Azidocoumarin Glycan Probes for Photoinduced Cross-Linking and In Situ Fluorescent LabelingClick to copy article linkArticle link copied!
- Nina JahnkeNina JahnkeDepartment of Organic and Macromolecular Chemistry, Heinrich Heine University Duesseldorf, Universitaetsstrasse 1, 40225 Duesseldorf, GermanyMore by Nina Jahnke
- Marc D. DriessenMarc D. DriessenFaculty of Medicine and University Hospital Cologne, Department of Oral, Maxillofacial and Plastic Surgery, University of Cologne, Kerpener Str. 62, 50937 Cologne, GermanyMore by Marc D. Driessen
- Georgia PartalidouGeorgia PartalidouInstitute for Macromolecular Chemistry, Albert Ludwig University of Freiburg, Stefan-Meier-Strasse 31, 79104 Freiburg, GermanyMore by Georgia Partalidou
- Simon PrzetakSimon PrzetakDepartment of Organic and Macromolecular Chemistry, Heinrich Heine University Duesseldorf, Universitaetsstrasse 1, 40225 Duesseldorf, GermanyMore by Simon Przetak
- Ulla I.M. Gerling-Driessen*Ulla I.M. Gerling-Driessen*Email: [email protected]Institute for Macromolecular Chemistry, Albert Ludwig University of Freiburg, Stefan-Meier-Strasse 31, 79104 Freiburg, GermanyMore by Ulla I.M. Gerling-Driessen
- Laura Hartmann*Laura Hartmann*Email: [email protected]Department of Organic and Macromolecular Chemistry, Heinrich Heine University Duesseldorf, Universitaetsstrasse 1, 40225 Duesseldorf, GermanyInstitute for Macromolecular Chemistry, Albert Ludwig University of Freiburg, Stefan-Meier-Strasse 31, 79104 Freiburg, GermanyFreiburg Center of Interactive Materials and Bioinspired Technologies (FIT), Georges-Köhler-Allee 105, 79110 Freiburg, GermanyFreiburg Materials Research Center (FMF), Stefan-Meier-Straße 21, 79104 Freiburg, GermanyMore by Laura Hartmann
Abstract
Photoinduced affinity labeling for cross-linking biomolecules in close spatial proximity has become a powerful strategy in life science studies to identify interaction partners in fundamental research as well as biomarkers in applied studies. Next-generation photo-cross-linkers additionally provide inducible fluorogenic properties to enable a visual read-out. Azido-substituted coumarin is nonfluorescent, but UV irradiation initiates the formation of a highly reactive nitrene radical that can act as a cross-linker while restoring the fluorescence activity of the coumarin chromophore. In this study, we present a 7-azidocoumarin derivative that is used as a suitable building block for solid-phase synthesis and demonstrates easy access to a variety of glycan-based photo affinity probes. Applications of photo-cross-linkers for glycans and their respective binding proteins are still rare. We show several azidocoumarin glycan-presenting probes and their selective targeting and covalent linking to lectins, accompanied by a turn-on fluorescence activity of the coumarin fluorophore. Selective recognition of specific target lectins from the presented glycan photo affinity probes is further demonstrated in complex biological environments, which now open opportunities for identifying and localizing both known and previously unidentified glycan receptors in cells, tissues, or patient samples.
This publication is licensed under
License Summary*
You are free to share(copy and redistribute) this article in any medium or format and to adapt(remix, transform, and build upon) the material for any purpose, even commercially within the parameters below:
Creative Commons (CC): This is a Creative Commons license.
Attribution (BY): Credit must be given to the creator.
*Disclaimer
This summary highlights only some of the key features and terms of the actual license. It is not a license and has no legal value. Carefully review the actual license before using these materials.
License Summary*
You are free to share(copy and redistribute) this article in any medium or format and to adapt(remix, transform, and build upon) the material for any purpose, even commercially within the parameters below:
Creative Commons (CC): This is a Creative Commons license.
Attribution (BY): Credit must be given to the creator.
*Disclaimer
This summary highlights only some of the key features and terms of the actual license. It is not a license and has no legal value. Carefully review the actual license before using these materials.
License Summary*
You are free to share(copy and redistribute) this article in any medium or format and to adapt(remix, transform, and build upon) the material for any purpose, even commercially within the parameters below:
Creative Commons (CC): This is a Creative Commons license.
Attribution (BY): Credit must be given to the creator.
*Disclaimer
This summary highlights only some of the key features and terms of the actual license. It is not a license and has no legal value. Carefully review the actual license before using these materials.
Introduction
Figure 1
Figure 1. Schematic overview of the study. (A) Use of the AzC building block in SPS to give easy access to novel fluorogenic glycan-based affinity probes for applications in PAL. (B) Schematic concept of selective lectin targeting of Man- and Gal-based AzC affinity probes. Photoactivated probes cross-link their targets, while unbound, activated probes are removed during workup procedures.
Results
Synthesis of the 7-Azidocoumarin Building Block and Solid-Phase Synthesis of AzC Glycan PAL Probes
Figure 2
Figure 2. (A) Synthesis of the AzC building block from the aminocoumarin precursor, (B) SPS strategy, and (C) chemical structure of the AzC glycan PAL probes. Synthesis conditions: coupling: 5 eq. building block, 5 eq. PyBOP, 10 eq. DIPEA in DMF for 1 h; deprotection: 25Vol% piperidine in DMF, twice for 10 min, once for 20 min; glycan conjugation: 2.5 eq. Man-N3 (or Gal-N3) in DMF, 1.25 eq. sodium ascorbate and 1.25 eq. copper sulfate in MiliQ overnight; deacetylation: 0.2 M sodium methanolate in methanol for 1 h; AzC conjugation: 5 eq. AzC building block, 5 eq. HOBt, 5 eq. DIC in DMF for 1 h; resin cleavage: 95Vol% TFA, 2.5Vol% TIPS and 2.5Vol% DCM for 1 h.
Selective Cross-Linking of AzC Glycan PAL Probes to Target Lectins
Figure 3
Figure 3. (A) Schematic presentation of ConA binding to AzCMan and nonbinding of the AzCGal probe. (B) MALDI-TOF-MS spectra of AzC glycan PAL probes with ConA before (black) and after (red) irradiation. (C) Overview of results from MALDI-TOF-MS spectra with (+) indicating a peak for ConA cross-linked to AzCMan and (−) reflecting that no peak for ConA-AzCMan or ConA-AzCGal conjugate was found. (D–F) MALDI-TOF-MS spectra of AzC glycan PAL probes with ConA in the presence of competing carbohydrates or proteins: (D) in the presence of D-Man, (E) in the presence of α-MeMan, or (F) in the presence of BSA.
Inducible Fluorogenic Properties of AzC Glycan PAL Probes
Figure 4
Figure 4. (A) Maximal fluorescence intensities of AzCMan and AzCGal probes at 10 μM concentration incubated with either ConA or BSA alone or a mixture of both proteins. Data represent the mean of four individual measurements. (B) Limit of detection for AzCMan and limit of selectivity for AzCGal. Data represent the mean of four individual measurements. (C) SDS–PAGE of ConA incubated with AzC glycan PAL probes after irradiation at 365 nm; the top panel shows AzC fluorescence and the lower panel shows Coomassie stain of the same gel. (D) SDS–PAGE of RCA120 incubated with AzC glycan PAL probes after irradiation at 365nm; the top panel shows AzC fluorescence and the lower panel shows Coomassie stain of the same gel. The complete SDS–PAGE images including the AzCManB probe are shown in the SI (Figure S37).
Stability of the Induced AzC Probe Fluorescence at Different pH Ranges
Evaluation of the Minimal Irradiation Time to Gain Stable Probe-Target Cross-Linking
Figure 5
Figure 5. (A) Coomassie-stained and fluorescence image of the SDS–PAGE containing ConA with AzCMan at different durations of irradiation; the top panel shows Coomassie stain; the lower panel shows AzC fluorescence of the same gel. (B) Quantification of the fluorescence intensities of the individual bands normalized against ConA (last lane on the gel). The quantification was calculated as the mean of two replicates of the time-dependent irradiation series applied on one SDS–PAGE (full image provided in the SI, Figure S36).
Selective Target Recognition in Complex Biological Environments
Figure 6
Figure 6. (A) SDS–PAGE showing MDA-MB 231 lysate with or without supplemented ConA that was incubated with various AzC glycan PAL probes and subsequently irradiated with UV light for 15 min. (B) Western blot of native rat kidney lysate that was incubated with AzCGal and AzCManB probes in the presence or absence of the respective carbohydrate inhibitors (α-MeMan and D-Gal) and subsequently irradiated with UV light for 15 min. (C–E) Microscope images (top: bright-field; bottom: fluorescence at 488 nm) of fixed MDA-MB-231 cells incubated with vehicle (C), AzCMan (D), or preactivated AzCMan (E) and subsequently irradiated at 365 nm for 15 min.
Conclusions
Methods
General
Nuclear Magnetic Resonance Spectroscopy (NMR)
High-Resolution Mass Spectrometry (HR-MS)
Matrix-Assisted Laser Desorption Ionization Time-of-Flight (MALDI-TOF) Mass Spectrometry (MALDI-TOF-MS)
Reversed-Phase High-Pressure Liquid Chromatography (RP-HPLC)
Synthesis of Building Blocks for Solid-Phase Synthesis
Synthesis of Azidocoumarin (AzC)
General Procedure for Solid-Phase Synthesis
Coupling and Fmoc Deprotection
Copper(I)-Catalyzed Alkyne–Azide Cycloaddition (CuAAC)
Removing Acetyl Protection Groups from Carbohydrates
Conjugation of Azidocoumarin
Cleavage from the Resin
Irradiation Experiments
General Procedure or Irradiation Experiments
Competition Assays with Carbohydrates
Competition Assays with BSA
ConA- and BSA-Fluorescence Measurements
Absorption and Fluorescence Measurements
pH-Dependent Fluorescence Measurements
Concentration-Dependent Fluorescence Measurements
Statistical Analysis LOD
General SDS–PAGE Procedure
Duration of Irradiation Measurements
Selective Binding of AzC Probes to Lectins ConA and RCA120
Selective ConA Binding of AzCMan Probes in Cell Lysate Background
Testing AzC Probes in Native Rat Kidney Lysate
Testing AzC Probes on Cells
Supporting Information
The Supporting Information is available free of charge at https://pubs.acs.org/doi/10.1021/acs.bioconjchem.5c00613.
Analytical data for the synthesized structures, additional data and figures showing the absorption and fluorescence spectra of AzC building blocks, full MALDI-TOF-MS and full fluorescence spectra, additional fluorescence spectra for the pH-stability study, additional SDS–PAGE images of irradiated and not irradiated lysates, and additional fluorescence microscopy images (PDF)
Terms & Conditions
Most electronic Supporting Information files are available without a subscription to ACS Web Editions. Such files may be downloaded by article for research use (if there is a public use license linked to the relevant article, that license may permit other uses). Permission may be obtained from ACS for other uses through requests via the RightsLink permission system: http://pubs.acs.org/page/copyright/permissions.html.
Acknowledgments
We acknowledge the support of the German Research Foundation (DFG) within the Collaborative Research Center 1208 “Identity and Dynamics of Membrane Systems”.
References
This article references 86 other publications.
- 1Jayachandran, B.; Parvin, T. N.; Alam, M. M.; Chanda, K.; Mm, B. Insights on Chemical Crosslinking Strategies for Proteins. Molecules 2022, 27, 8124, DOI: 10.3390/molecules27238124Google ScholarThere is no corresponding record for this reference.
- 2Liu, J.; Yang, B.; Wang, L. Residue selective crosslinking of proteins through photoactivatable or proximity-enabled reactivity. Curr. Opin. Chem. Biol. 2023, 74, 102285 DOI: 10.1016/j.cbpa.2023.102285Google ScholarThere is no corresponding record for this reference.
- 3Kotzyba-Hibert, F.; Kapfer, I.; Goeldner, M. Recent trends in photoaffinity labeling. Angew. Chem., Int. Ed. 1995, 34, 1296– 1312, DOI: 10.1002/anie.199512961Google ScholarThere is no corresponding record for this reference.
- 4Bayley, H.; Knowles, J. R. Photoaffinity labeling. Methods Enzymol. 1977, 46, 69– 114, DOI: 10.1016/s0076-6879(77)46012-9Google ScholarThere is no corresponding record for this reference.
- 5Yu, S.-H.; Wands, A. M.; Kohler, J. J. Photoaffinity probes for studying carbohydrate biology. J. Carbohydr. Chem. 2012, 31, 325– 352, DOI: 10.1080/07328303.2012.676118Google ScholarThere is no corresponding record for this reference.
- 6Dorman, G.; Nakamura, H.; Pulsipher, A.; Prestwich, G. D. The life of pi star: exploring the exciting and forbidden worlds of the benzophenone photophore. Chem. Rev. 2016, 116, 15284– 15398, DOI: 10.1021/acs.chemrev.6b00342Google ScholarThere is no corresponding record for this reference.
- 7Galardy, R. E.; Craig, L. C.; Jamieson, J. D.; Printz, M. P. Photoaffinity labeling of peptide hormone binding sites. J. Biol. Chem. 1974, 249, 3510– 3518, DOI: 10.1016/S0021-9258(19)42601-XGoogle ScholarThere is no corresponding record for this reference.
- 8Fleet, G.; Porter, R.; Knowles, J. Affinity labelling of antibodies with aryl nitrene as reactive group. Nature 1969, 224, 511– 512, DOI: 10.1038/224511a0Google ScholarThere is no corresponding record for this reference.
- 9Kym, P. R.; Carlson, K. E.; Katzenellenbogen, J. A. Evaluation of a highly efficient aryl azide photoaffinity labeling reagent for the progesterone receptor. Bioconjugate Chem. 1995, 6, 115– 122, DOI: 10.1021/bc00031a014Google ScholarThere is no corresponding record for this reference.
- 10Smith, R. A.; Knowles, J. R. Aryldiazirines. Potential reagents for photolabeling of biological receptor sites. J. Am. Chem. Soc. 1973, 95, 5072– 5073, DOI: 10.1021/ja00796a062Google ScholarThere is no corresponding record for this reference.
- 11Mishra, P. K.; Yoo, C. M.; Hong, E.; Rhee, H. W. Photo-crosslinking: an emerging chemical tool for investigating molecular networks in live cells. ChemBioChem. 2020, 21, 924– 932, DOI: 10.1002/cbic.201900600Google ScholarThere is no corresponding record for this reference.
- 12Leyva, E.; Platz, M. S.; Persy, G.; Wirz, J. Photochemistry of phenyl azide: the role of singlet and triplet phenylnitrene as transient intermediates. J. Am. Chem. Soc. 1986, 108, 3783– 3790, DOI: 10.1021/ja00273a037Google ScholarThere is no corresponding record for this reference.
- 13Walrant, A.; Sachon, E. Photoaffinity labeling coupled to MS to identify peptide biological partners: Secondary reactions, for better or for worse?. Mass Spectrom. Rev. 2025, 44, 715– 756, DOI: 10.1002/mas.21880Google ScholarThere is no corresponding record for this reference.
- 14Murale, D. P.; Hong, S. C.; Haque, M. M.; Lee, J. S. Photo-affinity labeling (PAL) in chemical proteomics: a handy tool to investigate protein-protein interactions (PPIs). Proteome Sci. 2017, 15, 14, DOI: 10.1186/s12953-017-0123-3Google ScholarThere is no corresponding record for this reference.
- 15Liang, T. Y.; Schuster, G. B. Photochemistry of 3- and 4-nitrophenyl azide: detection and characterization of reactive intermediates. J. Am. Chem. Soc. 1987, 109, 7803– 7810, DOI: 10.1021/ja00259a032Google ScholarThere is no corresponding record for this reference.
- 16Wentrup, C. Nitrenes, carbenes, diradicals, and ylides. Interconversions of reactive intermediates. Acc. Chem. Res. 2011, 44, 393– 404, DOI: 10.1021/ar700198zGoogle ScholarThere is no corresponding record for this reference.
- 17Chowdhry, V.; Westheimer, F. Photoaffinity labeling of biological systems. Annu. Rev. Biochem. 1979, 48, 293– 325, DOI: 10.1146/annurev.bi.48.070179.001453Google ScholarThere is no corresponding record for this reference.
- 18Wu, H.; Kohler, J. Photocrosslinking probes for capture of carbohydrate interactions. Curr. Opin. Chem. Biol. 2019, 53, 173– 182, DOI: 10.1016/j.cbpa.2019.09.002Google ScholarThere is no corresponding record for this reference.
- 19Jahn, O.; Eckart, K.; Tezval, H.; Spiess, J. Characterization of peptide–protein interactions using photoaffinity labeling and LC/MS. Anal. Bioanal. Chem. 2004, 378, 1031– 1036, DOI: 10.1007/s00216-003-2353-8Google ScholarThere is no corresponding record for this reference.
- 20Robinette, D.; Neamati, N.; Tomer, K. B.; Borchers, C. H. Photoaffinity labeling combined with mass spectrometric approaches as a tool for structural proteomics. Expert review of proteomics 2006, 3, 399– 408, DOI: 10.1586/14789450.3.4.399Google ScholarThere is no corresponding record for this reference.
- 21Tulloch, L. B.; Menzies, S. K.; Fraser, A. L.; Gould, E. R.; King, E. F.; Zacharova, M. K.; Florence, G. J.; Smith, T. K. Photo-affinity labelling and biochemical analyses identify the target of trypanocidal simplified natural product analogues. PLoS Neglected Tropical Diseases 2017, 11, e0005886 DOI: 10.1371/journal.pntd.0005886Google ScholarThere is no corresponding record for this reference.
- 22Smith, E.; Collins, I. Photoaffinity labeling in target-and binding-site identification. Future medicinal chemistry 2015, 7, 159– 183, DOI: 10.4155/fmc.14.152Google ScholarThere is no corresponding record for this reference.
- 23Park, J.; Koh, M.; Koo, J. Y.; Lee, S.; Park, S. B. Investigation of specific binding proteins to photoaffinity linkers for efficient deconvolution of target protein. ACS Chem. Biol. 2016, 11, 44– 52, DOI: 10.1021/acschembio.5b00671Google ScholarThere is no corresponding record for this reference.
- 24Wagner, S.; Hauck, D.; Hoffmann, M.; Sommer, R.; Joachim, I.; Müller, R.; Imberty, A.; Varrot, A.; Titz, A. Covalent Lectin Inhibition and Application in Bacterial Biofilm Imaging. Angew. Chem., Int. Ed. 2017, 56, 16559– 16564, DOI: 10.1002/anie.201709368Google ScholarThere is no corresponding record for this reference.
- 25Wang, Y.; Torres-García, D.; Mostert, T. P.; Reinalda, L.; Van Kasteren, S. I. A Bioorthogonal Dual Fluorogenic Probe for the Live-Cell Monitoring of Nutrient Uptake by Mammalian Cells. Angew. Chem., Int. Ed. 2024, 63, e202401733 DOI: 10.1002/anie.202401733Google ScholarThere is no corresponding record for this reference.
- 26Yu, A.; He, X.; Shen, T.; Yu, X.; Mao, W.; Chi, W.; Liu, X.; Wu, H. Design strategies for tetrazine fluorogenic probes for bioorthogonal imaging. Chem. Soc. Rev. 2025, 54, 2984, DOI: 10.1039/D3CS00520HGoogle ScholarThere is no corresponding record for this reference.
- 27Jeong, H.; Wu, X.; Lee, J.-S.; Yoon, J. Recent advances in enzyme-activated NIR fluorescent probes for biological applications. TrAC Trends in Analytical Chemistry 2023, 168, 117335 DOI: 10.1016/j.trac.2023.117335Google ScholarThere is no corresponding record for this reference.
- 28Cosco, E. D.; Bogyo, M. Recent advances in ratiometric fluorescence imaging of enzyme activity in vivo. Curr. Opin. Chem. Biol. 2024, 80, 102441 DOI: 10.1016/j.cbpa.2024.102441Google ScholarThere is no corresponding record for this reference.
- 29Scott, J. I.; Deng, Q.; Vendrell, M. Near-infrared fluorescent probes for the detection of cancer-associated proteases. ACS Chem. Biol. 2021, 16, 1304– 1317, DOI: 10.1021/acschembio.1c00223Google ScholarThere is no corresponding record for this reference.
- 30Saleem, M.; Hanif, M.; Bonne, S.; Zeeshan, M.; Khan, S.; Rafiq, M.; Tahir, T.; Lu, C.; Cai, R. Turn-On Fluorescence Probe for Cancer-Related γ-Glutamyltranspeptidase Detection. Molecules 2024, 29, 4776, DOI: 10.3390/molecules29194776Google ScholarThere is no corresponding record for this reference.
- 31Tsao, K. K.; Imai, S.; Chang, M.; Hario, S.; Terai, T.; Campbell, R. E. The best of both worlds: Chemigenetic fluorescent sensors for biological imaging. Cell Chemical Biology 2024, 31, 1652– 1664, DOI: 10.1016/j.chembiol.2024.08.002Google ScholarThere is no corresponding record for this reference.
- 32Nadal-Bufi, F.; Salomon, P. L.; de Moliner, F.; Sarris, K. A.; Wang, Z.; Wills, R. D.; Marin, V. L.; Shi, X.; Zhou, K.; Wang, Z.; Xu, Z.; McPherson, M. J.; Marvin, C. C.; Hobson, A. D.; Vendrell, M. Fluorogenic Platform for Real-Time Imaging of Subcellular Payload Release in Antibody–Drug Conjugates. J. Am. Chem. Soc. 2025, 147, 7578– 7587, DOI: 10.1021/jacs.4c16842Google ScholarThere is no corresponding record for this reference.
- 33Kellner, S.; Seidu-Larry, S.; Burhenne, J.; Motorin, Y.; Helm, M. A multifunctional bioconjugate module for versatile photoaffinity labeling and click chemistry of RNA. Nucleic Acids Res. 2011, 39, 7348– 7360, DOI: 10.1093/nar/gkr449Google ScholarThere is no corresponding record for this reference.
- 34Morimoto, S.; Tomohiro, T.; Maruyama, N.; Hatanaka, Y. Photoaffinity casting of a coumarin flag for rapid identification of ligand-binding sites within protein. Chem. Commun. 2013, 49, 1811– 1813, DOI: 10.1039/c3cc38594aGoogle ScholarThere is no corresponding record for this reference.
- 35Tomohiro, T.; Yamamoto, A.; Tatsumi, Y.; Hatanaka, Y. [3-(Trifluoromethyl)-3 H-diazirin-3-yl] coumarin as a carbene-generating photocross-linker with masked fluorogenic beacon. Chem. Commun. 2013, 49, 11551– 11553, DOI: 10.1039/c3cc45780jGoogle ScholarThere is no corresponding record for this reference.
- 36Singha, M.; Roy, S.; Pandey, S. D.; Bag, S. S.; Bhattacharya, P.; Das, M.; Ghosh, A. S.; Ray, D.; Basak, A. Use of azidonaphthalimide carboxylic acids as fluorescent templates with a built-in photoreactive group and a flexible linker simplifies protein labeling studies: applications in selective tagging of HCAII and penicillin binding proteins. Chem. Commun. 2017, 53, 13015– 13018, DOI: 10.1039/C7CC08209FGoogle ScholarThere is no corresponding record for this reference.
- 37Képiró, M.; Várkuti, B. H.; Rauscher, A. A.; Kellermayer, M. S.; Varga, M.; Málnási-Csizmadia, A. Molecular tattoo: subcellular confinement of drug effects. Chemistry & biology 2015, 22, 548– 558, DOI: 10.1016/j.chembiol.2015.03.013Google ScholarThere is no corresponding record for this reference.
- 38Dai, S.-Y.; Yang, D. A visible and near-infrared light activatable diazocoumarin probe for fluorogenic protein labeling in living cells. J. Am. Chem. Soc. 2020, 142, 17156– 17166, DOI: 10.1021/jacs.0c08068Google ScholarThere is no corresponding record for this reference.
- 39Bousch, C.; Vreulz, B.; Kansal, K.; El-Husseini, A.; Cecioni, S. Fluorogenic Photo-Crosslinking of Glycan-Binding Protein Recognition Using a Fluorinated Azido-Coumarin Fucoside. Angew. Chem., Int. Ed. 2023, 62, e202314248 DOI: 10.1002/anie.202314248Google ScholarThere is no corresponding record for this reference.
- 40Vreulz, B.; De Crozals, D.; Cecioni, S. A trifunctional probe for generation of fluorogenic glycan-photocrosslinker conjugates. RSC chemical biology 2025, 6, 1779– 1786, DOI: 10.1039/D5CB00206KGoogle ScholarThere is no corresponding record for this reference.
- 41Brown, S. A. Biochemistry of The Coumarins, in Biochemistry of Plant Phenolics, Swain, T.; Harbone, J. B.; Van Sumere, C. F., Eds., Springer US: Boston, MA, 1979; pp 249– 286.Google ScholarThere is no corresponding record for this reference.
- 42Musa, M. A.; Cooperwood, J. S.; Khan, M. O. F. A review of coumarin derivatives in pharmacotherapy of breast cancer. Curr. Med. Chem. 2008, 15 (26), 2664– 2679, DOI: 10.2174/092986708786242877Google ScholarThere is no corresponding record for this reference.
- 43Stringlis, I. A.; de Jonge, R.; Pieterse, C. M. J. The Age of Coumarins in Plant–Microbe Interactions. Plant Cell Physiol. 2019, 60, 1405– 1419, DOI: 10.1093/pcp/pcz076Google ScholarThere is no corresponding record for this reference.
- 44Sun, X.-y.; Liu, T.; Sun, J.; Wang, X.-j. Synthesis and application of coumarin fluorescence probes. RSC Adv. 2020, 10, 10826– 10847, DOI: 10.1039/C9RA10290FGoogle ScholarThere is no corresponding record for this reference.
- 45Tasior, M.; Kim, D.; Singha, S.; Krzeszewski, M.; Ahn, K. H.; Gryko, D. T. π-Expanded coumarins: synthesis, optical properties and applications. Journal of Materials Chemistry C 2015, 3, 1421– 1446, DOI: 10.1039/C4TC02665AGoogle ScholarThere is no corresponding record for this reference.
- 46Szwaczko, K. Coumarins synthesis and transformation via C–H bond activation─A review. Inorganics 2022, 10, 23, DOI: 10.3390/inorganics10020023Google ScholarThere is no corresponding record for this reference.
- 47Kang, D.; Ahn, K.; Hong, S. Site-Selective C– H Bond Functionalization of Chromones and Coumarins. Asian Journal of Organic Chemistry 2018, 7, 1136– 1150, DOI: 10.1002/ajoc.201800128Google ScholarThere is no corresponding record for this reference.
- 48Tod, M.; Prevot, M.; Poulou, M.; Farinotti, R.; Chalom, J.; Mahuzier, G. Chromatographic and luminescence properties of a 7-aminocoumarin derivative with peroxyoxalate chemiexcitation. Anal. Chim. Acta 1989, 223, 309– 317, DOI: 10.1016/S0003-2670(00)84096-9Google ScholarThere is no corresponding record for this reference.
- 49Murase, T.; Yoshihara, T.; Yamada, K.; Tobita, S. Fluorescent peptides labeled with environment-sensitive 7-aminocoumarins and their interactions with lipid bilayer membranes and living cells. Bull. Chem. Soc. Jpn. 2013, 86, 510– 519, DOI: 10.1246/bcsj.20120314Google ScholarThere is no corresponding record for this reference.
- 50Nowakowska, M.; Smoluch, M.; Sendor, D. The effect of cyclodextrins on the photochemical stability of 7-amino-4-methylcoumarin in aqueous solution. J. Incl. Phenom. Macrocycl. Chem. 2001, 40, 213– 219, DOI: 10.1023/A:1011820513256Google ScholarThere is no corresponding record for this reference.
- 51Raju, B. B.; Costa, S. M. Excited-state behavior of 7-diethylaminocoumarin dyes in AOT reversed micelles: size effects. J. Phys. Chem. B 1999, 103, 4309– 4317, DOI: 10.1021/jp984527yGoogle ScholarThere is no corresponding record for this reference.
- 52Gandioso, A.; Palau, M.; Bresolí-Obach, R.; Galindo, A.; Rovira, A.; Bosch, M.; Nonell, S.; Marchán, V. High photostability in nonconventional coumarins with far-red/NIR emission through azetidinyl substitution. Journal of Organic Chemistry 2018, 83, 11519– 11531, DOI: 10.1021/acs.joc.8b01422Google ScholarThere is no corresponding record for this reference.
- 53Rovira, A.; Pujals, M.; Gandioso, A.; López-Corrales, M.; Bosch, M.; Marchán, V. Modulating photostability and mitochondria selectivity in far-red/NIR emitting coumarin fluorophores through replacement of pyridinium by pyrimidinium. Journal of Organic Chemistry 2020, 85, 6086– 6097, DOI: 10.1021/acs.joc.0c00570Google ScholarThere is no corresponding record for this reference.
- 54Lord, S. J.; Lee, H.-l. D.; Samuel, R.; Weber, R.; Liu, N.; Conley, N. R.; Thompson, M. A.; Twieg, R. J.; Moerner, W. Azido push– pull fluorogens photoactivate to produce bright fluorescent labels. J. Phys. Chem. B 2010, 114, 14157– 14167, DOI: 10.1021/jp907080rGoogle ScholarThere is no corresponding record for this reference.
- 55De-La-Cuesta, J.; González, E.; Pomposo, J. A. Advances in fluorescent single-chain nanoparticles. Molecules 2017, 22, 1819, DOI: 10.3390/molecules22111819Google ScholarThere is no corresponding record for this reference.
- 56Sivakumar, K.; Xie, F.; Cash, B. M.; Long, S.; Barnhill, H. N.; Wang, Q. A fluorogenic 1, 3-dipolar cycloaddition reaction of 3-azidocoumarins and acetylenes. Org. Lett. 2004, 6, 4603– 4606, DOI: 10.1021/ol047955xGoogle ScholarThere is no corresponding record for this reference.
- 57Porell, R. N.; Follmar, J. L.; Purcell, S. C.; Timm, B.; Laubach, L. K.; Kozirovskiy, D.; Thacker, B. E.; Glass, C. A.; Gordts, P. L. S. M.; Godula, K. Biologically Derived Neoproteoglycans for Profiling Protein–Glycosaminoglycan Interactions. ACS Chem. Biol. 2022, 17, 1534– 1542, DOI: 10.1021/acschembio.2c00205Google ScholarThere is no corresponding record for this reference.
- 58Chalansonnet, V.; Lowe, J.; Orenga, S.; Perry, J. D.; Robinson, S. N.; Stanforth, S. P.; Sykes, H. E.; Truong, T. V. Fluorogenic 7-azidocoumarin and 3/4-azidophthalimide derivatives as indicators of reductase activity in microorganisms. Bioorg. Med. Chem. Lett. 2019, 29, 2354– 2357, DOI: 10.1016/j.bmcl.2019.06.010Google ScholarThere is no corresponding record for this reference.
- 59Hill, S. A.; Gerke, C.; Hartmann, L. Recent Developments in Solid-Phase Strategies towards Synthetic, Sequence-Defined Macromolecules. Chem.─Asian J. 2018, 13, 3611– 3622, DOI: 10.1002/asia.201801171Google ScholarThere is no corresponding record for this reference.
- 60Blawitzki, L.-C.; Bartels, N.; Bonda, L.; Schmidt, S.; Monzel, C.; Hartmann, L. Glycomacromolecules to Tailor Crowded and Heteromultivalent Glycocalyx Mimetics. Biomacromolecules 2024, 25, 5979– 5994, DOI: 10.1021/acs.biomac.4c00646Google ScholarThere is no corresponding record for this reference.
- 61Jäck, N.; Hemming, A.; Hartmann, L. Synthesis of Dual-Responsive Amphiphilic Glycomacromolecules: Controlled Release of Glycan Ligands via pH and UV Stimuli. Macromol. Rapid Commun. 2024, 45, 2400439 DOI: 10.1002/marc.202400439Google ScholarThere is no corresponding record for this reference.
- 62Hoffmann, M.; Snyder, N. L.; Hartmann, L. Glycosaminoglycan Mimetic Precision Glycomacromolecules with Sequence-Defined Sulfation and Rigidity Patterns. Biomacromolecules 2022, 23, 4004– 4014, DOI: 10.1021/acs.biomac.2c00829Google ScholarThere is no corresponding record for this reference.
- 63Pǎunescu, E.; Louise, L.; Jean, L.; Romieu, A.; Renard, P.-Y. A versatile access to new halogenated 7-azidocoumarins for photoaffinity labeling: Synthesis and photophysical properties. Dyes Pigm. 2011, 91, 427– 434, DOI: 10.1016/j.dyepig.2011.05.008Google ScholarThere is no corresponding record for this reference.
- 64Yang, Q.; Váňa, J.; Klán, P. The complex photochemistry of coumarin-3-carboxylic acid in acetonitrile and methanol. Photochemical & Photobiological Sciences 2022, 21, 1481– 1495, DOI: 10.1007/s43630-022-00238-8Google ScholarThere is no corresponding record for this reference.
- 65Ponader, D.; Wojcik, F.; Beceren-Braun, F.; Dernedde, J.; Hartmann, L. Sequence-defined glycopolymer segments presenting mannose: synthesis and lectin binding affinity. Biomacromolecules 2012, 13, 1845– 1852, DOI: 10.1021/bm300331zGoogle ScholarThere is no corresponding record for this reference.
- 66Wojcik, F.; Lel, S.; O’Brien, A. G.; Seeberger, P. H.; Hartmann, L. Synthesis of homo-and heteromultivalent carbohydrate-functionalized oligo (amidoamines) using novel glyco-building blocks. Beilstein journal of organic chemistry 2013, 9, 2395– 2403, DOI: 10.3762/bjoc.9.276Google ScholarThere is no corresponding record for this reference.
- 67Ponader, D.; Maffre, P.; Aretz, J.; Pussak, D.; Ninnemann, N. M.; Schmidt, S.; Seeberger, P. H.; Rademacher, C.; Nienhaus, G. U.; Hartmann, L. Carbohydrate-Lectin Recognition of Sequence-Defined Heteromultivalent Glycooligomers. J. Am. Chem. Soc. 2014, 136, 2008, DOI: 10.1021/ja411582tGoogle ScholarThere is no corresponding record for this reference.
- 68Cummings, R. D. The mannose receptor ligands and the macrophage glycome. Curr. Opin. Struct. Biol. 2022, 75, 102394 DOI: 10.1016/j.sbi.2022.102394Google ScholarThere is no corresponding record for this reference.
- 69Coombs, P. J.; Taylor, M. E.; Drickamer, K. Two categories of mammalian galactose-binding receptors distinguished by glycan array profiling. Glycobiology 2006, 16, 1C– 7C, DOI: 10.1093/glycob/cwj126Google ScholarThere is no corresponding record for this reference.
- 70Weatherman, R. V.; Mortell, K. H.; Chervenak, M.; Kiessling, L. L.; Toone, E. J. Specificity of C-glycoside complexation by mannose/glucose specific lectins. Biochemistry 1996, 35, 3619– 3624, DOI: 10.1021/bi951916zGoogle ScholarThere is no corresponding record for this reference.
- 71Itakura, Y.; Nakamura-Tsuruta, S.; Kominami, J.; Sharon, N.; Kasai, K.-i.; Hirabayashi, J. Systematic Comparison of Oligosaccharide Specificity of Ricinus communis Agglutinin I and Erythrina Lectins: a Search by Frontal Affinity Chromatography†. J. Biochem. 2007, 142, 459– 469, DOI: 10.1093/jb/mvm153Google ScholarThere is no corresponding record for this reference.
- 72Wang, Y.; Yu, G.; Han, Z.; Yang, B.; Hu, Y.; Zhao, X.; Wu, J.; Lv, Y.; Chai, W. Specificities of Ricinus communis agglutinin 120 interaction with sulfated galactose. FEBS Lett. 2011, 585, 3927– 3934, DOI: 10.1016/j.febslet.2011.10.035Google ScholarThere is no corresponding record for this reference.
- 73Bojar, D.; Meche, L.; Meng, G.; Eng, W.; Smith, D. F.; Cummings, R. D.; Mahal, L. K. A Useful Guide to Lectin Binding: Machine-Learning Directed Annotation of 57 Unique Lectin Specificities. ACS Chem. Biol. 2022, 17, 2993– 3012, DOI: 10.1021/acschembio.1c00689Google ScholarThere is no corresponding record for this reference.
- 74Maljaars, C. E. P.; Halkes, K. M.; de Oude, W. L.; Haseley, S. R.; Upton, P. J.; McDonnell, M. B.; Kamerling, J. P. Affinity Determination of Ricinus communis Agglutinin Ligands Identified from Combinatorial O- and S-,N-Glycopeptide Libraries. J. Comb. Chem. 2006, 8, 812– 819, DOI: 10.1021/cc060019jGoogle ScholarThere is no corresponding record for this reference.
- 75Schuster, G. B.; Platz, M. S. Photochemistry of phenyl azide. Advances in photochemistry 1992, 17, 69– 143, DOI: 10.1002/9780470133484.ch2Google ScholarThere is no corresponding record for this reference.
- 76Reddy, V. S.; Rao, V. Modes of binding of α (1–2) linked manno-oligosaccharides to concanavalin A. Int. J. Biol. Macromol. 1992, 14, 185– 192, DOI: 10.1016/S0141-8130(05)80025-XGoogle ScholarThere is no corresponding record for this reference.
- 77Jeyachandran, Y. L.; Mielczarski, J. A.; Mielczarski, E.; Rai, B. Efficiency of blocking of non-specific interaction of different proteins by BSA adsorbed on hydrophobic and hydrophilic surfaces. J. Colloid Interface Sci. 2010, 341, 136– 142, DOI: 10.1016/j.jcis.2009.09.007Google ScholarThere is no corresponding record for this reference.
- 78Al-Husseini, J. K.; Stanton, N. J.; Selassie, C. R.; Johal, M. S. The binding of drug molecules to serum albumin: The effect of drug hydrophobicity on binding strength and protein desolvation. Langmuir 2019, 35, 17054– 17060, DOI: 10.1021/acs.langmuir.9b02318Google ScholarThere is no corresponding record for this reference.
- 79Smith, E. A.; Thomas, W. D.; Kiessling, L. L.; Corn, R. M. Surface Plasmon Resonance Imaging Studies of Protein-Carbohydrate Interactions. J. Am. Chem. Soc. 2003, 125, 6140– 6148, DOI: 10.1021/ja034165uGoogle ScholarThere is no corresponding record for this reference.
- 80Parera Pera, N.; Branderhorst, H. M.; Kooij, R.; Maierhofer, C.; van der Kaaden, M.; Liskamp, R. M. J.; Wittmann, V.; Ruijtenbeek, R.; Pieters, R. J. Rapid Screening of Lectins for Multivalency Effects with a Glycodendrimer Microarray. ChemBioChem. 2010, 11, 1896– 1904, DOI: 10.1002/cbic.201000340Google ScholarThere is no corresponding record for this reference.
- 81Feldhof, M. I.; Sperzel, S.; Bonda, L.; Boye, S.; Braunschweig, A. B.; Gerling-Driessen, U. I. M.; Hartmann, L. Thiol-selective native grafting from polymerization for the generation of protein–polymer conjugates. Chem. Sci. 2024, 15, 16768– 16777, DOI: 10.1039/D4SC04818KGoogle ScholarThere is no corresponding record for this reference.
- 82Olmsted, I. R.; Kussrow, A.; Bornhop, D. J. Comparison of Free-Solution and Surface-Immobilized Molecular Interactions Using a Single Platform. Anal. Chem. 2012, 84, 10817– 10822, DOI: 10.1021/ac302933hGoogle ScholarThere is no corresponding record for this reference.
- 83Ramesh, D.; Srinivasan, M. Studies on ring opening of coumarins. Curr. Sci. 1984, 53, 369– 371Google ScholarThere is no corresponding record for this reference.
- 84Mała, P.; Siebs, E.; Meiers, J.; Rox, K.; Varrot, A.; Imberty, A.; Titz, A. Discovery of N-β-l-Fucosyl Amides as High-Affinity Ligands for the Pseudomonas aeruginosa Lectin LecB. J. Med. Chem. 2022, 65, 14180– 14200, DOI: 10.1021/acs.jmedchem.2c01373Google ScholarThere is no corresponding record for this reference.
- 85Hayes, W.; Osborn, H. M.; Osborne, S. D.; Rastall, R. A.; Romagnoli, B. One-pot synthesis of multivalent arrays of mannose mono-and disaccharides. Tetrahedron 2003, 59, 7983– 7996, DOI: 10.1016/j.tet.2003.08.011Google ScholarThere is no corresponding record for this reference.
- 86Wu, L.; Sampson, N. S. Fucose, Mannose, and β-N-Acetylglucosamine Glycopolymers Initiate the Mouse Sperm Acrosome Reaction through Convergent Signaling Pathways. ACS Chem. Biol. 2014, 9, 468– 475, DOI: 10.1021/cb400550jGoogle ScholarThere is no corresponding record for this reference.
Cited By
This article has not yet been cited by other publications.
Article Views
Altmetric
Citations
Article Views are the COUNTER-compliant sum of full text article downloads since November 2008 (both PDF and HTML) across all institutions and individuals. These metrics are regularly updated to reflect usage leading up to the last few days.
Citations are the number of other articles citing this article, calculated by Crossref and updated daily. Find more information about Crossref citation counts.
The Altmetric Attention Score is a quantitative measure of the attention that a research article has received online. Clicking on the donut icon will load a page at altmetric.com with additional details about the score and the social media presence for the given article. Find more information on the Altmetric Attention Score and how the score is calculated.
Recommended Articles
Abstract
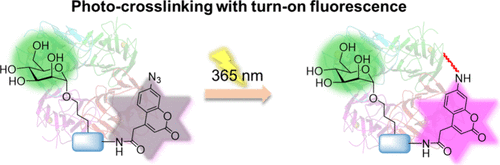
Figure 1

Figure 1. Schematic overview of the study. (A) Use of the AzC building block in SPS to give easy access to novel fluorogenic glycan-based affinity probes for applications in PAL. (B) Schematic concept of selective lectin targeting of Man- and Gal-based AzC affinity probes. Photoactivated probes cross-link their targets, while unbound, activated probes are removed during workup procedures.
Figure 2

Figure 2. (A) Synthesis of the AzC building block from the aminocoumarin precursor, (B) SPS strategy, and (C) chemical structure of the AzC glycan PAL probes. Synthesis conditions: coupling: 5 eq. building block, 5 eq. PyBOP, 10 eq. DIPEA in DMF for 1 h; deprotection: 25Vol% piperidine in DMF, twice for 10 min, once for 20 min; glycan conjugation: 2.5 eq. Man-N3 (or Gal-N3) in DMF, 1.25 eq. sodium ascorbate and 1.25 eq. copper sulfate in MiliQ overnight; deacetylation: 0.2 M sodium methanolate in methanol for 1 h; AzC conjugation: 5 eq. AzC building block, 5 eq. HOBt, 5 eq. DIC in DMF for 1 h; resin cleavage: 95Vol% TFA, 2.5Vol% TIPS and 2.5Vol% DCM for 1 h.
Figure 3

Figure 3. (A) Schematic presentation of ConA binding to AzCMan and nonbinding of the AzCGal probe. (B) MALDI-TOF-MS spectra of AzC glycan PAL probes with ConA before (black) and after (red) irradiation. (C) Overview of results from MALDI-TOF-MS spectra with (+) indicating a peak for ConA cross-linked to AzCMan and (−) reflecting that no peak for ConA-AzCMan or ConA-AzCGal conjugate was found. (D–F) MALDI-TOF-MS spectra of AzC glycan PAL probes with ConA in the presence of competing carbohydrates or proteins: (D) in the presence of D-Man, (E) in the presence of α-MeMan, or (F) in the presence of BSA.
Figure 4
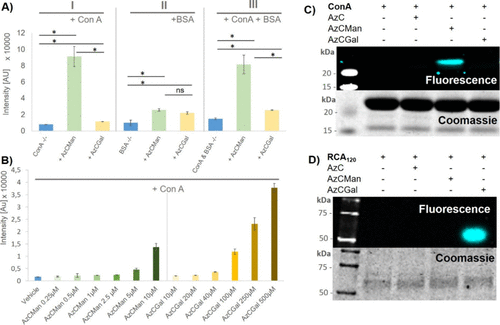
Figure 4. (A) Maximal fluorescence intensities of AzCMan and AzCGal probes at 10 μM concentration incubated with either ConA or BSA alone or a mixture of both proteins. Data represent the mean of four individual measurements. (B) Limit of detection for AzCMan and limit of selectivity for AzCGal. Data represent the mean of four individual measurements. (C) SDS–PAGE of ConA incubated with AzC glycan PAL probes after irradiation at 365 nm; the top panel shows AzC fluorescence and the lower panel shows Coomassie stain of the same gel. (D) SDS–PAGE of RCA120 incubated with AzC glycan PAL probes after irradiation at 365nm; the top panel shows AzC fluorescence and the lower panel shows Coomassie stain of the same gel. The complete SDS–PAGE images including the AzCManB probe are shown in the SI (Figure S37).
Figure 5

Figure 5. (A) Coomassie-stained and fluorescence image of the SDS–PAGE containing ConA with AzCMan at different durations of irradiation; the top panel shows Coomassie stain; the lower panel shows AzC fluorescence of the same gel. (B) Quantification of the fluorescence intensities of the individual bands normalized against ConA (last lane on the gel). The quantification was calculated as the mean of two replicates of the time-dependent irradiation series applied on one SDS–PAGE (full image provided in the SI, Figure S36).
Figure 6
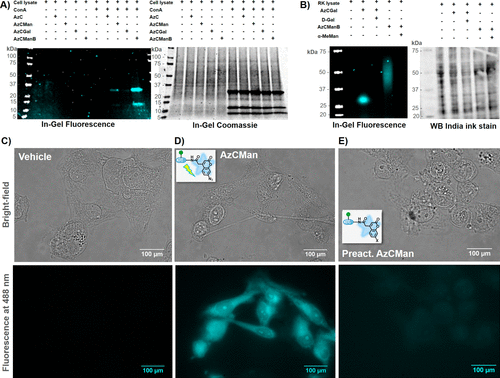
Figure 6. (A) SDS–PAGE showing MDA-MB 231 lysate with or without supplemented ConA that was incubated with various AzC glycan PAL probes and subsequently irradiated with UV light for 15 min. (B) Western blot of native rat kidney lysate that was incubated with AzCGal and AzCManB probes in the presence or absence of the respective carbohydrate inhibitors (α-MeMan and D-Gal) and subsequently irradiated with UV light for 15 min. (C–E) Microscope images (top: bright-field; bottom: fluorescence at 488 nm) of fixed MDA-MB-231 cells incubated with vehicle (C), AzCMan (D), or preactivated AzCMan (E) and subsequently irradiated at 365 nm for 15 min.
References
This article references 86 other publications.
- 1Jayachandran, B.; Parvin, T. N.; Alam, M. M.; Chanda, K.; Mm, B. Insights on Chemical Crosslinking Strategies for Proteins. Molecules 2022, 27, 8124, DOI: 10.3390/molecules27238124There is no corresponding record for this reference.
- 2Liu, J.; Yang, B.; Wang, L. Residue selective crosslinking of proteins through photoactivatable or proximity-enabled reactivity. Curr. Opin. Chem. Biol. 2023, 74, 102285 DOI: 10.1016/j.cbpa.2023.102285There is no corresponding record for this reference.
- 3Kotzyba-Hibert, F.; Kapfer, I.; Goeldner, M. Recent trends in photoaffinity labeling. Angew. Chem., Int. Ed. 1995, 34, 1296– 1312, DOI: 10.1002/anie.199512961There is no corresponding record for this reference.
- 4Bayley, H.; Knowles, J. R. Photoaffinity labeling. Methods Enzymol. 1977, 46, 69– 114, DOI: 10.1016/s0076-6879(77)46012-9There is no corresponding record for this reference.
- 5Yu, S.-H.; Wands, A. M.; Kohler, J. J. Photoaffinity probes for studying carbohydrate biology. J. Carbohydr. Chem. 2012, 31, 325– 352, DOI: 10.1080/07328303.2012.676118There is no corresponding record for this reference.
- 6Dorman, G.; Nakamura, H.; Pulsipher, A.; Prestwich, G. D. The life of pi star: exploring the exciting and forbidden worlds of the benzophenone photophore. Chem. Rev. 2016, 116, 15284– 15398, DOI: 10.1021/acs.chemrev.6b00342There is no corresponding record for this reference.
- 7Galardy, R. E.; Craig, L. C.; Jamieson, J. D.; Printz, M. P. Photoaffinity labeling of peptide hormone binding sites. J. Biol. Chem. 1974, 249, 3510– 3518, DOI: 10.1016/S0021-9258(19)42601-XThere is no corresponding record for this reference.
- 8Fleet, G.; Porter, R.; Knowles, J. Affinity labelling of antibodies with aryl nitrene as reactive group. Nature 1969, 224, 511– 512, DOI: 10.1038/224511a0There is no corresponding record for this reference.
- 9Kym, P. R.; Carlson, K. E.; Katzenellenbogen, J. A. Evaluation of a highly efficient aryl azide photoaffinity labeling reagent for the progesterone receptor. Bioconjugate Chem. 1995, 6, 115– 122, DOI: 10.1021/bc00031a014There is no corresponding record for this reference.
- 10Smith, R. A.; Knowles, J. R. Aryldiazirines. Potential reagents for photolabeling of biological receptor sites. J. Am. Chem. Soc. 1973, 95, 5072– 5073, DOI: 10.1021/ja00796a062There is no corresponding record for this reference.
- 11Mishra, P. K.; Yoo, C. M.; Hong, E.; Rhee, H. W. Photo-crosslinking: an emerging chemical tool for investigating molecular networks in live cells. ChemBioChem. 2020, 21, 924– 932, DOI: 10.1002/cbic.201900600There is no corresponding record for this reference.
- 12Leyva, E.; Platz, M. S.; Persy, G.; Wirz, J. Photochemistry of phenyl azide: the role of singlet and triplet phenylnitrene as transient intermediates. J. Am. Chem. Soc. 1986, 108, 3783– 3790, DOI: 10.1021/ja00273a037There is no corresponding record for this reference.
- 13Walrant, A.; Sachon, E. Photoaffinity labeling coupled to MS to identify peptide biological partners: Secondary reactions, for better or for worse?. Mass Spectrom. Rev. 2025, 44, 715– 756, DOI: 10.1002/mas.21880There is no corresponding record for this reference.
- 14Murale, D. P.; Hong, S. C.; Haque, M. M.; Lee, J. S. Photo-affinity labeling (PAL) in chemical proteomics: a handy tool to investigate protein-protein interactions (PPIs). Proteome Sci. 2017, 15, 14, DOI: 10.1186/s12953-017-0123-3There is no corresponding record for this reference.
- 15Liang, T. Y.; Schuster, G. B. Photochemistry of 3- and 4-nitrophenyl azide: detection and characterization of reactive intermediates. J. Am. Chem. Soc. 1987, 109, 7803– 7810, DOI: 10.1021/ja00259a032There is no corresponding record for this reference.
- 16Wentrup, C. Nitrenes, carbenes, diradicals, and ylides. Interconversions of reactive intermediates. Acc. Chem. Res. 2011, 44, 393– 404, DOI: 10.1021/ar700198zThere is no corresponding record for this reference.
- 17Chowdhry, V.; Westheimer, F. Photoaffinity labeling of biological systems. Annu. Rev. Biochem. 1979, 48, 293– 325, DOI: 10.1146/annurev.bi.48.070179.001453There is no corresponding record for this reference.
- 18Wu, H.; Kohler, J. Photocrosslinking probes for capture of carbohydrate interactions. Curr. Opin. Chem. Biol. 2019, 53, 173– 182, DOI: 10.1016/j.cbpa.2019.09.002There is no corresponding record for this reference.
- 19Jahn, O.; Eckart, K.; Tezval, H.; Spiess, J. Characterization of peptide–protein interactions using photoaffinity labeling and LC/MS. Anal. Bioanal. Chem. 2004, 378, 1031– 1036, DOI: 10.1007/s00216-003-2353-8There is no corresponding record for this reference.
- 20Robinette, D.; Neamati, N.; Tomer, K. B.; Borchers, C. H. Photoaffinity labeling combined with mass spectrometric approaches as a tool for structural proteomics. Expert review of proteomics 2006, 3, 399– 408, DOI: 10.1586/14789450.3.4.399There is no corresponding record for this reference.
- 21Tulloch, L. B.; Menzies, S. K.; Fraser, A. L.; Gould, E. R.; King, E. F.; Zacharova, M. K.; Florence, G. J.; Smith, T. K. Photo-affinity labelling and biochemical analyses identify the target of trypanocidal simplified natural product analogues. PLoS Neglected Tropical Diseases 2017, 11, e0005886 DOI: 10.1371/journal.pntd.0005886There is no corresponding record for this reference.
- 22Smith, E.; Collins, I. Photoaffinity labeling in target-and binding-site identification. Future medicinal chemistry 2015, 7, 159– 183, DOI: 10.4155/fmc.14.152There is no corresponding record for this reference.
- 23Park, J.; Koh, M.; Koo, J. Y.; Lee, S.; Park, S. B. Investigation of specific binding proteins to photoaffinity linkers for efficient deconvolution of target protein. ACS Chem. Biol. 2016, 11, 44– 52, DOI: 10.1021/acschembio.5b00671There is no corresponding record for this reference.
- 24Wagner, S.; Hauck, D.; Hoffmann, M.; Sommer, R.; Joachim, I.; Müller, R.; Imberty, A.; Varrot, A.; Titz, A. Covalent Lectin Inhibition and Application in Bacterial Biofilm Imaging. Angew. Chem., Int. Ed. 2017, 56, 16559– 16564, DOI: 10.1002/anie.201709368There is no corresponding record for this reference.
- 25Wang, Y.; Torres-García, D.; Mostert, T. P.; Reinalda, L.; Van Kasteren, S. I. A Bioorthogonal Dual Fluorogenic Probe for the Live-Cell Monitoring of Nutrient Uptake by Mammalian Cells. Angew. Chem., Int. Ed. 2024, 63, e202401733 DOI: 10.1002/anie.202401733There is no corresponding record for this reference.
- 26Yu, A.; He, X.; Shen, T.; Yu, X.; Mao, W.; Chi, W.; Liu, X.; Wu, H. Design strategies for tetrazine fluorogenic probes for bioorthogonal imaging. Chem. Soc. Rev. 2025, 54, 2984, DOI: 10.1039/D3CS00520HThere is no corresponding record for this reference.
- 27Jeong, H.; Wu, X.; Lee, J.-S.; Yoon, J. Recent advances in enzyme-activated NIR fluorescent probes for biological applications. TrAC Trends in Analytical Chemistry 2023, 168, 117335 DOI: 10.1016/j.trac.2023.117335There is no corresponding record for this reference.
- 28Cosco, E. D.; Bogyo, M. Recent advances in ratiometric fluorescence imaging of enzyme activity in vivo. Curr. Opin. Chem. Biol. 2024, 80, 102441 DOI: 10.1016/j.cbpa.2024.102441There is no corresponding record for this reference.
- 29Scott, J. I.; Deng, Q.; Vendrell, M. Near-infrared fluorescent probes for the detection of cancer-associated proteases. ACS Chem. Biol. 2021, 16, 1304– 1317, DOI: 10.1021/acschembio.1c00223There is no corresponding record for this reference.
- 30Saleem, M.; Hanif, M.; Bonne, S.; Zeeshan, M.; Khan, S.; Rafiq, M.; Tahir, T.; Lu, C.; Cai, R. Turn-On Fluorescence Probe for Cancer-Related γ-Glutamyltranspeptidase Detection. Molecules 2024, 29, 4776, DOI: 10.3390/molecules29194776There is no corresponding record for this reference.
- 31Tsao, K. K.; Imai, S.; Chang, M.; Hario, S.; Terai, T.; Campbell, R. E. The best of both worlds: Chemigenetic fluorescent sensors for biological imaging. Cell Chemical Biology 2024, 31, 1652– 1664, DOI: 10.1016/j.chembiol.2024.08.002There is no corresponding record for this reference.
- 32Nadal-Bufi, F.; Salomon, P. L.; de Moliner, F.; Sarris, K. A.; Wang, Z.; Wills, R. D.; Marin, V. L.; Shi, X.; Zhou, K.; Wang, Z.; Xu, Z.; McPherson, M. J.; Marvin, C. C.; Hobson, A. D.; Vendrell, M. Fluorogenic Platform for Real-Time Imaging of Subcellular Payload Release in Antibody–Drug Conjugates. J. Am. Chem. Soc. 2025, 147, 7578– 7587, DOI: 10.1021/jacs.4c16842There is no corresponding record for this reference.
- 33Kellner, S.; Seidu-Larry, S.; Burhenne, J.; Motorin, Y.; Helm, M. A multifunctional bioconjugate module for versatile photoaffinity labeling and click chemistry of RNA. Nucleic Acids Res. 2011, 39, 7348– 7360, DOI: 10.1093/nar/gkr449There is no corresponding record for this reference.
- 34Morimoto, S.; Tomohiro, T.; Maruyama, N.; Hatanaka, Y. Photoaffinity casting of a coumarin flag for rapid identification of ligand-binding sites within protein. Chem. Commun. 2013, 49, 1811– 1813, DOI: 10.1039/c3cc38594aThere is no corresponding record for this reference.
- 35Tomohiro, T.; Yamamoto, A.; Tatsumi, Y.; Hatanaka, Y. [3-(Trifluoromethyl)-3 H-diazirin-3-yl] coumarin as a carbene-generating photocross-linker with masked fluorogenic beacon. Chem. Commun. 2013, 49, 11551– 11553, DOI: 10.1039/c3cc45780jThere is no corresponding record for this reference.
- 36Singha, M.; Roy, S.; Pandey, S. D.; Bag, S. S.; Bhattacharya, P.; Das, M.; Ghosh, A. S.; Ray, D.; Basak, A. Use of azidonaphthalimide carboxylic acids as fluorescent templates with a built-in photoreactive group and a flexible linker simplifies protein labeling studies: applications in selective tagging of HCAII and penicillin binding proteins. Chem. Commun. 2017, 53, 13015– 13018, DOI: 10.1039/C7CC08209FThere is no corresponding record for this reference.
- 37Képiró, M.; Várkuti, B. H.; Rauscher, A. A.; Kellermayer, M. S.; Varga, M.; Málnási-Csizmadia, A. Molecular tattoo: subcellular confinement of drug effects. Chemistry & biology 2015, 22, 548– 558, DOI: 10.1016/j.chembiol.2015.03.013There is no corresponding record for this reference.
- 38Dai, S.-Y.; Yang, D. A visible and near-infrared light activatable diazocoumarin probe for fluorogenic protein labeling in living cells. J. Am. Chem. Soc. 2020, 142, 17156– 17166, DOI: 10.1021/jacs.0c08068There is no corresponding record for this reference.
- 39Bousch, C.; Vreulz, B.; Kansal, K.; El-Husseini, A.; Cecioni, S. Fluorogenic Photo-Crosslinking of Glycan-Binding Protein Recognition Using a Fluorinated Azido-Coumarin Fucoside. Angew. Chem., Int. Ed. 2023, 62, e202314248 DOI: 10.1002/anie.202314248There is no corresponding record for this reference.
- 40Vreulz, B.; De Crozals, D.; Cecioni, S. A trifunctional probe for generation of fluorogenic glycan-photocrosslinker conjugates. RSC chemical biology 2025, 6, 1779– 1786, DOI: 10.1039/D5CB00206KThere is no corresponding record for this reference.
- 41Brown, S. A. Biochemistry of The Coumarins, in Biochemistry of Plant Phenolics, Swain, T.; Harbone, J. B.; Van Sumere, C. F., Eds., Springer US: Boston, MA, 1979; pp 249– 286.There is no corresponding record for this reference.
- 42Musa, M. A.; Cooperwood, J. S.; Khan, M. O. F. A review of coumarin derivatives in pharmacotherapy of breast cancer. Curr. Med. Chem. 2008, 15 (26), 2664– 2679, DOI: 10.2174/092986708786242877There is no corresponding record for this reference.
- 43Stringlis, I. A.; de Jonge, R.; Pieterse, C. M. J. The Age of Coumarins in Plant–Microbe Interactions. Plant Cell Physiol. 2019, 60, 1405– 1419, DOI: 10.1093/pcp/pcz076There is no corresponding record for this reference.
- 44Sun, X.-y.; Liu, T.; Sun, J.; Wang, X.-j. Synthesis and application of coumarin fluorescence probes. RSC Adv. 2020, 10, 10826– 10847, DOI: 10.1039/C9RA10290FThere is no corresponding record for this reference.
- 45Tasior, M.; Kim, D.; Singha, S.; Krzeszewski, M.; Ahn, K. H.; Gryko, D. T. π-Expanded coumarins: synthesis, optical properties and applications. Journal of Materials Chemistry C 2015, 3, 1421– 1446, DOI: 10.1039/C4TC02665AThere is no corresponding record for this reference.
- 46Szwaczko, K. Coumarins synthesis and transformation via C–H bond activation─A review. Inorganics 2022, 10, 23, DOI: 10.3390/inorganics10020023There is no corresponding record for this reference.
- 47Kang, D.; Ahn, K.; Hong, S. Site-Selective C– H Bond Functionalization of Chromones and Coumarins. Asian Journal of Organic Chemistry 2018, 7, 1136– 1150, DOI: 10.1002/ajoc.201800128There is no corresponding record for this reference.
- 48Tod, M.; Prevot, M.; Poulou, M.; Farinotti, R.; Chalom, J.; Mahuzier, G. Chromatographic and luminescence properties of a 7-aminocoumarin derivative with peroxyoxalate chemiexcitation. Anal. Chim. Acta 1989, 223, 309– 317, DOI: 10.1016/S0003-2670(00)84096-9There is no corresponding record for this reference.
- 49Murase, T.; Yoshihara, T.; Yamada, K.; Tobita, S. Fluorescent peptides labeled with environment-sensitive 7-aminocoumarins and their interactions with lipid bilayer membranes and living cells. Bull. Chem. Soc. Jpn. 2013, 86, 510– 519, DOI: 10.1246/bcsj.20120314There is no corresponding record for this reference.
- 50Nowakowska, M.; Smoluch, M.; Sendor, D. The effect of cyclodextrins on the photochemical stability of 7-amino-4-methylcoumarin in aqueous solution. J. Incl. Phenom. Macrocycl. Chem. 2001, 40, 213– 219, DOI: 10.1023/A:1011820513256There is no corresponding record for this reference.
- 51Raju, B. B.; Costa, S. M. Excited-state behavior of 7-diethylaminocoumarin dyes in AOT reversed micelles: size effects. J. Phys. Chem. B 1999, 103, 4309– 4317, DOI: 10.1021/jp984527yThere is no corresponding record for this reference.
- 52Gandioso, A.; Palau, M.; Bresolí-Obach, R.; Galindo, A.; Rovira, A.; Bosch, M.; Nonell, S.; Marchán, V. High photostability in nonconventional coumarins with far-red/NIR emission through azetidinyl substitution. Journal of Organic Chemistry 2018, 83, 11519– 11531, DOI: 10.1021/acs.joc.8b01422There is no corresponding record for this reference.
- 53Rovira, A.; Pujals, M.; Gandioso, A.; López-Corrales, M.; Bosch, M.; Marchán, V. Modulating photostability and mitochondria selectivity in far-red/NIR emitting coumarin fluorophores through replacement of pyridinium by pyrimidinium. Journal of Organic Chemistry 2020, 85, 6086– 6097, DOI: 10.1021/acs.joc.0c00570There is no corresponding record for this reference.
- 54Lord, S. J.; Lee, H.-l. D.; Samuel, R.; Weber, R.; Liu, N.; Conley, N. R.; Thompson, M. A.; Twieg, R. J.; Moerner, W. Azido push– pull fluorogens photoactivate to produce bright fluorescent labels. J. Phys. Chem. B 2010, 114, 14157– 14167, DOI: 10.1021/jp907080rThere is no corresponding record for this reference.
- 55De-La-Cuesta, J.; González, E.; Pomposo, J. A. Advances in fluorescent single-chain nanoparticles. Molecules 2017, 22, 1819, DOI: 10.3390/molecules22111819There is no corresponding record for this reference.
- 56Sivakumar, K.; Xie, F.; Cash, B. M.; Long, S.; Barnhill, H. N.; Wang, Q. A fluorogenic 1, 3-dipolar cycloaddition reaction of 3-azidocoumarins and acetylenes. Org. Lett. 2004, 6, 4603– 4606, DOI: 10.1021/ol047955xThere is no corresponding record for this reference.
- 57Porell, R. N.; Follmar, J. L.; Purcell, S. C.; Timm, B.; Laubach, L. K.; Kozirovskiy, D.; Thacker, B. E.; Glass, C. A.; Gordts, P. L. S. M.; Godula, K. Biologically Derived Neoproteoglycans for Profiling Protein–Glycosaminoglycan Interactions. ACS Chem. Biol. 2022, 17, 1534– 1542, DOI: 10.1021/acschembio.2c00205There is no corresponding record for this reference.
- 58Chalansonnet, V.; Lowe, J.; Orenga, S.; Perry, J. D.; Robinson, S. N.; Stanforth, S. P.; Sykes, H. E.; Truong, T. V. Fluorogenic 7-azidocoumarin and 3/4-azidophthalimide derivatives as indicators of reductase activity in microorganisms. Bioorg. Med. Chem. Lett. 2019, 29, 2354– 2357, DOI: 10.1016/j.bmcl.2019.06.010There is no corresponding record for this reference.
- 59Hill, S. A.; Gerke, C.; Hartmann, L. Recent Developments in Solid-Phase Strategies towards Synthetic, Sequence-Defined Macromolecules. Chem.─Asian J. 2018, 13, 3611– 3622, DOI: 10.1002/asia.201801171There is no corresponding record for this reference.
- 60Blawitzki, L.-C.; Bartels, N.; Bonda, L.; Schmidt, S.; Monzel, C.; Hartmann, L. Glycomacromolecules to Tailor Crowded and Heteromultivalent Glycocalyx Mimetics. Biomacromolecules 2024, 25, 5979– 5994, DOI: 10.1021/acs.biomac.4c00646There is no corresponding record for this reference.
- 61Jäck, N.; Hemming, A.; Hartmann, L. Synthesis of Dual-Responsive Amphiphilic Glycomacromolecules: Controlled Release of Glycan Ligands via pH and UV Stimuli. Macromol. Rapid Commun. 2024, 45, 2400439 DOI: 10.1002/marc.202400439There is no corresponding record for this reference.
- 62Hoffmann, M.; Snyder, N. L.; Hartmann, L. Glycosaminoglycan Mimetic Precision Glycomacromolecules with Sequence-Defined Sulfation and Rigidity Patterns. Biomacromolecules 2022, 23, 4004– 4014, DOI: 10.1021/acs.biomac.2c00829There is no corresponding record for this reference.
- 63Pǎunescu, E.; Louise, L.; Jean, L.; Romieu, A.; Renard, P.-Y. A versatile access to new halogenated 7-azidocoumarins for photoaffinity labeling: Synthesis and photophysical properties. Dyes Pigm. 2011, 91, 427– 434, DOI: 10.1016/j.dyepig.2011.05.008There is no corresponding record for this reference.
- 64Yang, Q.; Váňa, J.; Klán, P. The complex photochemistry of coumarin-3-carboxylic acid in acetonitrile and methanol. Photochemical & Photobiological Sciences 2022, 21, 1481– 1495, DOI: 10.1007/s43630-022-00238-8There is no corresponding record for this reference.
- 65Ponader, D.; Wojcik, F.; Beceren-Braun, F.; Dernedde, J.; Hartmann, L. Sequence-defined glycopolymer segments presenting mannose: synthesis and lectin binding affinity. Biomacromolecules 2012, 13, 1845– 1852, DOI: 10.1021/bm300331zThere is no corresponding record for this reference.
- 66Wojcik, F.; Lel, S.; O’Brien, A. G.; Seeberger, P. H.; Hartmann, L. Synthesis of homo-and heteromultivalent carbohydrate-functionalized oligo (amidoamines) using novel glyco-building blocks. Beilstein journal of organic chemistry 2013, 9, 2395– 2403, DOI: 10.3762/bjoc.9.276There is no corresponding record for this reference.
- 67Ponader, D.; Maffre, P.; Aretz, J.; Pussak, D.; Ninnemann, N. M.; Schmidt, S.; Seeberger, P. H.; Rademacher, C.; Nienhaus, G. U.; Hartmann, L. Carbohydrate-Lectin Recognition of Sequence-Defined Heteromultivalent Glycooligomers. J. Am. Chem. Soc. 2014, 136, 2008, DOI: 10.1021/ja411582tThere is no corresponding record for this reference.
- 68Cummings, R. D. The mannose receptor ligands and the macrophage glycome. Curr. Opin. Struct. Biol. 2022, 75, 102394 DOI: 10.1016/j.sbi.2022.102394There is no corresponding record for this reference.
- 69Coombs, P. J.; Taylor, M. E.; Drickamer, K. Two categories of mammalian galactose-binding receptors distinguished by glycan array profiling. Glycobiology 2006, 16, 1C– 7C, DOI: 10.1093/glycob/cwj126There is no corresponding record for this reference.
- 70Weatherman, R. V.; Mortell, K. H.; Chervenak, M.; Kiessling, L. L.; Toone, E. J. Specificity of C-glycoside complexation by mannose/glucose specific lectins. Biochemistry 1996, 35, 3619– 3624, DOI: 10.1021/bi951916zThere is no corresponding record for this reference.
- 71Itakura, Y.; Nakamura-Tsuruta, S.; Kominami, J.; Sharon, N.; Kasai, K.-i.; Hirabayashi, J. Systematic Comparison of Oligosaccharide Specificity of Ricinus communis Agglutinin I and Erythrina Lectins: a Search by Frontal Affinity Chromatography†. J. Biochem. 2007, 142, 459– 469, DOI: 10.1093/jb/mvm153There is no corresponding record for this reference.
- 72Wang, Y.; Yu, G.; Han, Z.; Yang, B.; Hu, Y.; Zhao, X.; Wu, J.; Lv, Y.; Chai, W. Specificities of Ricinus communis agglutinin 120 interaction with sulfated galactose. FEBS Lett. 2011, 585, 3927– 3934, DOI: 10.1016/j.febslet.2011.10.035There is no corresponding record for this reference.
- 73Bojar, D.; Meche, L.; Meng, G.; Eng, W.; Smith, D. F.; Cummings, R. D.; Mahal, L. K. A Useful Guide to Lectin Binding: Machine-Learning Directed Annotation of 57 Unique Lectin Specificities. ACS Chem. Biol. 2022, 17, 2993– 3012, DOI: 10.1021/acschembio.1c00689There is no corresponding record for this reference.
- 74Maljaars, C. E. P.; Halkes, K. M.; de Oude, W. L.; Haseley, S. R.; Upton, P. J.; McDonnell, M. B.; Kamerling, J. P. Affinity Determination of Ricinus communis Agglutinin Ligands Identified from Combinatorial O- and S-,N-Glycopeptide Libraries. J. Comb. Chem. 2006, 8, 812– 819, DOI: 10.1021/cc060019jThere is no corresponding record for this reference.
- 75Schuster, G. B.; Platz, M. S. Photochemistry of phenyl azide. Advances in photochemistry 1992, 17, 69– 143, DOI: 10.1002/9780470133484.ch2There is no corresponding record for this reference.
- 76Reddy, V. S.; Rao, V. Modes of binding of α (1–2) linked manno-oligosaccharides to concanavalin A. Int. J. Biol. Macromol. 1992, 14, 185– 192, DOI: 10.1016/S0141-8130(05)80025-XThere is no corresponding record for this reference.
- 77Jeyachandran, Y. L.; Mielczarski, J. A.; Mielczarski, E.; Rai, B. Efficiency of blocking of non-specific interaction of different proteins by BSA adsorbed on hydrophobic and hydrophilic surfaces. J. Colloid Interface Sci. 2010, 341, 136– 142, DOI: 10.1016/j.jcis.2009.09.007There is no corresponding record for this reference.
- 78Al-Husseini, J. K.; Stanton, N. J.; Selassie, C. R.; Johal, M. S. The binding of drug molecules to serum albumin: The effect of drug hydrophobicity on binding strength and protein desolvation. Langmuir 2019, 35, 17054– 17060, DOI: 10.1021/acs.langmuir.9b02318There is no corresponding record for this reference.
- 79Smith, E. A.; Thomas, W. D.; Kiessling, L. L.; Corn, R. M. Surface Plasmon Resonance Imaging Studies of Protein-Carbohydrate Interactions. J. Am. Chem. Soc. 2003, 125, 6140– 6148, DOI: 10.1021/ja034165uThere is no corresponding record for this reference.
- 80Parera Pera, N.; Branderhorst, H. M.; Kooij, R.; Maierhofer, C.; van der Kaaden, M.; Liskamp, R. M. J.; Wittmann, V.; Ruijtenbeek, R.; Pieters, R. J. Rapid Screening of Lectins for Multivalency Effects with a Glycodendrimer Microarray. ChemBioChem. 2010, 11, 1896– 1904, DOI: 10.1002/cbic.201000340There is no corresponding record for this reference.
- 81Feldhof, M. I.; Sperzel, S.; Bonda, L.; Boye, S.; Braunschweig, A. B.; Gerling-Driessen, U. I. M.; Hartmann, L. Thiol-selective native grafting from polymerization for the generation of protein–polymer conjugates. Chem. Sci. 2024, 15, 16768– 16777, DOI: 10.1039/D4SC04818KThere is no corresponding record for this reference.
- 82Olmsted, I. R.; Kussrow, A.; Bornhop, D. J. Comparison of Free-Solution and Surface-Immobilized Molecular Interactions Using a Single Platform. Anal. Chem. 2012, 84, 10817– 10822, DOI: 10.1021/ac302933hThere is no corresponding record for this reference.
- 83Ramesh, D.; Srinivasan, M. Studies on ring opening of coumarins. Curr. Sci. 1984, 53, 369– 371There is no corresponding record for this reference.
- 84Mała, P.; Siebs, E.; Meiers, J.; Rox, K.; Varrot, A.; Imberty, A.; Titz, A. Discovery of N-β-l-Fucosyl Amides as High-Affinity Ligands for the Pseudomonas aeruginosa Lectin LecB. J. Med. Chem. 2022, 65, 14180– 14200, DOI: 10.1021/acs.jmedchem.2c01373There is no corresponding record for this reference.
- 85Hayes, W.; Osborn, H. M.; Osborne, S. D.; Rastall, R. A.; Romagnoli, B. One-pot synthesis of multivalent arrays of mannose mono-and disaccharides. Tetrahedron 2003, 59, 7983– 7996, DOI: 10.1016/j.tet.2003.08.011There is no corresponding record for this reference.
- 86Wu, L.; Sampson, N. S. Fucose, Mannose, and β-N-Acetylglucosamine Glycopolymers Initiate the Mouse Sperm Acrosome Reaction through Convergent Signaling Pathways. ACS Chem. Biol. 2014, 9, 468– 475, DOI: 10.1021/cb400550jThere is no corresponding record for this reference.
Supporting Information
Supporting Information
The Supporting Information is available free of charge at https://pubs.acs.org/doi/10.1021/acs.bioconjchem.5c00613.
Analytical data for the synthesized structures, additional data and figures showing the absorption and fluorescence spectra of AzC building blocks, full MALDI-TOF-MS and full fluorescence spectra, additional fluorescence spectra for the pH-stability study, additional SDS–PAGE images of irradiated and not irradiated lysates, and additional fluorescence microscopy images (PDF)
Terms & Conditions
Most electronic Supporting Information files are available without a subscription to ACS Web Editions. Such files may be downloaded by article for research use (if there is a public use license linked to the relevant article, that license may permit other uses). Permission may be obtained from ACS for other uses through requests via the RightsLink permission system: http://pubs.acs.org/page/copyright/permissions.html.



