corresponding record for this reference.
- 23Gross, M. L. Charge-remote fragmentations: method, mechanism and applications. Int. J. Mass Spectrom. Ion Processes 1992, 118–119, 137– 165, DOI: 10.1016/0168-1176(92)85060-DGoogle ScholarThere is no corresponding record for this reference.
- 24Hunter, E. P. L.; Lias, S. G. Evaluated Gas Phase Basicities and Proton Affinities of Molecules: An Update. J. Phys. Chem. Ref. Data 1998, 27, 413– 656, DOI: 10.1063/1.556018Google ScholarThere is no corresponding record for this reference.
- 25Yu, F.; Zong, B.; Ji, L.; Sun, P.; Jia, D.; Wang, R. Free Fatty Acids and Free Fatty Acid Receptors: Role in Regulating Arterial Function. Int. J. Mol. Sci. 2024, 25, 7853, DOI: 10.3390/ijms25147853Google ScholarThere is no corresponding record for this reference.
- 26Yan, Y.; Wang, Z.; Greenwald, J.; Kothapalli, K. S. D.; Park, H. G.; Liu, R.; Mendralla, E.; Lawrence, P.; Wang, X.; Brenna, J. T. BCFA suppresses LPS induced IL-8 mRNA expression in human intestinal epithelial cells. Prostag. Leukotr. Ess. 2017, 116, 27– 31, DOI: 10.1016/j.plefa.2016.12.001Google ScholarThere is no corresponding record for this reference.
- 27Katare, P. B.; Tingstad, R. H.; Beajani, S. T.; Indseth, J. P.; Telle-Hansen, V. H.; Myhrstad, M. C. W.; Rustan, A. C.; Eide, L.; Witczak, O.; Aas, V. Divergent effects of monomethyl branched-chain fatty acids on energy metabolism and insulin signaling in human myotubes. J. Lipid Res. 2025, 66, 100764– 100775, DOI: 10.1016/j.jlr.2025.100764Google ScholarThere is no corresponding record for this reference.
- 28Chen, D.; Zhao, S.; Ma, G.; Li, L.; Li, L. An Analytical Strategy for Reliable Metabolome Analysis of Clinical Leftover Sera Using Timed Aliquoting. Anal. Chem. 2025, 97, 25655– 25663, DOI: 10.1021/acs.analchem.5c04801Google ScholarThere is no corresponding record for this reference.
- 29Menzel, J. P.; Young, R. S. E.; Benfield, A. H.; Scott, J. S.; Wongsomboon, P.; Cudlman, L.; Cvačka, J.; Butler, L. M.; Henriques, S. T.; Poad, B. L. J.; Blanksby, S. J. Ozone-enabled fatty acid discovery reveals unexpected diversity in the human lipidome. Nat. Commun. 2023, 14, 3940, DOI: 10.1038/s41467-023-39617-9Google ScholarThere is no corresponding record for this reference.
- 30Roy, R.; Roseblade, A.; Rawling, T. Expansion of the structure-activity relationship of branched chain fatty acids: Effect of unsaturation and branching group size on anticancer activity. Chem. Phys. Lipids 2020, 232, 104952– 104959, DOI: 10.1016/j.chemphyslip.2020.104952Google ScholarThere is no corresponding record for this reference.
- 31Gozdzik, P.; Magkos, F.; Sledzinski, T.; Mika, A. Monomethyl branched-chain fatty acids: Health effects and biological mechanisms. Prog. Lipid Res. 2023, 90, 101226– 101236, DOI: 10.1016/j.plipres.2023.101226Google ScholarThere is no corresponding record for this reference.
- 32Nagarajan, S. R.; Butler, L. M.; Hoy, A. J. The diversity and breadth of cancer cell fatty acid metabolism. Cancer Metab. 2021, 9, 2– 29, DOI: 10.1186/s40170-020-00237-2Google ScholarThere is no corresponding record for this reference.
- 33Wongtangtintharn, S.; Oku, H.; Iwasaki, H.; Toda, T. Effect of Branched-Chain Fatty Acids on Fatty Acid Biosynthesis of Human Breast Cancer Cells. J. Nutr. Sci. Vitaminol. 2004, 50, 137– 143, DOI: 10.3177/jnsv.50.137Google ScholarThere is no corresponding record for this reference.
Cited By
This article has not yet been cited by other publications.
Article Views
Altmetric
Citations
Article Views are the COUNTER-compliant sum of full text article downloads since November 2008 (both PDF and HTML) across all institutions and individuals. These metrics are regularly updated to reflect usage leading up to the last few days.
Citations are the number of other articles citing this article, calculated by Crossref and updated daily. Find more information about Crossref citation counts.
The Altmetric Attention Score is a quantitative measure of the attention that a research article has received online. Clicking on the donut icon will load a page at altmetric.com with additional details about the score and the social media presence for the given article. Find more information on the Altmetric Attention Score and how the score is calculated.
Recommended Articles
Abstract
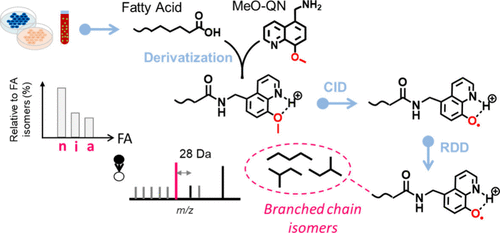
Figure 1
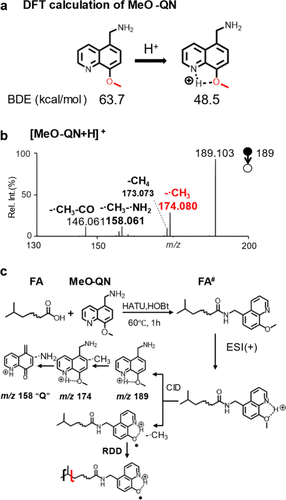
Figure 1. (a) BDE of the C–O bond in MeO-QN before and after protonation. (b) MS2 CID spectrum of [MeO-QN+H]+. (c) Derivatization of FA by MeO-QN and proposed fragmentation pathways of derivatized FA ([FA# + H]+) under CID.
Figure 2

Figure 2. Ion-trap CID spectra of MeO-QN derivatized FAs and observed fragmentation sites of protonated FA-QN-O•. (a) [n-17:0#+H]+, (b) [i-17:0#+H]+, (c) [a-17:0#+H]+, and (d) [FA 16:0;3Me,7Me,11Me,15Me#+H]+.
Figure 3
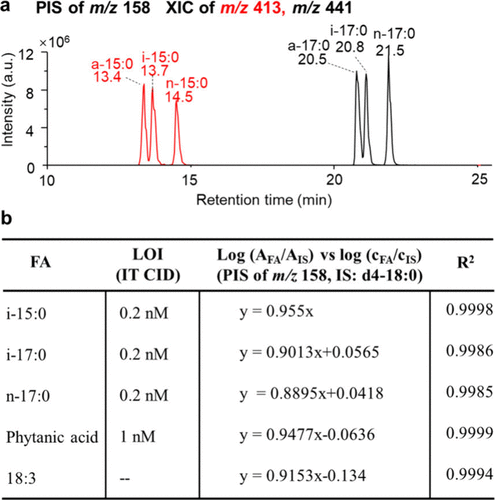
Figure 3. (a) XIC from PIS of m/z 158 for equimolar mixtures of FA 15:0 and FA 17:0 isomers. (b) Calibration curves and limit of identification (LOI) for various fatty acids employing PIS of m/z 158 (IS: FA d4–18:0).
Scheme 1
 Scheme 1. Workflow for the Analysis of Fatty Acids with Precision at the Branched-Chain Level Using QN-O• RDD
Scheme 1. Workflow for the Analysis of Fatty Acids with Precision at the Branched-Chain Level Using QN-O• RDDFigure 4
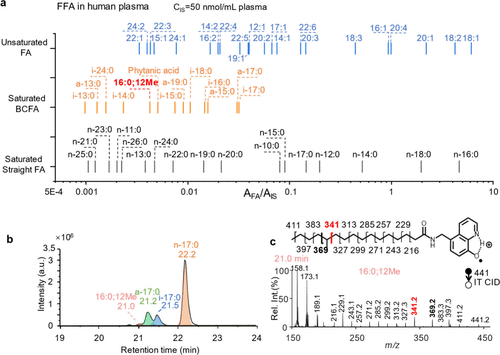
Figure 4. (a) Profiling of FFAs in human plasma (IS: FA 18:0-D4, 50 μ M). (b) XIC of [FA 17:0#+H]+ (m/z 441) in human plasma. (c) Ion trap CID spectrum of [FA 17:0#+H]+ eluted at 21.0 min in panel b.
Figure 5

Figure 5. (a) Volcano plot of TFA at the sum composition level in breast cancer cells and (b) comparisons of BCFA isomer compositions in breast cancer cells (MDA-MB-468) to normal breast epithelial cells (MCF-10A).
References
This article references 33 other publications.
- 1Rustan, A. C.; Drevon, C. A., Fatty Acids: Structures and Properties. In Encycl. Life Sci. , 2005. DOI: 10.1038/npg.els.0003894 .There is no corresponding record for this reference.
- 2Currie, E.; Schulze, A.; Zechner, R.; Walther, T. C.; Farese, R. V. Cellular Fatty Acid Metabolism and Cancer. Cell Metab 2013, 18, 153– 161, DOI: 10.1016/j.cmet.2013.05.017There is no corresponding record for this reference.
- 3Taormina, V. M.; Unger, A. L.; Schiksnis, M. R.; Torres-Gonzalez, M.; Kraft, J. Branched-Chain Fatty Acids─An Underexplored Class of Dairy-Derived Fatty Acids. Nutrients 2020, 12, 2875– 2890, DOI: 10.3390/nu12092875There is no corresponding record for this reference.
- 4Martínez-Montoro, J. I.; Núñez-Sánchez, M. Á.; Martinez-Sanchez, M. A.; Balaguer-Román, A.; Fernández-Ruiz, V. E.; Ferrer-Gómez, M.; Sledzinski, T.; Frutos, M. D.; Fernández-García, J. C.; Mika, A.; Ramos-Molina, B. Hepatic and serum branched-chain fatty acid profile in patients with nonalcoholic fatty liver disease: A case–control study. Obesity 2023, 31, 1064– 1074, DOI: 10.1002/oby.23711There is no corresponding record for this reference.
- 5Mika, A.; Stepnowski, P.; Kaska, L.; Proczko, M.; Wisniewski, P.; Sledzinski, M.; Sledzinski, T. A comprehensive study of serum odd- and branched-chain fatty acids in patients with excess weight. Obesity 2016, 24, 1669– 1676, DOI: 10.1002/oby.21560There is no corresponding record for this reference.
- 6Wang, Z.; Yang, T.; Brenna, J. T.; Wang, D. H. Fatty acid isomerism: analysis and selected biological functions. Food Funct 2024, 15, 1071– 1088, DOI: 10.1039/D3FO03716AThere is no corresponding record for this reference.
- 7Ran-Ressler, R. R.; Lawrence, P.; Brenna, J. T. Structural characterization of saturated branched chain fatty acid methyl esters by collisional dissociation of molecular ions generated by electron ionization. J. Lipid Res. 2012, 53, 195– 203, DOI: 10.1194/jlr.D020651There is no corresponding record for this reference.
- 8Wang, D. H.; Wang, Z.; Brenna, J. T. Gas Chromatography Chemical Ionization Mass Spectrometry and Tandem Mass Spectrometry for Identification and Straightforward Quantification of Branched Chain Fatty Acids in Foods. J. Agric. Food. Chem. 2020, 68, 4973– 4980, DOI: 10.1021/acs.jafc.0c01075There is no corresponding record for this reference.
- 9Wang, D. H.; Wang, Z.; Chen, R.; Kothapalli, K. S. D.; Brenna, J. T. Very Long-Chain Branched-Chain Fatty Acids in Chia Seeds: Implications for Human Use. J. Agric. Food. Chem. 2020, 68, 13871– 13878, DOI: 10.1021/acs.jafc.0c05612There is no corresponding record for this reference.
- 10Kerwin, J. L.; Wiens, A. M.; Ericsson, L. H. Identification of fatty acids by electrospray mass spectrometry and tandem mass spectrometry. J. Mass Spectrom. 1996, 31, 184– 192, DOI: 10.1002/(SICI)1096-9888(199602)31:2<184::AID-JMS283>3.0.CO;2-2There is no corresponding record for this reference.
- 11Randolph, C. E.; Beveridge, C. H.; Iyer, S.; Blanksby, S. J.; McLuckey, S. A.; Chopra, G. Identification of Monomethyl Branched-Chain Lipids by a Combination of Liquid Chromatography Tandem Mass Spectrometry and Charge-Switching Chemistries. J. Am. Soc. Mass Spectrom. 2022, 33, 2156– 2164, DOI: 10.1021/jasms.2c00225There is no corresponding record for this reference.
- 12Wang, M.; Han, R. H.; Han, X. Fatty Acidomics: Global Analysis of Lipid Species Containing a Carboxyl Group with a Charge-Remote Fragmentation-Assisted Approach. Anal. Chem. 2013, 85, 9312– 9320, DOI: 10.1021/ac402078pThere is no corresponding record for this reference.
- 13Randolph, C. E.; Foreman, D. J.; Betancourt, S. K.; Blanksby, S. J.; McLuckey, S. A. Gas-Phase Ion/Ion Reactions Involving Tris-Phenanthroline Alkaline Earth Metal Complexes as Charge Inversion Reagents for the Identification of Fatty Acids. Anal. Chem. 2018, 90, 12861– 12869, DOI: 10.1021/acs.analchem.8b03441There is no corresponding record for this reference.
- 14Shenault, D. S. M.; Fabijanczuk, K. C.; Murtada, R.; Finn, S.; Gonzalez, L. E.; Gao, J.; McLuckey, S. A. Gas-Phase Ion/Ion Reactions to Enable Radical-Directed Dissociation of Fatty Acid Ions: Application to Localization of Methyl Branching. Anal. Chem. 2024, 96, 3389– 3401, DOI: 10.1021/acs.analchem.3c04510There is no corresponding record for this reference.
- 15Pham, H. T.; Ly, T.; Trevitt, A. J.; Mitchell, T. W.; Blanksby, S. J. Differentiation of complex lipid isomers by radical-directed dissociation mass spectrometry. Anal. Chem. 2012, 84, 7525– 7532, DOI: 10.1021/ac301652aThere is no corresponding record for this reference.
- 16Narreddula, V. R.; Boase, N. R.; Ailuri, R.; Marshall, D. L.; Poad, B. L. J.; Kelso, M. J.; Trevitt, A. J.; Mitchell, T. W.; Blanksby, S. J. Introduction of a Fixed-Charge, Photolabile Derivative for Enhanced Structural Elucidation of Fatty Acids. Anal. Chem. 2019, 91, 9901– 9909, DOI: 10.1021/acs.analchem.9b01566There is no corresponding record for this reference.
- 17Nsiah, S. T.; Fabijanczuk, K. C.; McLuckey, S. A. Structural characterization of fatty acid anions via gas-phase charge inversion using Mg(tri-butyl-terpyridine)22+ reagent ions. Rapid Commun. Mass Spectrom. 2024, 38, e9741 DOI: 10.1002/rcm.9741There is no corresponding record for this reference.
- 18Jian, R.; Zhao, X.; Lin, Q.; Xia, Y. Profiling of branched-chain fatty acids via nitroxide radical-directed dissociation integrated on an LC-MS/MS workflow. Analyst 2022, 147, 2115– 2123, DOI: 10.1039/D2AN00266CThere is no corresponding record for this reference.
- 19Liebisch, G.; Fahy, E.; Aoki, J.; Dennis, E. A.; Durand, T.; Ejsing, C. S.; Fedorova, M.; Feussner, I.; Griffiths, W. J.; Kofeler, H.; Merrill, A. H., Jr.; Murphy, R. C.; O’Donnell, V. B.; Oskolkova, O.; Subramaniam, S.; Wakelam, M. J. O.; Spener, F. Update on LIPID MAPS classification, nomenclature, and shorthand notation for MS-derived lipid structures. J. Lipid Res. 2020, 61, 1539– 1555, DOI: 10.1194/jlr.S120001025There is no corresponding record for this reference.
- 20Quehenberger, O.; Armando, A. M.; Dennis, E. A. High sensitivity quantitative lipidomics analysis of fatty acids in biological samples by gas chromatography–mass spectrometry. BBA-Mol. Cell Biol. L. 2011, 1811, 648– 656, DOI: 10.1016/j.bbalip.2011.07.006There is no corresponding record for this reference.
- 21Matyash, V.; Liebisch, G.; Kurzchalia, T. V.; Shevchenko, A.; Schwudke, D. Lipid extraction by methyl-tert-butyl ether for high-throughput lipidomics. J. Lipid Res. 2008, 49, 1137– 1146, DOI: 10.1194/jlr.D700041-JLR200There is no corresponding record for this reference.
- 22Carpino, L. A.; El-Faham, A.; Minor, C. A.; Albericio, F. Advantageous Applications of Azabenzotriazole (Triazo1opyridine)-based Coupling Reagents to Solid-phase Peptide Synthesis. J. Chem. Soc., Chem. Commun. 1994, 2, 201– 203, DOI: 10.1039/c39940000201There is no corresponding record for this reference.
- 23Gross, M. L. Charge-remote fragmentations: method, mechanism and applications. Int. J. Mass Spectrom. Ion Processes 1992, 118–119, 137– 165, DOI: 10.1016/0168-1176(92)85060-DThere is no corresponding record for this reference.
- 24Hunter, E. P. L.; Lias, S. G. Evaluated Gas Phase Basicities and Proton Affinities of Molecules: An Update. J. Phys. Chem. Ref. Data 1998, 27, 413– 656, DOI: 10.1063/1.556018There is no corresponding record for this reference.
- 25Yu, F.; Zong, B.; Ji, L.; Sun, P.; Jia, D.; Wang, R. Free Fatty Acids and Free Fatty Acid Receptors: Role in Regulating Arterial Function. Int. J. Mol. Sci. 2024, 25, 7853, DOI: 10.3390/ijms25147853There is no corresponding record for this reference.
- 26Yan, Y.; Wang, Z.; Greenwald, J.; Kothapalli, K. S. D.; Park, H. G.; Liu, R.; Mendralla, E.; Lawrence, P.; Wang, X.; Brenna, J. T. BCFA suppresses LPS induced IL-8 mRNA expression in human intestinal epithelial cells. Prostag. Leukotr. Ess. 2017, 116, 27– 31, DOI: 10.1016/j.plefa.2016.12.001There is no corresponding record for this reference.
- 27Katare, P. B.; Tingstad, R. H.; Beajani, S. T.; Indseth, J. P.; Telle-Hansen, V. H.; Myhrstad, M. C. W.; Rustan, A. C.; Eide, L.; Witczak, O.; Aas, V. Divergent effects of monomethyl branched-chain fatty acids on energy metabolism and insulin signaling in human myotubes. J. Lipid Res. 2025, 66, 100764– 100775, DOI: 10.1016/j.jlr.2025.100764There is no corresponding record for this reference.
- 28Chen, D.; Zhao, S.; Ma, G.; Li, L.; Li, L. An Analytical Strategy for Reliable Metabolome Analysis of Clinical Leftover Sera Using Timed Aliquoting. Anal. Chem. 2025, 97, 25655– 25663, DOI: 10.1021/acs.analchem.5c04801There is no corresponding record for this reference.
- 29Menzel, J. P.; Young, R. S. E.; Benfield, A. H.; Scott, J. S.; Wongsomboon, P.; Cudlman, L.; Cvačka, J.; Butler, L. M.; Henriques, S. T.; Poad, B. L. J.; Blanksby, S. J. Ozone-enabled fatty acid discovery reveals unexpected diversity in the human lipidome. Nat. Commun. 2023, 14, 3940, DOI: 10.1038/s41467-023-39617-9There is no corresponding record for this reference.
- 30Roy, R.; Roseblade, A.; Rawling, T. Expansion of the structure-activity relationship of branched chain fatty acids: Effect of unsaturation and branching group size on anticancer activity. Chem. Phys. Lipids 2020, 232, 104952– 104959, DOI: 10.1016/j.chemphyslip.2020.104952There is no corresponding record for this reference.
- 31Gozdzik, P.; Magkos, F.; Sledzinski, T.; Mika, A. Monomethyl branched-chain fatty acids: Health effects and biological mechanisms. Prog. Lipid Res. 2023, 90, 101226– 101236, DOI: 10.1016/j.plipres.2023.101226There is no corresponding record for this reference.
- 32Nagarajan, S. R.; Butler, L. M.; Hoy, A. J. The diversity and breadth of cancer cell fatty acid metabolism. Cancer Metab. 2021, 9, 2– 29, DOI: 10.1186/s40170-020-00237-2There is no corresponding record for this reference.
- 33Wongtangtintharn, S.; Oku, H.; Iwasaki, H.; Toda, T. Effect of Branched-Chain Fatty Acids on Fatty Acid Biosynthesis of Human Breast Cancer Cells. J. Nutr. Sci. Vitaminol. 2004, 50, 137– 143, DOI: 10.3177/jnsv.50.137There is no corresponding record for this reference.
Supporting Information
Supporting Information
The Supporting Information is available free of charge at https://pubs.acs.org/doi/10.1021/jasms.6c00001.
Experimental details; figures of optimized MeO-QN structure through DFT calculations, mass spectra of MeO-QN derivatized FAs, and background contaminations, tables of QTRAP 4500 parameters and gas basicity of functional groups (PDF)
Terms & Conditions
Most electronic Supporting Information files are available without a subscription to ACS Web Editions. Such files may be downloaded by article for research use (if there is a public use license linked to the relevant article, that license may permit other uses). Permission may be obtained from ACS for other uses through requests via the RightsLink permission system: http://pubs.acs.org/page/copyright/permissions.html.


