ChipNMR: Hyperpolarized NMR for Noninvasive Metabolic Flux Analysis in Perfused Microfluidic ChipsClick to copy article linkArticle link copied!
- Thomas B. Wareham MathiassenThomas B. Wareham MathiassenCenter for Hyperpolarization in Magnetic Resonance, Department of Health Technology, Technical University of Denmark, Ørsteds Plads 349, 2800 Kgs. Lyngby, DenmarkMore by Thomas B. Wareham Mathiassen
- Juan D. Sánchez-HerediaJuan D. Sánchez-HerediaDepartment of Information Technologies and Communications, Technical University of Cartagena (UPCT), 302020 Cartagena, SpainMore by Juan D. Sánchez-Heredia
- Ke-Chuan WangKe-Chuan WangCenter for Hyperpolarization in Magnetic Resonance, Department of Health Technology, Technical University of Denmark, Ørsteds Plads 349, 2800 Kgs. Lyngby, DenmarkMore by Ke-Chuan Wang
- Cajsa R. HauptCajsa R. HauptCenter for Hyperpolarization in Magnetic Resonance, Department of Health Technology, Technical University of Denmark, Ørsteds Plads 349, 2800 Kgs. Lyngby, DenmarkMore by Cajsa R. Haupt
- Magnus KarlssonMagnus KarlssonCenter for Hyperpolarization in Magnetic Resonance, Department of Health Technology, Technical University of Denmark, Ørsteds Plads 349, 2800 Kgs. Lyngby, DenmarkMore by Magnus Karlsson
- Alexander JönssonAlexander JönssonDepartment of Micro and Nanotechnology, Technical University of Denmark, 2800 Kgs Lyngby, DenmarkMore by Alexander Jönsson
- Martin DufvaMartin DufvaDepartment of Micro and Nanotechnology, Technical University of Denmark, 2800 Kgs Lyngby, DenmarkMore by Martin Dufva
- Roland ThuenauerRoland ThuenauerTechnology Platform Light Microscopy (TPLM), University of Hamburg (UHH); Advanced Light and Fluorescence Microscopy (ALFM) Facility, Centre for Structural Systems Biology (CSSB) Hamburg; Leibniz Institute of Virology (LIV), Notkestrasse 85, 22607 Hamburg, GermanyMore by Roland Thuenauer
- Pernille Rose Jensen*Pernille Rose Jensen*Email: [email protected]Center for Hyperpolarization in Magnetic Resonance, Department of Health Technology, Technical University of Denmark, Ørsteds Plads 349, 2800 Kgs. Lyngby, DenmarkMore by Pernille Rose Jensen
Abstract
Dissolution dynamic nuclear polarization NMR Spectroscopy (dDNP-NMR) has become a transformative tool for metabolic studies by significantly enhancing signal sensitivity more than 3 orders of magnitude compared to traditional NMR. However, NMR detection probes are optimized for round narrow glass tubes typically 5 mm in diameter, which impose constraints on their utility for metabolic studies of adherent cells. Here, we present a novel NMR probe head integrated with a custom microfluidic chip that facilitates real-time monitoring of hyperpolarized substrate conversion from adherent cells. This system enables metabolic flux analysis in a controlled, in vitro environment, as demonstrated by tracking the conversion of [1-13C]pyruvate to [1-13C]lactate in HeLa cells over 48 h. The custom microfluidic chip design is modular and adaptable allowing expansion to two-chamber chips, demonstrating its potential in applications for more complex cellular models, such as organ-on-a-chip systems.
This publication is licensed for personal use by The American Chemical Society.
Introduction
Experimental Section
RF Coil Design and Creation
Figure 1
Figure 1. RF Coil design. (A) exploded view of the NMR probe head, showing the two RF coils, the 3D-printed support, and the microfluidic chip. (B) Cross-section (Z = 0 plane) of the simulated B1+ field distribution generated by Coil 1 at the 13C frequency (per 1 W accepted power). (C) Photograph of the fabricated NMR probe with the chip inserted.
Chip Fabrication
Figure 2
Figure 2. Fluidic chip production and assembly. (A) visual representation of the chip components and their assembly process. The stacked components are shown in order from top to bottom: 1 mm PETG, 0.75 mm SEBS, track-etched polycarbonate membrane (8 μm pore size, 18 μm thickness, 5% porosity), 0.25 mm SEBS, and 0.3 mm PETG. (B) 3D-printed inlets/injection ports, designed using SolidWorks, shown with female mini Luer connectors made from BioMed clear resin. (C) Fully assembled microfluidic chip.
Chip Sterilization and Treatment
Cell Culturing and Chip Seeding
Cell Staining and Microscopy
dDNP-NMR Experiments
Cell Counting
Data Analysis
Statistical Analysis
Results and Discussion
NMR Coil Design and Functionality
Figure 3
Figure 3. Performance evaluation. (A) single pyruvate spectrum, SNR of the highest pyruvate peak was 5000, (B) substrate integral over time (n = 3) and (C) comparative substrate integrals (AUC = 24 ± 0.6) following injection of 10 mM hyperpolarized [1-13C] pyruvate, CV for the AUC was 2.5%. T1 of pyruvate was 52 ± 2 s.
Microfluidic Chip as a Cell Culturing Device
Figure 4
Figure 4. One-chamber chip cell growth. (A) one-chamber chip seeded with 3 × 106 HeLa cells, cultured for 3 h, and imaged with bright-field microscopy (bottom left, BF 20×) and fluorescence microscopy of cells stained with Calcein AM (bottom right, Calcein AM 20×). Scale bar: 100 μm. (B) Cell counts in chips after 3, 24, and 48 h. The mean ± SD cell counts (in millions) were 1.90 ± 0.26 at 3 h, 2.70 ± 0.61 at 24 h, and 4.46 ± 0.69 at 48 h. Statistical analysis (one-way ANOVA) showed a significant difference (p < 0.01) in cell count between 3 and 48 h. N = 3 for each time point.
Detection of Metabolic Activity via dDNP-NMR
Figure 5
Figure 5. One chamber chip hyperpolarized 13C NMR. (A) spectrum with the highest lactate signal (spectrum 9), showing pyruvate C2 at 205 ppm, lactate C1 at 182.3 ppm, pyruvate hydrate C1 at 178.4 ppm, and pyruvate C1 at 171 ppm. (SNR of highest lactate peak was approximately 50) (B) time-series spectra displaying the production of [1-13C] lactate in one-chamber chips at 3, 24, and 48 h postseeding, following the injection of 10 mM hyperpolarized [1-13C] pyruvate. (C) Rate constants normalized to cell counts, yielding mean ± SD values (s–1) of (4.12 ± 0.53) × 10–4 at 3 h, (3.76 ± 0.64) × 10–4 at 24 h, and (4.78 ± 0.53) × 10–4 at 48 h. Statistical analysis (one-way ANOVA) showed no significant difference (NS) between normalized rate constants. N = 3 for each time point.
Comparative Analysis of One-Chamber and Two-Chamber Chips
Figure 6
Figure 6. Two-chamber chip cell growth and lactate production. (A) two-chamber chip seeded with 3 × 106 HeLa cells, using a flow rate of 0.200 mL/min in the lower chamber, and no flow in the top chamber, for 24 h at 37 °C and 5% CO2. Bright-field microscopy image (bottom left, BF 20×) and fluorescence microscopy image of cells stained with Calcein AM (bottom right, Calcein AM 20×). Scale bar: 100 μm. (B) Time-series spectra showing [1-13C] lactate accumulation in the two-chamber chip seeded with 3 × 106 HeLa cells, following injection of 10 mM hyperpolarized [1-13C] pyruvate. The mean ± SD rate constant for lactate production was 2.55 ± 0.28 × 10–4 (n = 2).
Conclusion
Data Availability
All data are available from the corresponding author upon request.
Supporting Information
The Supporting Information is available free of charge at https://pubs.acs.org/doi/10.1021/acs.analchem.5c04058.
Return loss curves for the chip probe, Simulated B1+ fields of 2H and 1H coils, Rate constants from dDNP-NMR measurements in microfluidic chips not normalized to cell count (PDF)
Terms & Conditions
Most electronic Supporting Information files are available without a subscription to ACS Web Editions. Such files may be downloaded by article for research use (if there is a public use license linked to the relevant article, that license may permit other uses). Permission may be obtained from ACS for other uses through requests via the RightsLink permission system: http://pubs.acs.org/page/copyright/permissions.html.
Acknowledgments
This research was funded by the Danish National Research Foundation (grant DNRF124), EU Interreg Öresund-Kattegat-Skagerrak project “Hanseatic Life Science Research Infrastructure Consortium” (HALRIC) and the Novo Nordisk Foundation (infrastructure grant NNF19OC0055825). JDSH contribution to this work was supported by Fundación Séneca (22401/SF/23).
References
This article references 54 other publications.
- 1Preclinical CRO Market Size, Share & Growth | Report [2030]. https://www.fortunebusinessinsights.com/preclinical-cro-market-106939 (accessed 2024–07–22).Google ScholarThere is no corresponding record for this reference.
- 2Helmlinger, G.; Al-Huniti, N.; Aksenov, S.; Peskov, K.; Hallow, K. M.; Chu, L.; Boulton, D.; Eriksson, U.; Hamrén, B.; Lambert, C.; Masson, E.; Tomkinson, H.; Stanski, D. Drug–disease modeling in the pharmaceutical industry─Where mechanistic systems pharmacology and statistical pharmacometrics meet. Eur. J. Pharm. Sci. 2017, 109, S39– S46, DOI: 10.1016/j.ejps.2017.05.028Google ScholarThere is no corresponding record for this reference.
- 3Kapałczyńska, M.; Kolenda, T.; Przybyła, W.; Zajączkowska, M.; Teresiak, A.; Filas, V.; Ibbs, M.; Bliźniak, R.; Łuczewski, Ł.; Lamperska, K. 2D and 3D cell cultures─A comparison of different types of cancer cell cultures. Arch. Med. Sci. 2018, 14 (4), 910– 919, DOI: 10.5114/aoms.2016.63743Google ScholarThere is no corresponding record for this reference.
- 4Zingales, V.; Esposito, M. R.; Torriero, N.; Taroncher, M.; Cimetta, E.; Ruiz, M. J. The growing importance of three-dimensional models and microphysiological systems in the assessment of mycotoxin toxicity. Toxins 2023, 15 (7), 422, DOI: 10.3390/toxins15070422Google ScholarThere is no corresponding record for this reference.
- 5Singh, D.; Mathur, A.; Arora, S.; Roy, S.; Mahindroo, N. Journey of organ-on-a-chip technology and its role in future healthcare scenario. Appl. Surf. Sci. Adv. 2022, 9, 100246, DOI: 10.1016/j.apsadv.2022.100246Google ScholarThere is no corresponding record for this reference.
- 6Moro, L. G.; Guarnier, L. P.; Azevedo, M. F.; Fracasso, J. A. R.; Lucio, M. A.; Castro, M. V. d.; Dias, M. L.; Lívero, F. A. d. R.; Ribeiro-Paes, J. T. A brief history of cell culture: From Harrison to organs-on-a-chip. Cells 2024, 13 (24), 2068, DOI: 10.3390/cells13242068Google ScholarThere is no corresponding record for this reference.
- 7Huh, D.; Matthews, B. D.; Mammoto, A.; Montoya-Zavala, M.; Hsin, H. Y.; Ingber, D. E. Reconstituting organ-level lung functions on a chip. Science 2010, 328 (5986), 1662– 1668, DOI: 10.1126/science.1188302Google ScholarThere is no corresponding record for this reference.
- 8Yan, J.; Li, Z.; Guo, J.; Liu, S.; Guo, J. Organ-on-a-chip: A new tool for in vitro research. Biosens. Bioelectron. 2022, 216, 114626, DOI: 10.1016/j.bios.2022.114626Google ScholarThere is no corresponding record for this reference.
- 9Ingber, D. E. Human organs-on-chips for disease modelling, drug development and personalized medicine. Nat. Rev. Genet. 2022, 23 (8), 467– 491, DOI: 10.1038/s41576-022-00466-9Google ScholarThere is no corresponding record for this reference.
- 10Ma, C.; Peng, Y.; Li, H.; Chen, W. Organ-on-a-chip: A new paradigm for drug development. Trends Pharmacol. Sci. 2021, 42 (2), 119– 133, DOI: 10.1016/j.tips.2020.11.009Google ScholarThere is no corresponding record for this reference.
- 11Leung, C. M.; de Haan, P.; Ronaldson-Bouchard, K.; Kim, G. A.; Ko, J.; Rho, H. S.; Chen, Z.; Habibovic, P.; Jeon, N. L.; Takayama, S.; Shuler, M. L.; Vunjak-Novakovic, G.; Frey, O.; Verpoorte, E.; Toh, Y. C. A guide to the organ-on-a-chip. Nat. Rev. Methods Primers 2022, 2, 33, DOI: 10.1038/s43586-022-00118-6Google ScholarThere is no corresponding record for this reference.
- 12Dorst, J.; Weydt, P.; Brenner, D. Metabolic alterations precede neurofilament changes in presymptomatic ALS gene carriers. EBioMedicine 2023, 90, 104521, DOI: 10.1016/j.ebiom.2023.104521Google ScholarThere is no corresponding record for this reference.
- 13Tang, C. C.; Poston, K. L.; Dhawan, V.; Eidelberg, D. Abnormalities in metabolic network activity precede the onset of motor symptoms in Parkinson’s disease. J. Neurosci. 2010, 30 (3), 1049– 1056, DOI: 10.1523/JNEUROSCI.4188-09.2010Google ScholarThere is no corresponding record for this reference.
- 14Duarte, J. M. N.; Schuck, P. F.; Wenk, G. L.; Ferreira, G. C. Metabolic disturbances in diseases with neurological involvement. Aging Dis 2014, 5 (4), 238– 255, DOI: 10.14336/AD.2014.0500238Google ScholarThere is no corresponding record for this reference.
- 15Hu, S.; Balakrishnan, A.; Bok, R. A. 13C-pyruvate imaging reveals alterations in glycolysis that precede c-Myc-induced tumor formation and regression. Cell Metab. 2011, 14 (1), 131– 142, DOI: 10.1016/j.cmet.2011.04.012Google ScholarThere is no corresponding record for this reference.
- 16Sánchez-Salazar, M. G. Continuous inline monitoring of glucose in an organ-on-chip using FreeStyle Libre glucometers. Front. Lab Chip Technol. 2024, 3, 1289437, DOI: 10.3389/frlct.2024.1289437Google ScholarThere is no corresponding record for this reference.
- 17Zirath, H. Every breath you take: Non-invasive real-time oxygen biosensing in two- and three-dimensional microfluidic cell models. Front. Physiol. 2018, 9, 815, DOI: 10.3389/fphys.2018.00815Google ScholarThere is no corresponding record for this reference.
- 18Prill, S. Real-time monitoring of oxygen uptake in hepatic bioreactors shows mitochondrial toxicity. Arch. Toxicol. 2016, 90 (5), 1181– 1191, DOI: 10.1007/s00204-015-1537-2Google ScholarThere is no corresponding record for this reference.
- 19Weltin, A. Cell culture monitoring for drug screening and cancer research. Lab Chip 2014, 14 (1), 138– 146, DOI: 10.1039/C3LC50759AGoogle ScholarThere is no corresponding record for this reference.
- 20Bavli, D. Real-time monitoring of metabolic function in liver-on-chip microdevices. Proc. Natl. Acad. Sci. U.S.A. 2016, 113 (16), E2231– E2240, DOI: 10.1073/pnas.1522556113Google ScholarThere is no corresponding record for this reference.
- 21Dornhof, J.; Kieninger, J.; Muralidharan, H.; Maurer, J.; Urban, G. A.; Weltin, A. Microfluidic organ-on-chip system for multi-analyte monitoring of metabolites in 3D cell cultures. Lab Chip 2022, 22 (2), 225– 239, DOI: 10.1039/D1LC00689DGoogle ScholarThere is no corresponding record for this reference.
- 22Ferrari, E.; Palma, C.; Vesentini, S.; Occhetta, P.; Rasponi, M. Integrating biosensors in organs-on-chip devices: A perspective on current strategies to monitor microphysiological systems. Biosensors 2020, 10 (9), 110, DOI: 10.3390/bios10090110Google ScholarThere is no corresponding record for this reference.
- 23Izadifar, Z.; Charrez, B.; Almeida, M.; Robben, S.; Pilobello, K.; van der Graaf-Mas, J.; Marquez, S. L.; Ferrante, T. C.; Shcherbina, K.; Gould, R.; LoGrande, N. T.; Sesay, A. M.; Ingber, D. E. Organ chips with integrated multifunctional sensors enable continuous metabolic monitoring at controlled oxygen levels. Biosens. Bioelectron. 2024, 265, 116683, DOI: 10.1016/j.bios.2024.116683Google ScholarThere is no corresponding record for this reference.
- 24Henry, O. Y. F.; Villenave, R.; Cronce, M. J.; Leineweber, W. D.; Benz, M. A.; Ingber, D. E. Organs-on-chips with integrated electrodes for transepithelial electrical resistance measurements of human epithelial barrier function. Lab Chip 2017, 17 (13), 2264– 2271, DOI: 10.1039/C7LC00155JGoogle ScholarThere is no corresponding record for this reference.
- 25Ardenkjær-Larsen, J. H.; Fridlund, B.; Gram, A.; Hansson, G.; Hansson, L.; Lerche, M. H.; Servin, R.; Thaning, M.; Golman, K. Increase in signal-to-noise ratio of > 10,000 times in liquid-state NMR. Proc. Natl. Acad. Sci. U.S.A. 2003, 100 (18), 10158– 10163, DOI: 10.1073/pnas.1733835100Google ScholarThere is no corresponding record for this reference.
- 26Eills, J.; Hale, W.; Utz, M. Synergies between hyperpolarized NMR and microfluidics: A review. Prog. Nucl. Magn. Reson. Spectrosc. 2022, 128, 44– 69, DOI: 10.1016/j.pnmrs.2021.09.001Google ScholarThere is no corresponding record for this reference.
- 27Barker, S. J.; Dagys, L.; Levitt, M. H.; Utz, M. Efficient parahydrogen-induced ̂13C hyperpolarization on a microfluidic device. J. Am. Chem. Soc. 2024, 146 (27), 18379– 18386, DOI: 10.1021/jacs.4c03271Google ScholarThere is no corresponding record for this reference.
- 28Rogers, G.; Barker, S.; Sharma, M.; Khakoo, S.; Utz, M. Operando NMR metabolomics of a microfluidic cell culture. J. Magn. Reson. 2023, 349, 107405, DOI: 10.1016/j.jmr.2023.107405Google ScholarThere is no corresponding record for this reference.
- 29Mangas-Florencio, L.; Herrero-Gómez, A.; Eills, J.; Azagra, M.; Batlló-Rius, M.; Marco-Rius, I. A DIY bioreactor for in situ metabolic tracking in 3D cell models via hyperpolarized 13C NMR spectroscopy. Anal. Chem. 2025, 97 (3), 1594– 1602, DOI: 10.1021/acs.analchem.4c04183Google ScholarThere is no corresponding record for this reference.
- 30Mathiassen, T. B. W.; Høgh, A. E.; Karlsson, M.; Katsikis, S.; Wang, K.; Pennestri, M.; Ardenkjær-Larsen, J. H.; Jensen, P. R. Hyperpolarized 13C NMR for longitudinal in-cell metabolism using a mobile 3D cell culture system. J. Magn. Reson. Open 2023, 16–17, 100131, DOI: 10.1016/j.jmro.2023.100131Google ScholarThere is no corresponding record for this reference.
- 31Tan, H. Y.; Trier, S.; Rahbek, U. L.; Dufva, M.; Kutter, J. P.; Andresen, T. L. A multi-chamber microfluidic intestinal barrier model using Caco-2 cells for drug transport studies. PLoS One 2018, 13 (5), e0197101 DOI: 10.1371/journal.pone.0197101Google ScholarThere is no corresponding record for this reference.
- 32Harris, T.; Eliyahu, G.; Frydman, L.; Degani, H. Kinetics of hyperpolarized ̂13C1-pyruvate transport and metabolism in living human breast cancer cells. Proc. Natl. Acad. Sci. U.S.A. 2009, 106 (43), 18131– 18136, DOI: 10.1073/pnas.0909049106Google ScholarThere is no corresponding record for this reference.
- 33Christensen, N. V.; Holm, R.; Sanchez, J. D.; Hansen, E. S. S.; Lerche, M. H.; Ardenkjær-Larsen, J. H.; Laustsen, C.; Bertelsen, L. B. A continuous-flow bioreactor system for high-throughput hyperpolarized metabolic flux analysis. NMR Biomed 2024, 37 (5), e5107 DOI: 10.1002/nbm.5107Google ScholarThere is no corresponding record for this reference.
- 34Christensen, N. V.; Laustsen, C.; Bertelsen, L. B. Differentiating leukemia subtypes based on metabolic signatures using hyperpolarized ̂13C NMR. NMR Biomed 2024, 37 (12), e5264 DOI: 10.1002/nbm.5264Google ScholarThere is no corresponding record for this reference.
- 35Massin, C.; Vincent, F.; Homsy, A.; Ehrmann, K.; Boero, G.; Besse, P. A.; Daridon, A.; Verpoorte, E.; de Rooij, N. F.; Popovic, R. S. Planar microcoil-based microfluidic NMR probes. J. Magn. Reson. 2003, 164 (2), 242– 255, DOI: 10.1016/S1090-7807(03)00151-4Google ScholarThere is no corresponding record for this reference.
- 36Wensink, H.; Benito-Lopez, F.; Hermes, D. C.; Verboom, W.; Gardeniers, H. J. G. E.; Reinhoudt, D. N.; van den Berg, A. Measuring reaction kinetics in a lab-on-a-chip by microcoil NMR. Lab Chip 2005, 5 (3), 280– 284, DOI: 10.1039/b414832kGoogle ScholarThere is no corresponding record for this reference.
- 37Liu, Y.; Mi, Y.; Mueller, T.; Kreibich, S.; Williams, E. G.; van Drogen, A.; Borel, C.; Frank, M.; Germain, P. L.; Bludau, I. Multi-omic measurements of heterogeneity in HeLa cells across laboratories. Nat. Biotechnol. 2019, 37 (3), 314– 322, DOI: 10.1038/s41587-019-0037-yGoogle ScholarThere is no corresponding record for this reference.
- 38Mason, J. A.; Davison-Versagli, C. A.; Leliaert, A. K.; Pape, D. J.; McCallister, C.; Zuo, J.; Durbin, S. M.; Buchheit, C. L.; Zhang, S.; Schafer, Z. T. Oncogenic Ras differentially regulates metabolism and anoikis in extracellular matrix-detached cells. Cell Death Differ. 2016, 23 (8), 1271– 1282, DOI: 10.1038/cdd.2016.15Google ScholarThere is no corresponding record for this reference.
- 39Buchheit, C. L.; Rayavarapu, R. R.; Schafer, Z. T. The regulation of cancer cell death and metabolism by extracellular matrix attachment. Semin. Cell Dev. Biol. 2012, 23 (4), 402– 411, DOI: 10.1016/j.semcdb.2012.04.007Google ScholarThere is no corresponding record for this reference.
- 40Schafer, Z. T.; Grassian, A. R.; Song, L.; Jiang, Z.; Gerhart-Hines, Z.; Irie, H. Y.; Gao, S.; Puigserver, P.; Brugge, J. S. Antioxidant and oncogene rescue of metabolic defects caused by loss of matrix attachment. Nature 2009, 461 (7260), 109– 113, DOI: 10.1038/nature08268Google ScholarThere is no corresponding record for this reference.
- 41Avivar-Valderas, A.; Salas, E.; Bobrovnikova-Marjon, E.; Diehl, J. A.; Nagi, C.; Debnath, J.; Aguirre-Ghiso, J. A. PERK integrates autophagy and oxidative stress responses to promote survival during extracellular matrix detachment. Mol. Cell. Biol. 2011, 31 (17), 3616– 3629, DOI: 10.1128/MCB.05164-11Google ScholarThere is no corresponding record for this reference.
- 42Hawk, M. A.; Schafer, Z. T. Mechanisms of redox metabolism and cancer cell survival during extracellular matrix detachment. J. Biol. Chem. 2018, 293 (20), 7531– 7538, DOI: 10.1074/jbc.TM117.000260Google ScholarThere is no corresponding record for this reference.
- 43Takakusagi, Y.; Takakusagi, K.; Inoue, K.; Saito, K.; Homma, Y.; Ichikawa, K. Hyperpolarized [1–13C]pyruvate NMR spectroscopy reveals transition of tumor energy metabolism in microscale multicellular spheroids. Sci. Rep. 2025, 15 (1), 3454, DOI: 10.1038/s41598-025-03454-1Google ScholarThere is no corresponding record for this reference.
- 44Nielsen, N.; Laustsen, C.; Bertelsen, L. B. ̂13C dynamic nuclear polarization for measuring metabolic flux in endothelial progenitor cells. Exp. Cell Res. 2016, 349 (1), 95– 100, DOI: 10.1016/j.yexcr.2016.10.002Google ScholarThere is no corresponding record for this reference.
- 45Harris, T.; Degani, H.; Frydman, L. Hyperpolarized 13C NMR studies of glucose metabolism in living breast cancer cell cultures. NMR Biomed 2013, 26 (12), 1831– 1843, DOI: 10.1002/nbm.3024Google ScholarThere is no corresponding record for this reference.
- 46Sriram, R.; Nguyen, J.; Santos, J. D. L.; Nguyen, L.; Sun, J.; Vigneron, S.; van Criekinge, M.; Kurhanewicz, J.; MacKenzie, J. D. Molecular detection of inflammation in cell models using hyperpolarized ̂13C-pyruvate. Theranostics 2018, 8 (12), 3400– 3411, DOI: 10.7150/thno.24322Google ScholarThere is no corresponding record for this reference.
- 47Keshari, K. R.; Kurhanewicz, J.; Jeffries, R. E.; Wilson, D. M.; Dewar, B. J.; van Criekinge, M.; Zierhut, M.; Vigneron, D. B.; Macdonald, J. M. Hyperpolarized ̂13C spectroscopy and an NMR-compatible bioreactor system for the investigation of real-time cellular metabolism. Magn. Reson. Med. 2010, 63 (2), 322– 329, DOI: 10.1002/mrm.22225Google ScholarThere is no corresponding record for this reference.
- 48Thuenauer, R.; Nicklaus, S.; Frensch, M.; Troendle, K.; Madl, J.; Römer, W. A microfluidic biochip for locally confined stimulation of cells within an epithelial monolayer. RSC Adv. 2018, 8 (14), 7839– 7846, DOI: 10.1039/C7RA11943GGoogle ScholarThere is no corresponding record for this reference.
- 49Thuenauer, R.; Rodriguez-Boulan, E.; Römer, W. Microfluidic approaches for epithelial cell layer culture and characterisation. Analyst 2014, 139 (13), 3206– 3218, DOI: 10.1039/C4AN00056KGoogle ScholarThere is no corresponding record for this reference.
- 50Kim, H. J.; Ingber, D. E. Gut-on-a-chip microenvironment induces human intestinal cells to undergo villus differentiation. Integr. Biol. 2013, 5 (9), 1130– 1140, DOI: 10.1039/c3ib40126jGoogle ScholarThere is no corresponding record for this reference.
- 51Chung, H. H.; Mireles, M.; Kwarta, B. J.; Gaborski, T. R. Use of porous membranes in tissue barrier and co-culture models. Lab Chip 2018, 18 (12), 1671– 1689, DOI: 10.1039/C7LC01248AGoogle ScholarThere is no corresponding record for this reference.
- 52Maschmeyer, I.; Lorenz, A. K.; Schimek, K.; Hasenberg, T.; Ramme, A. P.; Hübner, J.; Lindner, M.; Drewell, C.; Bauer, S.; Thomas, A.; Sambo, N. S.; Sonntag, F.; Lauster, R.; Marx, U. A four-organ-chip for interconnected long-term co-culture of human intestine, liver, skin, and kidney equivalents. Lab Chip 2015, 15 (12), 2688– 2699, DOI: 10.1039/C5LC00392JGoogle ScholarThere is no corresponding record for this reference.
- 53Ferruzza, S.; Rossi, C.; Scarino, M. L.; Sambuy, Y. A protocol for differentiation of human intestinal Caco-2 cells in asymmetric serum-containing medium. Toxicol. In Vitro 2012, 26 (8), 1252– 1255, DOI: 10.1016/j.tiv.2012.01.008Google ScholarThere is no corresponding record for this reference.
- 54Kopp, B.; Khawam, A.; Di Perna, K.; Lenart, D.; Vinette, M.; Silva, R.; Zanoni, T. B.; Rore, C.; Guenigault, G.; Richardson, E.; Kostrzewski, T.; Boswell, A.; Van, P.; Valentine, C.; Salk, J.; Hamel, A. Liver-on-chip model and application in predictive genotoxicity and mutagenicity of drugs. Mutat. Res. Genet. Toxicol. Environ. Mutagen. 2024, 896, 503762, DOI: 10.1016/j.mrgentox.2024.503762Google ScholarThere is no corresponding record for this reference.
Cited By
This article has not yet been cited by other publications.
Article Views
Altmetric
Citations
Article Views are the COUNTER-compliant sum of full text article downloads since November 2008 (both PDF and HTML) across all institutions and individuals. These metrics are regularly updated to reflect usage leading up to the last few days.
Citations are the number of other articles citing this article, calculated by Crossref and updated daily. Find more information about Crossref citation counts.
The Altmetric Attention Score is a quantitative measure of the attention that a research article has received online. Clicking on the donut icon will load a page at altmetric.com with additional details about the score and the social media presence for the given article. Find more information on the Altmetric Attention Score and how the score is calculated.
Recommended Articles
Abstract

Figure 1
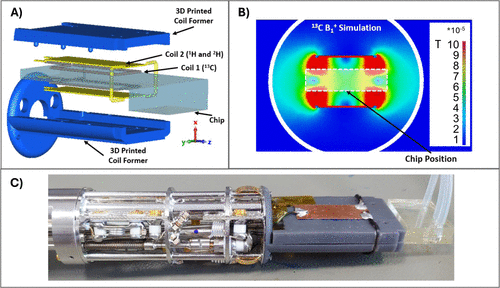
Figure 1. RF Coil design. (A) exploded view of the NMR probe head, showing the two RF coils, the 3D-printed support, and the microfluidic chip. (B) Cross-section (Z = 0 plane) of the simulated B1+ field distribution generated by Coil 1 at the 13C frequency (per 1 W accepted power). (C) Photograph of the fabricated NMR probe with the chip inserted.
Figure 2
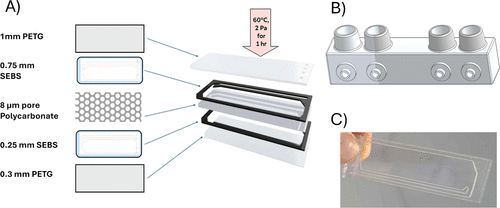
Figure 2. Fluidic chip production and assembly. (A) visual representation of the chip components and their assembly process. The stacked components are shown in order from top to bottom: 1 mm PETG, 0.75 mm SEBS, track-etched polycarbonate membrane (8 μm pore size, 18 μm thickness, 5% porosity), 0.25 mm SEBS, and 0.3 mm PETG. (B) 3D-printed inlets/injection ports, designed using SolidWorks, shown with female mini Luer connectors made from BioMed clear resin. (C) Fully assembled microfluidic chip.
Figure 3
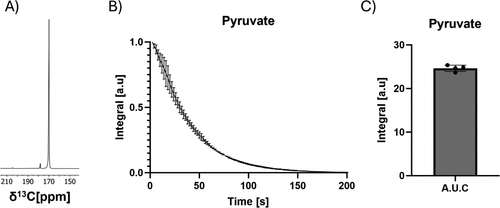
Figure 3. Performance evaluation. (A) single pyruvate spectrum, SNR of the highest pyruvate peak was 5000, (B) substrate integral over time (n = 3) and (C) comparative substrate integrals (AUC = 24 ± 0.6) following injection of 10 mM hyperpolarized [1-13C] pyruvate, CV for the AUC was 2.5%. T1 of pyruvate was 52 ± 2 s.
Figure 4

Figure 4. One-chamber chip cell growth. (A) one-chamber chip seeded with 3 × 106 HeLa cells, cultured for 3 h, and imaged with bright-field microscopy (bottom left, BF 20×) and fluorescence microscopy of cells stained with Calcein AM (bottom right, Calcein AM 20×). Scale bar: 100 μm. (B) Cell counts in chips after 3, 24, and 48 h. The mean ± SD cell counts (in millions) were 1.90 ± 0.26 at 3 h, 2.70 ± 0.61 at 24 h, and 4.46 ± 0.69 at 48 h. Statistical analysis (one-way ANOVA) showed a significant difference (p < 0.01) in cell count between 3 and 48 h. N = 3 for each time point.
Figure 5

Figure 5. One chamber chip hyperpolarized 13C NMR. (A) spectrum with the highest lactate signal (spectrum 9), showing pyruvate C2 at 205 ppm, lactate C1 at 182.3 ppm, pyruvate hydrate C1 at 178.4 ppm, and pyruvate C1 at 171 ppm. (SNR of highest lactate peak was approximately 50) (B) time-series spectra displaying the production of [1-13C] lactate in one-chamber chips at 3, 24, and 48 h postseeding, following the injection of 10 mM hyperpolarized [1-13C] pyruvate. (C) Rate constants normalized to cell counts, yielding mean ± SD values (s–1) of (4.12 ± 0.53) × 10–4 at 3 h, (3.76 ± 0.64) × 10–4 at 24 h, and (4.78 ± 0.53) × 10–4 at 48 h. Statistical analysis (one-way ANOVA) showed no significant difference (NS) between normalized rate constants. N = 3 for each time point.
Figure 6

Figure 6. Two-chamber chip cell growth and lactate production. (A) two-chamber chip seeded with 3 × 106 HeLa cells, using a flow rate of 0.200 mL/min in the lower chamber, and no flow in the top chamber, for 24 h at 37 °C and 5% CO2. Bright-field microscopy image (bottom left, BF 20×) and fluorescence microscopy image of cells stained with Calcein AM (bottom right, Calcein AM 20×). Scale bar: 100 μm. (B) Time-series spectra showing [1-13C] lactate accumulation in the two-chamber chip seeded with 3 × 106 HeLa cells, following injection of 10 mM hyperpolarized [1-13C] pyruvate. The mean ± SD rate constant for lactate production was 2.55 ± 0.28 × 10–4 (n = 2).
References
This article references 54 other publications.
- 1Preclinical CRO Market Size, Share & Growth | Report [2030]. https://www.fortunebusinessinsights.com/preclinical-cro-market-106939 (accessed 2024–07–22).There is no corresponding record for this reference.
- 2Helmlinger, G.; Al-Huniti, N.; Aksenov, S.; Peskov, K.; Hallow, K. M.; Chu, L.; Boulton, D.; Eriksson, U.; Hamrén, B.; Lambert, C.; Masson, E.; Tomkinson, H.; Stanski, D. Drug–disease modeling in the pharmaceutical industry─Where mechanistic systems pharmacology and statistical pharmacometrics meet. Eur. J. Pharm. Sci. 2017, 109, S39– S46, DOI: 10.1016/j.ejps.2017.05.028There is no corresponding record for this reference.
- 3Kapałczyńska, M.; Kolenda, T.; Przybyła, W.; Zajączkowska, M.; Teresiak, A.; Filas, V.; Ibbs, M.; Bliźniak, R.; Łuczewski, Ł.; Lamperska, K. 2D and 3D cell cultures─A comparison of different types of cancer cell cultures. Arch. Med. Sci. 2018, 14 (4), 910– 919, DOI: 10.5114/aoms.2016.63743There is no corresponding record for this reference.
- 4Zingales, V.; Esposito, M. R.; Torriero, N.; Taroncher, M.; Cimetta, E.; Ruiz, M. J. The growing importance of three-dimensional models and microphysiological systems in the assessment of mycotoxin toxicity. Toxins 2023, 15 (7), 422, DOI: 10.3390/toxins15070422There is no corresponding record for this reference.
- 5Singh, D.; Mathur, A.; Arora, S.; Roy, S.; Mahindroo, N. Journey of organ-on-a-chip technology and its role in future healthcare scenario. Appl. Surf. Sci. Adv. 2022, 9, 100246, DOI: 10.1016/j.apsadv.2022.100246There is no corresponding record for this reference.
- 6Moro, L. G.; Guarnier, L. P.; Azevedo, M. F.; Fracasso, J. A. R.; Lucio, M. A.; Castro, M. V. d.; Dias, M. L.; Lívero, F. A. d. R.; Ribeiro-Paes, J. T. A brief history of cell culture: From Harrison to organs-on-a-chip. Cells 2024, 13 (24), 2068, DOI: 10.3390/cells13242068There is no corresponding record for this reference.
- 7Huh, D.; Matthews, B. D.; Mammoto, A.; Montoya-Zavala, M.; Hsin, H. Y.; Ingber, D. E. Reconstituting organ-level lung functions on a chip. Science 2010, 328 (5986), 1662– 1668, DOI: 10.1126/science.1188302There is no corresponding record for this reference.
- 8Yan, J.; Li, Z.; Guo, J.; Liu, S.; Guo, J. Organ-on-a-chip: A new tool for in vitro research. Biosens. Bioelectron. 2022, 216, 114626, DOI: 10.1016/j.bios.2022.114626There is no corresponding record for this reference.
- 9Ingber, D. E. Human organs-on-chips for disease modelling, drug development and personalized medicine. Nat. Rev. Genet. 2022, 23 (8), 467– 491, DOI: 10.1038/s41576-022-00466-9There is no corresponding record for this reference.
- 10Ma, C.; Peng, Y.; Li, H.; Chen, W. Organ-on-a-chip: A new paradigm for drug development. Trends Pharmacol. Sci. 2021, 42 (2), 119– 133, DOI: 10.1016/j.tips.2020.11.009There is no corresponding record for this reference.
- 11Leung, C. M.; de Haan, P.; Ronaldson-Bouchard, K.; Kim, G. A.; Ko, J.; Rho, H. S.; Chen, Z.; Habibovic, P.; Jeon, N. L.; Takayama, S.; Shuler, M. L.; Vunjak-Novakovic, G.; Frey, O.; Verpoorte, E.; Toh, Y. C. A guide to the organ-on-a-chip. Nat. Rev. Methods Primers 2022, 2, 33, DOI: 10.1038/s43586-022-00118-6There is no corresponding record for this reference.
- 12Dorst, J.; Weydt, P.; Brenner, D. Metabolic alterations precede neurofilament changes in presymptomatic ALS gene carriers. EBioMedicine 2023, 90, 104521, DOI: 10.1016/j.ebiom.2023.104521There is no corresponding record for this reference.
- 13Tang, C. C.; Poston, K. L.; Dhawan, V.; Eidelberg, D. Abnormalities in metabolic network activity precede the onset of motor symptoms in Parkinson’s disease. J. Neurosci. 2010, 30 (3), 1049– 1056, DOI: 10.1523/JNEUROSCI.4188-09.2010There is no corresponding record for this reference.
- 14Duarte, J. M. N.; Schuck, P. F.; Wenk, G. L.; Ferreira, G. C. Metabolic disturbances in diseases with neurological involvement. Aging Dis 2014, 5 (4), 238– 255, DOI: 10.14336/AD.2014.0500238There is no corresponding record for this reference.
- 15Hu, S.; Balakrishnan, A.; Bok, R. A. 13C-pyruvate imaging reveals alterations in glycolysis that precede c-Myc-induced tumor formation and regression. Cell Metab. 2011, 14 (1), 131– 142, DOI: 10.1016/j.cmet.2011.04.012There is no corresponding record for this reference.
- 16Sánchez-Salazar, M. G. Continuous inline monitoring of glucose in an organ-on-chip using FreeStyle Libre glucometers. Front. Lab Chip Technol. 2024, 3, 1289437, DOI: 10.3389/frlct.2024.1289437There is no corresponding record for this reference.
- 17Zirath, H. Every breath you take: Non-invasive real-time oxygen biosensing in two- and three-dimensional microfluidic cell models. Front. Physiol. 2018, 9, 815, DOI: 10.3389/fphys.2018.00815There is no corresponding record for this reference.
- 18Prill, S. Real-time monitoring of oxygen uptake in hepatic bioreactors shows mitochondrial toxicity. Arch. Toxicol. 2016, 90 (5), 1181– 1191, DOI: 10.1007/s00204-015-1537-2There is no corresponding record for this reference.
- 19Weltin, A. Cell culture monitoring for drug screening and cancer research. Lab Chip 2014, 14 (1), 138– 146, DOI: 10.1039/C3LC50759AThere is no corresponding record for this reference.
- 20Bavli, D. Real-time monitoring of metabolic function in liver-on-chip microdevices. Proc. Natl. Acad. Sci. U.S.A. 2016, 113 (16), E2231– E2240, DOI: 10.1073/pnas.1522556113There is no corresponding record for this reference.
- 21Dornhof, J.; Kieninger, J.; Muralidharan, H.; Maurer, J.; Urban, G. A.; Weltin, A. Microfluidic organ-on-chip system for multi-analyte monitoring of metabolites in 3D cell cultures. Lab Chip 2022, 22 (2), 225– 239, DOI: 10.1039/D1LC00689DThere is no corresponding record for this reference.
- 22Ferrari, E.; Palma, C.; Vesentini, S.; Occhetta, P.; Rasponi, M. Integrating biosensors in organs-on-chip devices: A perspective on current strategies to monitor microphysiological systems. Biosensors 2020, 10 (9), 110, DOI: 10.3390/bios10090110There is no corresponding record for this reference.
- 23Izadifar, Z.; Charrez, B.; Almeida, M.; Robben, S.; Pilobello, K.; van der Graaf-Mas, J.; Marquez, S. L.; Ferrante, T. C.; Shcherbina, K.; Gould, R.; LoGrande, N. T.; Sesay, A. M.; Ingber, D. E. Organ chips with integrated multifunctional sensors enable continuous metabolic monitoring at controlled oxygen levels. Biosens. Bioelectron. 2024, 265, 116683, DOI: 10.1016/j.bios.2024.116683There is no corresponding record for this reference.
- 24Henry, O. Y. F.; Villenave, R.; Cronce, M. J.; Leineweber, W. D.; Benz, M. A.; Ingber, D. E. Organs-on-chips with integrated electrodes for transepithelial electrical resistance measurements of human epithelial barrier function. Lab Chip 2017, 17 (13), 2264– 2271, DOI: 10.1039/C7LC00155JThere is no corresponding record for this reference.
- 25Ardenkjær-Larsen, J. H.; Fridlund, B.; Gram, A.; Hansson, G.; Hansson, L.; Lerche, M. H.; Servin, R.; Thaning, M.; Golman, K. Increase in signal-to-noise ratio of > 10,000 times in liquid-state NMR. Proc. Natl. Acad. Sci. U.S.A. 2003, 100 (18), 10158– 10163, DOI: 10.1073/pnas.1733835100There is no corresponding record for this reference.
- 26Eills, J.; Hale, W.; Utz, M. Synergies between hyperpolarized NMR and microfluidics: A review. Prog. Nucl. Magn. Reson. Spectrosc. 2022, 128, 44– 69, DOI: 10.1016/j.pnmrs.2021.09.001There is no corresponding record for this reference.
- 27Barker, S. J.; Dagys, L.; Levitt, M. H.; Utz, M. Efficient parahydrogen-induced ̂13C hyperpolarization on a microfluidic device. J. Am. Chem. Soc. 2024, 146 (27), 18379– 18386, DOI: 10.1021/jacs.4c03271There is no corresponding record for this reference.
- 28Rogers, G.; Barker, S.; Sharma, M.; Khakoo, S.; Utz, M. Operando NMR metabolomics of a microfluidic cell culture. J. Magn. Reson. 2023, 349, 107405, DOI: 10.1016/j.jmr.2023.107405There is no corresponding record for this reference.
- 29Mangas-Florencio, L.; Herrero-Gómez, A.; Eills, J.; Azagra, M.; Batlló-Rius, M.; Marco-Rius, I. A DIY bioreactor for in situ metabolic tracking in 3D cell models via hyperpolarized 13C NMR spectroscopy. Anal. Chem. 2025, 97 (3), 1594– 1602, DOI: 10.1021/acs.analchem.4c04183There is no corresponding record for this reference.
- 30Mathiassen, T. B. W.; Høgh, A. E.; Karlsson, M.; Katsikis, S.; Wang, K.; Pennestri, M.; Ardenkjær-Larsen, J. H.; Jensen, P. R. Hyperpolarized 13C NMR for longitudinal in-cell metabolism using a mobile 3D cell culture system. J. Magn. Reson. Open 2023, 16–17, 100131, DOI: 10.1016/j.jmro.2023.100131There is no corresponding record for this reference.
- 31Tan, H. Y.; Trier, S.; Rahbek, U. L.; Dufva, M.; Kutter, J. P.; Andresen, T. L. A multi-chamber microfluidic intestinal barrier model using Caco-2 cells for drug transport studies. PLoS One 2018, 13 (5), e0197101 DOI: 10.1371/journal.pone.0197101There is no corresponding record for this reference.
- 32Harris, T.; Eliyahu, G.; Frydman, L.; Degani, H. Kinetics of hyperpolarized ̂13C1-pyruvate transport and metabolism in living human breast cancer cells. Proc. Natl. Acad. Sci. U.S.A. 2009, 106 (43), 18131– 18136, DOI: 10.1073/pnas.0909049106There is no corresponding record for this reference.
- 33Christensen, N. V.; Holm, R.; Sanchez, J. D.; Hansen, E. S. S.; Lerche, M. H.; Ardenkjær-Larsen, J. H.; Laustsen, C.; Bertelsen, L. B. A continuous-flow bioreactor system for high-throughput hyperpolarized metabolic flux analysis. NMR Biomed 2024, 37 (5), e5107 DOI: 10.1002/nbm.5107There is no corresponding record for this reference.
- 34Christensen, N. V.; Laustsen, C.; Bertelsen, L. B. Differentiating leukemia subtypes based on metabolic signatures using hyperpolarized ̂13C NMR. NMR Biomed 2024, 37 (12), e5264 DOI: 10.1002/nbm.5264There is no corresponding record for this reference.
- 35Massin, C.; Vincent, F.; Homsy, A.; Ehrmann, K.; Boero, G.; Besse, P. A.; Daridon, A.; Verpoorte, E.; de Rooij, N. F.; Popovic, R. S. Planar microcoil-based microfluidic NMR probes. J. Magn. Reson. 2003, 164 (2), 242– 255, DOI: 10.1016/S1090-7807(03)00151-4There is no corresponding record for this reference.
- 36Wensink, H.; Benito-Lopez, F.; Hermes, D. C.; Verboom, W.; Gardeniers, H. J. G. E.; Reinhoudt, D. N.; van den Berg, A. Measuring reaction kinetics in a lab-on-a-chip by microcoil NMR. Lab Chip 2005, 5 (3), 280– 284, DOI: 10.1039/b414832kThere is no corresponding record for this reference.
- 37Liu, Y.; Mi, Y.; Mueller, T.; Kreibich, S.; Williams, E. G.; van Drogen, A.; Borel, C.; Frank, M.; Germain, P. L.; Bludau, I. Multi-omic measurements of heterogeneity in HeLa cells across laboratories. Nat. Biotechnol. 2019, 37 (3), 314– 322, DOI: 10.1038/s41587-019-0037-yThere is no corresponding record for this reference.
- 38Mason, J. A.; Davison-Versagli, C. A.; Leliaert, A. K.; Pape, D. J.; McCallister, C.; Zuo, J.; Durbin, S. M.; Buchheit, C. L.; Zhang, S.; Schafer, Z. T. Oncogenic Ras differentially regulates metabolism and anoikis in extracellular matrix-detached cells. Cell Death Differ. 2016, 23 (8), 1271– 1282, DOI: 10.1038/cdd.2016.15There is no corresponding record for this reference.
- 39Buchheit, C. L.; Rayavarapu, R. R.; Schafer, Z. T. The regulation of cancer cell death and metabolism by extracellular matrix attachment. Semin. Cell Dev. Biol. 2012, 23 (4), 402– 411, DOI: 10.1016/j.semcdb.2012.04.007There is no corresponding record for this reference.
- 40Schafer, Z. T.; Grassian, A. R.; Song, L.; Jiang, Z.; Gerhart-Hines, Z.; Irie, H. Y.; Gao, S.; Puigserver, P.; Brugge, J. S. Antioxidant and oncogene rescue of metabolic defects caused by loss of matrix attachment. Nature 2009, 461 (7260), 109– 113, DOI: 10.1038/nature08268There is no corresponding record for this reference.
- 41Avivar-Valderas, A.; Salas, E.; Bobrovnikova-Marjon, E.; Diehl, J. A.; Nagi, C.; Debnath, J.; Aguirre-Ghiso, J. A. PERK integrates autophagy and oxidative stress responses to promote survival during extracellular matrix detachment. Mol. Cell. Biol. 2011, 31 (17), 3616– 3629, DOI: 10.1128/MCB.05164-11There is no corresponding record for this reference.
- 42Hawk, M. A.; Schafer, Z. T. Mechanisms of redox metabolism and cancer cell survival during extracellular matrix detachment. J. Biol. Chem. 2018, 293 (20), 7531– 7538, DOI: 10.1074/jbc.TM117.000260There is no corresponding record for this reference.
- 43Takakusagi, Y.; Takakusagi, K.; Inoue, K.; Saito, K.; Homma, Y.; Ichikawa, K. Hyperpolarized [1–13C]pyruvate NMR spectroscopy reveals transition of tumor energy metabolism in microscale multicellular spheroids. Sci. Rep. 2025, 15 (1), 3454, DOI: 10.1038/s41598-025-03454-1There is no corresponding record for this reference.
- 44Nielsen, N.; Laustsen, C.; Bertelsen, L. B. ̂13C dynamic nuclear polarization for measuring metabolic flux in endothelial progenitor cells. Exp. Cell Res. 2016, 349 (1), 95– 100, DOI: 10.1016/j.yexcr.2016.10.002There is no corresponding record for this reference.
- 45Harris, T.; Degani, H.; Frydman, L. Hyperpolarized 13C NMR studies of glucose metabolism in living breast cancer cell cultures. NMR Biomed 2013, 26 (12), 1831– 1843, DOI: 10.1002/nbm.3024There is no corresponding record for this reference.
- 46Sriram, R.; Nguyen, J.; Santos, J. D. L.; Nguyen, L.; Sun, J.; Vigneron, S.; van Criekinge, M.; Kurhanewicz, J.; MacKenzie, J. D. Molecular detection of inflammation in cell models using hyperpolarized ̂13C-pyruvate. Theranostics 2018, 8 (12), 3400– 3411, DOI: 10.7150/thno.24322There is no corresponding record for this reference.
- 47Keshari, K. R.; Kurhanewicz, J.; Jeffries, R. E.; Wilson, D. M.; Dewar, B. J.; van Criekinge, M.; Zierhut, M.; Vigneron, D. B.; Macdonald, J. M. Hyperpolarized ̂13C spectroscopy and an NMR-compatible bioreactor system for the investigation of real-time cellular metabolism. Magn. Reson. Med. 2010, 63 (2), 322– 329, DOI: 10.1002/mrm.22225There is no corresponding record for this reference.
- 48Thuenauer, R.; Nicklaus, S.; Frensch, M.; Troendle, K.; Madl, J.; Römer, W. A microfluidic biochip for locally confined stimulation of cells within an epithelial monolayer. RSC Adv. 2018, 8 (14), 7839– 7846, DOI: 10.1039/C7RA11943GThere is no corresponding record for this reference.
- 49Thuenauer, R.; Rodriguez-Boulan, E.; Römer, W. Microfluidic approaches for epithelial cell layer culture and characterisation. Analyst 2014, 139 (13), 3206– 3218, DOI: 10.1039/C4AN00056KThere is no corresponding record for this reference.
- 50Kim, H. J.; Ingber, D. E. Gut-on-a-chip microenvironment induces human intestinal cells to undergo villus differentiation. Integr. Biol. 2013, 5 (9), 1130– 1140, DOI: 10.1039/c3ib40126jThere is no corresponding record for this reference.
- 51Chung, H. H.; Mireles, M.; Kwarta, B. J.; Gaborski, T. R. Use of porous membranes in tissue barrier and co-culture models. Lab Chip 2018, 18 (12), 1671– 1689, DOI: 10.1039/C7LC01248AThere is no corresponding record for this reference.
- 52Maschmeyer, I.; Lorenz, A. K.; Schimek, K.; Hasenberg, T.; Ramme, A. P.; Hübner, J.; Lindner, M.; Drewell, C.; Bauer, S.; Thomas, A.; Sambo, N. S.; Sonntag, F.; Lauster, R.; Marx, U. A four-organ-chip for interconnected long-term co-culture of human intestine, liver, skin, and kidney equivalents. Lab Chip 2015, 15 (12), 2688– 2699, DOI: 10.1039/C5LC00392JThere is no corresponding record for this reference.
- 53Ferruzza, S.; Rossi, C.; Scarino, M. L.; Sambuy, Y. A protocol for differentiation of human intestinal Caco-2 cells in asymmetric serum-containing medium. Toxicol. In Vitro 2012, 26 (8), 1252– 1255, DOI: 10.1016/j.tiv.2012.01.008There is no corresponding record for this reference.
- 54Kopp, B.; Khawam, A.; Di Perna, K.; Lenart, D.; Vinette, M.; Silva, R.; Zanoni, T. B.; Rore, C.; Guenigault, G.; Richardson, E.; Kostrzewski, T.; Boswell, A.; Van, P.; Valentine, C.; Salk, J.; Hamel, A. Liver-on-chip model and application in predictive genotoxicity and mutagenicity of drugs. Mutat. Res. Genet. Toxicol. Environ. Mutagen. 2024, 896, 503762, DOI: 10.1016/j.mrgentox.2024.503762There is no corresponding record for this reference.
Supporting Information
Supporting Information
The Supporting Information is available free of charge at https://pubs.acs.org/doi/10.1021/acs.analchem.5c04058.
Return loss curves for the chip probe, Simulated B1+ fields of 2H and 1H coils, Rate constants from dDNP-NMR measurements in microfluidic chips not normalized to cell count (PDF)
Terms & Conditions
Most electronic Supporting Information files are available without a subscription to ACS Web Editions. Such files may be downloaded by article for research use (if there is a public use license linked to the relevant article, that license may permit other uses). Permission may be obtained from ACS for other uses through requests via the RightsLink permission system: http://pubs.acs.org/page/copyright/permissions.html.



