
Decoding the Penicillium italicum–Citrus Interaction: Untargeted Metabolomics Sheds Light on a Neglected Postharvest PathogenClick to copy article linkArticle link copied!
- Evandro SilvaEvandro SilvaInstituto de Química, Universidade Estadual de Campinas, CEP 13083-970 Campinas, São Paulo, BrazilInstituto de Química de São Carlos, Universidade de São Paulo, CP 780, CEP 13560-970 São Carlos, São Paulo, BrazilMore by Evandro Silva
- Aline Midori KanashiroAline Midori KanashiroInstituto de Química, Universidade Estadual de Campinas, CEP 13083-970 Campinas, São Paulo, BrazilMore by Aline Midori Kanashiro
- José Rodrigo Ferreira MacielJosé Rodrigo Ferreira MacielInstituto de Química, Universidade Estadual de Campinas, CEP 13083-970 Campinas, São Paulo, BrazilMore by José Rodrigo Ferreira Maciel
- Rodolfo Dantas Lima JuniorRodolfo Dantas Lima JuniorInstituto de Química, Universidade Estadual de Campinas, CEP 13083-970 Campinas, São Paulo, BrazilMore by Rodolfo Dantas Lima Junior
- Maria Antonia Fraga BotelhoMaria Antonia Fraga BotelhoInstituto de Química, Universidade Estadual de Campinas, CEP 13083-970 Campinas, São Paulo, BrazilMore by Maria Antonia Fraga Botelho
- Alana Kelyene PereiraAlana Kelyene PereiraInstituto de Química, Universidade Estadual de Campinas, CEP 13083-970 Campinas, São Paulo, BrazilMore by Alana Kelyene Pereira
- Stephanie Nemesio da SilvaStephanie Nemesio da SilvaInstituto de Química, Universidade Estadual de Campinas, CEP 13083-970 Campinas, São Paulo, BrazilMore by Stephanie Nemesio da Silva
- Jonas Henrique CostaJonas Henrique CostaInstituto de Química, Universidade Estadual de Campinas, CEP 13083-970 Campinas, São Paulo, BrazilMore by Jonas Henrique Costa
- João Guilherme de Moraes PontesJoão Guilherme de Moraes PontesInstituto de Química, Universidade Estadual de Campinas, CEP 13083-970 Campinas, São Paulo, BrazilMore by João Guilherme de Moraes Pontes
- Amanda Ferreira da SilvaAmanda Ferreira da SilvaInstituto de Química, Universidade Estadual de Campinas, CEP 13083-970 Campinas, São Paulo, BrazilMore by Amanda Ferreira da Silva
- Igor Dias JurbergIgor Dias JurbergInstituto de Química, Universidade Estadual de Campinas, CEP 13083-970 Campinas, São Paulo, BrazilMore by Igor Dias Jurberg
- Roberto G. S. Berlinck*Roberto G. S. Berlinck*Email: [email protected]Instituto de Química de São Carlos, Universidade de São Paulo, CP 780, CEP 13560-970 São Carlos, São Paulo, BrazilMore by Roberto G. S. Berlinck
- Taicia Pacheco Fill*Taicia Pacheco Fill*Email: [email protected]Instituto de Química, Universidade Estadual de Campinas, CEP 13083-970 Campinas, São Paulo, BrazilMore by Taicia Pacheco Fill
Abstract
Penicillium italicum, the causal agent of citrus blue mold, is a major postharvest pathogen that reduces fruit quality and global citrus productivity. Understanding the molecular basis of infection is crucial to reveal virulence mechanisms, host defense responses, and potential targets for disease control. Here, we investigated the metabolic profile of theCitrus sinensis–P. italicum interaction using mass spectrometry-based metabolomics and desorption electrospray ionization mass spectrometry imaging. Key differentialP. italicum-derived metabolites were identified, including 12,13-dehydroprolyltryptophanyldiketopiperazine, deoxybrevianamide E, dehydrodeoxybrevianamide E, deoxyisoaustamide, and brevianamide F. To assess its biological role, brevianamide F was chemically synthesized and tested against citrus-associated endophytes. It selectively inhibitedDiaporthe sp., suggesting thatP. italicummay utilize this compound as an antimicrobial strategy to modulate the endophytic community during infection. These results provide the first insights into the natural products involved inP. italicumassociation with citrus and point to potential alternative strategies for controlling blue mold disease.
This publication is licensed under
License Summary*
You are free to share(copy and redistribute) this article in any medium or format and to adapt(remix, transform, and build upon) the material for any purpose, even commercially within the parameters below:
Creative Commons (CC): This is a Creative Commons license.
Attribution (BY): Credit must be given to the creator.
*Disclaimer
This summary highlights only some of the key features and terms of the actual license. It is not a license and has no legal value. Carefully review the actual license before using these materials.
License Summary*
You are free to share(copy and redistribute) this article in any medium or format and to adapt(remix, transform, and build upon) the material for any purpose, even commercially within the parameters below:
Creative Commons (CC): This is a Creative Commons license.
Attribution (BY): Credit must be given to the creator.
*Disclaimer
This summary highlights only some of the key features and terms of the actual license. It is not a license and has no legal value. Carefully review the actual license before using these materials.
License Summary*
You are free to share(copy and redistribute) this article in any medium or format and to adapt(remix, transform, and build upon) the material for any purpose, even commercially within the parameters below:
Creative Commons (CC): This is a Creative Commons license.
Attribution (BY): Credit must be given to the creator.
*Disclaimer
This summary highlights only some of the key features and terms of the actual license. It is not a license and has no legal value. Carefully review the actual license before using these materials.
1. Introduction
2. Material and Methods
2.1. Fungal Strain
2.2. Fruit Inoculation withP. italicum and Sample Collection
2.3. Metabolite Extraction and Untargeted Metabolomics Analysis
2.4. Data Processing and Statistical Analyses
2.5. Molecular Networking and Metabolite Annotation
2.6. In Vitro Secondary Metabolites Produced by P. italicum
2.7. Mass Spectrometry Imaging Analyses
2.8. Isolation and Identification of Endophytic Fungi fromCitrus sinensis
2.9. Coculture Growth Conditions and Extraction of Secondary Metabolites
2.10. Antifungal Assays
2.11. Confocal Microscopy Analysis of Diaporthesp. Growth in Response to Brevianamide F
2.12. Isolation, Preparation, and Analysis of Marfey’s Derivatives Reaction
3. Results
3.1. Blue Mold Disease Symptoms and LC–HRMS Analysis of Blue Mold Disease
Figure 1
Figure 1. Metabolic changes in oranges infected with P. italicum at 10 days post-inoculation. (Left) Representative images of healthy (control) andP. italicum-infected oranges at 10 days post-inoculation, showing extensive fungal colonization. Scale: 2 cm. (Right) PCA of metabolic profiles from control (green) and infected (red) fruit samples at 10 days post-inoculation. A clear separation is observed between the two groups, with PC1 accounting for 40.0% and PC2 for 11.6% of the total variance, indicating significant infection-induced metabolic reprogramming.
3.2. Multivariate Data Analysis
| ID | scan | metabolite annotation | molecular formula | [M + H]+ theoretical mass (m/z) | [M + H]+ measured mass (m/z) | mass accuracy (ppm) | in vivo control inoculated | in vitro | VIP | GNPS library accession | |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 558 | asparagine | C4H8N2O3 | 133.0608 | 133.0607 | –0.08 | X | X | n.d | 1.00 | CCMSLIB00006120683 |
| 2 | 1534 | 4-hydroxycinnamyl alcohol | C9H10O2 | 151.0754 | 151.0753 | –0.07 | X | X | n.d | 1.51 | supplementary data |
| 3 | 836 | phenylalanine | C9H11NO2 | 166.0863 | 166.0862 | –0.06 | X | X | n.d | CCMSLIB00003135371 | |
| 4 | 687 | synephrine | C9H13NO2 | 168.1019 | 168.1018 | –0.06 | X | X | n.d | CCMSLIB00004691344 | |
| 5 | 2371 | indolelactic acid | C11H11NO3 | 206.0811 | 206.0810 | –0.05 | X | X | n.d | 1.50 | CCMSLIB00006684092 |
| 6 | 6090 | nootkatone | C15H22O | 219.1743 | 219.1742 | –0.05 | X | X | n.d | CCMSLIB00005763621 | |
| 7 | 1081 | feruloyl putrescine | C14H20N2O3 | 265.1547 | 265.1546 | –0.04 | X | X | n.d | CCMSLIB00005748443 | |
| 8 | 1356 | 3′,5,7-trihydroxyflavanone | C15H12O5 | 273.0757 | 273.0755 | –0.07 | X | X | n.d | CCMSLIB00006411935 | |
| 9 | 3778 | naringenin | C15H12O5 | 273.0757 | 273.0756 | –0.04 | X | X | n.d | 1.46 | CCMSLIB00010105222 |
| 10 | 3978 | hesperetin | C16H14O6 | 303.0850 | 303.0861 | 0.36 | X | X | n.d | 1.48 | CCMSLIB00006374511 |
| 11 | 6944 | tetramethyl-O-scutellarein | C19H18O6 | 343.1176 | 343.1173 | –0.09 | X | X | n.d | CCMSLIB00006422351 | |
| 12 | 5337 | tangeretin | C20H20O7 | 373.1282 | 373.1281 | –0.03 | X | X | n.d | 1.36 | CCMSLIB00012320217 |
| 13 | 4959 | nobiletin | C21H22O8 | 403.1387 | 403.1386 | –0.02 | X | X | n.d | 1.45 | CCMSLIB00012349534 |
| 14 | 2719 | hesperetin 7-O-glucoside | C22H24O8 | 465.1324 | 465.1389 | 1.40 | X | X | n.d | supplementary data | |
| 15 | 2248 | diosmin | C28H32O15 | 609.1814 | 609.1810 | –0.07 | X | X | n.d | 1.37 | CCMSLIB00012176442 |
| 16 | 2410 | hesperidin | C28H35O15 | 611.1970 | 611.1966 | –0.07 | X | X | n.d | 1.46 | CCMSLIB00012176443 |
| 17 | 2215 | 12,13-dehydroprolyltryptophanyldiketopiperazin | C16H15N3O2 | 282.1237 | 282.1236 | –0.04 | n.d | X | X | 1.49 | supplementary data |
| 18 | 4265 | brevianamide F | C16H17N3O2 | 284.1394 | 284.1392 | –0.07 | n.d | X | X | 1.44 | |
| 19 | 4570 | deoxyisoaustamide | C21H21N3O2 | 348.1706 | 348.1707 | 0.03 | n.d | X | X | 1.38 | CCMSLIB00012438440 |
| 20 | 4280 | dehydrodeoxybrevianamide E | C21H23N3O2 | 350.1863 | 350.1862 | –0.03 | n.d | X | X | 1.50 | supplementary data |
| 21 | 4506 | deoxybrevianamide E | C21H25N3O2 | 352.2020 | 352.2013 | –0.20 | n.d | X | X | 1.34 | supplementary data |
| 22 | 4012 | brevianamide A | C21H23N3O3 | 366.1817 | 366.1808 | –0.25 | n.d | X | X | 1.03 | supplementary data |
Differentially expressed metabolites with VIP values ≥1 obtained from the PLS-DA model; X: detected, n.d: not detected.
3.3. Molecular Networking of theP. italicum–Citrus Interaction
Figure 2
Figure 2. Molecular network cluster of diketopiperazine alkaloids identified inP. italicum-infected oranges. Blue nodes represent metabolites exclusively detected in infected samples. Nodes with a polygonal shape indicate spectral matches with GNPS library compounds. Several interconnected nodes form a cluster with high cosine similarity, suggesting structurally related diketopiperazines. Annotated compounds include brevianamide F, deoxybrevianamide E, dehydrodeoxybrevianamide E, deoxyisoaustamide, and 12,13-dehydroprolyltryptophanyldiketopiperazine. This cluster highlights the metabolic specialization associated withP. italicum infection. Edge thickness corresponds to the cosine score, with thicker edges indicating higher spectral similarity.
3.4. In Vivo Production and Spatial Mapping of Secondary Metabolites duringP. italicum Infection
Figure 3
Figure 3. Spatial distribution of brevianamide F (18) and deoxyisoaustamide (19) in P. italicum-infected oranges using DESI-MSI. Ion images show metabolite localization in control and infected samples at 6 days post-inoculation. The respective m/z values were not detected in the control orange peels, while strong signals were observed in infected tissues, particularly at the fungal infection site. The color scale indicates relative ion intensity (from low in blue to high in red). Observed mass errors are −0.39 ppm for brevianamide F and 3.87 ppm for deoxyisoaustamide.
3.5. In Vitro Production of Diketopiperazine Alkaloids byP. italicum
3.6. Isolation and Identification of Endophytic Fungi Community from Citrus sinensis
3.7. Coculture Assays between Endophytes andP. italicum
Figure 4
Figure 4. Confrontation assay betweenP. italicum and endophytic fungi. (Upper): coculture assays showing the interaction zones betweenP. italicum andDiaporthe sp. (left) orP. italicum andColletotrichum sp. (right). Dashed green lines highlight the confrontation zone. (Lower): diketopiperazine alkaloids detected in the confrontation zone through LC–HRMS analysis and structure annotation: deoxyisoaustamide, brevianamide F, 12,13-dehydroprolyltryptophanyldiketopiperazine, deoxybrevianamide E, and dehydrodeoxybrevianamide E.
3.8. Brevianamide F Antifungal Assays with Citrus Endophytes
Figure 5
Figure 5. Antifungal activity of secondary metabolites againstDiaporthe sp. (Left): mycelial growth of Diaporthe sp. on PDA plates after 7 days of incubation under four treatments: control (PDA only), negative control (ethanol/water 1:1, v/v), brevianamide F (0.3 mg/mL), and imazalil (0.3 mg/mL, positive control). (Right, upper): quantification of fungal growth area (cm2). Data represent mean ± standard deviation (n = 3). Statistical analysis was performed using one-way ANOVA (p < 0.0001), followed by Tukey’s post hoc test. Treatments labeled with the same letter are not significantly different, while those with different letters indicate statistically significant differences. (Right, lower): chemical structures of brevianamide F and imazalil used in the treatments. Brevianamide F inhibited mycelial growth by 51.92% compared to the control.
3.9. Confocal Microscopy Reveals the Inhibitory Effect of Brevianamide F on Diaporthe sp. Hyphal Development
Figure 6
Figure 6. Effect of brevianamide F on fungal hyphal morphology. Representative images ofDiaporthe sp. grown on PDA plates supplemented with brevianamide F (upper row) or control PDA (lower row). Left panels show colony morphology, while middle and right panels display confocal microscopy images of the hyphal structure stained with a fluorescent dye. Treatment with brevianamide F results in notable alterations in hyphal organization and density compared to the untreated control.
4. Discussion
Data Availability
All data generated or analyzed during this study are included in this published article and its Supporting Information.
Supporting Information
The Supporting Information is available free of charge at https://pubs.acs.org/doi/10.1021/acs.jafc.5c07618.
Synthesis and NMR analysis of brevianamide F; LC–MS TICs of orange peel extracts; PCA and PLS-DA plots of metabolomic data; MS/MS molecular network showing metabolite differences between healthy and infected samples; in vitro LC–MS/MS confirmation of diketopiperazines; FDAA derivatization HPLC-UV chromatograms for proline stereochemistry; FDAA derivatization HPLC-UV chromatograms for tryptophan stereochemistry; EICs for d- and l-tryptophan FDAA derivatives and brevianamide F; dual-culture assays with endophytic fungi; ITS sequences of isolates; phylogenetic dendrograms of Colletotrichum sp. and Diaporthe sp. isolates; and MIC assay of brevianamide F against Diaporthe sp. (PDF)
Terms & Conditions
Most electronic Supporting Information files are available without a subscription to ACS Web Editions. Such files may be downloaded by article for research use (if there is a public use license linked to the relevant article, that license may permit other uses). Permission may be obtained from ACS for other uses through requests via the RightsLink permission system: http://pubs.acs.org/page/copyright/permissions.html.
Acknowledgments
This work was funded by the National Council for Scientific and Technological Development (CNPq), the Coordination for the Improvement of Higher Education Personnel (CAPES)─Finance Code 001, and the São Paulo Research Foundation (FAPESP, grant numbers 2019/01235-8 to I.D.J., 2019/17721-9 to R.G.S.B., 2022/03594-8 to E.S., 2023/03831-2 to E.S., and 2022/02992-0 to T.P.F.). R.D.L.J. and J.R.F.M. acknowledge the research scholarship (142013/2024-2; 161466/2021-4), and R.G.S.B. also acknowledges CNPq for the senior research scholarship (304247/2021-9).
References
This article references 71 other publications.
- 1Wang, Z.; Sui, Y.; Li, J.; Tian, X.; Wang, Q. Biological control of postharvest fungal decays in citrus: a review. Crit. Rev. Food Sci. Nutr. 2022, 62 (4), 861– 870, DOI: 10.1080/10408398.2020.1829542Google ScholarThere is no corresponding record for this reference.
- 2François, G. A.; de Moraes Pontes, J. G.; Pereira, A. K.; Fill, T. P. Exploring the citrus sour rot pathogen: biochemical aspects, virulence factors, and strategies for disease management – a review. Fungal Biol. Rev. 2022, 41, 70– 83, DOI: 10.1016/j.fbr.2022.03.003Google ScholarThere is no corresponding record for this reference.
- 3Bhatta, U. K. Alternative management approaches of citrus diseases caused by Penicillium digitatum (green mold) and Penicillium italicum (blue mold). Front. Plant Sci. 2022, 12, 833328, DOI: 10.3389/fpls.2021.833328Google ScholarThere is no corresponding record for this reference.
- 4Khamsaw, P.; Sangta, J.; Chaiwan, P.; Rachtanapun, P.; Sirilun, S.; Sringarm, K.; Thanakkasaranee, S.; Sommano, S. R. Bio-circular perspective of citrus fruit loss caused by pathogens: occurrences, active ingredient recovery and applications. Horticulturae 2022, 8 (8), 748, DOI: 10.3390/horticulturae8080748Google ScholarThere is no corresponding record for this reference.
- 5Moraes Bazioli, J.; Belinato, J. R.; Costa, J. H.; Akiyama, D. Y.; Pontes, J. G. d. M.; Kupper, K. C.; Augusto, F.; de Carvalho, J. E.; Fill, T. P. Biological control of citrus postharvest phytopathogens. Toxins 2019, 11 (8), 460, DOI: 10.3390/toxins11080460Google ScholarThere is no corresponding record for this reference.
- 6Costa, J. H.; Bazioli, J. M.; de Moraes Pontes, J. G.; Fill, T. P. Penicillium digitatum infection mechanisms in citrus: what do we know so far?. Fungal Biol. 2019, 123 (8), 584– 593, DOI: 10.1016/j.funbio.2019.05.004Google ScholarThere is no corresponding record for this reference.
- 7Kanashiro, A. M.; Akiyama, D. Y.; Kupper, K. C.; Fill, T. P. Penicillium italicum: an underexplored postharvest pathogen. Front. Microbiol. 2020, 11, 606852, DOI: 10.3389/fmicb.2020.606852Google ScholarThere is no corresponding record for this reference.
- 8Plaza, P.; Usall, J.; Teixidó, N.; Viñas, I. Effect of water activity and temperature on germination and growth of Penicillium digitatum, Penicillium italicum and Geotrichum candidum. J. Appl. Microbiol. 2003, 94 (4), 549– 554, DOI: 10.1046/j.1365-2672.2003.01909.xGoogle ScholarThere is no corresponding record for this reference.
- 9Latifa, A.; Idriss, T.; Hassan, B.; Amine, S. M.; El Hassane, B.; Aoumar Abd, A. B. Effects of organic acids and salts on the development of Penicillium italicum: the causal agent of citrus blue mold. Plant Pathol. J. 2011, 10 (3), 99– 107, DOI: 10.3923/ppj.2011.99.107Google ScholarThere is no corresponding record for this reference.
- 10Li, X.; Yang, S.; Zhang, M.; Yang, Y.; Peng, L. Identification of pathogenicity-related effector proteins and the role of Piwsc1 in the virulence of Penicillium italicum on citrus fruits. J. Fungi 2022, 8 (6), 646, DOI: 10.3390/jof8060646Google ScholarThere is no corresponding record for this reference.
- 11Hershenhorn, J. Polygalacturonases associated with infection of Valencia orange by Penicillium italicum. Phytopathology 1990, 80 (12), 1374, DOI: 10.1094/Phyto-80-1374Google ScholarThere is no corresponding record for this reference.
- 12Ballester, A.-R.; Marcet-Houben, M.; Levin, E.; Sela, N.; Selma-Lázaro, C.; Carmona, L.; Wisniewski, M.; Droby, S.; González-Candelas, L.; Gabaldón, T. Genome, transcriptome, and functional analyses of Penicillium expansum provide new insights into secondary metabolism and pathogenicity. Mol. Plant-Microbe Interact. 2015, 28 (3), 232– 248, DOI: 10.1094/MPMI-09-14-0261-FIGoogle ScholarThere is no corresponding record for this reference.
- 13Ropars, J.; de la Vega, R. C. R.; López-Villavicencio, M.; Gouzy, J.; Dupont, J.; Swennen, D.; Dumas, E.; Giraud, T.; Branca, A. Diversity and mechanisms of genomic adaptation in Penicillium. Aspergillus and Penicillium in the Post-genomic Era; Caister Academic Press: Norfolk, UK, 2016; pp 27– 42.Google ScholarThere is no corresponding record for this reference.
- 14Błaszczyk, U.; Wyrzykowska, S.; Gąstoł, M. Application of bioactive coatings with killer yeasts to control post-harvest apple decay caused by Botrytis cinerea and Penicillium italicum. Foods 2022, 11 (13), 1868, DOI: 10.3390/foods11131868Google ScholarThere is no corresponding record for this reference.
- 15Chen, C.; Peng, X.; Wan, C.; Zhang, Y.; Gan, Z.; Zeng, J.; Kai, W.; Chen, J. Lignin biosynthesis pathway and redox balance act synergistically in conferring resistance against Penicillium italicum infection in 7-demethoxytylophorine-treated Navel orange. J. Agric. Food Chem. 2022, 70 (26), 8111– 8123, DOI: 10.1021/acs.jafc.2c02348Google ScholarThere is no corresponding record for this reference.
- 16Kahramanoğlu, I. ̇.; Nisar, M. F.; Chen, C.; Usanmaz, S.; Chen, J.; Wan, C. Light: an alternative method for physical control of postharvest rotting caused by fungi of citrus fruit. J. Food Qual. 2020, 2020, 1– 12, DOI: 10.1155/2020/8821346Google ScholarThere is no corresponding record for this reference.
- 17Jeong, M.-A.; Jeong, R.-D. Applications of ionizing radiation for the control of postharvest diseases in fresh produce: recent advances. Plant Pathol. 2018, 67 (1), 18– 29, DOI: 10.1111/ppa.12739Google ScholarThere is no corresponding record for this reference.
- 18Trabelsi, D.; Hamdane, A. M.; Said, M. B.; Abdrrabba, M. Chemical composition and antifungal activity of essential oils from flowers, leaves and peels of Tunisian Citrus aurantium against Penicillium digitatum and Penicillium italicum. J. Essent. Oil Bear. Plants 2016, 19 (7), 1660– 1674, DOI: 10.1080/0972060X.2016.1141069Google ScholarThere is no corresponding record for this reference.
- 19Vázquez-González, Y.; Ragazzo-Sánchez, J. A.; Calderón-Santoyo, M. Characterization and antifungal activity of jackfruit (Artocarpus heterophyllus Lam.) leaf extract obtained using conventional and emerging technologies. Food Chem. 2020, 330, 127211, DOI: 10.1016/j.foodchem.2020.127211Google ScholarThere is no corresponding record for this reference.
- 20Osonga, F. J.; Eshun, G.; Kalra, S.; Yazgan, I.; Sakhaee, L.; Ontman, R.; Jiang, S.; Sadik, O. A. Influence of particle size and shapes on the antifungal activities of greener nanostructured copper against Penicillium italicum. ACS Agric. Sci. Technol. 2022, 2 (1), 42– 56, DOI: 10.1021/acsagscitech.1c00102Google ScholarThere is no corresponding record for this reference.
- 21Papoutsis, K.; Mathioudakis, M. M.; Hasperué, J. H.; Ziogas, V. Non-chemical treatments for preventing the postharvest fungal rotting of citrus caused by Penicillium digitatum (green mold) and Penicillium italicum (blue mold). Trends Food Sci. Technol. 2019, 86, 479– 491, DOI: 10.1016/j.tifs.2019.02.053Google ScholarThere is no corresponding record for this reference.
- 22Talibi, I.; Boubaker, H.; Boudyach, E. H.; Ait Ben Aoumar, A. Alternative methods for the control of postharvest citrus diseases. J. Appl. Microbiol. 2014, 117 (1), 1– 17, DOI: 10.1111/jam.12495Google ScholarThere is no corresponding record for this reference.
- 23Pontes, J. G. M.; Fernandes, L. S.; dos Santos, R. V.; Tasic, L.; Fill, T. P. Virulence factors in the phytopathogen–host interactions: an overview. J. Agric. Food Chem. 2020, 68 (29), 7555– 7570, DOI: 10.1021/acs.jafc.0c02389Google ScholarThere is no corresponding record for this reference.
- 24Silva, E.; Dantas, R.; Barbosa, J. C.; Berlinck, R. G. S.; Fill, T. Metabolomics approach to understand molecular mechanisms involved in fungal pathogen–citrus pathosystems. Mol. Omics 2024, 20 (3), 154– 168, DOI: 10.1039/D3MO00182BGoogle ScholarThere is no corresponding record for this reference.
- 25Adusumilli, R.; Mallick, P. Data conversion with ProteoWizard MSConvert. Computational Methods for Mass Spectrometry Proteomics; Hamid, J., Ed.; Humana Press: New York, NY, 2017; pp 339– 368.Google ScholarThere is no corresponding record for this reference.
- 26Wang, M.; Carver, J. J.; Phelan, V. V.; Sanchez, L. M.; Garg, N.; Peng, Y.; Nguyen, D. D.; Watrous, J.; Kapono, C. A.; Luzzatto-Knaan, T.; Porto, C.; Bouslimani, A.; Melnik, A. V.; Meehan, M. J.; Liu, W.-T.; Crüsemann, M.; Boudreau, P. D.; Esquenazi, E.; Sandoval-Calderón, M.; Kersten, R. D.; Pace, L. A.; Quinn, R. A.; Duncan, K. R.; Hsu, C.-C.; Floros, D. J.; Gavilan, R. G.; Kleigrewe, K.; Northen, T.; Dutton, R. J.; Parrot, D.; Carlson, E. E.; Aigle, B.; Michelsen, C. F.; Jelsbak, L.; Sohlenkamp, C.; Pevzner, P.; Edlund, A.; McLean, J.; Piel, J.; Murphy, B. T.; Gerwick, L.; Liaw, C.-C.; Yang, Y.-L.; Humpf, H.-U.; Maansson, M.; Keyzers, R. A.; Sims, A. C.; Johnson, A. R.; Sidebottom, A. M.; Sedio, B. E.; Klitgaard, A.; Larson, C. B.; Boya P, C. A.; Torres-Mendoza, D.; Gonzalez, D. J.; Silva, D. B.; Marques, L. M.; Demarque, D. P.; Pociute, E.; O’Neill, E. C.; Briand, E.; Helfrich, E. J. N.; Granatosky, E. A.; Glukhov, E.; Ryffel, F.; Houson, H.; Mohimani, H.; Kharbush, J. J.; Zeng, Y.; Vorholt, J. A.; Kurita, K. L.; Charusanti, P.; McPhail, K. L.; Nielsen, K. F.; Vuong, L.; Elfeki, M.; Traxler, M. F.; Engene, N.; Koyama, N.; Vining, O. B.; Baric, R.; Silva, R. R.; Mascuch, S. J.; Tomasi, S.; Jenkins, S.; Macherla, V.; Hoffman, T.; Agarwal, V.; Williams, P. G.; Dai, J.; Neupane, R.; Gurr, J.; Rodríguez, A. M. C.; Lamsa, A.; Zhang, C.; Dorrestein, K.; Duggan, B. M.; Almaliti, J.; Allard, P.-M.; Phapale, P.; Nothias, L.-F.; Alexandrov, T.; Litaudon, M.; Wolfender, J.-L.; Kyle, J. E.; Metz, T. O.; Peryea, T.; Nguyen, D.-T.; VanLeer, D.; Shinn, P.; Jadhav, A.; Müller, R.; Waters, K. M.; Shi, W.; Liu, X.; Zhang, L.; Knight, R.; Jensen, P. R.; Palsson, B. Ø.; Pogliano, K.; Linington, R. G.; Gutiérrez, M.; Lopes, N. P.; Gerwick, W. H.; Moore, B. S.; Dorrestein, P. C.; Bandeira, N. Sharing and Community Curation of Mass Spectrometry Data with Global Natural Products Social Molecular Networking. Nat. Biotechnol. 2016, 34 (8), 828– 837, DOI: 10.1038/nbt.3597Google ScholarThere is no corresponding record for this reference.
- 27Shannon, P.; Markiel, A.; Ozier, O.; Baliga, N. S.; Wang, J. T.; Ramage, D.; Amin, N.; Schwikowski, B.; Ideker, T. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 2003, 13 (11), 2498– 2504, DOI: 10.1101/gr.1239303Google ScholarThere is no corresponding record for this reference.
- 28Kieffer, M. E.; Chuang, K. V.; Reisman, S. E. Copper-catalyzed diastereoselective arylation of tryptophan derivatives: total synthesis of (+)-naseseazines A and B. J. Am. Chem. Soc. 2013, 135 (15), 5557– 5560, DOI: 10.1021/ja4023557Google ScholarThere is no corresponding record for this reference.
- 29Godinho, B. T. V.; Santos, I. ́. A. F. M.; Gomes, E. A.; Piccoli, R. H.; Cardoso, P. G. Endophytic fungi community in Eremanthus erythropappus tree from anthropogenic and natural areas of Minas Gerais. Cerne 2019, 25 (3), 283– 293, DOI: 10.1590/01047760201925032642Google ScholarThere is no corresponding record for this reference.
- 30Aamir, S. A rapid and efficient method of fungal genomic DNA extraction, suitable for PCR-based molecular methods. Plant Pathol. Quar. 2015, 5 (2), 74– 81, DOI: 10.5943/ppq/5/2/6Google ScholarThere is no corresponding record for this reference.
- 31Thompson, J. The CLUSTAL_X Windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res. 1997, 25 (24), 4876– 4882, DOI: 10.1093/nar/25.24.4876Google ScholarThere is no corresponding record for this reference.
- 32Tamura, K.; Stecher, G.; Peterson, D.; Filipski, A.; Kumar, S. MEGA6: molecular evolutionary genetics analysis version 6.0. Mol. Biol. Evol. 2013, 30 (12), 2725– 2729, DOI: 10.1093/molbev/mst197Google ScholarThere is no corresponding record for this reference.
- 33Kimura, M. A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J. Mol. Evol. 1980, 16 (2), 111– 120, DOI: 10.1007/BF01731581Google ScholarThere is no corresponding record for this reference.
- 34Saitou, N.; Nei, M. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 1987, 4 (4), 406– 425, DOI: 10.1093/oxfordjournals.molbev.a040454Google ScholarThere is no corresponding record for this reference.
- 35Costa, J. H.; Wassano, C. I.; Angolini, C. F. F.; Scherlach, K.; Hertweck, C.; Pacheco Fill, T. Antifungal potential of secondary metabolites involved in the interaction between citrus pathogens. Sci. Rep. 2019, 9 (1), 18647, DOI: 10.1038/s41598-019-55204-9Google ScholarThere is no corresponding record for this reference.
- 36Xu, W.-F.; Mao, N.; Xue, X.-J.; Qi, Y.-X.; Wei, M.-Y.; Wang, C.-Y.; Shao, C.-L. Structures and absolute configurations of diketopiperazine alkaloids chrysopiperazines A–C from the gorgonian-derived Penicillium chrysogenum fungus. Mar. Drugs 2019, 17 (5), 250, DOI: 10.3390/md17050250Google ScholarThere is no corresponding record for this reference.
- 37Caballero, E.; Avendaño, C.; Menéndez, J. C. Brief total synthesis of the cell cycle inhibitor tryprostatin B and related preparation of its alanine analogue. J. Org. Chem. 2003, 68 (18), 6944– 6951, DOI: 10.1021/jo034703lGoogle ScholarThere is no corresponding record for this reference.
- 38Hawkins, J. P.; Geddes, B. A.; Oresnik, I. J. Common dyes used to determine bacterial polysaccharides on agar are affected by medium acidification. Can. J. Microbiol. 2017, 63 (6), 559– 562, DOI: 10.1139/cjm-2016-0743Google ScholarThere is no corresponding record for this reference.
- 39Costa, J. H.; Bazioli, J. M.; de Vilhena Araújo, E.; Vendramini, P. H.; de Freitas Porto, M. C.; Eberlin, M. N.; Souza-Neto, J. A.; Fill, T. P. Monitoring indole alkaloid production by Penicillium digitatum during infection process in citrus by mass spectrometry imaging and molecular networking. Fungal Biol. 2019, 123 (8), 594– 600, DOI: 10.1016/j.funbio.2019.03.002Google ScholarThere is no corresponding record for this reference.
- 40Silva, E.; Belinato, J. R.; Porto, C.; Nunes, E.; Guimarães, F.; Meyer, M. C.; Pilau, E. J. Soybean metabolomics based in mass spectrometry: decoding the plant’s signaling and defense responses under biotic stress. J. Agric. Food Chem. 2021, 69 (26), 7257– 7267, DOI: 10.1021/acs.jafc.0c07758Google ScholarThere is no corresponding record for this reference.
- 41Liu, X.; Lin, C.; Ma, X.; Tan, Y.; Wang, J.; Zeng, M. Functional characterization of a flavonoid glycosyltransferase in sweet orange (Citrus sinensis). Front. Plant Sci. 2018, 9, 166, DOI: 10.3389/fpls.2018.00166Google ScholarThere is no corresponding record for this reference.
- 42Nogata, Y.; Sakamoto, K.; Shiratsuchi, H.; Ishii, T.; Yano, M.; Ohta, H. Flavonoid composition of fruit tissues of Citrus species. Biosci., Biotechnol., Biochem. 2006, 70 (1), 178– 192, DOI: 10.1271/bbb.70.178Google ScholarThere is no corresponding record for this reference.
- 43Gattuso, G.; Barreca, D.; Gargiulli, C.; Leuzzi, U.; Caristi, C. Flavonoid composition of citrus juices. Molecules 2007, 12 (8), 1641– 1673, DOI: 10.3390/12081641Google ScholarThere is no corresponding record for this reference.
- 44Dixon, R. A.; Paiva, N. L. Stress-induced phenylpropanoid metabolism. Plant Cell 1995, 7 (7), 1085– 1097, DOI: 10.1105/tpc.7.7.1085Google ScholarThere is no corresponding record for this reference.
- 45Zhao, S.; Nan, Y.; Yao, R.; Wang, L.; Zeng, X.; Aadil, R. M.; Shabbir, M. A. Antibacterial activity and transcriptomic analysis of hesperetin against Alicyclobacillus acidoterrestris vegetative cells. Foods 2023, 12 (17), 3276, DOI: 10.3390/foods12173276Google ScholarThere is no corresponding record for this reference.
- 46Ferrer, J.-L.; Austin, M. B.; Stewart, C.; Noel, J. P. Structure and function of enzymes involved in the biosynthesis of phenylpropanoids. Plant Physiol. Biochem. 2008, 46 (3), 356– 370, DOI: 10.1016/j.plaphy.2007.12.009Google ScholarThere is no corresponding record for this reference.
- 47Treutter, D. Significance of flavonoids in plant resistance: a review. Environ. Chem. Lett. 2006, 4 (3), 147– 157, DOI: 10.1007/s10311-006-0068-8Google ScholarThere is no corresponding record for this reference.
- 48Agati, G.; Azzarello, E.; Pollastri, S.; Tattini, M. Flavonoids as antioxidants in plants: location and functional significance. Plant Sci. 2012, 196, 67– 76, DOI: 10.1016/j.plantsci.2012.07.014Google ScholarThere is no corresponding record for this reference.
- 49Nehela, Y.; Killiny, N. The unknown soldier in citrus plants: polyamines-based defensive mechanisms against biotic and abiotic stresses and their relationship with other stress-associated metabolites. Plant Signaling Behav. 2020, 15 (6), 1761080, DOI: 10.1080/15592324.2020.1761080Google ScholarThere is no corresponding record for this reference.
- 50Kyselka, J.; Bleha, R.; Dragoun, M.; Bialasová, K.; Horáčková, S. ̌.; Schätz, M.; Sluková, M.; Filip, V.; Synytsya, A. Antifungal polyamides of hydroxycinnamic acids from sunflower bee pollen. J. Agric. Food Chem. 2018, 66 (42), 11018– 11026, DOI: 10.1021/acs.jafc.8b03976Google ScholarThere is no corresponding record for this reference.
- 51Khan, M. K.; Zill-E-Huma; Dangles, O. A comprehensive review on flavanones, the major citrus polyphenols. J. Food Compos. Anal. 2014, 33 (1), 85– 104, DOI: 10.1016/j.jfca.2013.11.004Google ScholarThere is no corresponding record for this reference.
- 52Xu, X.; Pu, R.; Li, Y.; Wu, Z.; Li, C.; Miao, X.; Yang, W. Chemical compositions of propolis from China and the United States and their antimicrobial activities against Penicillium notatum. Molecules 2019, 24 (19), 3576, DOI: 10.3390/molecules24193576Google ScholarThere is no corresponding record for this reference.
- 53Yang, S. Z.; Peng, L. T.; Su, X. J.; Chen, F.; Cheng, Y. J.; Fan, G.; Pan, S. Y. Bioassay-guided isolation and identification of antifungal components from propolis against Penicillium italicum. Food Chem. 2011, 127 (1), 210– 215, DOI: 10.1016/j.foodchem.2010.12.011Google ScholarThere is no corresponding record for this reference.
- 54Huo, R.; Zhang, J.; Niu, S.; Liu, L. New prenylated indole diketopiperazine alkaloids and polyketides from the mangrove-derived fungus Penicillium sp. Front. Mar. Sci. 2022, 9, 1097594, DOI: 10.3389/fmars.2022.1097594Google ScholarThere is no corresponding record for this reference.
- 55Ma, Y.-M.; Liang, X.-A.; Kong, Y.; Jia, B. Structural diversity and biological activities of indole diketopiperazine alkaloids from fungi. J. Agric. Food Chem. 2016, 64 (35), 6659– 6671, DOI: 10.1021/acs.jafc.6b01772Google ScholarThere is no corresponding record for this reference.
- 56Kozlovsky, A. G.; Vinokurova, N. G.; Adanin, V. M.; Burkhardt, G.; Dahse, H.-M.; Gräfe, U. New diketopiperazine alkaloids from Penicillium fellutanum. J. Nat. Prod. 2000, 63 (5), 698– 700, DOI: 10.1021/np9903853Google ScholarThere is no corresponding record for this reference.
- 57Bräse, S.; Encinas, A.; Keck, J.; Nising, C. F. Chemistry and biology of mycotoxins and related fungal metabolites. Chem. Rev. 2009, 109 (9), 3903– 3990, DOI: 10.1021/cr050001fGoogle ScholarThere is no corresponding record for this reference.
- 58Asiri, I. A. M.; Badr, J. M.; Youssef, D. T. A. Penicillivinacine, antimigratory diketopiperazine alkaloid from the marine-derived fungus Penicillium vinaceum. Phytochem. Lett. 2015, 13, 53– 58, DOI: 10.1016/j.phytol.2015.05.014Google ScholarThere is no corresponding record for this reference.
- 59Liu, Y.-X.; Ma, S.-G.; Wang, X.-J.; Zhao, N.; Qu, J.; Yu, S.-S.; Dai, J.-G.; Wang, Y.-H.; Si, Y.-K. Diketopiperazine alkaloids produced by the endophytic fungus Aspergillus fumigatus from the stem of Erythrophloeum fordii Oliv. Helv. Chim. Acta 2012, 95 (8), 1401– 1408, DOI: 10.1002/hlca.201100521Google ScholarThere is no corresponding record for this reference.
- 60Danial, A. M.; Medina, A.; Sulyok, M.; Magan, N. Efficacy of metabolites of a Streptomyces strain (AS1) to control growth and mycotoxin production by Penicillium verrucosum, Fusarium verticillioides, and Aspergillus fumigatus in culture. Mycotoxin Res. 2020, 36 (2), 225– 234, DOI: 10.1007/s12550-020-00388-7Google ScholarThere is no corresponding record for this reference.
- 61Hubka, V.; Nováková, A.; Jurjević, Z. ̌.; Sklenář, F.; Frisvad, J. C.; Houbraken, J.; Arendrup, M. C.; Jørgensen, K. M.; Siqueira, J. P. Z.; Gené, J.; Kolařík, M. Polyphasic data support the splitting of Aspergillus candidus into two species; proposal of Aspergillus dobrogensis sp. nov. Int. J. Syst. Evol. Microbiol. 2018, 68 (4), 995– 1011, DOI: 10.1099/ijsem.0.002583Google ScholarThere is no corresponding record for this reference.
- 62Frisvad, J. C.; Samson, R. A. Polyphasic taxonomy of Penicillium subgenus Penicillium: a guide to identification of food and air-borne terverticillate penicillia and their mycotoxins. Stud. Mycol. 2004, 49 (1), 1– 174Google ScholarThere is no corresponding record for this reference.
- 63Scott, P. M.; Kennedy, B. P. C.; Harwig, J.; Chen, Y.-K. Formation of Diketopiperazines by Penicillium italicum Isolated from Oranges. Appl. Microbiol. 1974, 28 (5), 892– 894, DOI: 10.1128/am.28.5.892-894.1974Google ScholarThere is no corresponding record for this reference.
- 64Baek, S. Y.; Shim, S. H. Isolation and structure determination of metabolites from cultures of Aspergillus protuberus. Korean J. Microbiol. 2013, 49 (2), 179– 183, DOI: 10.7845/kjm.2013.3026Google ScholarThere is no corresponding record for this reference.
- 65Godfrey, R. C.; Green, N. J.; Nichol, G. S.; Lawrence, A. L. Total synthesis of brevianamide A. Nat. Chem. 2020, 12 (7), 615– 619, DOI: 10.1038/s41557-020-0442-3Google ScholarThere is no corresponding record for this reference.
- 66Sakhri, A.; Chaouche, N. K.; Catania, M. R.; Ritieni, A.; Santini, A. Chemical composition of Aspergillus creber extract and evaluation of its antimicrobial and antioxidant activities. Pol. J. Microbiol. 2019, 68 (3), 309– 316, DOI: 10.33073/pjm-2019-033Google ScholarThere is no corresponding record for this reference.
- 67Lee, S.; Sperry, J. Isolation and biological activity of azocine and azocane alkaloids. Bioorg. Med. Chem. 2022, 54, 116560, DOI: 10.1016/j.bmc.2021.116560Google ScholarThere is no corresponding record for this reference.
- 68Quang, T. H.; Lee, D.-S.; Sohn, J. H.; Kim, Y.-C.; Oh, H. New deoxyisoaustamide derivative from the marine-derived fungus Penicillium sp. JF-72. Bull. Korean Chem. Soc. 2013, 34 (10), 3109– 3112, DOI: 10.5012/bkcs.2013.34.10.3109Google ScholarThere is no corresponding record for this reference.
- 69Zhuravleva, O. I.; Antonov, A. S.; Trang, V. T. D.; Pivkin, M. V.; Khudyakova, Y. V.; Denisenko, V. A.; Popov, R. S.; Kim, N. Y.; Yurchenko, E. A.; Gerasimenko, A. V.; Udovenko, A. A.; Amsberg, G. v.; Dyshlovoy, S. A.; Afiyatullov, S. S. New deoxyisoaustamide derivatives from the coral-derived fungus Penicillium dimorphosporum KMM 4689. Mar. Drugs 2021, 19 (1), 32, DOI: 10.3390/md19010032Google ScholarThere is no corresponding record for this reference.
- 70Coleman, J. J.; Mylonakis, E. Efflux in Fungi: La Pièce de Résistance. PLoS Pathog. 2009, 5 (6), e1000486 DOI: 10.1371/journal.ppat.1000486Google ScholarThere is no corresponding record for this reference.
- 71Chen, X.; Pan, H.-X.; Tang, G.-L. Newly Discovered Mechanisms of Antibiotic Self-Resistance with Multiple Enzymes Acting at Different Locations and Stages. Antibiotics 2023, 12 (1), 35, DOI: 10.3390/antibiotics12010035Google ScholarThere is no corresponding record for this reference.
Cited By
This article is cited by 1 publications.
- Evandro Silva, Julio César Barbosa, Adrián González-Gárcia, Roberto G.S. Berlinck, Ana-Rosa Ballester, Luis González-Candelas, Taicia Fill. Decoding virulence in Penicillium italicum: A functional link between NRPS-derived cyclic peptides and citrus infection. Postharvest Biology and Technology 2026, 234 , 114069. https://doi.org/10.1016/j.postharvbio.2025.114069
Article Views are the COUNTER-compliant sum of full text article downloads since November 2008 (both PDF and HTML) across all institutions and individuals. These metrics are regularly updated to reflect usage leading up to the last few days.
Citations are the number of other articles citing this article, calculated by Crossref and updated daily. Find more information about Crossref citation counts.
The Altmetric Attention Score is a quantitative measure of the attention that a research article has received online. Clicking on the donut icon will load a page at altmetric.com with additional details about the score and the social media presence for the given article. Find more information on the Altmetric Attention Score and how the score is calculated.
Recommended Articles
Abstract
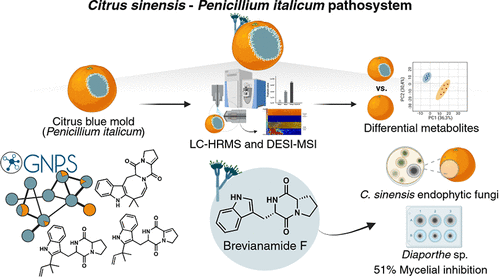
Figure 1
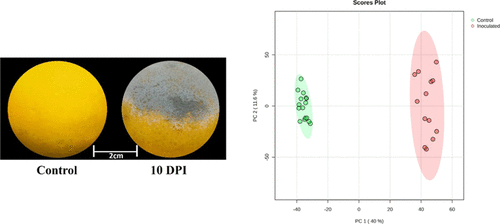
Figure 1. Metabolic changes in oranges infected with P. italicum at 10 days post-inoculation. (Left) Representative images of healthy (control) andP. italicum-infected oranges at 10 days post-inoculation, showing extensive fungal colonization. Scale: 2 cm. (Right) PCA of metabolic profiles from control (green) and infected (red) fruit samples at 10 days post-inoculation. A clear separation is observed between the two groups, with PC1 accounting for 40.0% and PC2 for 11.6% of the total variance, indicating significant infection-induced metabolic reprogramming.
Figure 2

Figure 2. Molecular network cluster of diketopiperazine alkaloids identified inP. italicum-infected oranges. Blue nodes represent metabolites exclusively detected in infected samples. Nodes with a polygonal shape indicate spectral matches with GNPS library compounds. Several interconnected nodes form a cluster with high cosine similarity, suggesting structurally related diketopiperazines. Annotated compounds include brevianamide F, deoxybrevianamide E, dehydrodeoxybrevianamide E, deoxyisoaustamide, and 12,13-dehydroprolyltryptophanyldiketopiperazine. This cluster highlights the metabolic specialization associated withP. italicum infection. Edge thickness corresponds to the cosine score, with thicker edges indicating higher spectral similarity.
Figure 3

Figure 3. Spatial distribution of brevianamide F (18) and deoxyisoaustamide (19) in P. italicum-infected oranges using DESI-MSI. Ion images show metabolite localization in control and infected samples at 6 days post-inoculation. The respective m/z values were not detected in the control orange peels, while strong signals were observed in infected tissues, particularly at the fungal infection site. The color scale indicates relative ion intensity (from low in blue to high in red). Observed mass errors are −0.39 ppm for brevianamide F and 3.87 ppm for deoxyisoaustamide.
Figure 4
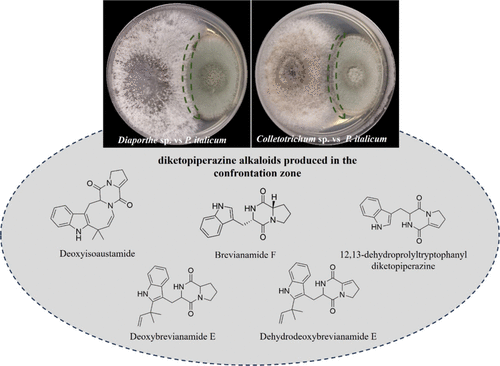
Figure 4. Confrontation assay betweenP. italicum and endophytic fungi. (Upper): coculture assays showing the interaction zones betweenP. italicum andDiaporthe sp. (left) orP. italicum andColletotrichum sp. (right). Dashed green lines highlight the confrontation zone. (Lower): diketopiperazine alkaloids detected in the confrontation zone through LC–HRMS analysis and structure annotation: deoxyisoaustamide, brevianamide F, 12,13-dehydroprolyltryptophanyldiketopiperazine, deoxybrevianamide E, and dehydrodeoxybrevianamide E.
Figure 5

Figure 5. Antifungal activity of secondary metabolites againstDiaporthe sp. (Left): mycelial growth of Diaporthe sp. on PDA plates after 7 days of incubation under four treatments: control (PDA only), negative control (ethanol/water 1:1, v/v), brevianamide F (0.3 mg/mL), and imazalil (0.3 mg/mL, positive control). (Right, upper): quantification of fungal growth area (cm2). Data represent mean ± standard deviation (n = 3). Statistical analysis was performed using one-way ANOVA (p < 0.0001), followed by Tukey’s post hoc test. Treatments labeled with the same letter are not significantly different, while those with different letters indicate statistically significant differences. (Right, lower): chemical structures of brevianamide F and imazalil used in the treatments. Brevianamide F inhibited mycelial growth by 51.92% compared to the control.
Figure 6

Figure 6. Effect of brevianamide F on fungal hyphal morphology. Representative images ofDiaporthe sp. grown on PDA plates supplemented with brevianamide F (upper row) or control PDA (lower row). Left panels show colony morphology, while middle and right panels display confocal microscopy images of the hyphal structure stained with a fluorescent dye. Treatment with brevianamide F results in notable alterations in hyphal organization and density compared to the untreated control.
References
This article references 71 other publications.
- 1Wang, Z.; Sui, Y.; Li, J.; Tian, X.; Wang, Q. Biological control of postharvest fungal decays in citrus: a review. Crit. Rev. Food Sci. Nutr. 2022, 62 (4), 861– 870, DOI: 10.1080/10408398.2020.1829542There is no corresponding record for this reference.
- 2François, G. A.; de Moraes Pontes, J. G.; Pereira, A. K.; Fill, T. P. Exploring the citrus sour rot pathogen: biochemical aspects, virulence factors, and strategies for disease management – a review. Fungal Biol. Rev. 2022, 41, 70– 83, DOI: 10.1016/j.fbr.2022.03.003There is no corresponding record for this reference.
- 3Bhatta, U. K. Alternative management approaches of citrus diseases caused by Penicillium digitatum (green mold) and Penicillium italicum (blue mold). Front. Plant Sci. 2022, 12, 833328, DOI: 10.3389/fpls.2021.833328There is no corresponding record for this reference.
- 4Khamsaw, P.; Sangta, J.; Chaiwan, P.; Rachtanapun, P.; Sirilun, S.; Sringarm, K.; Thanakkasaranee, S.; Sommano, S. R. Bio-circular perspective of citrus fruit loss caused by pathogens: occurrences, active ingredient recovery and applications. Horticulturae 2022, 8 (8), 748, DOI: 10.3390/horticulturae8080748There is no corresponding record for this reference.
- 5Moraes Bazioli, J.; Belinato, J. R.; Costa, J. H.; Akiyama, D. Y.; Pontes, J. G. d. M.; Kupper, K. C.; Augusto, F.; de Carvalho, J. E.; Fill, T. P. Biological control of citrus postharvest phytopathogens. Toxins 2019, 11 (8), 460, DOI: 10.3390/toxins11080460There is no corresponding record for this reference.
- 6Costa, J. H.; Bazioli, J. M.; de Moraes Pontes, J. G.; Fill, T. P. Penicillium digitatum infection mechanisms in citrus: what do we know so far?. Fungal Biol. 2019, 123 (8), 584– 593, DOI: 10.1016/j.funbio.2019.05.004There is no corresponding record for this reference.
- 7Kanashiro, A. M.; Akiyama, D. Y.; Kupper, K. C.; Fill, T. P. Penicillium italicum: an underexplored postharvest pathogen. Front. Microbiol. 2020, 11, 606852, DOI: 10.3389/fmicb.2020.606852There is no corresponding record for this reference.
- 8Plaza, P.; Usall, J.; Teixidó, N.; Viñas, I. Effect of water activity and temperature on germination and growth of Penicillium digitatum, Penicillium italicum and Geotrichum candidum. J. Appl. Microbiol. 2003, 94 (4), 549– 554, DOI: 10.1046/j.1365-2672.2003.01909.xThere is no corresponding record for this reference.
- 9Latifa, A.; Idriss, T.; Hassan, B.; Amine, S. M.; El Hassane, B.; Aoumar Abd, A. B. Effects of organic acids and salts on the development of Penicillium italicum: the causal agent of citrus blue mold. Plant Pathol. J. 2011, 10 (3), 99– 107, DOI: 10.3923/ppj.2011.99.107There is no corresponding record for this reference.
- 10Li, X.; Yang, S.; Zhang, M.; Yang, Y.; Peng, L. Identification of pathogenicity-related effector proteins and the role of Piwsc1 in the virulence of Penicillium italicum on citrus fruits. J. Fungi 2022, 8 (6), 646, DOI: 10.3390/jof8060646There is no corresponding record for this reference.
- 11Hershenhorn, J. Polygalacturonases associated with infection of Valencia orange by Penicillium italicum. Phytopathology 1990, 80 (12), 1374, DOI: 10.1094/Phyto-80-1374There is no corresponding record for this reference.
- 12Ballester, A.-R.; Marcet-Houben, M.; Levin, E.; Sela, N.; Selma-Lázaro, C.; Carmona, L.; Wisniewski, M.; Droby, S.; González-Candelas, L.; Gabaldón, T. Genome, transcriptome, and functional analyses of Penicillium expansum provide new insights into secondary metabolism and pathogenicity. Mol. Plant-Microbe Interact. 2015, 28 (3), 232– 248, DOI: 10.1094/MPMI-09-14-0261-FIThere is no corresponding record for this reference.
- 13Ropars, J.; de la Vega, R. C. R.; López-Villavicencio, M.; Gouzy, J.; Dupont, J.; Swennen, D.; Dumas, E.; Giraud, T.; Branca, A. Diversity and mechanisms of genomic adaptation in Penicillium. Aspergillus and Penicillium in the Post-genomic Era; Caister Academic Press: Norfolk, UK, 2016; pp 27– 42.There is no corresponding record for this reference.
- 14Błaszczyk, U.; Wyrzykowska, S.; Gąstoł, M. Application of bioactive coatings with killer yeasts to control post-harvest apple decay caused by Botrytis cinerea and Penicillium italicum. Foods 2022, 11 (13), 1868, DOI: 10.3390/foods11131868There is no corresponding record for this reference.
- 15Chen, C.; Peng, X.; Wan, C.; Zhang, Y.; Gan, Z.; Zeng, J.; Kai, W.; Chen, J. Lignin biosynthesis pathway and redox balance act synergistically in conferring resistance against Penicillium italicum infection in 7-demethoxytylophorine-treated Navel orange. J. Agric. Food Chem. 2022, 70 (26), 8111– 8123, DOI: 10.1021/acs.jafc.2c02348There is no corresponding record for this reference.
- 16Kahramanoğlu, I. ̇.; Nisar, M. F.; Chen, C.; Usanmaz, S.; Chen, J.; Wan, C. Light: an alternative method for physical control of postharvest rotting caused by fungi of citrus fruit. J. Food Qual. 2020, 2020, 1– 12, DOI: 10.1155/2020/8821346There is no corresponding record for this reference.
- 17Jeong, M.-A.; Jeong, R.-D. Applications of ionizing radiation for the control of postharvest diseases in fresh produce: recent advances. Plant Pathol. 2018, 67 (1), 18– 29, DOI: 10.1111/ppa.12739There is no corresponding record for this reference.
- 18Trabelsi, D.; Hamdane, A. M.; Said, M. B.; Abdrrabba, M. Chemical composition and antifungal activity of essential oils from flowers, leaves and peels of Tunisian Citrus aurantium against Penicillium digitatum and Penicillium italicum. J. Essent. Oil Bear. Plants 2016, 19 (7), 1660– 1674, DOI: 10.1080/0972060X.2016.1141069There is no corresponding record for this reference.
- 19Vázquez-González, Y.; Ragazzo-Sánchez, J. A.; Calderón-Santoyo, M. Characterization and antifungal activity of jackfruit (Artocarpus heterophyllus Lam.) leaf extract obtained using conventional and emerging technologies. Food Chem. 2020, 330, 127211, DOI: 10.1016/j.foodchem.2020.127211There is no corresponding record for this reference.
- 20Osonga, F. J.; Eshun, G.; Kalra, S.; Yazgan, I.; Sakhaee, L.; Ontman, R.; Jiang, S.; Sadik, O. A. Influence of particle size and shapes on the antifungal activities of greener nanostructured copper against Penicillium italicum. ACS Agric. Sci. Technol. 2022, 2 (1), 42– 56, DOI: 10.1021/acsagscitech.1c00102There is no corresponding record for this reference.
- 21Papoutsis, K.; Mathioudakis, M. M.; Hasperué, J. H.; Ziogas, V. Non-chemical treatments for preventing the postharvest fungal rotting of citrus caused by Penicillium digitatum (green mold) and Penicillium italicum (blue mold). Trends Food Sci. Technol. 2019, 86, 479– 491, DOI: 10.1016/j.tifs.2019.02.053There is no corresponding record for this reference.
- 22Talibi, I.; Boubaker, H.; Boudyach, E. H.; Ait Ben Aoumar, A. Alternative methods for the control of postharvest citrus diseases. J. Appl. Microbiol. 2014, 117 (1), 1– 17, DOI: 10.1111/jam.12495There is no corresponding record for this reference.
- 23Pontes, J. G. M.; Fernandes, L. S.; dos Santos, R. V.; Tasic, L.; Fill, T. P. Virulence factors in the phytopathogen–host interactions: an overview. J. Agric. Food Chem. 2020, 68 (29), 7555– 7570, DOI: 10.1021/acs.jafc.0c02389There is no corresponding record for this reference.
- 24Silva, E.; Dantas, R.; Barbosa, J. C.; Berlinck, R. G. S.; Fill, T. Metabolomics approach to understand molecular mechanisms involved in fungal pathogen–citrus pathosystems. Mol. Omics 2024, 20 (3), 154– 168, DOI: 10.1039/D3MO00182BThere is no corresponding record for this reference.
- 25Adusumilli, R.; Mallick, P. Data conversion with ProteoWizard MSConvert. Computational Methods for Mass Spectrometry Proteomics; Hamid, J., Ed.; Humana Press: New York, NY, 2017; pp 339– 368.There is no corresponding record for this reference.
- 26Wang, M.; Carver, J. J.; Phelan, V. V.; Sanchez, L. M.; Garg, N.; Peng, Y.; Nguyen, D. D.; Watrous, J.; Kapono, C. A.; Luzzatto-Knaan, T.; Porto, C.; Bouslimani, A.; Melnik, A. V.; Meehan, M. J.; Liu, W.-T.; Crüsemann, M.; Boudreau, P. D.; Esquenazi, E.; Sandoval-Calderón, M.; Kersten, R. D.; Pace, L. A.; Quinn, R. A.; Duncan, K. R.; Hsu, C.-C.; Floros, D. J.; Gavilan, R. G.; Kleigrewe, K.; Northen, T.; Dutton, R. J.; Parrot, D.; Carlson, E. E.; Aigle, B.; Michelsen, C. F.; Jelsbak, L.; Sohlenkamp, C.; Pevzner, P.; Edlund, A.; McLean, J.; Piel, J.; Murphy, B. T.; Gerwick, L.; Liaw, C.-C.; Yang, Y.-L.; Humpf, H.-U.; Maansson, M.; Keyzers, R. A.; Sims, A. C.; Johnson, A. R.; Sidebottom, A. M.; Sedio, B. E.; Klitgaard, A.; Larson, C. B.; Boya P, C. A.; Torres-Mendoza, D.; Gonzalez, D. J.; Silva, D. B.; Marques, L. M.; Demarque, D. P.; Pociute, E.; O’Neill, E. C.; Briand, E.; Helfrich, E. J. N.; Granatosky, E. A.; Glukhov, E.; Ryffel, F.; Houson, H.; Mohimani, H.; Kharbush, J. J.; Zeng, Y.; Vorholt, J. A.; Kurita, K. L.; Charusanti, P.; McPhail, K. L.; Nielsen, K. F.; Vuong, L.; Elfeki, M.; Traxler, M. F.; Engene, N.; Koyama, N.; Vining, O. B.; Baric, R.; Silva, R. R.; Mascuch, S. J.; Tomasi, S.; Jenkins, S.; Macherla, V.; Hoffman, T.; Agarwal, V.; Williams, P. G.; Dai, J.; Neupane, R.; Gurr, J.; Rodríguez, A. M. C.; Lamsa, A.; Zhang, C.; Dorrestein, K.; Duggan, B. M.; Almaliti, J.; Allard, P.-M.; Phapale, P.; Nothias, L.-F.; Alexandrov, T.; Litaudon, M.; Wolfender, J.-L.; Kyle, J. E.; Metz, T. O.; Peryea, T.; Nguyen, D.-T.; VanLeer, D.; Shinn, P.; Jadhav, A.; Müller, R.; Waters, K. M.; Shi, W.; Liu, X.; Zhang, L.; Knight, R.; Jensen, P. R.; Palsson, B. Ø.; Pogliano, K.; Linington, R. G.; Gutiérrez, M.; Lopes, N. P.; Gerwick, W. H.; Moore, B. S.; Dorrestein, P. C.; Bandeira, N. Sharing and Community Curation of Mass Spectrometry Data with Global Natural Products Social Molecular Networking. Nat. Biotechnol. 2016, 34 (8), 828– 837, DOI: 10.1038/nbt.3597There is no corresponding record for this reference.
- 27Shannon, P.; Markiel, A.; Ozier, O.; Baliga, N. S.; Wang, J. T.; Ramage, D.; Amin, N.; Schwikowski, B.; Ideker, T. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 2003, 13 (11), 2498– 2504, DOI: 10.1101/gr.1239303There is no corresponding record for this reference.
- 28Kieffer, M. E.; Chuang, K. V.; Reisman, S. E. Copper-catalyzed diastereoselective arylation of tryptophan derivatives: total synthesis of (+)-naseseazines A and B. J. Am. Chem. Soc. 2013, 135 (15), 5557– 5560, DOI: 10.1021/ja4023557There is no corresponding record for this reference.
- 29Godinho, B. T. V.; Santos, I. ́. A. F. M.; Gomes, E. A.; Piccoli, R. H.; Cardoso, P. G. Endophytic fungi community in Eremanthus erythropappus tree from anthropogenic and natural areas of Minas Gerais. Cerne 2019, 25 (3), 283– 293, DOI: 10.1590/01047760201925032642There is no corresponding record for this reference.
- 30Aamir, S. A rapid and efficient method of fungal genomic DNA extraction, suitable for PCR-based molecular methods. Plant Pathol. Quar. 2015, 5 (2), 74– 81, DOI: 10.5943/ppq/5/2/6There is no corresponding record for this reference.
- 31Thompson, J. The CLUSTAL_X Windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res. 1997, 25 (24), 4876– 4882, DOI: 10.1093/nar/25.24.4876There is no corresponding record for this reference.
- 32Tamura, K.; Stecher, G.; Peterson, D.; Filipski, A.; Kumar, S. MEGA6: molecular evolutionary genetics analysis version 6.0. Mol. Biol. Evol. 2013, 30 (12), 2725– 2729, DOI: 10.1093/molbev/mst197There is no corresponding record for this reference.
- 33Kimura, M. A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J. Mol. Evol. 1980, 16 (2), 111– 120, DOI: 10.1007/BF01731581There is no corresponding record for this reference.
- 34Saitou, N.; Nei, M. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 1987, 4 (4), 406– 425, DOI: 10.1093/oxfordjournals.molbev.a040454There is no corresponding record for this reference.
- 35Costa, J. H.; Wassano, C. I.; Angolini, C. F. F.; Scherlach, K.; Hertweck, C.; Pacheco Fill, T. Antifungal potential of secondary metabolites involved in the interaction between citrus pathogens. Sci. Rep. 2019, 9 (1), 18647, DOI: 10.1038/s41598-019-55204-9There is no corresponding record for this reference.
- 36Xu, W.-F.; Mao, N.; Xue, X.-J.; Qi, Y.-X.; Wei, M.-Y.; Wang, C.-Y.; Shao, C.-L. Structures and absolute configurations of diketopiperazine alkaloids chrysopiperazines A–C from the gorgonian-derived Penicillium chrysogenum fungus. Mar. Drugs 2019, 17 (5), 250, DOI: 10.3390/md17050250There is no corresponding record for this reference.
- 37Caballero, E.; Avendaño, C.; Menéndez, J. C. Brief total synthesis of the cell cycle inhibitor tryprostatin B and related preparation of its alanine analogue. J. Org. Chem. 2003, 68 (18), 6944– 6951, DOI: 10.1021/jo034703lThere is no corresponding record for this reference.
- 38Hawkins, J. P.; Geddes, B. A.; Oresnik, I. J. Common dyes used to determine bacterial polysaccharides on agar are affected by medium acidification. Can. J. Microbiol. 2017, 63 (6), 559– 562, DOI: 10.1139/cjm-2016-0743There is no corresponding record for this reference.
- 39Costa, J. H.; Bazioli, J. M.; de Vilhena Araújo, E.; Vendramini, P. H.; de Freitas Porto, M. C.; Eberlin, M. N.; Souza-Neto, J. A.; Fill, T. P. Monitoring indole alkaloid production by Penicillium digitatum during infection process in citrus by mass spectrometry imaging and molecular networking. Fungal Biol. 2019, 123 (8), 594– 600, DOI: 10.1016/j.funbio.2019.03.002There is no corresponding record for this reference.
- 40Silva, E.; Belinato, J. R.; Porto, C.; Nunes, E.; Guimarães, F.; Meyer, M. C.; Pilau, E. J. Soybean metabolomics based in mass spectrometry: decoding the plant’s signaling and defense responses under biotic stress. J. Agric. Food Chem. 2021, 69 (26), 7257– 7267, DOI: 10.1021/acs.jafc.0c07758There is no corresponding record for this reference.
- 41Liu, X.; Lin, C.; Ma, X.; Tan, Y.; Wang, J.; Zeng, M. Functional characterization of a flavonoid glycosyltransferase in sweet orange (Citrus sinensis). Front. Plant Sci. 2018, 9, 166, DOI: 10.3389/fpls.2018.00166There is no corresponding record for this reference.
- 42Nogata, Y.; Sakamoto, K.; Shiratsuchi, H.; Ishii, T.; Yano, M.; Ohta, H. Flavonoid composition of fruit tissues of Citrus species. Biosci., Biotechnol., Biochem. 2006, 70 (1), 178– 192, DOI: 10.1271/bbb.70.178There is no corresponding record for this reference.
- 43Gattuso, G.; Barreca, D.; Gargiulli, C.; Leuzzi, U.; Caristi, C. Flavonoid composition of citrus juices. Molecules 2007, 12 (8), 1641– 1673, DOI: 10.3390/12081641There is no corresponding record for this reference.
- 44Dixon, R. A.; Paiva, N. L. Stress-induced phenylpropanoid metabolism. Plant Cell 1995, 7 (7), 1085– 1097, DOI: 10.1105/tpc.7.7.1085There is no corresponding record for this reference.
- 45Zhao, S.; Nan, Y.; Yao, R.; Wang, L.; Zeng, X.; Aadil, R. M.; Shabbir, M. A. Antibacterial activity and transcriptomic analysis of hesperetin against Alicyclobacillus acidoterrestris vegetative cells. Foods 2023, 12 (17), 3276, DOI: 10.3390/foods12173276There is no corresponding record for this reference.
- 46Ferrer, J.-L.; Austin, M. B.; Stewart, C.; Noel, J. P. Structure and function of enzymes involved in the biosynthesis of phenylpropanoids. Plant Physiol. Biochem. 2008, 46 (3), 356– 370, DOI: 10.1016/j.plaphy.2007.12.009There is no corresponding record for this reference.
- 47Treutter, D. Significance of flavonoids in plant resistance: a review. Environ. Chem. Lett. 2006, 4 (3), 147– 157, DOI: 10.1007/s10311-006-0068-8There is no corresponding record for this reference.
- 48Agati, G.; Azzarello, E.; Pollastri, S.; Tattini, M. Flavonoids as antioxidants in plants: location and functional significance. Plant Sci. 2012, 196, 67– 76, DOI: 10.1016/j.plantsci.2012.07.014There is no corresponding record for this reference.
- 49Nehela, Y.; Killiny, N. The unknown soldier in citrus plants: polyamines-based defensive mechanisms against biotic and abiotic stresses and their relationship with other stress-associated metabolites. Plant Signaling Behav. 2020, 15 (6), 1761080, DOI: 10.1080/15592324.2020.1761080There is no corresponding record for this reference.
- 50Kyselka, J.; Bleha, R.; Dragoun, M.; Bialasová, K.; Horáčková, S. ̌.; Schätz, M.; Sluková, M.; Filip, V.; Synytsya, A. Antifungal polyamides of hydroxycinnamic acids from sunflower bee pollen. J. Agric. Food Chem. 2018, 66 (42), 11018– 11026, DOI: 10.1021/acs.jafc.8b03976There is no corresponding record for this reference.
- 51Khan, M. K.; Zill-E-Huma; Dangles, O. A comprehensive review on flavanones, the major citrus polyphenols. J. Food Compos. Anal. 2014, 33 (1), 85– 104, DOI: 10.1016/j.jfca.2013.11.004There is no corresponding record for this reference.
- 52Xu, X.; Pu, R.; Li, Y.; Wu, Z.; Li, C.; Miao, X.; Yang, W. Chemical compositions of propolis from China and the United States and their antimicrobial activities against Penicillium notatum. Molecules 2019, 24 (19), 3576, DOI: 10.3390/molecules24193576There is no corresponding record for this reference.
- 53Yang, S. Z.; Peng, L. T.; Su, X. J.; Chen, F.; Cheng, Y. J.; Fan, G.; Pan, S. Y. Bioassay-guided isolation and identification of antifungal components from propolis against Penicillium italicum. Food Chem. 2011, 127 (1), 210– 215, DOI: 10.1016/j.foodchem.2010.12.011There is no corresponding record for this reference.
- 54Huo, R.; Zhang, J.; Niu, S.; Liu, L. New prenylated indole diketopiperazine alkaloids and polyketides from the mangrove-derived fungus Penicillium sp. Front. Mar. Sci. 2022, 9, 1097594, DOI: 10.3389/fmars.2022.1097594There is no corresponding record for this reference.
- 55Ma, Y.-M.; Liang, X.-A.; Kong, Y.; Jia, B. Structural diversity and biological activities of indole diketopiperazine alkaloids from fungi. J. Agric. Food Chem. 2016, 64 (35), 6659– 6671, DOI: 10.1021/acs.jafc.6b01772There is no corresponding record for this reference.
- 56Kozlovsky, A. G.; Vinokurova, N. G.; Adanin, V. M.; Burkhardt, G.; Dahse, H.-M.; Gräfe, U. New diketopiperazine alkaloids from Penicillium fellutanum. J. Nat. Prod. 2000, 63 (5), 698– 700, DOI: 10.1021/np9903853There is no corresponding record for this reference.
- 57Bräse, S.; Encinas, A.; Keck, J.; Nising, C. F. Chemistry and biology of mycotoxins and related fungal metabolites. Chem. Rev. 2009, 109 (9), 3903– 3990, DOI: 10.1021/cr050001fThere is no corresponding record for this reference.
- 58Asiri, I. A. M.; Badr, J. M.; Youssef, D. T. A. Penicillivinacine, antimigratory diketopiperazine alkaloid from the marine-derived fungus Penicillium vinaceum. Phytochem. Lett. 2015, 13, 53– 58, DOI: 10.1016/j.phytol.2015.05.014There is no corresponding record for this reference.
- 59Liu, Y.-X.; Ma, S.-G.; Wang, X.-J.; Zhao, N.; Qu, J.; Yu, S.-S.; Dai, J.-G.; Wang, Y.-H.; Si, Y.-K. Diketopiperazine alkaloids produced by the endophytic fungus Aspergillus fumigatus from the stem of Erythrophloeum fordii Oliv. Helv. Chim. Acta 2012, 95 (8), 1401– 1408, DOI: 10.1002/hlca.201100521There is no corresponding record for this reference.
- 60Danial, A. M.; Medina, A.; Sulyok, M.; Magan, N. Efficacy of metabolites of a Streptomyces strain (AS1) to control growth and mycotoxin production by Penicillium verrucosum, Fusarium verticillioides, and Aspergillus fumigatus in culture. Mycotoxin Res. 2020, 36 (2), 225– 234, DOI: 10.1007/s12550-020-00388-7There is no corresponding record for this reference.
- 61Hubka, V.; Nováková, A.; Jurjević, Z. ̌.; Sklenář, F.; Frisvad, J. C.; Houbraken, J.; Arendrup, M. C.; Jørgensen, K. M.; Siqueira, J. P. Z.; Gené, J.; Kolařík, M. Polyphasic data support the splitting of Aspergillus candidus into two species; proposal of Aspergillus dobrogensis sp. nov. Int. J. Syst. Evol. Microbiol. 2018, 68 (4), 995– 1011, DOI: 10.1099/ijsem.0.002583There is no corresponding record for this reference.
- 62Frisvad, J. C.; Samson, R. A. Polyphasic taxonomy of Penicillium subgenus Penicillium: a guide to identification of food and air-borne terverticillate penicillia and their mycotoxins. Stud. Mycol. 2004, 49 (1), 1– 174There is no corresponding record for this reference.
- 63Scott, P. M.; Kennedy, B. P. C.; Harwig, J.; Chen, Y.-K. Formation of Diketopiperazines by Penicillium italicum Isolated from Oranges. Appl. Microbiol. 1974, 28 (5), 892– 894, DOI: 10.1128/am.28.5.892-894.1974There is no corresponding record for this reference.
- 64Baek, S. Y.; Shim, S. H. Isolation and structure determination of metabolites from cultures of Aspergillus protuberus. Korean J. Microbiol. 2013, 49 (2), 179– 183, DOI: 10.7845/kjm.2013.3026There is no corresponding record for this reference.
- 65Godfrey, R. C.; Green, N. J.; Nichol, G. S.; Lawrence, A. L. Total synthesis of brevianamide A. Nat. Chem. 2020, 12 (7), 615– 619, DOI: 10.1038/s41557-020-0442-3There is no corresponding record for this reference.
- 66Sakhri, A.; Chaouche, N. K.; Catania, M. R.; Ritieni, A.; Santini, A. Chemical composition of Aspergillus creber extract and evaluation of its antimicrobial and antioxidant activities. Pol. J. Microbiol. 2019, 68 (3), 309– 316, DOI: 10.33073/pjm-2019-033There is no corresponding record for this reference.
- 67Lee, S.; Sperry, J. Isolation and biological activity of azocine and azocane alkaloids. Bioorg. Med. Chem. 2022, 54, 116560, DOI: 10.1016/j.bmc.2021.116560There is no corresponding record for this reference.
- 68Quang, T. H.; Lee, D.-S.; Sohn, J. H.; Kim, Y.-C.; Oh, H. New deoxyisoaustamide derivative from the marine-derived fungus Penicillium sp. JF-72. Bull. Korean Chem. Soc. 2013, 34 (10), 3109– 3112, DOI: 10.5012/bkcs.2013.34.10.3109There is no corresponding record for this reference.
- 69Zhuravleva, O. I.; Antonov, A. S.; Trang, V. T. D.; Pivkin, M. V.; Khudyakova, Y. V.; Denisenko, V. A.; Popov, R. S.; Kim, N. Y.; Yurchenko, E. A.; Gerasimenko, A. V.; Udovenko, A. A.; Amsberg, G. v.; Dyshlovoy, S. A.; Afiyatullov, S. S. New deoxyisoaustamide derivatives from the coral-derived fungus Penicillium dimorphosporum KMM 4689. Mar. Drugs 2021, 19 (1), 32, DOI: 10.3390/md19010032There is no corresponding record for this reference.
- 70Coleman, J. J.; Mylonakis, E. Efflux in Fungi: La Pièce de Résistance. PLoS Pathog. 2009, 5 (6), e1000486 DOI: 10.1371/journal.ppat.1000486There is no corresponding record for this reference.
- 71Chen, X.; Pan, H.-X.; Tang, G.-L. Newly Discovered Mechanisms of Antibiotic Self-Resistance with Multiple Enzymes Acting at Different Locations and Stages. Antibiotics 2023, 12 (1), 35, DOI: 10.3390/antibiotics12010035There is no corresponding record for this reference.
Supporting Information
Supporting Information
The Supporting Information is available free of charge at https://pubs.acs.org/doi/10.1021/acs.jafc.5c07618.
Synthesis and NMR analysis of brevianamide F; LC–MS TICs of orange peel extracts; PCA and PLS-DA plots of metabolomic data; MS/MS molecular network showing metabolite differences between healthy and infected samples; in vitro LC–MS/MS confirmation of diketopiperazines; FDAA derivatization HPLC-UV chromatograms for proline stereochemistry; FDAA derivatization HPLC-UV chromatograms for tryptophan stereochemistry; EICs for d- and l-tryptophan FDAA derivatives and brevianamide F; dual-culture assays with endophytic fungi; ITS sequences of isolates; phylogenetic dendrograms of Colletotrichum sp. and Diaporthe sp. isolates; and MIC assay of brevianamide F against Diaporthe sp. (PDF)
Terms & Conditions
Most electronic Supporting Information files are available without a subscription to ACS Web Editions. Such files may be downloaded by article for research use (if there is a public use license linked to the relevant article, that license may permit other uses). Permission may be obtained from ACS for other uses through requests via the RightsLink permission system: http://pubs.acs.org/page/copyright/permissions.html.


