Lipid Nanoparticles with Side-Chain Polymer Coating for Targeted mRNA Delivery through Nanobody AttachmentClick to copy article linkArticle link copied!
- On Ting ChoyOn Ting ChoyAustralian Institute for Bioengineering and Nanotechnology, The University of Queensland, Brisbane, Queensland 4072, AustraliaMore by On Ting Choy
- Nicholas L. FletcherNicholas L. FletcherAustralian Institute for Bioengineering and Nanotechnology, The University of Queensland, Brisbane, Queensland 4072, AustraliaARC Research Hub for Advanced Manufacture of Targeted Radiopharmaceuticals, The University of Queensland, Brisbane, Queensland 4072, AustraliaCentre for Advanced Imaging, The University of Queensland, Brisbane, Queensland 4072, AustraliaMore by Nicholas L. Fletcher
- Pie HudaPie HudaAustralian Institute for Bioengineering and Nanotechnology, The University of Queensland, Brisbane, Queensland 4072, AustraliaARC Research Hub for Advanced Manufacture of Targeted Radiopharmaceuticals, The University of Queensland, Brisbane, Queensland 4072, AustraliaCentre for Advanced Imaging, The University of Queensland, Brisbane, Queensland 4072, AustraliaMore by Pie Huda
- Craig A. BellCraig A. BellAustralian Institute for Bioengineering and Nanotechnology, The University of Queensland, Brisbane, Queensland 4072, AustraliaCentre for Advanced Imaging, The University of Queensland, Brisbane, Queensland 4072, AustraliaMore by Craig A. Bell
- David J. OwenDavid J. OwenAustralian Institute for Bioengineering and Nanotechnology, The University of Queensland, Brisbane, Queensland 4072, AustraliaARC Research Hub for Advanced Manufacture of Targeted Radiopharmaceuticals, The University of Queensland, Brisbane, Queensland 4072, AustraliaMore by David J. Owen
- Andrew K. Whittaker*Andrew K. Whittaker*Email: [email protected]Australian Institute for Bioengineering and Nanotechnology, The University of Queensland, Brisbane, Queensland 4072, AustraliaAustralian Research Council Centre of Excellence for Green Electrochemical Transformation of Carbon Dioxide, The University of Queensland, Brisbane, Queensland 4072, AustraliaMore by Andrew K. Whittaker
Abstract
The use of mRNA therapies has innovated the clinical progress of cancer immunotherapy. However, current immunotherapeutic approaches are unable to achieve site- or immune-cell-specific delivery, resulting in adverse immune responses in off-target tissues. In addition, the commercial lipid nanoparticle (LNP) formulations with a poly(ethylene glycol) coating generally undergo significant hepatic accumulation during clearance. To promote site- and immune-cell-specific delivery of therapeutic mRNA-LNPs, we investigated several bioconjugation approaches to attach targeting antibodies onto the surface of polymer-functionalized mRNA-LNPs. Building on our previous work, side-chain sulfoxide polymer–lipid conjugate PMSEA-DSPE was used to incorporate a low-fouling polymeric LNP coating. trans-Cyclooctene functionality was incorporated within PMSEA-DSPE end groups to allow conjugation to the tetrazine-functionalized nanobody 9G8 for EGFR targeting. Bioconjugation methods were compared, including direct conjugation and post-insertion. The results showed that 9G8-attached PMSEA mRNA-LNPs prepared via direct conjugation significantly enhanced cell association and in vitro transfection efficiency with an EGFR-positive cell line, demonstrating the potency of active targeting for mRNA-LNP platforms with side-chain polymer coatings.
This publication is licensed for personal use by The American Chemical Society.
Introduction
Scheme 1
Results and Discussion
Preparation and Characterization of tCO-Functionalized PMSEA-Lipid Conjugates
Figure 1
Figure 1. (a) UV–vis spectra of PMSEA-DSPE and COOH-PMSEA-DSPE; the trithiocarbonyl peak at 300 – 310 nm was evident in PMSEA-DSPE. (b) 1H NMR (500 MHz, CDCl3) of COOH-PMSEA-DSPE with the peak comparison of PMSEA-DSPE. The pink arrows indicate the presence of the methyl group, while the orange arrow indicates the proton in carboxyl groups.
Figure 2
Figure 2. 1H NMR 2D DOSY spectrum (500 MHz, CDCl3) of tCO-PMSEA-DSPE, showing successful tCO attachment to the PMSEA-lipid conjugate and the absence of free tCO residues.
Tz-9G8 Nanobody Attachment of PMSEA mRNA-LNPs Via Different Preparation Methods
Figure 3
Figure 3. (a) Demonstration of two preparation methods to conjugate the Tz-9G8 nanobody to PMSEA mRNA-LNPs, including direct conjugation and post-insertion. (b) Gel electrophoresis of 9G8 and LNP samples prepared by either direct conjugation or post-insertion; (i) gel image with Coomassie Blue staining and (ii) gel analysis by ImageJ. (c) Z-average (d.nm) and PDI measured by DLS, statistical data is presented as the mean ± SD (n = 3).
Figure 4
Figure 4. In vitro characterization of 9G8-PMSEA mRNA-LNPs prepared with different conjugation methods. (a) Cellular association of MDA-MB-468 incubated with PMSEA FLuc mRNA-LNPs with or without 9G8 attachment for 1 h and (b) luciferase expression of MDA-MB-468 incubated with PMSEA FLuc mRNA-LNPs with or without 9G8 attachment for 24 h. Statistical data is presented as the mean ± SD (n = 3 for flow cytometry; n = 4 for luciferase assay), ***p < 0.001, **p < 0.01, *p < 0.05, and ns p > 0.05 with 95% of confidence level from Welch’s unpaired t test.
Effect of 9G8 Nanobody Density on PMSEA mRNA-LNPs
Figure 5
Figure 5. Physical and in vitro characterization of 9G8-PMSEA mRNA-LNPs with various molar ratios of tCO-PMSEA-DSPE (0 – 1 mol %) prepared by the direct conjugation method. (a) Gel image of 9G8-PMSEA mRNA-LNPs stained with Coomassie Blue. (b) Z-average and PDI of PMSEA mRNA-LNPs before and after conjugation, measured by DLS. (c) Flow cytometry analysis of 9G8 attachment on mRNA-LNPs via titration with various concentrations of AF488-labeled EGFR; (i) mean fluorescence intensity (MFI) of AF488 and (ii) half-maximum binding at equilibrium (Kd) calculated with the Saturation Binding model based on the titration results (R2 = 0.9459). (d) Cellular association of MDA-MB-468 incubated with PMSEA FLuc mRNA-LNPs with or without 9G8 for 1 h; and (e) luciferase expression of MDA-MB-468 incubated with PMSEA FLuc mRNA-LNPs with or without 9G8 for 24 h. Statistical data is presented as the mean ± SD (n = 3 for DLS and cell association assay; n = 4 for luciferase assay), ***p < 0.001, **p < 0.01, *p < 0.05, and ns p > 0.05 with 95% of confidence level from one-way ANOVA.
In Vitro Transfection Efficiency of PMSEA mRNA-LNPs Via Targeting
Figure 6
Figure 6. eGFP expression of MDA-MB-468 incubated with PMSEA eGFP mRNA-LNPs with or without 9G8 attachment for 24 h. (a) Mean fluorescence intensity (MFI) of eGFP quantified by flow cytometry. Statistical data is presented as the mean ± SD (n = 3), ***p < 0.001, **p < 0.01, *p < 0.05, and ns p > 0.05 with 95% of confidence level from Welch’s unpaired t test. (b) Confocal images of eGFP transfection and cellular association with DiD-labeled eGFP mRNA-LNPs for 24 h with Hoechst staining MDA-MB-468 nuclei; scale bars are 20 μm).
Conclusions
Supporting Information
The Supporting Information is available free of charge at https://pubs.acs.org/doi/10.1021/acs.bioconjchem.6c00045.
Experiment procedure (materials and methods), 1H NMR spectra of COOH-PMSEA-DSPE and tCO-PMSEA-DSPE, flow cytometry data of mRNA-LNPs, physical characterization of mRNA-LNPs (PDF)
Terms & Conditions
Most electronic Supporting Information files are available without a subscription to ACS Web Editions. Such files may be downloaded by article for research use (if there is a public use license linked to the relevant article, that license may permit other uses). Permission may be obtained from ACS for other uses through requests via the RightsLink permission system: http://pubs.acs.org/page/copyright/permissions.html.
Acknowledgments
The authors acknowledge the financial support from the Australian Research Council Discovery grants (DP210101496) and the Advance Queensland Industry Research Fellowship Scheme (AQIRF094-2023RD6; NLF). This work used the Queensland node of the NCRIS-enabled Australian National Fabrication Facility (ANFF). The Australian National Fabrication Facility is acknowledged for access to some items of equipment. Protein Expression Facility (PEF) and BASE facilities, The University of Queensland were acknowledged for the scientific and technical assistance. BASE is supported by Therapeutic Innovation Australia (TIA). TIA is supported by the Australian Government through the National Collaborative Research Infrastructure Strategy (NCRIS) program. The authors also acknowledge Biorender.com, which was used to create Scheme 1, Figure 3 and Figure 5 in this article.
References
This article references 27 other publications.
- 1Pardi, N.; Hogan, M. J.; Porter, F. W.; Weissman, D. mRNA vaccines ─ a new era in vaccinology. Nat. Rev. Drug Discovery 2018, 17 (4), 261– 279, DOI: 10.1038/nrd.2017.243Google ScholarThere is no corresponding record for this reference.
- 2Barbier, A. J.; Jiang, A. Y.; Zhang, P.; Wooster, R.; Anderson, D. G. The clinical progress of mRNA vaccines and immunotherapies. Nat. Biotechnol. 2022, 40 (6), 840– 854, DOI: 10.1038/s41587-022-01294-2Google ScholarThere is no corresponding record for this reference.
- 3Shi, Y.; Shi, M.; Wang, Y.; You, J. Progress and prospects of mRNA-based drugs in pre-clinical and clinical applications. Signal Transduction Targeted Ther. 2024, 9 (1), 322, DOI: 10.1038/s41392-024-02002-zGoogle ScholarThere is no corresponding record for this reference.
- 4Burris, H. A.; Patel, M. R.; Cho, D. C.; Clarke, J. M.; Gutierrez, M.; Zaks, T. Z.; Frederick, J.; Hopson, K.; Mody, K.; Binanti-Berube, A.; Robert-Tissot, C.; Goldstein, B.; Breton, B.; Sun, J.; Zhong, S.; Pruitt, S. K.; Keating, K.; Meehan, R. S.; Gainor, J. F. A phase I multicenter study to assess the safety, tolerability, and immunogenicity of mRNA-4157 alone in patients with resected solid tumors and in combination with pembrolizumab in patients with unresectable solid tumors. J. Clin. Oncol. 2019, 37 (15sup), 2523– 2523, DOI: 10.1200/JCO.2019.37.15_suppl.2523Google ScholarThere is no corresponding record for this reference.
- 5Weber, J. S.; Carlino, M. S.; Khattak, A.; Meniawy, T.; Ansstas, G.; Taylor, M. H.; Kim, K. B.; McKean, M.; Long, G. V.; Sullivan, R. J.; Faries, M.; Tran, T. T.; Cowey, C. L.; Pecora, A.; Shaheen, M.; Segar, J.; Medina, T.; Atkinson, V.; Gibney, G. T.; Luke, J. J.; Thomas, S.; Buchbinder, E. I.; Healy, J. A.; Huang, M.; Morrissey, M.; Feldman, I.; Sehgal, V.; Robert-Tissot, C.; Hou, P.; Zhu, L.; Brown, M.; Aanur, P.; Meehan, R. S.; Zaks, T. Individualised neoantigen therapy mRNA-4157 (V940) plus pembrolizumab versus pembrolizumab monotherapy in resected melanoma (KEYNOTE-942): a randomised, phase 2b study. Lancet 2024, 403 (10427), 632– 644, DOI: 10.1016/S0140-6736(23)02268-7Google ScholarThere is no corresponding record for this reference.
- 6Weber, J. S.; Luke, J. J.; Carlino, M. S.; Khattak, M. A.; Meehan, R. S.; Brown, M.; Zhang, J.; Krepler, C.; Duic, J. P.; Long, G. V. INTerpath-001: Pembrolizumab with V940 (mRNA-4157) versus pembrolizumab with placebo for adjuvant treatment of high-risk stage II-IV melanoma. J. Clin. Oncol. 2024, 42 (16sup), TPS9616– TPS9616, DOI: 10.1200/JCO.2024.42.16_suppl.TPS9616Google ScholarThere is no corresponding record for this reference.
- 7Hou, X.; Zaks, T.; Langer, R.; Dong, Y. Lipid nanoparticles for mRNA delivery. Nature Reviews Materials 2021, 6 (12), 1078– 1094, DOI: 10.1038/s41578-021-00358-0Google ScholarThere is no corresponding record for this reference.
- 8Akinc, A.; Querbes, W.; De, S.; Qin, J.; Frank-Kamenetsky, M.; Jayaprakash, K. N.; Jayaraman, M.; Rajeev, K. G.; Cantley, W. L.; Dorkin, J. R.; Butler, J. S.; Qin, L.; Racie, T.; Sprague, A.; Fava, E.; Zeigerer, A.; Hope, M. J.; Zerial, M.; Sah, D. W. Y.; Fitzgerald, K.; Tracy, M. A.; Manoharan, M.; Koteliansky, V.; Fougerolles, A. d.; Maier, M. A. Targeted Delivery of RNAi Therapeutics With Endogenous and Exogenous Ligand-Based Mechanisms. Mol. Ther. 2010, 18 (7), 1357– 1364, DOI: 10.1038/mt.2010.85Google ScholarThere is no corresponding record for this reference.
- 9Pardi, N.; Tuyishime, S.; Muramatsu, H.; Kariko, K.; Mui, B. L.; Tam, Y. K.; Madden, T. D.; Hope, M. J.; Weissman, D. Expression kinetics of nucleoside-modified mRNA delivered in lipid nanoparticles to mice by various routes. J. Controlled Release 2015, 217, 345– 351, DOI: 10.1016/j.jconrel.2015.08.007Google ScholarThere is no corresponding record for this reference.
- 10Hay, K. A.; Hanafi, L. A.; Li, D.; Gust, J.; Liles, W. C.; Wurfel, M. M.; López, J. A.; Chen, J.; Chung, D.; Harju-Baker, S.; Cherian, S.; Chen, X.; Riddell, S. R.; Maloney, D. G.; Turtle, C. J. Kinetics and biomarkers of severe cytokine release syndrome after CD19 chimeric antigen receptor-modified T-cell therapy. Blood 2017, 130 (21), 2295– 2306, DOI: 10.1182/blood-2017-06-793141Google ScholarThere is no corresponding record for this reference.
- 11De Martin, E.; Michot, J.-M.; Papouin, B.; Champiat, S.; Mateus, C.; Lambotte, O.; Roche, B.; Antonini, T. M.; Coilly, A.; Laghouati, S.; Robert, C.; Marabelle, A.; Guettier, C.; Samuel, D. Characterization of liver injury induced by cancer immunotherapy using immune checkpoint inhibitors. Journal of Hepatology 2018, 68 (6), 1181– 1190, DOI: 10.1016/j.jhep.2018.01.033Google ScholarThere is no corresponding record for this reference.
- 12Kularatne, R. N.; Crist, R. M.; Stern, S. T. The Future of Tissue-Targeted Lipid Nanoparticle-Mediated Nucleic Acid Delivery. Pharmaceuticals (Basel) 2022, 15 (7), 897, DOI: 10.3390/ph15070897Google ScholarThere is no corresponding record for this reference.
- 13Chen, B.-M.; Cheng, T.-L.; Roffler, S. R. Polyethylene Glycol Immunogenicity: Theoretical, Clinical, and Practical Aspects of Anti-Polyethylene Glycol Antibodies. ACS Nano 2021, 15 (9), 14022– 14048, DOI: 10.1021/acsnano.1c05922Google ScholarThere is no corresponding record for this reference.
- 14Xiao, Y.; Lian, X.; Sun, Y.; Sung, Y.-C.; Vaidya, A.; Chen, Z.; Gupta, A.; Chatterjee, S.; Zheng, L.; Guerrero, E.; Wang, X.; Farbiak, L.; Yang, Y.; Diamond, M. I.; Leal, C.; McDonald, J. G.; Siegwart, D. J. High-density brush-shaped polymer lipids reduce anti-PEG antibody binding for repeated administration of mRNA therapeutics. Nat. Mater. 2025, 24, 1840, DOI: 10.1038/s41563-024-02116-3Google ScholarThere is no corresponding record for this reference.
- 15Hassanel, D. N. B. P.; Pilkington, E. H.; Ju, Y.; Kent, S. J.; Pouton, C. W.; Truong, N. P. Replacing poly(ethylene glycol) with RAFT lipopolymers in mRNA lipid nanoparticle systems for effective gene delivery. Int. J. Pharm. 2024, 665, 124695 DOI: 10.1016/j.ijpharm.2024.124695Google ScholarThere is no corresponding record for this reference.
- 16He, X.; Payne, T. J.; Takanashi, A.; Fang, Y.; Kerai, S. D.; Morrow, J. P.; Al-Wassiti, H.; Pouton, C. W.; Kempe, K. Tailored Monoacyl Poly(2-oxazoline)- and Poly(2-oxazine)-Lipids as PEG-Lipid Alternatives for Stabilization and Delivery of mRNA-Lipid Nanoparticles. Biomacromolecules 2024, 25 (7), 4591– 4603, DOI: 10.1021/acs.biomac.4c00651Google ScholarThere is no corresponding record for this reference.
- 17Choy, O. T.; Fletcher, N. L.; Fu, C.; Vu, M. N.; Ediriweera, G. R.; Humphries, J.; Hu, M.; Kent, S. J.; Cai, R.; Chen, C.; Whittaker, A. K. PEG-free Lipid Nanoparticles for mRNA Delivery: Superhydrophilic Sulfoxide Polymer Coatings. Biomacromolecules 2026, 27, 624, DOI: 10.1021/acs.biomac.5c01899Google ScholarThere is no corresponding record for this reference.
- 18Ramishetti, S.; Kedmi, R.; Goldsmith, M.; Leonard, F.; Sprague, A. G.; Godin, B.; Gozin, M.; Cullis, P. R.; Dykxhoorn, D. M.; Peer, D. Systemic Gene Silencing in Primary T Lymphocytes Using Targeted Lipid Nanoparticles. ACS Nano 2015, 9 (7), 6706– 6716, DOI: 10.1021/acsnano.5b02796Google ScholarThere is no corresponding record for this reference.
- 19Tombácz, I.; Laczkó, D.; Shahnawaz, H.; Muramatsu, H.; Natesan, A.; Yadegari, A.; Papp, T. E.; Alameh, M.-G.; Shuvaev, V.; Mui, B. L.; Tam, Y. K.; Muzykantov, V.; Pardi, N.; Weissman, D.; Parhiz, H. Highly efficient CD4+ T cell targeting and genetic recombination using engineered CD4+ cell-homing mRNA-LNPs. Mol. Ther. 2021, 29 (11), 3293– 3304, DOI: 10.1016/j.ymthe.2021.06.004Google ScholarThere is no corresponding record for this reference.
- 20Chen, M. Z.; Yuen, D.; McLeod, V. M.; Yong, K. W.; Smyth, C. H.; Herling, B. R.; Payne, T. J.; Fabb, S. A.; Belousoff, M. J.; Algarni, A.; Sexton, P. M.; Porter, C. J. H.; Pouton, C. W.; Johnston, A. P. R. A versatile antibody capture system drives specific in vivo delivery of mRNA-loaded lipid nanoparticles. Nat. Nanotechnol. 2025, 20, 1273, DOI: 10.1038/s41565-025-01954-9Google ScholarThere is no corresponding record for this reference.
- 21Dietmair, B.; Humphries, J.; Mercer, T. R.; Thurecht, K. J.; Howard, C. B.; Cheetham, S. W. Targeted mRNA delivery with bispecific antibodies that tether LNPs to cell surface markers. Mol. Ther Nucleic Acids 2025, 36 (2), 102520 DOI: 10.1016/j.omtn.2025.102520Google ScholarThere is no corresponding record for this reference.
- 22Huda, P.; Humphries, J.; Fletcher, N. L.; Howard, C. B.; Thurecht, K. J.; Bell, C. A. Click-on Antibody Fragments for Customisable Targeted Nanomedicines – Site-specific Tetrazine and Azide Functionalisation through Non-canonical Amino Acid incorporation. Chemistry Methods 2024, 4 (2), e202300036 DOI: 10.1002/cmtd.202300036Google ScholarThere is no corresponding record for this reference.
- 23Choy, O. T.; Fletcher, N. L.; Fu, C.; Vu, M. N.; Ediriweera, G. R.; Humphries, J.; Hu, M.; Kent, S. J.; Cai, R.; Chen, C.; Whittaker, A. K. PEG-free Lipid Nanoparticles for mRNA Delivery: Superhydrophilic Sulfoxide Polymer Coatings. Biomacromolecules 2026, 27 (1), 624– 639, DOI: 10.1021/acs.biomac.5c01899Google ScholarThere is no corresponding record for this reference.
- 24Chong, Y. K.; Moad, G.; Rizzardo, E.; Thang, S. H. Thiocarbonylthio End Group Removal from RAFT-Synthesized Polymers by Radical-Induced Reduction. Macromolecules 2007, 40 (13), 4446– 4455, DOI: 10.1021/ma062919uGoogle ScholarThere is no corresponding record for this reference.
- 25Li, Q.; Chan, C.; Peterson, N.; Hanna, R. N.; Alfaro, A.; Allen, K. L.; Wu, H.; Dall’Acqua, W. F.; Borrok, M. J.; Santos, J. L. Engineering Caveolae-Targeted Lipid Nanoparticles To Deliver mRNA to the Lungs. ACS Chem. Biol. 2020, 15 (4), 830– 836, DOI: 10.1021/acschembio.0c00003Google ScholarThere is no corresponding record for this reference.
- 26Kedmi, R.; Veiga, N.; Ramishetti, S.; Goldsmith, M.; Rosenblum, D.; Dammes, N.; Hazan-Halevy, I.; Nahary, L.; Leviatan-Ben-Arye, S.; Harlev, M.; Behlke, M.; Benhar, I.; Lieberman, J.; Peer, D. A modular platform for targeted RNAi therapeutics. Nat. Nanotechnol. 2018, 13 (3), 214– 219, DOI: 10.1038/s41565-017-0043-5Google ScholarThere is no corresponding record for this reference.
- 27Escudé Martinez de Castilla, P.; Verdi, V.; de Voogt, W.; Estapé Sentí, M.; Koekman, A. C.; Rietveld, J.; van Kempen, S.; Yang, Q.; van Merris, J.; Jenster, G.; van Royen, M. E.; Fens, M. H.; Kooijmans, S. A. A.; van Weerden, W. M.; van Niel, G.; Vader, P.; Schiffelers, R. M. Nanobody-Decorated Lipid Nanoparticles for Enhanced mRNA Delivery to Tumors In Vivo. Advanced Healthcare Materials 2025, 14 (24), 2500605 DOI: 10.1002/adhm.202500605Google ScholarThere is no corresponding record for this reference.
Cited By
This article has not yet been cited by other publications.
Article Views
Altmetric
Citations
Article Views are the COUNTER-compliant sum of full text article downloads since November 2008 (both PDF and HTML) across all institutions and individuals. These metrics are regularly updated to reflect usage leading up to the last few days.
Citations are the number of other articles citing this article, calculated by Crossref and updated daily. Find more information about Crossref citation counts.
The Altmetric Attention Score is a quantitative measure of the attention that a research article has received online. Clicking on the donut icon will load a page at altmetric.com with additional details about the score and the social media presence for the given article. Find more information on the Altmetric Attention Score and how the score is calculated.
Recommended Articles
Abstract
Scheme 1
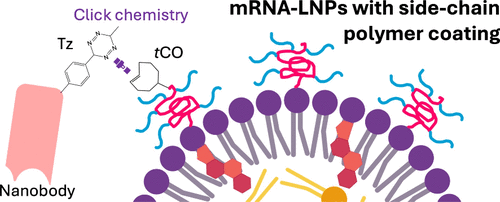
 Scheme 1. Schematic Diagram of Nanobody-LNP Conjugation Via trans-Cyclooctene (tCO)-Functionalization of PMSEA-Lipid Conjugate and Attachment of Tz-9G8 Nanobody to the tCO Group on PMSEA mRNA-LNPs
Scheme 1. Schematic Diagram of Nanobody-LNP Conjugation Via trans-Cyclooctene (tCO)-Functionalization of PMSEA-Lipid Conjugate and Attachment of Tz-9G8 Nanobody to the tCO Group on PMSEA mRNA-LNPsFigure 1

Figure 1. (a) UV–vis spectra of PMSEA-DSPE and COOH-PMSEA-DSPE; the trithiocarbonyl peak at 300 – 310 nm was evident in PMSEA-DSPE. (b) 1H NMR (500 MHz, CDCl3) of COOH-PMSEA-DSPE with the peak comparison of PMSEA-DSPE. The pink arrows indicate the presence of the methyl group, while the orange arrow indicates the proton in carboxyl groups.
Figure 2

Figure 2. 1H NMR 2D DOSY spectrum (500 MHz, CDCl3) of tCO-PMSEA-DSPE, showing successful tCO attachment to the PMSEA-lipid conjugate and the absence of free tCO residues.
Figure 3

Figure 3. (a) Demonstration of two preparation methods to conjugate the Tz-9G8 nanobody to PMSEA mRNA-LNPs, including direct conjugation and post-insertion. (b) Gel electrophoresis of 9G8 and LNP samples prepared by either direct conjugation or post-insertion; (i) gel image with Coomassie Blue staining and (ii) gel analysis by ImageJ. (c) Z-average (d.nm) and PDI measured by DLS, statistical data is presented as the mean ± SD (n = 3).
Figure 4
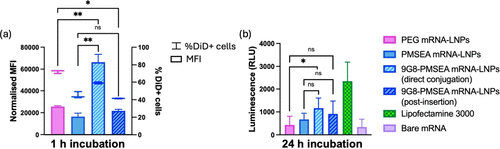
Figure 4. In vitro characterization of 9G8-PMSEA mRNA-LNPs prepared with different conjugation methods. (a) Cellular association of MDA-MB-468 incubated with PMSEA FLuc mRNA-LNPs with or without 9G8 attachment for 1 h and (b) luciferase expression of MDA-MB-468 incubated with PMSEA FLuc mRNA-LNPs with or without 9G8 attachment for 24 h. Statistical data is presented as the mean ± SD (n = 3 for flow cytometry; n = 4 for luciferase assay), ***p < 0.001, **p < 0.01, *p < 0.05, and ns p > 0.05 with 95% of confidence level from Welch’s unpaired t test.
Figure 5
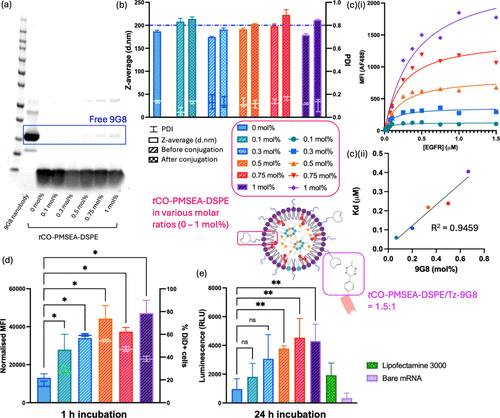
Figure 5. Physical and in vitro characterization of 9G8-PMSEA mRNA-LNPs with various molar ratios of tCO-PMSEA-DSPE (0 – 1 mol %) prepared by the direct conjugation method. (a) Gel image of 9G8-PMSEA mRNA-LNPs stained with Coomassie Blue. (b) Z-average and PDI of PMSEA mRNA-LNPs before and after conjugation, measured by DLS. (c) Flow cytometry analysis of 9G8 attachment on mRNA-LNPs via titration with various concentrations of AF488-labeled EGFR; (i) mean fluorescence intensity (MFI) of AF488 and (ii) half-maximum binding at equilibrium (Kd) calculated with the Saturation Binding model based on the titration results (R2 = 0.9459). (d) Cellular association of MDA-MB-468 incubated with PMSEA FLuc mRNA-LNPs with or without 9G8 for 1 h; and (e) luciferase expression of MDA-MB-468 incubated with PMSEA FLuc mRNA-LNPs with or without 9G8 for 24 h. Statistical data is presented as the mean ± SD (n = 3 for DLS and cell association assay; n = 4 for luciferase assay), ***p < 0.001, **p < 0.01, *p < 0.05, and ns p > 0.05 with 95% of confidence level from one-way ANOVA.
Figure 6
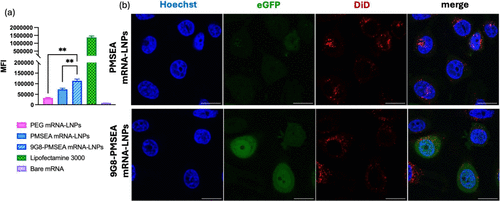
Figure 6. eGFP expression of MDA-MB-468 incubated with PMSEA eGFP mRNA-LNPs with or without 9G8 attachment for 24 h. (a) Mean fluorescence intensity (MFI) of eGFP quantified by flow cytometry. Statistical data is presented as the mean ± SD (n = 3), ***p < 0.001, **p < 0.01, *p < 0.05, and ns p > 0.05 with 95% of confidence level from Welch’s unpaired t test. (b) Confocal images of eGFP transfection and cellular association with DiD-labeled eGFP mRNA-LNPs for 24 h with Hoechst staining MDA-MB-468 nuclei; scale bars are 20 μm).
References
This article references 27 other publications.
- 1Pardi, N.; Hogan, M. J.; Porter, F. W.; Weissman, D. mRNA vaccines ─ a new era in vaccinology. Nat. Rev. Drug Discovery 2018, 17 (4), 261– 279, DOI: 10.1038/nrd.2017.243There is no corresponding record for this reference.
- 2Barbier, A. J.; Jiang, A. Y.; Zhang, P.; Wooster, R.; Anderson, D. G. The clinical progress of mRNA vaccines and immunotherapies. Nat. Biotechnol. 2022, 40 (6), 840– 854, DOI: 10.1038/s41587-022-01294-2There is no corresponding record for this reference.
- 3Shi, Y.; Shi, M.; Wang, Y.; You, J. Progress and prospects of mRNA-based drugs in pre-clinical and clinical applications. Signal Transduction Targeted Ther. 2024, 9 (1), 322, DOI: 10.1038/s41392-024-02002-zThere is no corresponding record for this reference.
- 4Burris, H. A.; Patel, M. R.; Cho, D. C.; Clarke, J. M.; Gutierrez, M.; Zaks, T. Z.; Frederick, J.; Hopson, K.; Mody, K.; Binanti-Berube, A.; Robert-Tissot, C.; Goldstein, B.; Breton, B.; Sun, J.; Zhong, S.; Pruitt, S. K.; Keating, K.; Meehan, R. S.; Gainor, J. F. A phase I multicenter study to assess the safety, tolerability, and immunogenicity of mRNA-4157 alone in patients with resected solid tumors and in combination with pembrolizumab in patients with unresectable solid tumors. J. Clin. Oncol. 2019, 37 (15sup), 2523– 2523, DOI: 10.1200/JCO.2019.37.15_suppl.2523There is no corresponding record for this reference.
- 5Weber, J. S.; Carlino, M. S.; Khattak, A.; Meniawy, T.; Ansstas, G.; Taylor, M. H.; Kim, K. B.; McKean, M.; Long, G. V.; Sullivan, R. J.; Faries, M.; Tran, T. T.; Cowey, C. L.; Pecora, A.; Shaheen, M.; Segar, J.; Medina, T.; Atkinson, V.; Gibney, G. T.; Luke, J. J.; Thomas, S.; Buchbinder, E. I.; Healy, J. A.; Huang, M.; Morrissey, M.; Feldman, I.; Sehgal, V.; Robert-Tissot, C.; Hou, P.; Zhu, L.; Brown, M.; Aanur, P.; Meehan, R. S.; Zaks, T. Individualised neoantigen therapy mRNA-4157 (V940) plus pembrolizumab versus pembrolizumab monotherapy in resected melanoma (KEYNOTE-942): a randomised, phase 2b study. Lancet 2024, 403 (10427), 632– 644, DOI: 10.1016/S0140-6736(23)02268-7There is no corresponding record for this reference.
- 6Weber, J. S.; Luke, J. J.; Carlino, M. S.; Khattak, M. A.; Meehan, R. S.; Brown, M.; Zhang, J.; Krepler, C.; Duic, J. P.; Long, G. V. INTerpath-001: Pembrolizumab with V940 (mRNA-4157) versus pembrolizumab with placebo for adjuvant treatment of high-risk stage II-IV melanoma. J. Clin. Oncol. 2024, 42 (16sup), TPS9616– TPS9616, DOI: 10.1200/JCO.2024.42.16_suppl.TPS9616There is no corresponding record for this reference.
- 7Hou, X.; Zaks, T.; Langer, R.; Dong, Y. Lipid nanoparticles for mRNA delivery. Nature Reviews Materials 2021, 6 (12), 1078– 1094, DOI: 10.1038/s41578-021-00358-0There is no corresponding record for this reference.
- 8Akinc, A.; Querbes, W.; De, S.; Qin, J.; Frank-Kamenetsky, M.; Jayaprakash, K. N.; Jayaraman, M.; Rajeev, K. G.; Cantley, W. L.; Dorkin, J. R.; Butler, J. S.; Qin, L.; Racie, T.; Sprague, A.; Fava, E.; Zeigerer, A.; Hope, M. J.; Zerial, M.; Sah, D. W. Y.; Fitzgerald, K.; Tracy, M. A.; Manoharan, M.; Koteliansky, V.; Fougerolles, A. d.; Maier, M. A. Targeted Delivery of RNAi Therapeutics With Endogenous and Exogenous Ligand-Based Mechanisms. Mol. Ther. 2010, 18 (7), 1357– 1364, DOI: 10.1038/mt.2010.85There is no corresponding record for this reference.
- 9Pardi, N.; Tuyishime, S.; Muramatsu, H.; Kariko, K.; Mui, B. L.; Tam, Y. K.; Madden, T. D.; Hope, M. J.; Weissman, D. Expression kinetics of nucleoside-modified mRNA delivered in lipid nanoparticles to mice by various routes. J. Controlled Release 2015, 217, 345– 351, DOI: 10.1016/j.jconrel.2015.08.007There is no corresponding record for this reference.
- 10Hay, K. A.; Hanafi, L. A.; Li, D.; Gust, J.; Liles, W. C.; Wurfel, M. M.; López, J. A.; Chen, J.; Chung, D.; Harju-Baker, S.; Cherian, S.; Chen, X.; Riddell, S. R.; Maloney, D. G.; Turtle, C. J. Kinetics and biomarkers of severe cytokine release syndrome after CD19 chimeric antigen receptor-modified T-cell therapy. Blood 2017, 130 (21), 2295– 2306, DOI: 10.1182/blood-2017-06-793141There is no corresponding record for this reference.
- 11De Martin, E.; Michot, J.-M.; Papouin, B.; Champiat, S.; Mateus, C.; Lambotte, O.; Roche, B.; Antonini, T. M.; Coilly, A.; Laghouati, S.; Robert, C.; Marabelle, A.; Guettier, C.; Samuel, D. Characterization of liver injury induced by cancer immunotherapy using immune checkpoint inhibitors. Journal of Hepatology 2018, 68 (6), 1181– 1190, DOI: 10.1016/j.jhep.2018.01.033There is no corresponding record for this reference.
- 12Kularatne, R. N.; Crist, R. M.; Stern, S. T. The Future of Tissue-Targeted Lipid Nanoparticle-Mediated Nucleic Acid Delivery. Pharmaceuticals (Basel) 2022, 15 (7), 897, DOI: 10.3390/ph15070897There is no corresponding record for this reference.
- 13Chen, B.-M.; Cheng, T.-L.; Roffler, S. R. Polyethylene Glycol Immunogenicity: Theoretical, Clinical, and Practical Aspects of Anti-Polyethylene Glycol Antibodies. ACS Nano 2021, 15 (9), 14022– 14048, DOI: 10.1021/acsnano.1c05922There is no corresponding record for this reference.
- 14Xiao, Y.; Lian, X.; Sun, Y.; Sung, Y.-C.; Vaidya, A.; Chen, Z.; Gupta, A.; Chatterjee, S.; Zheng, L.; Guerrero, E.; Wang, X.; Farbiak, L.; Yang, Y.; Diamond, M. I.; Leal, C.; McDonald, J. G.; Siegwart, D. J. High-density brush-shaped polymer lipids reduce anti-PEG antibody binding for repeated administration of mRNA therapeutics. Nat. Mater. 2025, 24, 1840, DOI: 10.1038/s41563-024-02116-3There is no corresponding record for this reference.
- 15Hassanel, D. N. B. P.; Pilkington, E. H.; Ju, Y.; Kent, S. J.; Pouton, C. W.; Truong, N. P. Replacing poly(ethylene glycol) with RAFT lipopolymers in mRNA lipid nanoparticle systems for effective gene delivery. Int. J. Pharm. 2024, 665, 124695 DOI: 10.1016/j.ijpharm.2024.124695There is no corresponding record for this reference.
- 16He, X.; Payne, T. J.; Takanashi, A.; Fang, Y.; Kerai, S. D.; Morrow, J. P.; Al-Wassiti, H.; Pouton, C. W.; Kempe, K. Tailored Monoacyl Poly(2-oxazoline)- and Poly(2-oxazine)-Lipids as PEG-Lipid Alternatives for Stabilization and Delivery of mRNA-Lipid Nanoparticles. Biomacromolecules 2024, 25 (7), 4591– 4603, DOI: 10.1021/acs.biomac.4c00651There is no corresponding record for this reference.
- 17Choy, O. T.; Fletcher, N. L.; Fu, C.; Vu, M. N.; Ediriweera, G. R.; Humphries, J.; Hu, M.; Kent, S. J.; Cai, R.; Chen, C.; Whittaker, A. K. PEG-free Lipid Nanoparticles for mRNA Delivery: Superhydrophilic Sulfoxide Polymer Coatings. Biomacromolecules 2026, 27, 624, DOI: 10.1021/acs.biomac.5c01899There is no corresponding record for this reference.
- 18Ramishetti, S.; Kedmi, R.; Goldsmith, M.; Leonard, F.; Sprague, A. G.; Godin, B.; Gozin, M.; Cullis, P. R.; Dykxhoorn, D. M.; Peer, D. Systemic Gene Silencing in Primary T Lymphocytes Using Targeted Lipid Nanoparticles. ACS Nano 2015, 9 (7), 6706– 6716, DOI: 10.1021/acsnano.5b02796There is no corresponding record for this reference.
- 19Tombácz, I.; Laczkó, D.; Shahnawaz, H.; Muramatsu, H.; Natesan, A.; Yadegari, A.; Papp, T. E.; Alameh, M.-G.; Shuvaev, V.; Mui, B. L.; Tam, Y. K.; Muzykantov, V.; Pardi, N.; Weissman, D.; Parhiz, H. Highly efficient CD4+ T cell targeting and genetic recombination using engineered CD4+ cell-homing mRNA-LNPs. Mol. Ther. 2021, 29 (11), 3293– 3304, DOI: 10.1016/j.ymthe.2021.06.004There is no corresponding record for this reference.
- 20Chen, M. Z.; Yuen, D.; McLeod, V. M.; Yong, K. W.; Smyth, C. H.; Herling, B. R.; Payne, T. J.; Fabb, S. A.; Belousoff, M. J.; Algarni, A.; Sexton, P. M.; Porter, C. J. H.; Pouton, C. W.; Johnston, A. P. R. A versatile antibody capture system drives specific in vivo delivery of mRNA-loaded lipid nanoparticles. Nat. Nanotechnol. 2025, 20, 1273, DOI: 10.1038/s41565-025-01954-9There is no corresponding record for this reference.
- 21Dietmair, B.; Humphries, J.; Mercer, T. R.; Thurecht, K. J.; Howard, C. B.; Cheetham, S. W. Targeted mRNA delivery with bispecific antibodies that tether LNPs to cell surface markers. Mol. Ther Nucleic Acids 2025, 36 (2), 102520 DOI: 10.1016/j.omtn.2025.102520There is no corresponding record for this reference.
- 22Huda, P.; Humphries, J.; Fletcher, N. L.; Howard, C. B.; Thurecht, K. J.; Bell, C. A. Click-on Antibody Fragments for Customisable Targeted Nanomedicines – Site-specific Tetrazine and Azide Functionalisation through Non-canonical Amino Acid incorporation. Chemistry Methods 2024, 4 (2), e202300036 DOI: 10.1002/cmtd.202300036There is no corresponding record for this reference.
- 23Choy, O. T.; Fletcher, N. L.; Fu, C.; Vu, M. N.; Ediriweera, G. R.; Humphries, J.; Hu, M.; Kent, S. J.; Cai, R.; Chen, C.; Whittaker, A. K. PEG-free Lipid Nanoparticles for mRNA Delivery: Superhydrophilic Sulfoxide Polymer Coatings. Biomacromolecules 2026, 27 (1), 624– 639, DOI: 10.1021/acs.biomac.5c01899There is no corresponding record for this reference.
- 24Chong, Y. K.; Moad, G.; Rizzardo, E.; Thang, S. H. Thiocarbonylthio End Group Removal from RAFT-Synthesized Polymers by Radical-Induced Reduction. Macromolecules 2007, 40 (13), 4446– 4455, DOI: 10.1021/ma062919uThere is no corresponding record for this reference.
- 25Li, Q.; Chan, C.; Peterson, N.; Hanna, R. N.; Alfaro, A.; Allen, K. L.; Wu, H.; Dall’Acqua, W. F.; Borrok, M. J.; Santos, J. L. Engineering Caveolae-Targeted Lipid Nanoparticles To Deliver mRNA to the Lungs. ACS Chem. Biol. 2020, 15 (4), 830– 836, DOI: 10.1021/acschembio.0c00003There is no corresponding record for this reference.
- 26Kedmi, R.; Veiga, N.; Ramishetti, S.; Goldsmith, M.; Rosenblum, D.; Dammes, N.; Hazan-Halevy, I.; Nahary, L.; Leviatan-Ben-Arye, S.; Harlev, M.; Behlke, M.; Benhar, I.; Lieberman, J.; Peer, D. A modular platform for targeted RNAi therapeutics. Nat. Nanotechnol. 2018, 13 (3), 214– 219, DOI: 10.1038/s41565-017-0043-5There is no corresponding record for this reference.
- 27Escudé Martinez de Castilla, P.; Verdi, V.; de Voogt, W.; Estapé Sentí, M.; Koekman, A. C.; Rietveld, J.; van Kempen, S.; Yang, Q.; van Merris, J.; Jenster, G.; van Royen, M. E.; Fens, M. H.; Kooijmans, S. A. A.; van Weerden, W. M.; van Niel, G.; Vader, P.; Schiffelers, R. M. Nanobody-Decorated Lipid Nanoparticles for Enhanced mRNA Delivery to Tumors In Vivo. Advanced Healthcare Materials 2025, 14 (24), 2500605 DOI: 10.1002/adhm.202500605There is no corresponding record for this reference.
Supporting Information
Supporting Information
The Supporting Information is available free of charge at https://pubs.acs.org/doi/10.1021/acs.bioconjchem.6c00045.
Experiment procedure (materials and methods), 1H NMR spectra of COOH-PMSEA-DSPE and tCO-PMSEA-DSPE, flow cytometry data of mRNA-LNPs, physical characterization of mRNA-LNPs (PDF)
Terms & Conditions
Most electronic Supporting Information files are available without a subscription to ACS Web Editions. Such files may be downloaded by article for research use (if there is a public use license linked to the relevant article, that license may permit other uses). Permission may be obtained from ACS for other uses through requests via the RightsLink permission system: http://pubs.acs.org/page/copyright/permissions.html.



