Native Chemical Ligation of Peptoid OligomersClick to copy article linkArticle link copied!
- Matthew R. SeraydarianMatthew R. SeraydarianDepartment of Chemistry, New York University, 100 Washington Square East, New York, New York 10003, United StatesThe Molecular Foundry, Lawrence Berkeley National Laboratory, 1 Cyclotron Road, Berkeley, California 94720, United StatesMore by Matthew R. Seraydarian
- Michael D. ConnollyMichael D. ConnollyThe Molecular Foundry, Lawrence Berkeley National Laboratory, 1 Cyclotron Road, Berkeley, California 94720, United StatesMore by Michael D. Connolly
- Ronald N. ZuckermannRonald N. ZuckermannThe Molecular Foundry, Lawrence Berkeley National Laboratory, 1 Cyclotron Road, Berkeley, California 94720, United StatesMore by Ronald N. Zuckermann
- Kent Kirshenbaum*Kent Kirshenbaum*Email: [email protected]Department of Chemistry, New York University, 100 Washington Square East, New York, New York 10003, United StatesMore by Kent Kirshenbaum
Abstract
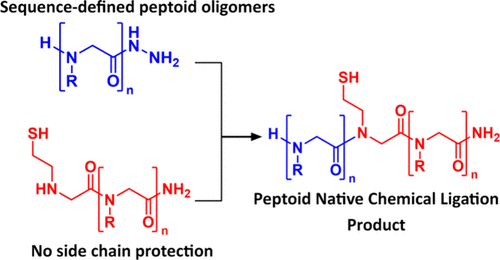
Bioorganic chemists are inspired by natural biopolymers to design peptidomimetic oligomers that can exhibit sequence-structure–function relationships. Biomimetic polymers can be synthesized to incorporate a specific sequence of nonbiological monomer units using a variety of iterative solution-phase or solid-phase reaction schemes. These protocols generally provide access to a vast diversity of oligomeric compounds but are limited with respect to their ability to attain protein-like chain lengths. This constraint can preclude access to sequence-defined synthetic macromolecules with sufficient sizes required to exhibit tertiary structure and other protein-mimetic attributes. In contrast, peptide chemists have overcome this limitation by developing convergent synthetic methods, such as native chemical ligation, to join individual, smaller peptide chains together to make larger peptides or full proteins. A similar convergent approach is needed to establish efficient synthetic routes to non-natural sequence-defined macromolecules. Herein, we adapt the peptide native chemical ligation method to peptoid oligomers, demonstrating how short chains can be conjoined to create sequence-defined peptoid macromolecules. Nanosheet-forming peptoid polymers with distinct surface loop display domains were generated by sequential ligation of several discrete fragments. This method provides a reliable convergent ligation route for sequence-defined polypeptoids that results in a native amide bond joining the fragments. We envision that this strategy will be useful in synthesizing peptoid-based proteomimetics that incorporate diverse chemical features.
This publication is licensed under
License Summary*
You are free to share(copy and redistribute) this article in any medium or format and to adapt(remix, transform, and build upon) the material for any purpose, even commercially within the parameters below:
Creative Commons (CC): This is a Creative Commons license.
Attribution (BY): Credit must be given to the creator.
*Disclaimer
This summary highlights only some of the key features and terms of the actual license. It is not a license and has no legal value. Carefully review the actual license before using these materials.
License Summary*
You are free to share(copy and redistribute) this article in any medium or format and to adapt(remix, transform, and build upon) the material for any purpose, even commercially within the parameters below:
Creative Commons (CC): This is a Creative Commons license.
Attribution (BY): Credit must be given to the creator.
*Disclaimer
This summary highlights only some of the key features and terms of the actual license. It is not a license and has no legal value. Carefully review the actual license before using these materials.
License Summary*
You are free to share(copy and redistribute) this article in any medium or format and to adapt(remix, transform, and build upon) the material for any purpose, even commercially within the parameters below:
Creative Commons (CC): This is a Creative Commons license.
Attribution (BY): Credit must be given to the creator.
*Disclaimer
This summary highlights only some of the key features and terms of the actual license. It is not a license and has no legal value. Carefully review the actual license before using these materials.
Special Issue
Published as part of Biochemistryspecial issue “Chemistry and Biology of Peptides”.
Introduction
Scheme 1
Scheme 2
Scheme 3
Materials and Methods
Synthesis of 3-(Tritylthio)-propylamine
General Strategy for Native Chemical Ligation of Peptoids
Radical Desulfurization
Peptoid Macrocyclization Using NCL Chemistry
Peptoid Nanosheet Formation
Fluorescence Imaging of Assembled Peptoid Nanosheets
Results and Discussion
Figure 1
Figure 1. (a) Peptoid 1 H-Npm-(Npl)3-Sar-NHNH2, the C-terminal hydrazide peptoid fragment used in first ligation (b) Peptoid 2 H-Nte-(Npl)3-NH2, the N-terminal thiol-bearing peptoid fragment used in first ligation (c) Sequence of peptoid 3 following a successful NCL reaction of peptoids 1 and 2. (d) Analytical HPLC traces of the crude ligation mixture overlaid with the traces of the pure starting material peptoids. The peak at ∼24 min contained the product. The large peak at ∼22 min contains the thiol additive MPAA, which is used in excess. (e) Positive ion mode mass spectrum of the selected product peak displaying the expected [M + H] and [M + Na] peaks.

| C-terminal hydrazide sequence (peptoid #) | N-terminal thiol sequence (peptoid #) | Ligated product sequence (peptoid #) calculated mass: observed mass |
|---|---|---|
| H-(Npl-Npm)2-NHNH2(4) | H-Nte-(Npl)3-NH2(5) | H-(Npl-Npm)2-Nte-(Npl)3-NH2(6) 924.2: 924.8 |
| H-Npm-(Nme)3-Sar-NHNH2(7) | H-Ntp-Npl-Npm-Nme-NH2(8) | H-Npm-(Nme)3-Sar-Ntp-Npl-Npm-Nme-NH2(9) 1025.3: 1024.8 |
| H-Npm-(Nme)3-Sar-NHNH2(7) | H-Nte-Npl-Nte-Npm-Nme-Npl-NH2(10) | H-Npm-(Nme)3-Sar-Nte-Npl-Nte-Npm-Nme-Npl-NH2(11) 1227.6: 1226.8 |
Scheme 4
Figure 2
Figure 2. Evidence of the formation a nanosheet-forming product from the ligation of peptoids 14 and 15 (a) and (b) Sequences of peptoids 14 and 15, respectively. The loop segment is depicted in blue. (c) Analytical HPLC traces of pure peptoids 14 and 15, and the purified ligation product of the two (d) Fluorescence microcopy imaging of micron-scale peptoid nanosheets formed from the ligation product of peptoids 14 and 15. Nanosheets were stained with Nile Red dye and imaged on a Zeiss LSM710 confocal microscope.
Figure 3
Figure 3. Formation of the expected product peptoid 18 from the ligation of peptoids 16 and 17. (a) and (b) Sequences of peptoids 16 and 17, respectively. (c) Sequence of ligation product peptoid 18. Loop segment is shown in blue. (d) Fluorescence microscopy imaging of peptoid nanosheets formed from peptoid 18. Nanosheets were stained with Nile Red dye on an inverted Zeiss Elyra 7 microscope.
Conclusions
Supporting Information
The Supporting Information is available free of charge at https://pubs.acs.org/doi/10.1021/acs.biochem.5c00833.
Analytical characterization of the reactions and compounds described above (PDF)
Terms & Conditions
Most electronic Supporting Information files are available without a subscription to ACS Web Editions. Such files may be downloaded by article for research use (if there is a public use license linked to the relevant article, that license may permit other uses). Permission may be obtained from ACS for other uses through requests via the RightsLink permission system: http://pubs.acs.org/page/copyright/permissions.html.
Acknowledgments
We would like to thank Shaoting Peng and Natalia Molchanova for their help with synthesizing and characterizing peptoid oligomers, and Behzad Rad for his help with imaging peptoid nanosheets. Work at the Molecular Foundry was supported by the Office of Science, Office of Basic Energy Sciences, of the U.S. Department of Energy under Contract No. DE-AC02-05CH11231 under proposal numbers MFP-09143 and MFP-10275.
References
This article references 59 other publications.
- 1Hakobyan, K.; Noble, B. B.; Xu, J. The current science of sequence-defined macromolecules. Prog. Polym. Sci. 2023, 147, 101754 DOI: 10.1016/j.progpolymsci.2023.101754Google ScholarThere is no corresponding record for this reference.
- 2Meier, M. A. R.; Barner-Kowollik, C. A New Class of Materials: Sequence-Defined Macromolecules and Their Emerging Applications. Adv. Mater. 2019, 31 (26), 1806027, DOI: 10.1002/adma.201806027Google ScholarThere is no corresponding record for this reference.
- 3Solleder, S. C.; Schneider, R. V.; Wetzel, K. S.; Boukis, A. C.; Meier, M. A. Recent progress in the design of monodisperse, sequence-defined macromolecules. Macromol. Rapid Commun. 2017, 38 (9), 1600711, DOI: 10.1002/marc.201600711Google ScholarThere is no corresponding record for this reference.
- 4Dill, K. A. Dominant forces in protein folding. Biochemistry 1990, 29 (31), 7133– 7155, DOI: 10.1021/bi00483a001Google ScholarThere is no corresponding record for this reference.
- 5Kresge, N.; Simoni, R. D.; Hill, R. L. The thermodynamic hypothesis of protein folding: the work of Christian Anfinsen. J. Biol. Chem. 2006, 281 (14), e11– e13, DOI: 10.1016/S0021-9258(19)56522-XGoogle ScholarThere is no corresponding record for this reference.
- 6Dill, K. A. Polymer principles and protein folding. Protein Sci. 1999, 8 (6), 1166– 1180, DOI: 10.1110/ps.8.6.1166Google ScholarThere is no corresponding record for this reference.
- 7Dill, K. A. Dominant forces in protein folding. Biochemistry 1990, 29 (31), 7133– 7155, DOI: 10.1021/bi00483a001Google ScholarThere is no corresponding record for this reference.
- 8Lee, B.-C.; Zuckermann, R. N.; Dill, K. A. Folding a nonbiological polymer into a compact multihelical structure. J. Am. Chem. Soc. 2005, 127 (31), 10999– 11009, DOI: 10.1021/ja0514904Google ScholarThere is no corresponding record for this reference.
- 9Burkoth, T. S.; Beausoleil, E.; Kaur, S.; Tang, D.; Cohen, F. E.; Zuckermann, R. N. Toward the synthesis of artificial proteins: the discovery of an amphiphilic helical peptoid assembly. Chemistry & Biology 2002, 9 (5), 647– 654, DOI: 10.1016/S1074-5521(02)00140-0Google ScholarThere is no corresponding record for this reference.
- 10Lee, B.-C.; Chu, T. K.; Dill, K. A.; Zuckermann, R. N. Biomimetic nanostructures: Creating a high-affinity zinc-binding site in a folded nonbiological polymer. J. Am. Chem. Soc. 2008, 130 (27), 8847– 8855, DOI: 10.1021/ja802125xGoogle ScholarThere is no corresponding record for this reference.
- 11Horne, W. S.; Johnson, L. M.; Ketas, T. J.; Klasse, P. J.; Lu, M.; Moore, J. P.; Gellman, S. H. Structural and biological mimicry of protein surface recognition by α/β-peptide foldamers. Proc. Natl. Acad. Sci. U. S. A. 2009, 106 (35), 14751– 14756, DOI: 10.1073/pnas.0902663106Google ScholarThere is no corresponding record for this reference.
- 12Giuliano, M. W.; Horne, W. S.; Gellman, S. H. An α/β-peptide helix bundle with a pure β3-amino acid core and a distinctive quaternary structure. J. Am. Chem. Soc. 2009, 131 (29), 9860– 9861, DOI: 10.1021/ja8099294Google ScholarThere is no corresponding record for this reference.
- 13Ziach, K.; Chollet, C.; Parissi, V.; Prabhakaran, P.; Marchivie, M.; Corvaglia, V.; Bose, P. P.; Laxmi-Reddy, K.; Godde, F.; Schmitter, J. M.; Chaignepain, S.; Pourquier, P.; Huc, I. Single helically folded aromatic oligoamides that mimic the charge surface of double-stranded B-DNA. Nat. Chem. 2018, 10 (5), 511– 518, DOI: 10.1038/s41557-018-0018-7Google ScholarThere is no corresponding record for this reference.
- 14Wang, P. S.; Schepartz, A. β-Peptide bundles: Design. Build. Analyze. Biosynthesize. Chem. Commun. 2016, 52 (47), 7420– 7432, DOI: 10.1039/C6CC01546HGoogle ScholarThere is no corresponding record for this reference.
- 15Cheng, R. P.; Gellman, S. H.; DeGrado, W. F. β-Peptides: from structure to function. Chem. Rev. 2001, 101 (10), 3219– 3232, DOI: 10.1021/cr000045iGoogle ScholarThere is no corresponding record for this reference.
- 16Hill, D. J.; Mio, M. J.; Prince, R. B.; Hughes, T. S.; Moore, J. S. A field guide to foldamers. Chem. Rev. 2001, 101 (12), 3893– 4012, DOI: 10.1021/cr990120tGoogle ScholarThere is no corresponding record for this reference.
- 17Zuckermann, R. N.; Kerr, J. M.; Kent, S. B.; Moos, W. H. Efficient method for the preparation of peptoids [oligo (N-substituted glycines)] by submonomer solid-phase synthesis. J. Am. Chem. Soc. 1992, 114 (26), 10646– 10647, DOI: 10.1021/ja00052a076Google ScholarThere is no corresponding record for this reference.
- 18Zuckermann, R. N. Peptoid origins. Peptide Science 2011, 96 (5), 545– 555, DOI: 10.1002/bip.21573Google ScholarThere is no corresponding record for this reference.
- 19Clapperton, A. M.; Hood, C.; Tran, H. Sequence-defined peptoids via iterative exponential growth. Chemical Science 2025, 16 (22), 9638– 9647, DOI: 10.1039/D5SC01296AGoogle ScholarThere is no corresponding record for this reference.
- 20Knight, A. S.; Zhou, E. Y.; Francis, M. B.; Zuckermann, R. N. Sequence programmable peptoid polymers for diverse materials applications. Adv. Mater. 2015, 27 (38), 5665– 5691, DOI: 10.1002/adma.201500275Google ScholarThere is no corresponding record for this reference.
- 21Gangloff, N.; Ulbricht, J.; Lorson, T.; Schlaad, H.; Luxenhofer, R. Peptoids and polypeptoids at the frontier of supra-and macromolecular engineering. Chem. Rev. 2016, 116 (4), 1753– 1802, DOI: 10.1021/acs.chemrev.5b00201Google ScholarThere is no corresponding record for this reference.
- 22Clapperton, A. M.; Babi, J.; Tran, H. A field guide to optimizing peptoid synthesis. ACS Polymers Au 2022, 2 (6), 417– 429, DOI: 10.1021/acspolymersau.2c00036Google ScholarThere is no corresponding record for this reference.
- 23Dawson, P. E.; Muir, T. W.; Clark-Lewis, I.; Kent, S. B. Synthesis of proteins by native chemical ligation. Science 1994, 266 (5186), 776– 779, DOI: 10.1126/science.7973629Google ScholarThere is no corresponding record for this reference.
- 24Agouridas, V.; El Mahdi, O.; Diemer, V.; Cargoet, M.; Monbaliu, J.-C. M.; Melnyk, O. Native chemical ligation and extended methods: mechanisms, catalysis, scope, and limitations. Chem. Rev. 2019, 119 (12), 7328– 7443, DOI: 10.1021/acs.chemrev.8b00712Google ScholarThere is no corresponding record for this reference.
- 25Shimko, J. C.; North, J. A.; Bruns, A. N.; Poirier, M. G.; Ottesen, J. J. Preparation of fully synthetic histone H3 reveals that acetyl-lysine 56 facilitates protein binding within nucleosomes. J. Mol. Biol. 2011, 408 (2), 187– 204, DOI: 10.1016/j.jmb.2011.01.003Google ScholarThere is no corresponding record for this reference.
- 26Wang, Y.; Xiang, J.; Jiang, H. Halide-Guided Oligo (aryl-triazole-amide)s Foldamers: Receptors for Multiple Halide Ions. Chem. - Eur. J. 2011, 17 (2), 613– 619, DOI: 10.1002/chem.201001560Google ScholarThere is no corresponding record for this reference.
- 27Maury, J.; Le Bailly, B. A.; Raftery, J.; Clayden, J. Conformational cooperativity between helical domains of differing geometry in oligoamide–oligourea foldamer chimeras. Chem. Commun. 2015, 51 (59), 11802– 11805, DOI: 10.1039/C5CC02995CGoogle ScholarThere is no corresponding record for this reference.
- 28Pendem, N.; Nelli, Y.; Cussol, L.; Didierjean, C.; Kauffmann, B.; Dolain, C.; Guichard, G. Synthesis and crystallographic characterization of helical hairpin oligourea foldamers. Chem. - Eur. J. 2023, 29 (39), e202300087 DOI: 10.1002/chem.202301615Google ScholarThere is no corresponding record for this reference.
- 29Sood, A.; Mandal, P. K.; Ottelé, J.; Wu, J.; Eleveld, M.; Hatai, J.; Pappas, C. G.; Huc, I.; Otto, S. Simultaneous Formation of a Foldamer and a Self-Replicator by Out-of-Equilibrium Dynamic Covalent Chemistry. J. Am. Chem. Soc. 2024, 146 (49), 33386– 33394, DOI: 10.1021/jacs.4c09111Google ScholarThere is no corresponding record for this reference.
- 30Liu, B.; Beatty, M. A.; Pappas, C. G.; Liu, K.; Ottelé, J.; Otto, S. Self-Sorting in Dynamic Combinatorial Libraries Leads to the Co-Existence of Foldamers and Self-Replicators. Angew. Chem., Int. Ed. 2021, 60 (24), 13569– 13573, DOI: 10.1002/anie.202101052Google ScholarThere is no corresponding record for this reference.
- 31Liu, B.; Wu, J.; Geerts, M.; Markovitch, O.; Pappas, C. G.; Liu, K.; Otto, S. Out-of-equilibrium self-replication allows selection for dynamic kinetic stability in a system of competing replicators. Angew. Chem., Int. Ed. 2022, 61 (18), e202117605 DOI: 10.1002/anie.202117605Google ScholarThere is no corresponding record for this reference.
- 32Jin, Y.; Mandal, P. K.; Wu, J.; Böcher, N.; Huc, I.; Otto, S. (Re-)directing oligomerization of a single building block into two specific dynamic covalent foldamers through pH. J. Am. Chem. Soc. 2023, 145 (5), 2822– 2829, DOI: 10.1021/jacs.2c09325Google ScholarThere is no corresponding record for this reference.
- 33Murnen, H. K.; Khokhlov, A. R.; Khalatur, P. G.; Segalman, R. A.; Zuckermann, R. N. Impact of hydrophobic sequence patterning on the coil-to-globule transition of protein-like polymers. Macromolecules 2012, 45 (12), 5229– 5236, DOI: 10.1021/ma300707tGoogle ScholarThere is no corresponding record for this reference.
- 34Yoo, B.; Kirshenbaum, K. Protease-mediated ligation of abiotic oligomers. J. Am. Chem. Soc. 2005, 127 (49), 17132– 17133, DOI: 10.1021/ja055105mGoogle ScholarThere is no corresponding record for this reference.
- 35Levine, P. M.; Craven, T. W.; Bonneau, R.; Kirshenbaum, K. Chemoselective fragment condensation between peptide and peptidomimetic oligomers. Organic & Biomolecular Chemistry 2013, 11 (25), 4142– 4146, DOI: 10.1039/c3ob40606gGoogle ScholarThere is no corresponding record for this reference.
- 36Levine, P. M.; Craven, T. W.; Bonneau, R.; Kirshenbaum, K. Semisynthesis of peptoid–protein hybrids by chemical ligation at serine. Org. Lett. 2014, 16 (2), 512– 515, DOI: 10.1021/ol4033978Google ScholarThere is no corresponding record for this reference.
- 37Pattabiraman, V. R.; Ogunkoya, A. O.; Bode, J. W. Chemical protein synthesis by chemoselective α-ketoacid–hydroxylamine (KAHA) ligations with 5-oxaproline. Angew. Chem., Int. Ed. 2012, 51 (21), 5114– 5118, DOI: 10.1002/anie.201200907Google ScholarThere is no corresponding record for this reference.
- 38Bode, J. W.; Fox, R. M.; Baucom, K. D. Chemoselective amide ligations by decarboxylative condensations of N-alkylhydroxylamines and α-ketoacids. Angew. Chem., Int. Ed. 2006, 45 (8), 1248– 1252, DOI: 10.1002/anie.200503991Google ScholarThere is no corresponding record for this reference.
- 39Li, X.; Lam, H. Y.; Zhang, Y.; Chan, C. K. Salicylaldehyde ester-induced chemoselective peptide ligations: enabling generation of natural peptidic linkages at the serine/threonine sites. Org. Lett. 2010, 12 (8), 1724– 1727, DOI: 10.1021/ol1003109Google ScholarThere is no corresponding record for this reference.
- 40Liu, H.; Li, X. Serine/threonine ligation: origin, mechanistic aspects, and applications. Acc. Chem. Res. 2018, 51 (7), 1643– 1655, DOI: 10.1021/acs.accounts.8b00151Google ScholarThere is no corresponding record for this reference.
- 41Huang, Y.-C.; Fang, G.-M.; Liu, L. Chemical synthesis of proteins using hydrazide intermediates. National Science Review 2016, 3 (1), 107– 116, DOI: 10.1093/nsr/nwv072Google ScholarThere is no corresponding record for this reference.
- 42Zheng, J.-S.; Tang, S.; Qi, Y.-K.; Wang, Z.-P.; Liu, L. Chemical synthesis of proteins using peptide hydrazides as thioester surrogates. Nat. Protoc 2013, 8 (12), 2483– 2495, DOI: 10.1038/nprot.2013.152Google ScholarThere is no corresponding record for this reference.
- 43Li, J.; Li, Y.; He, Q.; Li, Y.; Li, H.; Liu, L. One-pot native chemical ligation of peptide hydrazides enables total synthesis of modified histones. Organic & Biomolecular Chemistry 2014, 12 (29), 5435– 5441, DOI: 10.1039/C4OB00715HGoogle ScholarThere is no corresponding record for this reference.
- 44Werner, H. M.; Estabrooks, S. K.; Preston, G. M.; Brodsky, J. L.; Horne, W. S. Exploring the functional consequences of protein backbone alteration in ubiquitin through native chemical ligation. ChemBioChem. 2019, 20 (18), 2346– 2350, DOI: 10.1002/cbic.201900225Google ScholarThere is no corresponding record for this reference.
- 45Bird, M. J.; Dawson, P. E. A shelf stable Fmoc hydrazine resin for the synthesis of peptide hydrazides. Peptide Sci. 2022, 114 (5), e24268 DOI: 10.1002/pep2.24268Google ScholarThere is no corresponding record for this reference.
- 46Chelushkin, P. S.; Polyanichko, K. V.; Leko, M. V.; Dorosh, M. Y.; Bruckdorfer, T.; Burov, S. V. Convenient method of peptide hydrazide synthesis using a new hydrazone resin. Tetrahedron Lett. 2015, 56 (4), 619– 622, DOI: 10.1016/j.tetlet.2014.12.056Google ScholarThere is no corresponding record for this reference.
- 47Wong, C. T.; Tung, C. L.; Li, X. Synthetic cysteine surrogates used in native chemical ligation. Molecular BioSystems 2012, 9 (5), 826– 833, DOI: 10.1039/C2MB25437AGoogle ScholarThere is no corresponding record for this reference.
- 48Chen, J.; Wan, Q.; Yuan, Y.; Zhu, J.; Danishefsky, S. J. Native chemical ligation at valine: a contribution to peptide and glycopeptide synthesis. Angew. Chem., Int. Ed. 2008, 47 (44), 8521– 8524, DOI: 10.1002/anie.200803523Google ScholarThere is no corresponding record for this reference.
- 49Tan, Y.; Li, J.; Jin, K.; Liu, J.; Chen, Z.; Yang, J.; Li, X. Cysteine/penicillamine ligation independent of terminal steric demands for chemical protein synthesis. Angew. Chem. 2020, 132 (31), 12841– 12845, DOI: 10.1002/ange.202003652Google ScholarThere is no corresponding record for this reference.
- 50Ajish Kumar, K.; Haj-Yahya, M.; Olschewski, D.; Lashuel, H. A.; Brik, A. Highly efficient and chemoselective peptide ubiquitylation. Angew. Chem. 2009, 121 (43), 8234– 8238, DOI: 10.1002/ange.200902936Google ScholarThere is no corresponding record for this reference.
- 51Dawson, P. E. Native chemical ligation combined with desulfurization and deselenization: a general strategy for chemical protein synthesis. Isr. J. Chem. 2011, 51 (8–9), 862– 867, DOI: 10.1002/ijch.201100128Google ScholarThere is no corresponding record for this reference.
- 52Jin, K.; Li, X. Advances in native chemical ligation–desulfurization: a powerful strategy for peptide and protein synthesis. Chem. - Chem. J. 2018, 24 (66), 17397– 17404, DOI: 10.1002/chem.201802067Google ScholarThere is no corresponding record for this reference.
- 53Robertson, E. J.; Battigelli, A.; Proulx, C.; Mannige, R. V.; Haxton, T. K.; Yun, L.; Whitelam, S.; Zuckermann, R. N. Design, synthesis, assembly, and engineering of peptoid nanosheets. Acc. Chem. Res. 2016, 49 (3), 379– 389, DOI: 10.1021/acs.accounts.5b00439Google ScholarThere is no corresponding record for this reference.
- 54Olivier, G. K.; Cho, A.; Sanii, B.; Connolly, M. D.; Tran, H.; Zuckermann, R. N. Antibody-mimetic peptoid nanosheets for molecular recognition. ACS Nano 2013, 7 (10), 9276– 9286, DOI: 10.1021/nn403899yGoogle ScholarThere is no corresponding record for this reference.
- 55Kim, J. H.; Kim, S. C.; Kline, M. A.; Grzincic, E. M.; Tresca, B. W.; Cardiel, J.; Karbaschi, M.; Dehigaspitiya, D. C.; Chen, Y.; Udumula, V.; Jian, T.; Murray, D. J.; Yun, L.; Connolly, M. D.; Liu, J.; Ren, G.; Chen, C. L.; Kirshenbaum, K.; Abate, A. R.; Zuckermann, R. N. Discovery of stable and selective antibody mimetics from combinatorial libraries of polyvalent, loop-functionalized peptoid nanosheets. ACS Nano 2020, 14 (1), 185– 195, DOI: 10.1021/acsnano.9b07498Google ScholarThere is no corresponding record for this reference.
- 56Battigelli, A.; Kim, J. H.; Dehigaspitiya, D. C.; Proulx, C.; Robertson, E. J.; Murray, D. J.; Rad, B.; Kirshenbaum, K.; Zuckermann, R. N. Glycosylated peptoid nanosheets as a multivalent scaffold for protein recognition. ACS Nano 2018, 12 (3), 2455– 2465, DOI: 10.1021/acsnano.7b08018Google ScholarThere is no corresponding record for this reference.
- 57Robertson, E. J.; Proulx, C.; Su, J. K.; Garcia, R. L.; Yoo, S.; Nehls, E. M.; Connolly, M. D.; Taravati, L.; Zuckermann, R. N. Molecular engineering of the peptoid nanosheet hydrophobic core. Langmuir 2016, 32 (45), 11946– 11957, DOI: 10.1021/acs.langmuir.6b02735Google ScholarThere is no corresponding record for this reference.
- 58Beard, J. C.; Swager, T. M. An organic chemist’s guide to N-nitrosamines: their structure, reactivity, and role as contaminants. Journal of Organic Chemistry 2021, 86 (3), 2037– 2057, DOI: 10.1021/acs.joc.0c02774Google ScholarThere is no corresponding record for this reference.
- 59Kubacka, W.; Libbey, L. M.; Scanlan, R. A. Formation and chemical characterization of some nitroso dipeptides N terminal in proline. J. Agric. Food Chem. 1984, 32 (2), 401– 404, DOI: 10.1021/jf00122a055Google ScholarThere is no corresponding record for this reference.
Cited By
This article has not yet been cited by other publications.
Article Views
Altmetric
Citations
Article Views are the COUNTER-compliant sum of full text article downloads since November 2008 (both PDF and HTML) across all institutions and individuals. These metrics are regularly updated to reflect usage leading up to the last few days.
Citations are the number of other articles citing this article, calculated by Crossref and updated daily. Find more information about Crossref citation counts.
The Altmetric Attention Score is a quantitative measure of the attention that a research article has received online. Clicking on the donut icon will load a page at altmetric.com with additional details about the score and the social media presence for the given article. Find more information on the Altmetric Attention Score and how the score is calculated.
Recommended Articles
Abstract
Scheme 1
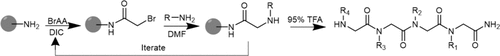 Scheme 1. General Peptoid Submonomer Synthesis Protocol
Scheme 1. General Peptoid Submonomer Synthesis ProtocolScheme 2
 Scheme 2. Canonical Peptide Native Chemical Ligation
Scheme 2. Canonical Peptide Native Chemical LigationScheme 3
 Scheme 3. C-Terminal Peptoid Hydrazide Oxidation, Thiolysis, and Ligation
Scheme 3. C-Terminal Peptoid Hydrazide Oxidation, Thiolysis, and LigationFigure 1

Figure 1. (a) Peptoid 1 H-Npm-(Npl)3-Sar-NHNH2, the C-terminal hydrazide peptoid fragment used in first ligation (b) Peptoid 2 H-Nte-(Npl)3-NH2, the N-terminal thiol-bearing peptoid fragment used in first ligation (c) Sequence of peptoid 3 following a successful NCL reaction of peptoids 1 and 2. (d) Analytical HPLC traces of the crude ligation mixture overlaid with the traces of the pure starting material peptoids. The peak at ∼24 min contained the product. The large peak at ∼22 min contains the thiol additive MPAA, which is used in excess. (e) Positive ion mode mass spectrum of the selected product peak displaying the expected [M + H] and [M + Na] peaks.
Scheme 4
 Scheme 4. NCL Chemistry as a Method for Peptoid Macrocyclization
Scheme 4. NCL Chemistry as a Method for Peptoid MacrocyclizationFigure 2
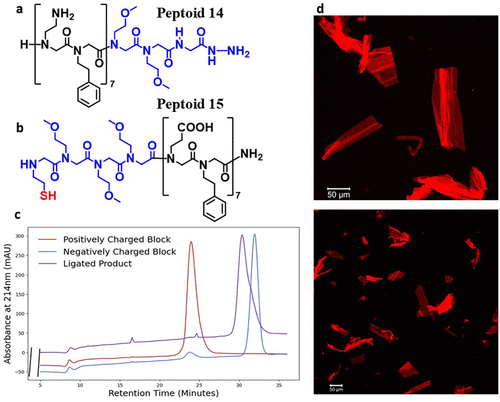
Figure 2. Evidence of the formation a nanosheet-forming product from the ligation of peptoids 14 and 15 (a) and (b) Sequences of peptoids 14 and 15, respectively. The loop segment is depicted in blue. (c) Analytical HPLC traces of pure peptoids 14 and 15, and the purified ligation product of the two (d) Fluorescence microcopy imaging of micron-scale peptoid nanosheets formed from the ligation product of peptoids 14 and 15. Nanosheets were stained with Nile Red dye and imaged on a Zeiss LSM710 confocal microscope.
Figure 3
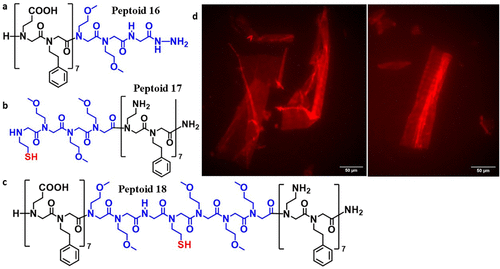
Figure 3. Formation of the expected product peptoid 18 from the ligation of peptoids 16 and 17. (a) and (b) Sequences of peptoids 16 and 17, respectively. (c) Sequence of ligation product peptoid 18. Loop segment is shown in blue. (d) Fluorescence microscopy imaging of peptoid nanosheets formed from peptoid 18. Nanosheets were stained with Nile Red dye on an inverted Zeiss Elyra 7 microscope.
References
This article references 59 other publications.
- 1Hakobyan, K.; Noble, B. B.; Xu, J. The current science of sequence-defined macromolecules. Prog. Polym. Sci. 2023, 147, 101754 DOI: 10.1016/j.progpolymsci.2023.101754There is no corresponding record for this reference.
- 2Meier, M. A. R.; Barner-Kowollik, C. A New Class of Materials: Sequence-Defined Macromolecules and Their Emerging Applications. Adv. Mater. 2019, 31 (26), 1806027, DOI: 10.1002/adma.201806027There is no corresponding record for this reference.
- 3Solleder, S. C.; Schneider, R. V.; Wetzel, K. S.; Boukis, A. C.; Meier, M. A. Recent progress in the design of monodisperse, sequence-defined macromolecules. Macromol. Rapid Commun. 2017, 38 (9), 1600711, DOI: 10.1002/marc.201600711There is no corresponding record for this reference.
- 4Dill, K. A. Dominant forces in protein folding. Biochemistry 1990, 29 (31), 7133– 7155, DOI: 10.1021/bi00483a001There is no corresponding record for this reference.
- 5Kresge, N.; Simoni, R. D.; Hill, R. L. The thermodynamic hypothesis of protein folding: the work of Christian Anfinsen. J. Biol. Chem. 2006, 281 (14), e11– e13, DOI: 10.1016/S0021-9258(19)56522-XThere is no corresponding record for this reference.
- 6Dill, K. A. Polymer principles and protein folding. Protein Sci. 1999, 8 (6), 1166– 1180, DOI: 10.1110/ps.8.6.1166There is no corresponding record for this reference.
- 7Dill, K. A. Dominant forces in protein folding. Biochemistry 1990, 29 (31), 7133– 7155, DOI: 10.1021/bi00483a001There is no corresponding record for this reference.
- 8Lee, B.-C.; Zuckermann, R. N.; Dill, K. A. Folding a nonbiological polymer into a compact multihelical structure. J. Am. Chem. Soc. 2005, 127 (31), 10999– 11009, DOI: 10.1021/ja0514904There is no corresponding record for this reference.
- 9Burkoth, T. S.; Beausoleil, E.; Kaur, S.; Tang, D.; Cohen, F. E.; Zuckermann, R. N. Toward the synthesis of artificial proteins: the discovery of an amphiphilic helical peptoid assembly. Chemistry & Biology 2002, 9 (5), 647– 654, DOI: 10.1016/S1074-5521(02)00140-0There is no corresponding record for this reference.
- 10Lee, B.-C.; Chu, T. K.; Dill, K. A.; Zuckermann, R. N. Biomimetic nanostructures: Creating a high-affinity zinc-binding site in a folded nonbiological polymer. J. Am. Chem. Soc. 2008, 130 (27), 8847– 8855, DOI: 10.1021/ja802125xThere is no corresponding record for this reference.
- 11Horne, W. S.; Johnson, L. M.; Ketas, T. J.; Klasse, P. J.; Lu, M.; Moore, J. P.; Gellman, S. H. Structural and biological mimicry of protein surface recognition by α/β-peptide foldamers. Proc. Natl. Acad. Sci. U. S. A. 2009, 106 (35), 14751– 14756, DOI: 10.1073/pnas.0902663106There is no corresponding record for this reference.
- 12Giuliano, M. W.; Horne, W. S.; Gellman, S. H. An α/β-peptide helix bundle with a pure β3-amino acid core and a distinctive quaternary structure. J. Am. Chem. Soc. 2009, 131 (29), 9860– 9861, DOI: 10.1021/ja8099294There is no corresponding record for this reference.
- 13Ziach, K.; Chollet, C.; Parissi, V.; Prabhakaran, P.; Marchivie, M.; Corvaglia, V.; Bose, P. P.; Laxmi-Reddy, K.; Godde, F.; Schmitter, J. M.; Chaignepain, S.; Pourquier, P.; Huc, I. Single helically folded aromatic oligoamides that mimic the charge surface of double-stranded B-DNA. Nat. Chem. 2018, 10 (5), 511– 518, DOI: 10.1038/s41557-018-0018-7There is no corresponding record for this reference.
- 14Wang, P. S.; Schepartz, A. β-Peptide bundles: Design. Build. Analyze. Biosynthesize. Chem. Commun. 2016, 52 (47), 7420– 7432, DOI: 10.1039/C6CC01546HThere is no corresponding record for this reference.
- 15Cheng, R. P.; Gellman, S. H.; DeGrado, W. F. β-Peptides: from structure to function. Chem. Rev. 2001, 101 (10), 3219– 3232, DOI: 10.1021/cr000045iThere is no corresponding record for this reference.
- 16Hill, D. J.; Mio, M. J.; Prince, R. B.; Hughes, T. S.; Moore, J. S. A field guide to foldamers. Chem. Rev. 2001, 101 (12), 3893– 4012, DOI: 10.1021/cr990120tThere is no corresponding record for this reference.
- 17Zuckermann, R. N.; Kerr, J. M.; Kent, S. B.; Moos, W. H. Efficient method for the preparation of peptoids [oligo (N-substituted glycines)] by submonomer solid-phase synthesis. J. Am. Chem. Soc. 1992, 114 (26), 10646– 10647, DOI: 10.1021/ja00052a076There is no corresponding record for this reference.
- 18Zuckermann, R. N. Peptoid origins. Peptide Science 2011, 96 (5), 545– 555, DOI: 10.1002/bip.21573There is no corresponding record for this reference.
- 19Clapperton, A. M.; Hood, C.; Tran, H. Sequence-defined peptoids via iterative exponential growth. Chemical Science 2025, 16 (22), 9638– 9647, DOI: 10.1039/D5SC01296AThere is no corresponding record for this reference.
- 20Knight, A. S.; Zhou, E. Y.; Francis, M. B.; Zuckermann, R. N. Sequence programmable peptoid polymers for diverse materials applications. Adv. Mater. 2015, 27 (38), 5665– 5691, DOI: 10.1002/adma.201500275There is no corresponding record for this reference.
- 21Gangloff, N.; Ulbricht, J.; Lorson, T.; Schlaad, H.; Luxenhofer, R. Peptoids and polypeptoids at the frontier of supra-and macromolecular engineering. Chem. Rev. 2016, 116 (4), 1753– 1802, DOI: 10.1021/acs.chemrev.5b00201There is no corresponding record for this reference.
- 22Clapperton, A. M.; Babi, J.; Tran, H. A field guide to optimizing peptoid synthesis. ACS Polymers Au 2022, 2 (6), 417– 429, DOI: 10.1021/acspolymersau.2c00036There is no corresponding record for this reference.
- 23Dawson, P. E.; Muir, T. W.; Clark-Lewis, I.; Kent, S. B. Synthesis of proteins by native chemical ligation. Science 1994, 266 (5186), 776– 779, DOI: 10.1126/science.7973629There is no corresponding record for this reference.
- 24Agouridas, V.; El Mahdi, O.; Diemer, V.; Cargoet, M.; Monbaliu, J.-C. M.; Melnyk, O. Native chemical ligation and extended methods: mechanisms, catalysis, scope, and limitations. Chem. Rev. 2019, 119 (12), 7328– 7443, DOI: 10.1021/acs.chemrev.8b00712There is no corresponding record for this reference.
- 25Shimko, J. C.; North, J. A.; Bruns, A. N.; Poirier, M. G.; Ottesen, J. J. Preparation of fully synthetic histone H3 reveals that acetyl-lysine 56 facilitates protein binding within nucleosomes. J. Mol. Biol. 2011, 408 (2), 187– 204, DOI: 10.1016/j.jmb.2011.01.003There is no corresponding record for this reference.
- 26Wang, Y.; Xiang, J.; Jiang, H. Halide-Guided Oligo (aryl-triazole-amide)s Foldamers: Receptors for Multiple Halide Ions. Chem. - Eur. J. 2011, 17 (2), 613– 619, DOI: 10.1002/chem.201001560There is no corresponding record for this reference.
- 27Maury, J.; Le Bailly, B. A.; Raftery, J.; Clayden, J. Conformational cooperativity between helical domains of differing geometry in oligoamide–oligourea foldamer chimeras. Chem. Commun. 2015, 51 (59), 11802– 11805, DOI: 10.1039/C5CC02995CThere is no corresponding record for this reference.
- 28Pendem, N.; Nelli, Y.; Cussol, L.; Didierjean, C.; Kauffmann, B.; Dolain, C.; Guichard, G. Synthesis and crystallographic characterization of helical hairpin oligourea foldamers. Chem. - Eur. J. 2023, 29 (39), e202300087 DOI: 10.1002/chem.202301615There is no corresponding record for this reference.
- 29Sood, A.; Mandal, P. K.; Ottelé, J.; Wu, J.; Eleveld, M.; Hatai, J.; Pappas, C. G.; Huc, I.; Otto, S. Simultaneous Formation of a Foldamer and a Self-Replicator by Out-of-Equilibrium Dynamic Covalent Chemistry. J. Am. Chem. Soc. 2024, 146 (49), 33386– 33394, DOI: 10.1021/jacs.4c09111There is no corresponding record for this reference.
- 30Liu, B.; Beatty, M. A.; Pappas, C. G.; Liu, K.; Ottelé, J.; Otto, S. Self-Sorting in Dynamic Combinatorial Libraries Leads to the Co-Existence of Foldamers and Self-Replicators. Angew. Chem., Int. Ed. 2021, 60 (24), 13569– 13573, DOI: 10.1002/anie.202101052There is no corresponding record for this reference.
- 31Liu, B.; Wu, J.; Geerts, M.; Markovitch, O.; Pappas, C. G.; Liu, K.; Otto, S. Out-of-equilibrium self-replication allows selection for dynamic kinetic stability in a system of competing replicators. Angew. Chem., Int. Ed. 2022, 61 (18), e202117605 DOI: 10.1002/anie.202117605There is no corresponding record for this reference.
- 32Jin, Y.; Mandal, P. K.; Wu, J.; Böcher, N.; Huc, I.; Otto, S. (Re-)directing oligomerization of a single building block into two specific dynamic covalent foldamers through pH. J. Am. Chem. Soc. 2023, 145 (5), 2822– 2829, DOI: 10.1021/jacs.2c09325There is no corresponding record for this reference.
- 33Murnen, H. K.; Khokhlov, A. R.; Khalatur, P. G.; Segalman, R. A.; Zuckermann, R. N. Impact of hydrophobic sequence patterning on the coil-to-globule transition of protein-like polymers. Macromolecules 2012, 45 (12), 5229– 5236, DOI: 10.1021/ma300707tThere is no corresponding record for this reference.
- 34Yoo, B.; Kirshenbaum, K. Protease-mediated ligation of abiotic oligomers. J. Am. Chem. Soc. 2005, 127 (49), 17132– 17133, DOI: 10.1021/ja055105mThere is no corresponding record for this reference.
- 35Levine, P. M.; Craven, T. W.; Bonneau, R.; Kirshenbaum, K. Chemoselective fragment condensation between peptide and peptidomimetic oligomers. Organic & Biomolecular Chemistry 2013, 11 (25), 4142– 4146, DOI: 10.1039/c3ob40606gThere is no corresponding record for this reference.
- 36Levine, P. M.; Craven, T. W.; Bonneau, R.; Kirshenbaum, K. Semisynthesis of peptoid–protein hybrids by chemical ligation at serine. Org. Lett. 2014, 16 (2), 512– 515, DOI: 10.1021/ol4033978There is no corresponding record for this reference.
- 37Pattabiraman, V. R.; Ogunkoya, A. O.; Bode, J. W. Chemical protein synthesis by chemoselective α-ketoacid–hydroxylamine (KAHA) ligations with 5-oxaproline. Angew. Chem., Int. Ed. 2012, 51 (21), 5114– 5118, DOI: 10.1002/anie.201200907There is no corresponding record for this reference.
- 38Bode, J. W.; Fox, R. M.; Baucom, K. D. Chemoselective amide ligations by decarboxylative condensations of N-alkylhydroxylamines and α-ketoacids. Angew. Chem., Int. Ed. 2006, 45 (8), 1248– 1252, DOI: 10.1002/anie.200503991There is no corresponding record for this reference.
- 39Li, X.; Lam, H. Y.; Zhang, Y.; Chan, C. K. Salicylaldehyde ester-induced chemoselective peptide ligations: enabling generation of natural peptidic linkages at the serine/threonine sites. Org. Lett. 2010, 12 (8), 1724– 1727, DOI: 10.1021/ol1003109There is no corresponding record for this reference.
- 40Liu, H.; Li, X. Serine/threonine ligation: origin, mechanistic aspects, and applications. Acc. Chem. Res. 2018, 51 (7), 1643– 1655, DOI: 10.1021/acs.accounts.8b00151There is no corresponding record for this reference.
- 41Huang, Y.-C.; Fang, G.-M.; Liu, L. Chemical synthesis of proteins using hydrazide intermediates. National Science Review 2016, 3 (1), 107– 116, DOI: 10.1093/nsr/nwv072There is no corresponding record for this reference.
- 42Zheng, J.-S.; Tang, S.; Qi, Y.-K.; Wang, Z.-P.; Liu, L. Chemical synthesis of proteins using peptide hydrazides as thioester surrogates. Nat. Protoc 2013, 8 (12), 2483– 2495, DOI: 10.1038/nprot.2013.152There is no corresponding record for this reference.
- 43Li, J.; Li, Y.; He, Q.; Li, Y.; Li, H.; Liu, L. One-pot native chemical ligation of peptide hydrazides enables total synthesis of modified histones. Organic & Biomolecular Chemistry 2014, 12 (29), 5435– 5441, DOI: 10.1039/C4OB00715HThere is no corresponding record for this reference.
- 44Werner, H. M.; Estabrooks, S. K.; Preston, G. M.; Brodsky, J. L.; Horne, W. S. Exploring the functional consequences of protein backbone alteration in ubiquitin through native chemical ligation. ChemBioChem. 2019, 20 (18), 2346– 2350, DOI: 10.1002/cbic.201900225There is no corresponding record for this reference.
- 45Bird, M. J.; Dawson, P. E. A shelf stable Fmoc hydrazine resin for the synthesis of peptide hydrazides. Peptide Sci. 2022, 114 (5), e24268 DOI: 10.1002/pep2.24268There is no corresponding record for this reference.
- 46Chelushkin, P. S.; Polyanichko, K. V.; Leko, M. V.; Dorosh, M. Y.; Bruckdorfer, T.; Burov, S. V. Convenient method of peptide hydrazide synthesis using a new hydrazone resin. Tetrahedron Lett. 2015, 56 (4), 619– 622, DOI: 10.1016/j.tetlet.2014.12.056There is no corresponding record for this reference.
- 47Wong, C. T.; Tung, C. L.; Li, X. Synthetic cysteine surrogates used in native chemical ligation. Molecular BioSystems 2012, 9 (5), 826– 833, DOI: 10.1039/C2MB25437AThere is no corresponding record for this reference.
- 48Chen, J.; Wan, Q.; Yuan, Y.; Zhu, J.; Danishefsky, S. J. Native chemical ligation at valine: a contribution to peptide and glycopeptide synthesis. Angew. Chem., Int. Ed. 2008, 47 (44), 8521– 8524, DOI: 10.1002/anie.200803523There is no corresponding record for this reference.
- 49Tan, Y.; Li, J.; Jin, K.; Liu, J.; Chen, Z.; Yang, J.; Li, X. Cysteine/penicillamine ligation independent of terminal steric demands for chemical protein synthesis. Angew. Chem. 2020, 132 (31), 12841– 12845, DOI: 10.1002/ange.202003652There is no corresponding record for this reference.
- 50Ajish Kumar, K.; Haj-Yahya, M.; Olschewski, D.; Lashuel, H. A.; Brik, A. Highly efficient and chemoselective peptide ubiquitylation. Angew. Chem. 2009, 121 (43), 8234– 8238, DOI: 10.1002/ange.200902936There is no corresponding record for this reference.
- 51Dawson, P. E. Native chemical ligation combined with desulfurization and deselenization: a general strategy for chemical protein synthesis. Isr. J. Chem. 2011, 51 (8–9), 862– 867, DOI: 10.1002/ijch.201100128There is no corresponding record for this reference.
- 52Jin, K.; Li, X. Advances in native chemical ligation–desulfurization: a powerful strategy for peptide and protein synthesis. Chem. - Chem. J. 2018, 24 (66), 17397– 17404, DOI: 10.1002/chem.201802067There is no corresponding record for this reference.
- 53Robertson, E. J.; Battigelli, A.; Proulx, C.; Mannige, R. V.; Haxton, T. K.; Yun, L.; Whitelam, S.; Zuckermann, R. N. Design, synthesis, assembly, and engineering of peptoid nanosheets. Acc. Chem. Res. 2016, 49 (3), 379– 389, DOI: 10.1021/acs.accounts.5b00439There is no corresponding record for this reference.
- 54Olivier, G. K.; Cho, A.; Sanii, B.; Connolly, M. D.; Tran, H.; Zuckermann, R. N. Antibody-mimetic peptoid nanosheets for molecular recognition. ACS Nano 2013, 7 (10), 9276– 9286, DOI: 10.1021/nn403899yThere is no corresponding record for this reference.
- 55Kim, J. H.; Kim, S. C.; Kline, M. A.; Grzincic, E. M.; Tresca, B. W.; Cardiel, J.; Karbaschi, M.; Dehigaspitiya, D. C.; Chen, Y.; Udumula, V.; Jian, T.; Murray, D. J.; Yun, L.; Connolly, M. D.; Liu, J.; Ren, G.; Chen, C. L.; Kirshenbaum, K.; Abate, A. R.; Zuckermann, R. N. Discovery of stable and selective antibody mimetics from combinatorial libraries of polyvalent, loop-functionalized peptoid nanosheets. ACS Nano 2020, 14 (1), 185– 195, DOI: 10.1021/acsnano.9b07498There is no corresponding record for this reference.
- 56Battigelli, A.; Kim, J. H.; Dehigaspitiya, D. C.; Proulx, C.; Robertson, E. J.; Murray, D. J.; Rad, B.; Kirshenbaum, K.; Zuckermann, R. N. Glycosylated peptoid nanosheets as a multivalent scaffold for protein recognition. ACS Nano 2018, 12 (3), 2455– 2465, DOI: 10.1021/acsnano.7b08018There is no corresponding record for this reference.
- 57Robertson, E. J.; Proulx, C.; Su, J. K.; Garcia, R. L.; Yoo, S.; Nehls, E. M.; Connolly, M. D.; Taravati, L.; Zuckermann, R. N. Molecular engineering of the peptoid nanosheet hydrophobic core. Langmuir 2016, 32 (45), 11946– 11957, DOI: 10.1021/acs.langmuir.6b02735There is no corresponding record for this reference.
- 58Beard, J. C.; Swager, T. M. An organic chemist’s guide to N-nitrosamines: their structure, reactivity, and role as contaminants. Journal of Organic Chemistry 2021, 86 (3), 2037– 2057, DOI: 10.1021/acs.joc.0c02774There is no corresponding record for this reference.
- 59Kubacka, W.; Libbey, L. M.; Scanlan, R. A. Formation and chemical characterization of some nitroso dipeptides N terminal in proline. J. Agric. Food Chem. 1984, 32 (2), 401– 404, DOI: 10.1021/jf00122a055There is no corresponding record for this reference.
Supporting Information
Supporting Information
The Supporting Information is available free of charge at https://pubs.acs.org/doi/10.1021/acs.biochem.5c00833.
Analytical characterization of the reactions and compounds described above (PDF)
Terms & Conditions
Most electronic Supporting Information files are available without a subscription to ACS Web Editions. Such files may be downloaded by article for research use (if there is a public use license linked to the relevant article, that license may permit other uses). Permission may be obtained from ACS for other uses through requests via the RightsLink permission system: http://pubs.acs.org/page/copyright/permissions.html.



