
Unveiling the Activation Mechanism of Glucagon-Like Peptide-1 Receptor by an Ago-Allosteric Modulator via Molecular Dynamics SimulationsClick to copy article linkArticle link copied!
- Yue Chen*Yue Chen*Email: [email protected]School of Biological Sciences, Nanyang Technological University, 60 Nanyang Drive, Singapore 637551, SingaporeMore by Yue Chen
- Junhao LiJunhao LiViva Biotech (Shanghai) Limited, 735 Ziping Road, Pudong New District, Shanghai 201318, P. R. ChinaMore by Junhao Li
- Lucie DelemotteLucie DelemotteScience for Life Laboratory, Department of Applied Physics, KTH Royal Institute of Technology, Stockholm SE-121 21, SwedenMore by Lucie Delemotte
- Yuguang Mu*Yuguang Mu*Email: [email protected]School of Biological Sciences, Nanyang Technological University, 60 Nanyang Drive, Singapore 637551, SingaporeMore by Yuguang Mu
Abstract
The glucagon-like peptide-1 receptor (GLP-1R) is a key therapeutic target for metabolic disorders, particularly type 2 diabetes and obesity. Although current treatments are effective, their unavoidable side effects continue to drive the search for novel therapeutic strategies. Ago-allosteric modulators (ago-PAMs), which act as agonists on their own while enhancing the affinity and efficacy of orthosteric agonists, represent a promising avenue to overcome limitations associated with traditional peptide-based therapies. However, the molecular mechanisms by which ago-PAMs modulate GLP-1R activation remain poorly understood. In this work, we selected compound 2, a validated ago-PAM of GLP-1R, as a probe to explore these mechanisms at the atomic level. Using molecular dynamics (MD) simulations, we elucidate how compound 2 stabilizes the active conformation of GLP-1R through allosteric binding and reveal distinct pathways by which it enhances the binding of both peptide and non-peptide orthosteric agonists. Enhanced sampling simulations further provided a comprehensive conformational landscape of GLP-1R activation, identifying two intermediate states that bridge inactive and active conformations. Compound 2 was found to bias the receptor toward active-like ensembles, consistent with its intrinsic agonist activity. Together, our findings provide mechanistic insights into ago-allosteric modulation of GLP-1R, offering useful information for the rational design of small-molecule modulators with improved therapeutic profiles.
This publication is licensed for personal use by The American Chemical Society.
Introduction
Results and Discussion
Compound 2 Binding Stabilizes the Active Conformation of GLP-1R
Figure 1
Figure 1. (A) Overall structure of GLP-1R shown in purple cartoon bound to compound 2 (brown sticks) (PDB ID: 7DUR (28)). (B) TM6 movement during 2-μs production simulations of apo (green) and compound 2-bound (orange) GLP-1R, quantified by the Cα distance between Y2503.53 and K3466.35. Representative snapshots sampled from each trajectory are superimposed onto the starting structure (purple) for comparison. (C) Residue-wise importance profiles derived from RF analysis highlight structural differences between apo and compound 2-bound GLP-1R conformations. (D) Distribution of significant structural features distinguishing apo and compound 2-bound GLP-1R states. Representative active cryo-EM structures of GLP-1R includes 7DUR, 7DUQ, 7E14. (28)
Binding Mode and Covalent Interaction of Compound 2 with GLP-1R
Figure 2
Figure 2. (A) Compound 2-binding mode at the GLP-1R. (B) Changes in interaction frequencies between compound 2 and nearby residues at the allosteric site. (C) Computed energy profiles for the reaction between Cys3476.36 and compound 2. Insets illustrate the key features of the reactant, transition, and product states for two possible pathways: one with Cys3476.36 in its protonated form and the other in its deprotonated form.
Compound 2 Enhances the Binding of Orthosteric Ligands via Allosteric Synergy
Figure 3
Figure 3. Compound 2 enhances the binding of GLP-1 or LY3502970 at the orthosteric site. (A) Overall structure of the compound 2-GLP-1-GLP-1R complex, highlighting the GLP-1 binding mode. (B) Changes in interaction frequencies between GLP-1 and nearby residues with and without compound 2. Solid bars represent the apo system, while dashed bars indicate the compound 2-bound system. (C) Overall structure of the compound 2-LY3502970-GLP-1R complex, highlighting the LY3502970 binding mode. (D) Changes in interaction frequencies between LY3502970 and nearby residues with and without compound 2. Solid bars represent the apo system, while dashed bars indicate the compound 2-bound system.
Dynamic Allosteric Networks Reveal Ligand-Specific Signaling in GLP-1R
Figure 4
Figure 4. Allosteric signal pathways from the intracellular to extracellular domains in GLP-1R. Optimal communication pathways were computed from C3476.36 (source node) to R36 of GLP-1 in the GLP-1-GLP-1R and compound 2-GLP-1-GLP-1R systems, and from C3476.36 to the centroid of LY3502970 in the LY3502970-GLP-1R and compound 2-LY3502970-GLP-1R systems. Residues involved in the allosteric pathways are shown as spheres and colored according to their community assignments. Connecting edges represent inter-residue communication, with line widths proportional to edge betweenness.
Activation Mechanism of GLP-1R Revealed by Enhanced Sampling Simulations
Figure 5
Figure 5. (A) Two-dimensional free energy landscapes of apo and compound 2-, GLP-1-, and LY3502970-bound GLP-1R systems, projected along the outward movement of TM6 (measured by the Cα distance between Y2503.35 and K3466.35) and the swing of the N-terminal α-helix (measured by the Cα distance between W33N-term and F3857.40). (B) Four representative conformations, inactive (cyan), intermediate 1 (I1, green), intermediate 2 (I2, pink), and active (orange), extracted from the string simulation trajectory of apo GLP-1R and superimposed on the inactive experimental structure (gray), corresponding to local energy minima in the free energy landscape.
Conclusions
Figure 6
Figure 6. Schematic illustration of ligand-mediated GLP-1R activation. In the absence of ligand, GLP-1R dynamically transitions between inactive and intermediate-like states. Binding of agonists to either the orthosteric or allosteric sites induces conformational rearrangements in both the extracellular and intracellular domains, promoting engagement of downstream transducers and driving the receptor toward its fully active state. SM: small molecule; PAM: positive allosteric modulator.
Methods
Simulation System Setup
Enhanced Sampling Simulation
Collective Variable Selection
String Method with Swarms of Trajectories
Free Energy Landscape Calculations
Quantum Chemical Calculations
Simulation Analysis
Demystifying
GetContacts
Dynamic Network Analysis
Data Availability
All data supporting the findings of this study are provided within the article and its supplementary information file. The source code used to perform the string method with swarms of trajectories is openly available at https://github.com/delemottelab/string-method-swarms-trajectories.
Supporting Information
The Supporting Information is available free of charge at https://pubs.acs.org/doi/10.1021/acs.jcim.6c00224.
Additional details on the simulation analysis of structural changes, comparison of ligand stability, transmembrane helix 6 displacement, sampling convergence estimation, energies from quantum chemical calculations, total simulation time for each system, and collective variables for enhanced sampling simulations in this study (PDF)
Terms & Conditions
Most electronic Supporting Information files are available without a subscription to ACS Web Editions. Such files may be downloaded by article for research use (if there is a public use license linked to the relevant article, that license may permit other uses). Permission may be obtained from ACS for other uses through requests via the RightsLink permission system: http://pubs.acs.org/page/copyright/permissions.html.
Acknowledgments
This work was supported by the Knut and Alice Wallenberg Foundation (2019.0130), the Science for Life Laboratory, and the Swedish Research Council (VR 2019-02433 and 2022-04305). The National Academic Infrastructure for Supercomputing in Sweden (NAISS) and in the Swedish Research Council through grant agreement no. 2022-06725 funded MD simulations.
References
This article references 79 other publications.
- 1Folli, F.; Finzi, G.; Manfrini, R.; Galli, A.; Casiraghi, F.; Centofanti, L.; Berra, C.; Fiorina, P.; Davalli, A.; La Rosa, S. Mechanisms of action of incretin receptor based dual- and tri-agonists in pancreatic islets. Am. J. Physiol. Endocrinol. Metab. 2023, 325 (5), E595–e609 DOI: 10.1152/ajpendo.00236.2023Google ScholarThere is no corresponding record for this reference.
- 2Drucker, D. J. GLP-1 physiology informs the pharmacotherapy of obesity. Mol. Metab. 2022, 57, 101351, DOI: 10.1016/j.molmet.2021.101351Google ScholarThere is no corresponding record for this reference.
- 3Gentilella, R.; Pechtner, V.; Corcos, A.; Consoli, A. Glucagon-like peptide-1 receptor agonists in type 2 diabetes treatment: Are they all the same?. Diabetes Metabolism Res. 2019, 35 (1), e3070 DOI: 10.1002/dmrr.3070Google ScholarThere is no corresponding record for this reference.
- 4Alavi, S. E.; Cabot, P. J.; Moyle, P. M. Glucagon-like peptide-1 receptor agonists and strategies to improve their efficiency. Mol. Pharmaceutics 2019, 16 (6), 2278– 2295, DOI: 10.1021/acs.molpharmaceut.9b00308Google ScholarThere is no corresponding record for this reference.
- 5Chen, B.; Yu, X.; Horvath-Diano, C.; Ortuño, M. J.; Tschöp, M. H.; Jastreboff, A. M.; Schneeberger, M. GLP-1 programs the neurovascular landscape. Cell Metab. 2024, 36 (10), P2173– 2189, DOI: 10.1016/j.cmet.2024.09.003Google ScholarThere is no corresponding record for this reference.
- 6Ussher, J. R.; Drucker, D. J. Glucagon-like peptide 1 receptor agonists: Cardiovascular benefits and mechanisms of action. Nat. Rev. Cardiol. 2023, 20 (7), 463– 474, DOI: 10.1038/s41569-023-00849-3Google ScholarThere is no corresponding record for this reference.
- 7Wong, C. K.; McLean, B. A.; Baggio, L. L.; Koehler, J. A.; Hammoud, R.; Rittig, N.; Yabut, J. M.; Seeley, R. J.; Brown, T. J.; Drucker, D. J. Central glucagon-like peptide 1 receptor activation inhibits Toll-like receptor agonist-induced inflammation. Cell Metab. 2024, 36 (1), P130– 143.E5, DOI: 10.1016/j.cmet.2023.11.009Google ScholarThere is no corresponding record for this reference.
- 8Sourris, K. C.; Ding, Y.; Maxwell, S. S.; Al-Sharea, A.; Kantharidis, P.; Mohan, M.; Rosado, C. J.; Penfold, S. A.; Haase, C.; Xu, Y. Glucagon-like peptide-1 receptor signaling modifies the extent of diabetic kidney disease through dampening the receptor for advanced glycation end products–induced inflammation. Kidney Int. 2024, 105 (1), 132– 149, DOI: 10.1016/j.kint.2023.09.029Google ScholarThere is no corresponding record for this reference.
- 9Zhou, Q.; Li, G.; Hang, K.; Li, J.; Yang, D.; Wang, M.-W. Weight Loss Blockbuster Development: A Role for Unimolecular Polypharmacology. Annu. Rev. Pharmacol. Toxicol. 2025, 65 (1), 191– 213, DOI: 10.1146/annurev-pharmtox-061324-011832Google ScholarThere is no corresponding record for this reference.
- 10Saraiva, J. F. K.; Franco, D. Oral GLP-1 analogue: Perspectives and impact on atherosclerosis in type 2 diabetic patients. Cardiovasc. Diabetol. 2021, 20 (1), 235, DOI: 10.1186/s12933-021-01417-0Google ScholarThere is no corresponding record for this reference.
- 11Meissner, W. G.; Remy, P.; Giordana, C.; Maltête, D.; Derkinderen, P.; Houéto, J. L.; Anheim, M.; Benatru, I.; Boraud, T.; Brefel-Courbon, C. Trial of Lixisenatide in Early Parkinson’s Disease. N. Engl. J. Med. 2024, 390, 1176– 1185, DOI: 10.1056/NEJMoa2312323Google ScholarThere is no corresponding record for this reference.
- 12Newsome, P. N.; Buchholtz, K.; Cusi, K.; Linder, M.; Okanoue, T.; Ratziu, V.; Sanyal, A. J.; Sejling, A.-S.; Harrison, S. A. A Placebo-Controlled Trial of Subcutaneous Semaglutide in Nonalcoholic Steatohepatitis. N. Engl. J. Med. 2021, 384 (12), 1113– 1124, DOI: 10.1056/NEJMoa2028395Google ScholarThere is no corresponding record for this reference.
- 13Mayo, K. E.; Miller, L. J.; Bataille, D.; Dalle, S.; Göke, B.; Thorens, B.; Drucker, D. J. International Union of Pharmacology. XXXV. The glucagon receptor family. Pharmacol. Rev. 2003, 55 (1), 167– 194, DOI: 10.1124/pr.55.1.6Google ScholarThere is no corresponding record for this reference.
- 14Zhang, Y.; Sun, B.; Feng, D.; Hu, H.; Chu, M.; Qu, Q.; Tarrasch, J. T.; Li, S.; Sun Kobilka, T.; Kobilka, B. K. Cryo-EM structure of the activated GLP-1 receptor in complex with a G protein. Nature 2017, 546 (7657), 248– 253, DOI: 10.1038/nature22394Google ScholarThere is no corresponding record for this reference.
- 15Zhou, Q.; Zhao, F.; Zhang, Y.; Yang, D.; Wang, M.-W. Structural pharmacology and mechanisms of GLP-1R signaling. Trends Pharmacol. Sci. 2025, 46 (5), P422– 436, DOI: 10.1016/j.tips.2025.03.003Google ScholarThere is no corresponding record for this reference.
- 16Secher, A.; Jelsing, J.; Baquero, A. F.; Hecksher-Sørensen, J.; Cowley, M. A.; Dalbøge, L. S.; Hansen, G.; Grove, K. L.; Pyke, C.; Raun, K. The arcuate nucleus mediates GLP-1 receptor agonist liraglutide-dependent weight loss. J. Clin. Invest. 2014, 124 (10), 4473– 4488, DOI: 10.1172/JCI75276Google ScholarThere is no corresponding record for this reference.
- 17Hall, S.; Isaacs, D.; Clements, J. N. Pharmacokinetics and clinical implications of semaglutide: A new glucagon-like peptide (GLP)-1 receptor agonist. Clin. Pharmacokinet. 2018, 57 (12), 1529– 1538, DOI: 10.1007/s40262-018-0668-zGoogle ScholarThere is no corresponding record for this reference.
- 18Neumiller, J. J. Incretin-based therapies. Med. Clin. 2015, 99, 107– 129, DOI: 10.1016/j.mcna.2014.08.013Google ScholarThere is no corresponding record for this reference.
- 19Buckley, S. T.; Bækdal, T. A.; Vegge, A.; Maarbjerg, S. J.; Pyke, C.; Ahnfelt-Rønne, J.; Madsen, K. G.; Schéele, S. G.; Alanentalo, T.; Kirk, R. K. Transcellular stomach absorption of a derivatized glucagon-like peptide-1 receptor agonist. Sci. Transl. Med. 2018, 10 (467), eaar7047 DOI: 10.1126/scitranslmed.aar7047Google ScholarThere is no corresponding record for this reference.
- 20Saxena, A. R.; Frias, J. P.; Brown, L. S.; Gorman, D. N.; Vasas, S.; Tsamandouras, N.; Birnbaum, M. J. Efficacy and safety of oral small molecule glucagon-like peptide 1 receptor agonist danuglipron for glycemic control among patients with type 2 diabetes: A randomized clinical trial. JAMA Netw. Open 2023, 6 (5), e2314493 DOI: 10.1001/jamanetworkopen.2023.14493Google ScholarThere is no corresponding record for this reference.
- 21Buckeridge, C.; Tsamandouras, N.; Carvajal-Gonzalez, S.; Brown, L. S.; Hernandez-Illas, M.; Saxena, A. R. Once-daily oral small-molecule glucagon-like peptide-1 receptor agonist lotiglipron (PF-07081532) for type 2 diabetes and obesity: Two randomized, placebo-controlled, multiple-ascending-dose phase 1 studies. Diabetes Obes. Metab. 2024, 26 (8), 3155– 3166, DOI: 10.1111/dom.15643Google ScholarThere is no corresponding record for this reference.
- 22Frias, J. P.; Hsia, S.; Eyde, S.; Liu, R.; Ma, X.; Konig, M.; Kazda, C.; Mather, K. J.; Haupt, A.; Pratt, E. Efficacy and safety of oral orforglipron in patients with type 2 diabetes: A multicentre, randomised, dose-response, phase 2 study. Lancet 2023, 402 (10400), P472– 483, DOI: 10.1016/S0140-6736(23)01302-8Google ScholarThere is no corresponding record for this reference.
- 23Smelcerovic, A.; Lazarevic, J.; Tomovic, K.; Anastasijevic, M.; Jukic, M.; Kocic, G.; Anderluh, M. An overview, advantages and therapeutic potential of nonpeptide positive allosteric modulators of glucagon-like peptide-1 receptor. ChemMedChem 2019, 14 (5), 514– 521, DOI: 10.1002/cmdc.201800699Google ScholarThere is no corresponding record for this reference.
- 24Bueno, A. B.; Sun, B.; Willard, F. S.; Feng, D.; Ho, J. D.; Wainscott, D. B.; Showalter, A. D.; Vieth, M.; Chen, Q.; Stutsman, C. Structural insights into probe-dependent positive allosterism of the GLP-1 receptor. Nat. Chem. Biol. 2020, 16 (10), 1105– 1110, DOI: 10.1038/s41589-020-0589-7Google ScholarThere is no corresponding record for this reference.
- 25Knudsen, L. B.; Kiel, D.; Teng, M.; Behrens, C.; Bhumralkar, D.; Kodra, J. T.; Holst, J. J.; Jeppesen, C. B.; Johnson, M. D.; de Jong, J. C. Small-molecule agonists for the glucagon-like peptide 1 receptor. Proc. Natl. Acad. Sci. U. S. A. 2007, 104 (3), 937– 942, DOI: 10.1073/pnas.0605701104Google ScholarThere is no corresponding record for this reference.
- 26Willard, F. S.; Ho, J. D.; Sloop, K. W. Discovery and pharmacology of the covalent GLP-1 receptor (GLP-1R) allosteric modulator BETP: A novel tool to probe GLP-1R pharmacology. Adv. Pharmacol. 2020, 88, 173– 191, DOI: 10.1016/bs.apha.2020.02.001Google ScholarThere is no corresponding record for this reference.
- 27Coopman, K.; Huang, Y.; Johnston, N.; Bradley, S. J.; Wilkinson, G. F.; Willars, G. B. Comparative effects of the endogenous agonist glucagon-like peptide-1 (GLP-1)-(7-36) amide and the small-molecule ago-allosteric agent “compound 2” at the GLP-1 receptor. J. Pharmacol. Exp. Ther. 2010, 334 (3), 795– 808, DOI: 10.1124/jpet.110.166009Google ScholarThere is no corresponding record for this reference.
- 28Cong, Z.; Chen, L.-N.; Ma, H.; Zhou, Q.; Zou, X.; Ye, C.; Dai, A.; Liu, Q.; Huang, W.; Sun, X. Molecular insights into ago-allosteric modulation of the human glucagon-like peptide-1 receptor. Nat. Commun. 2021, 12 (1), 3763, DOI: 10.1038/s41467-021-24058-zGoogle ScholarThere is no corresponding record for this reference.
- 29Hilger, D.; Masureel, M.; Kobilka, B. K. Structure and dynamics of GPCR signaling complexes. Nat. Struct. Mol. Biol. 2018, 25 (1), 4– 12, DOI: 10.1038/s41594-017-0011-7Google ScholarThere is no corresponding record for this reference.
- 30Latorraca, N. R.; Venkatakrishnan, A. J.; Dror, R. O. GPCR dynamics: Structures in motion. Chem. Rev. 2017, 117, 139– 155, DOI: 10.1021/acs.chemrev.6b00177Google ScholarThere is no corresponding record for this reference.
- 31Stevens, R. C.; Cherezov, V.; Katritch, V.; Abagyan, R.; Kuhn, P.; Rosen, H.; Wüthrich, K. The GPCR Network: A large-scale collaboration to determine human GPCR structure and function. Nat. Rev. Drug Discovery 2013, 12 (1), 25– 34, DOI: 10.1038/nrd3859Google ScholarThere is no corresponding record for this reference.
- 32Zhao, L.-H.; Yin, Y.; Yang, D.; Liu, B.; Hou, L.; Wang, X.; Pal, K.; Jiang, Y.; Feng, Y.; Cai, X. Differential requirement of the extracellular domain in activation of class BG protein-coupled receptors. J. Biol. Chem. 2016, 291 (29), 15119– 15130, DOI: 10.1074/jbc.M116.726620Google ScholarThere is no corresponding record for this reference.
- 33Teng, M.; Johnson, M. D.; Thomas, C.; Kiel, D.; Lakis, J. N.; Kercher, T.; Aytes, S.; Kostrowicki, J.; Bhumralkar, D.; Truesdale, L. Small molecule ago-allosteric modulators of the human glucagon-like peptide-1 (hGLP-1) receptor. Bioorg. Med. Chem. Lett. 2007, 17 (19), 5472– 5478, DOI: 10.1016/j.bmcl.2007.06.086Google ScholarThere is no corresponding record for this reference.
- 34Takhar, S.; Gyomorey, S.; Su, R. C.; Mathi, S. K.; Li, X.; Wheeler, M. B. The third cytoplasmic domain of the GLP-1 [7-36 amide] receptor is required for coupling to the adenylyl cyclase system. Endocrinology 1996, 137 (5), 2175– 2178, DOI: 10.1210/en.137.5.2175Google ScholarThere is no corresponding record for this reference.
- 35De Graaf, C.; Donnelly, D.; Wootten, D.; Lau, J.; Sexton, P. M.; Miller, L. J.; Ahn, J.-M.; Liao, J.; Fletcher, M. M.; Yang, D. Glucagon-Like Peptide-1 and Its Class B G Protein–Coupled Receptors: A Long March to Therapeutic Successes. Pharmacol. Rev. 2016, 68 (4), 954– 1013, DOI: 10.1124/pr.115.011395Google ScholarThere is no corresponding record for this reference.
- 36Cee, V. J.; Volak, L. P.; Chen, Y.; Bartberger, M. D.; Tegley, C.; Arvedson, T.; McCarter, J.; Tasker, A. S.; Fotsch, C. Systematic Study of the Glutathione (GSH) Reactivity of N-Arylacrylamides: 1. Effects of Aryl Substitution. J. Med. Chem. 2015, 58 (23), 9171– 9178, DOI: 10.1021/acs.jmedchem.5b01018Google ScholarThere is no corresponding record for this reference.
- 37Lonsdale, R.; Burgess, J.; Colclough, N.; Davies, N. L.; Lenz, E. M.; Orton, A. L.; Ward, R. A. Expanding the Armory: Predicting and Tuning Covalent Warhead Reactivity. J. Chem. Inf. Model. 2017, 57 (12), 3124– 3137, DOI: 10.1021/acs.jcim.7b00553Google ScholarThere is no corresponding record for this reference.
- 38Waku, T.; Shiraki, T.; Oyama, T.; Fujimoto, Y.; Maebara, K.; Kamiya, N.; Jingami, H.; Morikawa, K. Structural Insight into PPARγ Activation Through Covalent Modification with Endogenous Fatty Acids. J. Mol. Biol. 2009, 385 (1), 188– 199, DOI: 10.1016/j.jmb.2008.10.039Google ScholarThere is no corresponding record for this reference.
- 39Berteotti, A.; Vacondio, F.; Lodola, A.; Bassi, M.; Silva, C.; Mor, M.; Cavalli, A. Predicting the reactivity of nitrile-carrying compounds with cysteine: A combined computational and experimental study. ACS Med. Chem. Lett. 2014, 5 (5), 501– 505, DOI: 10.1021/ml400489bGoogle ScholarThere is no corresponding record for this reference.
- 40Underwood, C. R.; Garibay, P.; Knudsen, L. B.; Hastrup, S.; Peters, G. H.; Rudolph, R.; Reedtz-Runge, S. Crystal structure of glucagon-like peptide-1 in complex with the extracellular domain of the glucagon-like peptide-1 receptor. J. Biol. Chem. 2010, 285 (1), P723– 730, DOI: 10.1074/jbc.M109.033829Google ScholarThere is no corresponding record for this reference.
- 41Coopman, K.; Wallis, R.; Robb, G.; Brown, A. J. H.; Wilkinson, G. F.; Timms, D.; Willars, G. B. Residues within the transmembrane domain of the glucagon-like peptide-1 receptor involved in ligand binding and receptor activation: Modelling the ligand-bound receptor. Mol. Endocrinol. 2011, 25 (10), 1804– 1818, DOI: 10.1210/me.2011-1160Google ScholarThere is no corresponding record for this reference.
- 42Yang, D.; de Graaf, C.; Yang, L.; Song, G.; Dai, A.; Cai, X.; Feng, Y.; Reedtz-Runge, S.; Hanson, M. A.; Yang, H. Structural determinants of binding the seven-transmembrane domain of the glucagon-like peptide-1 receptor (GLP-1R). J. Biol. Chem. 2016, 291 (25), P12991– 13004, DOI: 10.1074/jbc.M116.721977Google ScholarThere is no corresponding record for this reference.
- 43Wootten, D.; Reynolds, C. A.; Smith, K. J.; Mobarec, J. C.; Koole, C.; Savage, E. E.; Pabreja, K.; Simms, J.; Sridhar, R.; Furness, S. G. B. The extracellular surface of the GLP-1 receptor is a molecular trigger for biased agonism. Cell 2016, 165 (7), P1632– 1643, DOI: 10.1016/j.cell.2016.05.023Google ScholarThere is no corresponding record for this reference.
- 44Kawai, T.; Sun, B.; Yoshino, H.; Feng, D.; Suzuki, Y.; Fukazawa, M.; Nagao, S.; Wainscott, D. B.; Showalter, A. D.; Droz, B. A. Structural basis for GLP-1 receptor activation by LY3502970, an orally active nonpeptide agonist. Proc. Natl. Acad. Sci. U. S. A. 2020, 117 (47), 29959– 29967, DOI: 10.1073/pnas.2014879117Google ScholarThere is no corresponding record for this reference.
- 45Cong, Z.; Liang, Y.-L.; Zhou, Q.; Darbalaei, S.; Zhao, F.; Feng, W.; Zhao, L.; Xu, H. E.; Yang, D.; Wang, M.-W. Structural perspective of class B1 GPCR signaling. Trends Pharmacol. Sci. 2022, 43 (4), P321– 334, DOI: 10.1016/j.tips.2022.01.002Google ScholarThere is no corresponding record for this reference.
- 46Dwivedi, H.; Baidya, M.; Shukla, A. K. GPCR Signaling: The Interplay of Gαi and β-arrestin. Curr. Biol. 2018, 28 (7), PR324– R327, DOI: 10.1016/j.cub.2018.02.027Google ScholarThere is no corresponding record for this reference.
- 47Zhang, H.; Luginina, A.; Mishin, A.; Baidya, M.; Shukla, A. K.; Cherezov, V. Structural insights into ligand recognition and activation of angiotensin receptors. Trends Pharmacol. Sci. 2021, 42 (7), P577– 587, DOI: 10.1016/j.tips.2021.04.006Google ScholarThere is no corresponding record for this reference.
- 48Deganutti, G.; Liang, Y.-L.; Zhang, X.; Khoshouei, M.; Clydesdale, L.; Belousoff, M. J.; Venugopal, H.; Truong, T. T.; Glukhova, A.; Keller, A. N. Dynamics of GLP-1R peptide agonist engagement are correlated with kinetics of G protein activation. Nat. Commun. 2022, 13 (1), 92, DOI: 10.1038/s41467-021-27760-0Google ScholarThere is no corresponding record for this reference.
- 49Zhang, M.; Chen, T.; Lu, X.; Lan, X.; Chen, Z.; Lu, S. G protein-coupled receptors (GPCRs): Advances in structures, mechanisms and drug discovery. Signal Transduction Targeted Ther. 2024, 9 (1), 88, DOI: 10.1038/s41392-024-01803-6Google ScholarThere is no corresponding record for this reference.
- 50Cary, B. P.; Deganutti, G.; Zhao, P.; Truong, T. T.; Piper, S. J.; Liu, X.; Belousoff, M. J.; Danev, R.; Sexton, P. M.; Wootten, D. Structural and functional diversity among agonist-bound states of the GLP-1 receptor. Nat. Chem. Biol. 2022, 18 (3), 256– 263, DOI: 10.1038/s41589-021-00945-wGoogle ScholarThere is no corresponding record for this reference.
- 51Russell, I. C.; Lee, D.; Wootten, D.; Sexton, P. M.; Bumbak, F. Cryoelectron microscopy as a tool for illuminating activation mechanisms of human class A orphan G protein-coupled receptors. Pharmacol. Rev. 2025, 77 (3), 100056, DOI: 10.1016/j.pharmr.2025.100056Google ScholarThere is no corresponding record for this reference.
- 52Kim, J.; Kim, J.; Choi, C.; Bae, J.; Choi, H.-J. Structural insights into GPCR signaling activated by peptide ligands: from molecular mechanism to therapeutic application. Exp. Mol. Med. 2025, 57, 1467– 1481, DOI: 10.1038/s12276-025-01497-yGoogle ScholarThere is no corresponding record for this reference.
- 53Lu, S.; He, X.; Yang, Z.; Chai, Z.; Zhou, S.; Wang, J.; Rehman, A. U.; Ni, D.; Pu, J.; Sun, J. Activation pathway of a G protein-coupled receptor uncovers conformational intermediates as targets for allosteric drug design. Nat. Commun. 2021, 12 (1), 4721, DOI: 10.1038/s41467-021-25020-9Google ScholarThere is no corresponding record for this reference.
- 54Bi, M.; Wang, X.; Wang, J.; Xu, J.; Sun, W.; Adediwura, V. A.; Miao, Y.; Cheng, Y.; Ye, L. Structure and function of a near fully-activated intermediate GPCR-Gαβγ complex. Nat. Commun. 2025, 16 (1), 1100, DOI: 10.1038/s41467-025-56434-4Google ScholarThere is no corresponding record for this reference.
- 55Fleetwood, O.; Matricon, P.; Carlsson, J.; Delemotte, L. Energy landscapes reveal agonist control of G protein-coupled receptor activation via microswitches. Biochemistry 2020, 59 (7), 880– 891, DOI: 10.1021/acs.biochem.9b00842Google ScholarThere is no corresponding record for this reference.
- 56McComas, S. E.; Reichenbach, T.; Mitrovic, D.; Alleva, C.; Bonaccorsi, M.; Delemotte, L.; Drew, D. Determinants of sugar-induced influx in the mammalian fructose transporter GLUT5. eLife 2023, 12, e84808 DOI: 10.7554/eLife.84808Google ScholarThere is no corresponding record for this reference.
- 57Pérez-Conesa, S.; Delemotte, L. Free energy landscapes of KcsA inactivation bioRxiv 2023 DOI: 10.1101/2023.04.05.535698Google ScholarThere is no corresponding record for this reference.
- 58Dror, R. O.; Arlow, D. H.; Maragakis, P.; Mildorf, T. J.; Pan, A. C.; Xu, H.; Borhani, D. W.; Shaw, D. E. Activation mechanism of the β2-adrenergic receptor. Proc. Natl. Acad. Sci. U. S. A. 2011, 108 (46), 18684– 18689, DOI: 10.1073/pnas.1110499108Google ScholarThere is no corresponding record for this reference.
- 59Keri, D.; Barth, P. Reprogramming G protein coupled receptor structure and function. Curr. Opin. Struct. Biol. 2018, 51, 187– 194, DOI: 10.1016/j.sbi.2018.07.008Google ScholarThere is no corresponding record for this reference.
- 60van der Velden, W. J. C.; Mukhaleva, E.; Vaidehi, N. Allosteric communication mechanism in the glucagon receptor. J. Biol. Chem. 2025, 301 (6), 108530, DOI: 10.1016/j.jbc.2025.108530Google ScholarThere is no corresponding record for this reference.
- 61Song, G.; Yang, D.; Wang, Y.; de Graaf, C.; Zhou, Q.; Jiang, S.; Liu, K.; Cai, X.; Dai, A.; Lin, G. Human GLP-1 receptor transmembrane domain structure in complex with allosteric modulators. Nature 2017, 546 (7657), 312– 315, DOI: 10.1038/nature22378Google ScholarThere is no corresponding record for this reference.
- 62Wu, F.; Yang, L.; Hang, K.; Laursen, M.; Wu, L.; Han, G. W.; Ren, Q.; Roed, N. K.; Lin, G.; Hanson, M. A. Full-length human GLP-1 receptor structure without orthosteric ligands. Nat. Commun. 2020, 11 (1), 1272, DOI: 10.1038/s41467-020-14934-5Google ScholarThere is no corresponding record for this reference.
- 63Webb, B.; Sali, A. Comparative protein structure modeling using MODELLER. Curr. Protoc. Bioinf. 2016, 54 (1), 5– 6, DOI: 10.1002/cpbi.3Google ScholarThere is no corresponding record for this reference.
- 64Wu, E. L.; Cheng, X.; Jo, S.; Rui, H.; Song, K. C.; Dávila-Contreras, E. M.; Qi, Y.; Lee, J.; Monje-Galvan, V.; Venable, R. M. CHARMM-GUI membrane builder toward realistic biological membrane simulations. J. Comput. Chem. 2014, 35, 1997– 2004, DOI: 10.1002/jcc.23702Google ScholarThere is no corresponding record for this reference.
- 65Huang, J.; Rauscher, S.; Nawrocki, G.; Ran, T.; Feig, M.; de Groot, B. L.; Grubmüller, H.; MacKerell, A. D., Jr CHARMM36m: An improved force field for folded and intrinsically disordered proteins. Nat. Methods 2017, 14 (1), 71– 73, DOI: 10.1038/nmeth.4067Google ScholarThere is no corresponding record for this reference.
- 66Hess, B. P-LINCS: A parallel linear constraint solver for molecular simulation. J. Chem. Theory Comput. 2008, 4 (1), 116– 122, DOI: 10.1021/ct700200bGoogle ScholarThere is no corresponding record for this reference.
- 67Darden, T.; York, D.; Pedersen, L. Particle mesh Ewald: An N log (N) method for Ewald sums in large systems. J. Chem. Phys. 1993, 98 (12), 10089– 10092, DOI: 10.1063/1.464397Google ScholarThere is no corresponding record for this reference.
- 68Fleetwood, O.; Kasimova, M. A.; Westerlund, A. M.; Delemotte, L. Molecular insights from conformational ensembles via machine learning. Biophys. J. 2020, 118 (3), 765– 780, DOI: 10.1016/j.bpj.2019.12.016Google ScholarThere is no corresponding record for this reference.
- 69Pan, A. C.; Sezer, D.; Roux, B. Finding transition pathways using the string method with swarms of trajectories. J. Phys. Chem. B 2008, 112 (11), 3432– 3440, DOI: 10.1021/jp0777059Google ScholarThere is no corresponding record for this reference.
- 70Hoffmann, M.; Scherer, M.; Hempel, T.; Mardt, A.; de Silva, B.; Husic, B. E.; Klus, S.; Wu, H.; Kutz, N.; Brunton, S. L. Deeptime: A Python library for machine learning dynamical models from time series data. Mach. learn.: Sci. Technol. 2021, 3, 015009, DOI: 10.1088/2632-2153/ac3de0Google ScholarThere is no corresponding record for this reference.
- 71Hub, J. S.; de Groot, B. L.; van der Spoel, D. g_wham─A Free Weighted Histogram Analysis Implementation Including Robust Error and Autocorrelation Estimates. J. Chem. Theory Comput. 2010, 6, 3713– 3720, DOI: 10.1021/ct100494zGoogle ScholarThere is no corresponding record for this reference.
- 72Frisch, M. J.; Trucks, G. W.; Schlegel, H. B.; Scuseria, G. E.; Robb, M. A.; Cheeseman, J. R.; Scalmani, G.; Barone, V.; Petersson, G. A.; Nakatsuji, H., Gaussian 16, Revision C.02; Gaussian Inc.: Wallingford CT, 2016.Google ScholarThere is no corresponding record for this reference.
- 73Chai, J.-D.; Head-Gordon, M. Systematic optimization of long-range corrected hybrid density functionals. J. Chem. Phys. 2008, 128, 084106, DOI: 10.1063/1.2834918Google ScholarThere is no corresponding record for this reference.
- 74Astore, M. A.; Pradhan, A. S.; Thiede, E. H.; Hanson, S. M. Protein dynamics underlying allosteric regulation. Curr. Opin. Struct. Biol. 2024, 84, 102768, DOI: 10.1016/j.sbi.2023.102768Google ScholarThere is no corresponding record for this reference.
- 75Bernetti, M.; Bosio, S.; Bresciani, V.; Falchi, F.; Masetti, M. Probing allosteric communication with combined molecular dynamics simulations and network analysis. Curr. Opin. Struct. Biol. 2024, 86, 102820, DOI: 10.1016/j.sbi.2024.102820Google ScholarThere is no corresponding record for this reference.
- 76Stone, J.; Developers, N.T.; Eargle, J.; Sethi, A.; Li, L.; Luthey-Schulten, Z., Dynamical Network Analysis; University of Illinois at Urbana-Champaign, 2012.Google ScholarThere is no corresponding record for this reference.
- 77Eargle, J.; Luthey-Schulten, Z. NetworkView: 3D display and analysis of protein· RNA interaction networks. Bioinformatics 2012, 28 (22), 3000– 3001, DOI: 10.1093/bioinformatics/bts546Google ScholarThere is no corresponding record for this reference.
- 78Floyd, R. W. Algorithm 97: Shortest path. Commun. ACM 1962, 5 (6), 345– 345, DOI: 10.1145/367766.368168Google ScholarThere is no corresponding record for this reference.
- 79Girvan, M.; Newman, M. E. J. Community structure in social and biological networks. Proc. Natl. Acad. Sci. U. S. A. 2002, 99 (12), 7821– 7826, DOI: 10.1073/pnas.122653799Google ScholarThere is no corresponding record for this reference.
Cited By
This article has not yet been cited by other publications.
Article Views
Altmetric
Citations
Article Views are the COUNTER-compliant sum of full text article downloads since November 2008 (both PDF and HTML) across all institutions and individuals. These metrics are regularly updated to reflect usage leading up to the last few days.
Citations are the number of other articles citing this article, calculated by Crossref and updated daily. Find more information about Crossref citation counts.
The Altmetric Attention Score is a quantitative measure of the attention that a research article has received online. Clicking on the donut icon will load a page at altmetric.com with additional details about the score and the social media presence for the given article. Find more information on the Altmetric Attention Score and how the score is calculated.
Recommended Articles
Abstract
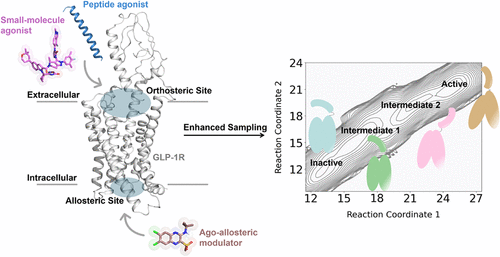
Figure 1
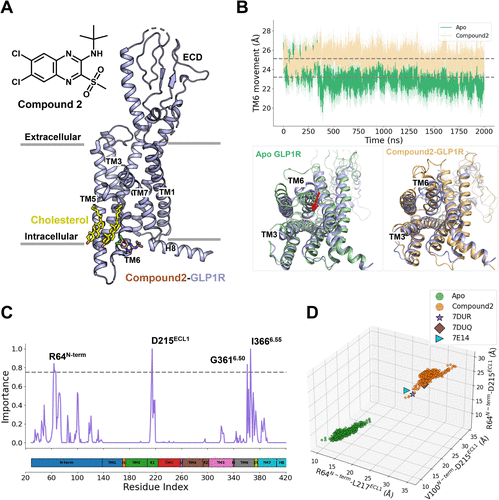
Figure 1. (A) Overall structure of GLP-1R shown in purple cartoon bound to compound 2 (brown sticks) (PDB ID: 7DUR (28)). (B) TM6 movement during 2-μs production simulations of apo (green) and compound 2-bound (orange) GLP-1R, quantified by the Cα distance between Y2503.53 and K3466.35. Representative snapshots sampled from each trajectory are superimposed onto the starting structure (purple) for comparison. (C) Residue-wise importance profiles derived from RF analysis highlight structural differences between apo and compound 2-bound GLP-1R conformations. (D) Distribution of significant structural features distinguishing apo and compound 2-bound GLP-1R states. Representative active cryo-EM structures of GLP-1R includes 7DUR, 7DUQ, 7E14. (28)
Figure 2
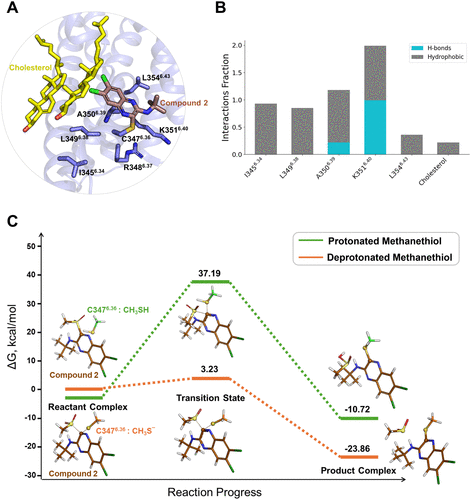
Figure 2. (A) Compound 2-binding mode at the GLP-1R. (B) Changes in interaction frequencies between compound 2 and nearby residues at the allosteric site. (C) Computed energy profiles for the reaction between Cys3476.36 and compound 2. Insets illustrate the key features of the reactant, transition, and product states for two possible pathways: one with Cys3476.36 in its protonated form and the other in its deprotonated form.
Figure 3
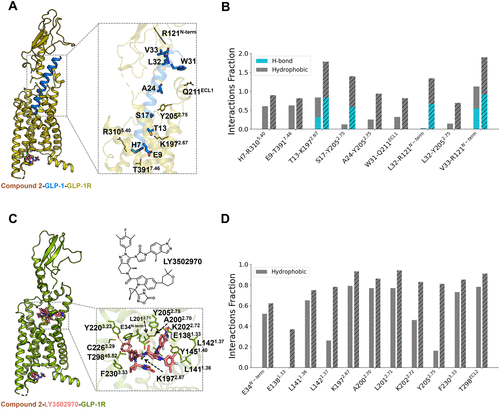
Figure 3. Compound 2 enhances the binding of GLP-1 or LY3502970 at the orthosteric site. (A) Overall structure of the compound 2-GLP-1-GLP-1R complex, highlighting the GLP-1 binding mode. (B) Changes in interaction frequencies between GLP-1 and nearby residues with and without compound 2. Solid bars represent the apo system, while dashed bars indicate the compound 2-bound system. (C) Overall structure of the compound 2-LY3502970-GLP-1R complex, highlighting the LY3502970 binding mode. (D) Changes in interaction frequencies between LY3502970 and nearby residues with and without compound 2. Solid bars represent the apo system, while dashed bars indicate the compound 2-bound system.
Figure 4
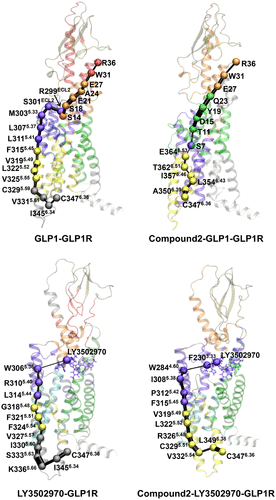
Figure 4. Allosteric signal pathways from the intracellular to extracellular domains in GLP-1R. Optimal communication pathways were computed from C3476.36 (source node) to R36 of GLP-1 in the GLP-1-GLP-1R and compound 2-GLP-1-GLP-1R systems, and from C3476.36 to the centroid of LY3502970 in the LY3502970-GLP-1R and compound 2-LY3502970-GLP-1R systems. Residues involved in the allosteric pathways are shown as spheres and colored according to their community assignments. Connecting edges represent inter-residue communication, with line widths proportional to edge betweenness.
Figure 5
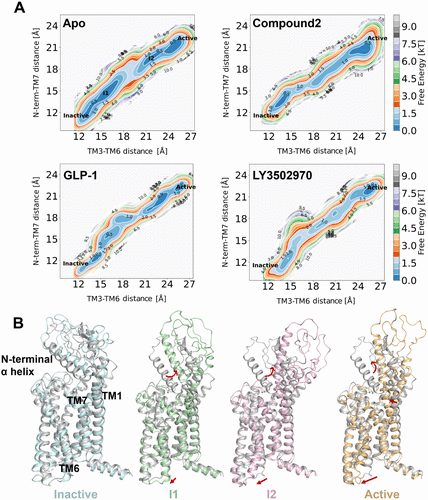
Figure 5. (A) Two-dimensional free energy landscapes of apo and compound 2-, GLP-1-, and LY3502970-bound GLP-1R systems, projected along the outward movement of TM6 (measured by the Cα distance between Y2503.35 and K3466.35) and the swing of the N-terminal α-helix (measured by the Cα distance between W33N-term and F3857.40). (B) Four representative conformations, inactive (cyan), intermediate 1 (I1, green), intermediate 2 (I2, pink), and active (orange), extracted from the string simulation trajectory of apo GLP-1R and superimposed on the inactive experimental structure (gray), corresponding to local energy minima in the free energy landscape.
Figure 6
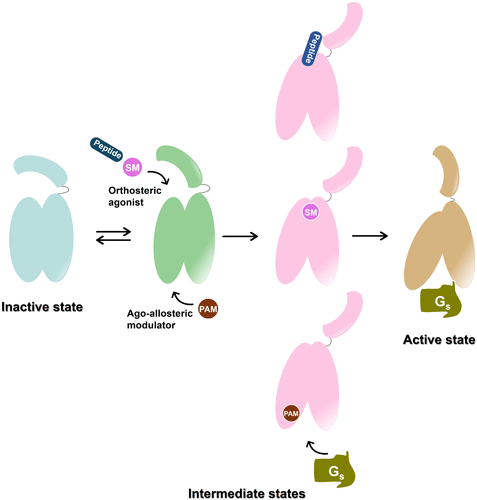
Figure 6. Schematic illustration of ligand-mediated GLP-1R activation. In the absence of ligand, GLP-1R dynamically transitions between inactive and intermediate-like states. Binding of agonists to either the orthosteric or allosteric sites induces conformational rearrangements in both the extracellular and intracellular domains, promoting engagement of downstream transducers and driving the receptor toward its fully active state. SM: small molecule; PAM: positive allosteric modulator.
References
This article references 79 other publications.
- 1Folli, F.; Finzi, G.; Manfrini, R.; Galli, A.; Casiraghi, F.; Centofanti, L.; Berra, C.; Fiorina, P.; Davalli, A.; La Rosa, S. Mechanisms of action of incretin receptor based dual- and tri-agonists in pancreatic islets. Am. J. Physiol. Endocrinol. Metab. 2023, 325 (5), E595–e609 DOI: 10.1152/ajpendo.00236.2023There is no corresponding record for this reference.
- 2Drucker, D. J. GLP-1 physiology informs the pharmacotherapy of obesity. Mol. Metab. 2022, 57, 101351, DOI: 10.1016/j.molmet.2021.101351There is no corresponding record for this reference.
- 3Gentilella, R.; Pechtner, V.; Corcos, A.; Consoli, A. Glucagon-like peptide-1 receptor agonists in type 2 diabetes treatment: Are they all the same?. Diabetes Metabolism Res. 2019, 35 (1), e3070 DOI: 10.1002/dmrr.3070There is no corresponding record for this reference.
- 4Alavi, S. E.; Cabot, P. J.; Moyle, P. M. Glucagon-like peptide-1 receptor agonists and strategies to improve their efficiency. Mol. Pharmaceutics 2019, 16 (6), 2278– 2295, DOI: 10.1021/acs.molpharmaceut.9b00308There is no corresponding record for this reference.
- 5Chen, B.; Yu, X.; Horvath-Diano, C.; Ortuño, M. J.; Tschöp, M. H.; Jastreboff, A. M.; Schneeberger, M. GLP-1 programs the neurovascular landscape. Cell Metab. 2024, 36 (10), P2173– 2189, DOI: 10.1016/j.cmet.2024.09.003There is no corresponding record for this reference.
- 6Ussher, J. R.; Drucker, D. J. Glucagon-like peptide 1 receptor agonists: Cardiovascular benefits and mechanisms of action. Nat. Rev. Cardiol. 2023, 20 (7), 463– 474, DOI: 10.1038/s41569-023-00849-3There is no corresponding record for this reference.
- 7Wong, C. K.; McLean, B. A.; Baggio, L. L.; Koehler, J. A.; Hammoud, R.; Rittig, N.; Yabut, J. M.; Seeley, R. J.; Brown, T. J.; Drucker, D. J. Central glucagon-like peptide 1 receptor activation inhibits Toll-like receptor agonist-induced inflammation. Cell Metab. 2024, 36 (1), P130– 143.E5, DOI: 10.1016/j.cmet.2023.11.009There is no corresponding record for this reference.
- 8Sourris, K. C.; Ding, Y.; Maxwell, S. S.; Al-Sharea, A.; Kantharidis, P.; Mohan, M.; Rosado, C. J.; Penfold, S. A.; Haase, C.; Xu, Y. Glucagon-like peptide-1 receptor signaling modifies the extent of diabetic kidney disease through dampening the receptor for advanced glycation end products–induced inflammation. Kidney Int. 2024, 105 (1), 132– 149, DOI: 10.1016/j.kint.2023.09.029There is no corresponding record for this reference.
- 9Zhou, Q.; Li, G.; Hang, K.; Li, J.; Yang, D.; Wang, M.-W. Weight Loss Blockbuster Development: A Role for Unimolecular Polypharmacology. Annu. Rev. Pharmacol. Toxicol. 2025, 65 (1), 191– 213, DOI: 10.1146/annurev-pharmtox-061324-011832There is no corresponding record for this reference.
- 10Saraiva, J. F. K.; Franco, D. Oral GLP-1 analogue: Perspectives and impact on atherosclerosis in type 2 diabetic patients. Cardiovasc. Diabetol. 2021, 20 (1), 235, DOI: 10.1186/s12933-021-01417-0There is no corresponding record for this reference.
- 11Meissner, W. G.; Remy, P.; Giordana, C.; Maltête, D.; Derkinderen, P.; Houéto, J. L.; Anheim, M.; Benatru, I.; Boraud, T.; Brefel-Courbon, C. Trial of Lixisenatide in Early Parkinson’s Disease. N. Engl. J. Med. 2024, 390, 1176– 1185, DOI: 10.1056/NEJMoa2312323There is no corresponding record for this reference.
- 12Newsome, P. N.; Buchholtz, K.; Cusi, K.; Linder, M.; Okanoue, T.; Ratziu, V.; Sanyal, A. J.; Sejling, A.-S.; Harrison, S. A. A Placebo-Controlled Trial of Subcutaneous Semaglutide in Nonalcoholic Steatohepatitis. N. Engl. J. Med. 2021, 384 (12), 1113– 1124, DOI: 10.1056/NEJMoa2028395There is no corresponding record for this reference.
- 13Mayo, K. E.; Miller, L. J.; Bataille, D.; Dalle, S.; Göke, B.; Thorens, B.; Drucker, D. J. International Union of Pharmacology. XXXV. The glucagon receptor family. Pharmacol. Rev. 2003, 55 (1), 167– 194, DOI: 10.1124/pr.55.1.6There is no corresponding record for this reference.
- 14Zhang, Y.; Sun, B.; Feng, D.; Hu, H.; Chu, M.; Qu, Q.; Tarrasch, J. T.; Li, S.; Sun Kobilka, T.; Kobilka, B. K. Cryo-EM structure of the activated GLP-1 receptor in complex with a G protein. Nature 2017, 546 (7657), 248– 253, DOI: 10.1038/nature22394There is no corresponding record for this reference.
- 15Zhou, Q.; Zhao, F.; Zhang, Y.; Yang, D.; Wang, M.-W. Structural pharmacology and mechanisms of GLP-1R signaling. Trends Pharmacol. Sci. 2025, 46 (5), P422– 436, DOI: 10.1016/j.tips.2025.03.003There is no corresponding record for this reference.
- 16Secher, A.; Jelsing, J.; Baquero, A. F.; Hecksher-Sørensen, J.; Cowley, M. A.; Dalbøge, L. S.; Hansen, G.; Grove, K. L.; Pyke, C.; Raun, K. The arcuate nucleus mediates GLP-1 receptor agonist liraglutide-dependent weight loss. J. Clin. Invest. 2014, 124 (10), 4473– 4488, DOI: 10.1172/JCI75276There is no corresponding record for this reference.
- 17Hall, S.; Isaacs, D.; Clements, J. N. Pharmacokinetics and clinical implications of semaglutide: A new glucagon-like peptide (GLP)-1 receptor agonist. Clin. Pharmacokinet. 2018, 57 (12), 1529– 1538, DOI: 10.1007/s40262-018-0668-zThere is no corresponding record for this reference.
- 18Neumiller, J. J. Incretin-based therapies. Med. Clin. 2015, 99, 107– 129, DOI: 10.1016/j.mcna.2014.08.013There is no corresponding record for this reference.
- 19Buckley, S. T.; Bækdal, T. A.; Vegge, A.; Maarbjerg, S. J.; Pyke, C.; Ahnfelt-Rønne, J.; Madsen, K. G.; Schéele, S. G.; Alanentalo, T.; Kirk, R. K. Transcellular stomach absorption of a derivatized glucagon-like peptide-1 receptor agonist. Sci. Transl. Med. 2018, 10 (467), eaar7047 DOI: 10.1126/scitranslmed.aar7047There is no corresponding record for this reference.
- 20Saxena, A. R.; Frias, J. P.; Brown, L. S.; Gorman, D. N.; Vasas, S.; Tsamandouras, N.; Birnbaum, M. J. Efficacy and safety of oral small molecule glucagon-like peptide 1 receptor agonist danuglipron for glycemic control among patients with type 2 diabetes: A randomized clinical trial. JAMA Netw. Open 2023, 6 (5), e2314493 DOI: 10.1001/jamanetworkopen.2023.14493There is no corresponding record for this reference.
- 21Buckeridge, C.; Tsamandouras, N.; Carvajal-Gonzalez, S.; Brown, L. S.; Hernandez-Illas, M.; Saxena, A. R. Once-daily oral small-molecule glucagon-like peptide-1 receptor agonist lotiglipron (PF-07081532) for type 2 diabetes and obesity: Two randomized, placebo-controlled, multiple-ascending-dose phase 1 studies. Diabetes Obes. Metab. 2024, 26 (8), 3155– 3166, DOI: 10.1111/dom.15643There is no corresponding record for this reference.
- 22Frias, J. P.; Hsia, S.; Eyde, S.; Liu, R.; Ma, X.; Konig, M.; Kazda, C.; Mather, K. J.; Haupt, A.; Pratt, E. Efficacy and safety of oral orforglipron in patients with type 2 diabetes: A multicentre, randomised, dose-response, phase 2 study. Lancet 2023, 402 (10400), P472– 483, DOI: 10.1016/S0140-6736(23)01302-8There is no corresponding record for this reference.
- 23Smelcerovic, A.; Lazarevic, J.; Tomovic, K.; Anastasijevic, M.; Jukic, M.; Kocic, G.; Anderluh, M. An overview, advantages and therapeutic potential of nonpeptide positive allosteric modulators of glucagon-like peptide-1 receptor. ChemMedChem 2019, 14 (5), 514– 521, DOI: 10.1002/cmdc.201800699There is no corresponding record for this reference.
- 24Bueno, A. B.; Sun, B.; Willard, F. S.; Feng, D.; Ho, J. D.; Wainscott, D. B.; Showalter, A. D.; Vieth, M.; Chen, Q.; Stutsman, C. Structural insights into probe-dependent positive allosterism of the GLP-1 receptor. Nat. Chem. Biol. 2020, 16 (10), 1105– 1110, DOI: 10.1038/s41589-020-0589-7There is no corresponding record for this reference.
- 25Knudsen, L. B.; Kiel, D.; Teng, M.; Behrens, C.; Bhumralkar, D.; Kodra, J. T.; Holst, J. J.; Jeppesen, C. B.; Johnson, M. D.; de Jong, J. C. Small-molecule agonists for the glucagon-like peptide 1 receptor. Proc. Natl. Acad. Sci. U. S. A. 2007, 104 (3), 937– 942, DOI: 10.1073/pnas.0605701104There is no corresponding record for this reference.
- 26Willard, F. S.; Ho, J. D.; Sloop, K. W. Discovery and pharmacology of the covalent GLP-1 receptor (GLP-1R) allosteric modulator BETP: A novel tool to probe GLP-1R pharmacology. Adv. Pharmacol. 2020, 88, 173– 191, DOI: 10.1016/bs.apha.2020.02.001There is no corresponding record for this reference.
- 27Coopman, K.; Huang, Y.; Johnston, N.; Bradley, S. J.; Wilkinson, G. F.; Willars, G. B. Comparative effects of the endogenous agonist glucagon-like peptide-1 (GLP-1)-(7-36) amide and the small-molecule ago-allosteric agent “compound 2” at the GLP-1 receptor. J. Pharmacol. Exp. Ther. 2010, 334 (3), 795– 808, DOI: 10.1124/jpet.110.166009There is no corresponding record for this reference.
- 28Cong, Z.; Chen, L.-N.; Ma, H.; Zhou, Q.; Zou, X.; Ye, C.; Dai, A.; Liu, Q.; Huang, W.; Sun, X. Molecular insights into ago-allosteric modulation of the human glucagon-like peptide-1 receptor. Nat. Commun. 2021, 12 (1), 3763, DOI: 10.1038/s41467-021-24058-zThere is no corresponding record for this reference.
- 29Hilger, D.; Masureel, M.; Kobilka, B. K. Structure and dynamics of GPCR signaling complexes. Nat. Struct. Mol. Biol. 2018, 25 (1), 4– 12, DOI: 10.1038/s41594-017-0011-7There is no corresponding record for this reference.
- 30Latorraca, N. R.; Venkatakrishnan, A. J.; Dror, R. O. GPCR dynamics: Structures in motion. Chem. Rev. 2017, 117, 139– 155, DOI: 10.1021/acs.chemrev.6b00177There is no corresponding record for this reference.
- 31Stevens, R. C.; Cherezov, V.; Katritch, V.; Abagyan, R.; Kuhn, P.; Rosen, H.; Wüthrich, K. The GPCR Network: A large-scale collaboration to determine human GPCR structure and function. Nat. Rev. Drug Discovery 2013, 12 (1), 25– 34, DOI: 10.1038/nrd3859There is no corresponding record for this reference.
- 32Zhao, L.-H.; Yin, Y.; Yang, D.; Liu, B.; Hou, L.; Wang, X.; Pal, K.; Jiang, Y.; Feng, Y.; Cai, X. Differential requirement of the extracellular domain in activation of class BG protein-coupled receptors. J. Biol. Chem. 2016, 291 (29), 15119– 15130, DOI: 10.1074/jbc.M116.726620There is no corresponding record for this reference.
- 33Teng, M.; Johnson, M. D.; Thomas, C.; Kiel, D.; Lakis, J. N.; Kercher, T.; Aytes, S.; Kostrowicki, J.; Bhumralkar, D.; Truesdale, L. Small molecule ago-allosteric modulators of the human glucagon-like peptide-1 (hGLP-1) receptor. Bioorg. Med. Chem. Lett. 2007, 17 (19), 5472– 5478, DOI: 10.1016/j.bmcl.2007.06.086There is no corresponding record for this reference.
- 34Takhar, S.; Gyomorey, S.; Su, R. C.; Mathi, S. K.; Li, X.; Wheeler, M. B. The third cytoplasmic domain of the GLP-1 [7-36 amide] receptor is required for coupling to the adenylyl cyclase system. Endocrinology 1996, 137 (5), 2175– 2178, DOI: 10.1210/en.137.5.2175There is no corresponding record for this reference.
- 35De Graaf, C.; Donnelly, D.; Wootten, D.; Lau, J.; Sexton, P. M.; Miller, L. J.; Ahn, J.-M.; Liao, J.; Fletcher, M. M.; Yang, D. Glucagon-Like Peptide-1 and Its Class B G Protein–Coupled Receptors: A Long March to Therapeutic Successes. Pharmacol. Rev. 2016, 68 (4), 954– 1013, DOI: 10.1124/pr.115.011395There is no corresponding record for this reference.
- 36Cee, V. J.; Volak, L. P.; Chen, Y.; Bartberger, M. D.; Tegley, C.; Arvedson, T.; McCarter, J.; Tasker, A. S.; Fotsch, C. Systematic Study of the Glutathione (GSH) Reactivity of N-Arylacrylamides: 1. Effects of Aryl Substitution. J. Med. Chem. 2015, 58 (23), 9171– 9178, DOI: 10.1021/acs.jmedchem.5b01018There is no corresponding record for this reference.
- 37Lonsdale, R.; Burgess, J.; Colclough, N.; Davies, N. L.; Lenz, E. M.; Orton, A. L.; Ward, R. A. Expanding the Armory: Predicting and Tuning Covalent Warhead Reactivity. J. Chem. Inf. Model. 2017, 57 (12), 3124– 3137, DOI: 10.1021/acs.jcim.7b00553There is no corresponding record for this reference.
- 38Waku, T.; Shiraki, T.; Oyama, T.; Fujimoto, Y.; Maebara, K.; Kamiya, N.; Jingami, H.; Morikawa, K. Structural Insight into PPARγ Activation Through Covalent Modification with Endogenous Fatty Acids. J. Mol. Biol. 2009, 385 (1), 188– 199, DOI: 10.1016/j.jmb.2008.10.039There is no corresponding record for this reference.
- 39Berteotti, A.; Vacondio, F.; Lodola, A.; Bassi, M.; Silva, C.; Mor, M.; Cavalli, A. Predicting the reactivity of nitrile-carrying compounds with cysteine: A combined computational and experimental study. ACS Med. Chem. Lett. 2014, 5 (5), 501– 505, DOI: 10.1021/ml400489bThere is no corresponding record for this reference.
- 40Underwood, C. R.; Garibay, P.; Knudsen, L. B.; Hastrup, S.; Peters, G. H.; Rudolph, R.; Reedtz-Runge, S. Crystal structure of glucagon-like peptide-1 in complex with the extracellular domain of the glucagon-like peptide-1 receptor. J. Biol. Chem. 2010, 285 (1), P723– 730, DOI: 10.1074/jbc.M109.033829There is no corresponding record for this reference.
- 41Coopman, K.; Wallis, R.; Robb, G.; Brown, A. J. H.; Wilkinson, G. F.; Timms, D.; Willars, G. B. Residues within the transmembrane domain of the glucagon-like peptide-1 receptor involved in ligand binding and receptor activation: Modelling the ligand-bound receptor. Mol. Endocrinol. 2011, 25 (10), 1804– 1818, DOI: 10.1210/me.2011-1160There is no corresponding record for this reference.
- 42Yang, D.; de Graaf, C.; Yang, L.; Song, G.; Dai, A.; Cai, X.; Feng, Y.; Reedtz-Runge, S.; Hanson, M. A.; Yang, H. Structural determinants of binding the seven-transmembrane domain of the glucagon-like peptide-1 receptor (GLP-1R). J. Biol. Chem. 2016, 291 (25), P12991– 13004, DOI: 10.1074/jbc.M116.721977There is no corresponding record for this reference.
- 43Wootten, D.; Reynolds, C. A.; Smith, K. J.; Mobarec, J. C.; Koole, C.; Savage, E. E.; Pabreja, K.; Simms, J.; Sridhar, R.; Furness, S. G. B. The extracellular surface of the GLP-1 receptor is a molecular trigger for biased agonism. Cell 2016, 165 (7), P1632– 1643, DOI: 10.1016/j.cell.2016.05.023There is no corresponding record for this reference.
- 44Kawai, T.; Sun, B.; Yoshino, H.; Feng, D.; Suzuki, Y.; Fukazawa, M.; Nagao, S.; Wainscott, D. B.; Showalter, A. D.; Droz, B. A. Structural basis for GLP-1 receptor activation by LY3502970, an orally active nonpeptide agonist. Proc. Natl. Acad. Sci. U. S. A. 2020, 117 (47), 29959– 29967, DOI: 10.1073/pnas.2014879117There is no corresponding record for this reference.
- 45Cong, Z.; Liang, Y.-L.; Zhou, Q.; Darbalaei, S.; Zhao, F.; Feng, W.; Zhao, L.; Xu, H. E.; Yang, D.; Wang, M.-W. Structural perspective of class B1 GPCR signaling. Trends Pharmacol. Sci. 2022, 43 (4), P321– 334, DOI: 10.1016/j.tips.2022.01.002There is no corresponding record for this reference.
- 46Dwivedi, H.; Baidya, M.; Shukla, A. K. GPCR Signaling: The Interplay of Gαi and β-arrestin. Curr. Biol. 2018, 28 (7), PR324– R327, DOI: 10.1016/j.cub.2018.02.027There is no corresponding record for this reference.
- 47Zhang, H.; Luginina, A.; Mishin, A.; Baidya, M.; Shukla, A. K.; Cherezov, V. Structural insights into ligand recognition and activation of angiotensin receptors. Trends Pharmacol. Sci. 2021, 42 (7), P577– 587, DOI: 10.1016/j.tips.2021.04.006There is no corresponding record for this reference.
- 48Deganutti, G.; Liang, Y.-L.; Zhang, X.; Khoshouei, M.; Clydesdale, L.; Belousoff, M. J.; Venugopal, H.; Truong, T. T.; Glukhova, A.; Keller, A. N. Dynamics of GLP-1R peptide agonist engagement are correlated with kinetics of G protein activation. Nat. Commun. 2022, 13 (1), 92, DOI: 10.1038/s41467-021-27760-0There is no corresponding record for this reference.
- 49Zhang, M.; Chen, T.; Lu, X.; Lan, X.; Chen, Z.; Lu, S. G protein-coupled receptors (GPCRs): Advances in structures, mechanisms and drug discovery. Signal Transduction Targeted Ther. 2024, 9 (1), 88, DOI: 10.1038/s41392-024-01803-6There is no corresponding record for this reference.
- 50Cary, B. P.; Deganutti, G.; Zhao, P.; Truong, T. T.; Piper, S. J.; Liu, X.; Belousoff, M. J.; Danev, R.; Sexton, P. M.; Wootten, D. Structural and functional diversity among agonist-bound states of the GLP-1 receptor. Nat. Chem. Biol. 2022, 18 (3), 256– 263, DOI: 10.1038/s41589-021-00945-wThere is no corresponding record for this reference.
- 51Russell, I. C.; Lee, D.; Wootten, D.; Sexton, P. M.; Bumbak, F. Cryoelectron microscopy as a tool for illuminating activation mechanisms of human class A orphan G protein-coupled receptors. Pharmacol. Rev. 2025, 77 (3), 100056, DOI: 10.1016/j.pharmr.2025.100056There is no corresponding record for this reference.
- 52Kim, J.; Kim, J.; Choi, C.; Bae, J.; Choi, H.-J. Structural insights into GPCR signaling activated by peptide ligands: from molecular mechanism to therapeutic application. Exp. Mol. Med. 2025, 57, 1467– 1481, DOI: 10.1038/s12276-025-01497-yThere is no corresponding record for this reference.
- 53Lu, S.; He, X.; Yang, Z.; Chai, Z.; Zhou, S.; Wang, J.; Rehman, A. U.; Ni, D.; Pu, J.; Sun, J. Activation pathway of a G protein-coupled receptor uncovers conformational intermediates as targets for allosteric drug design. Nat. Commun. 2021, 12 (1), 4721, DOI: 10.1038/s41467-021-25020-9There is no corresponding record for this reference.
- 54Bi, M.; Wang, X.; Wang, J.; Xu, J.; Sun, W.; Adediwura, V. A.; Miao, Y.; Cheng, Y.; Ye, L. Structure and function of a near fully-activated intermediate GPCR-Gαβγ complex. Nat. Commun. 2025, 16 (1), 1100, DOI: 10.1038/s41467-025-56434-4There is no corresponding record for this reference.
- 55Fleetwood, O.; Matricon, P.; Carlsson, J.; Delemotte, L. Energy landscapes reveal agonist control of G protein-coupled receptor activation via microswitches. Biochemistry 2020, 59 (7), 880– 891, DOI: 10.1021/acs.biochem.9b00842There is no corresponding record for this reference.
- 56McComas, S. E.; Reichenbach, T.; Mitrovic, D.; Alleva, C.; Bonaccorsi, M.; Delemotte, L.; Drew, D. Determinants of sugar-induced influx in the mammalian fructose transporter GLUT5. eLife 2023, 12, e84808 DOI: 10.7554/eLife.84808There is no corresponding record for this reference.
- 57Pérez-Conesa, S.; Delemotte, L. Free energy landscapes of KcsA inactivation bioRxiv 2023 DOI: 10.1101/2023.04.05.535698There is no corresponding record for this reference.
- 58Dror, R. O.; Arlow, D. H.; Maragakis, P.; Mildorf, T. J.; Pan, A. C.; Xu, H.; Borhani, D. W.; Shaw, D. E. Activation mechanism of the β2-adrenergic receptor. Proc. Natl. Acad. Sci. U. S. A. 2011, 108 (46), 18684– 18689, DOI: 10.1073/pnas.1110499108There is no corresponding record for this reference.
- 59Keri, D.; Barth, P. Reprogramming G protein coupled receptor structure and function. Curr. Opin. Struct. Biol. 2018, 51, 187– 194, DOI: 10.1016/j.sbi.2018.07.008There is no corresponding record for this reference.
- 60van der Velden, W. J. C.; Mukhaleva, E.; Vaidehi, N. Allosteric communication mechanism in the glucagon receptor. J. Biol. Chem. 2025, 301 (6), 108530, DOI: 10.1016/j.jbc.2025.108530There is no corresponding record for this reference.
- 61Song, G.; Yang, D.; Wang, Y.; de Graaf, C.; Zhou, Q.; Jiang, S.; Liu, K.; Cai, X.; Dai, A.; Lin, G. Human GLP-1 receptor transmembrane domain structure in complex with allosteric modulators. Nature 2017, 546 (7657), 312– 315, DOI: 10.1038/nature22378There is no corresponding record for this reference.
- 62Wu, F.; Yang, L.; Hang, K.; Laursen, M.; Wu, L.; Han, G. W.; Ren, Q.; Roed, N. K.; Lin, G.; Hanson, M. A. Full-length human GLP-1 receptor structure without orthosteric ligands. Nat. Commun. 2020, 11 (1), 1272, DOI: 10.1038/s41467-020-14934-5There is no corresponding record for this reference.
- 63Webb, B.; Sali, A. Comparative protein structure modeling using MODELLER. Curr. Protoc. Bioinf. 2016, 54 (1), 5– 6, DOI: 10.1002/cpbi.3There is no corresponding record for this reference.
- 64Wu, E. L.; Cheng, X.; Jo, S.; Rui, H.; Song, K. C.; Dávila-Contreras, E. M.; Qi, Y.; Lee, J.; Monje-Galvan, V.; Venable, R. M. CHARMM-GUI membrane builder toward realistic biological membrane simulations. J. Comput. Chem. 2014, 35, 1997– 2004, DOI: 10.1002/jcc.23702There is no corresponding record for this reference.
- 65Huang, J.; Rauscher, S.; Nawrocki, G.; Ran, T.; Feig, M.; de Groot, B. L.; Grubmüller, H.; MacKerell, A. D., Jr CHARMM36m: An improved force field for folded and intrinsically disordered proteins. Nat. Methods 2017, 14 (1), 71– 73, DOI: 10.1038/nmeth.4067There is no corresponding record for this reference.
- 66Hess, B. P-LINCS: A parallel linear constraint solver for molecular simulation. J. Chem. Theory Comput. 2008, 4 (1), 116– 122, DOI: 10.1021/ct700200bThere is no corresponding record for this reference.
- 67Darden, T.; York, D.; Pedersen, L. Particle mesh Ewald: An N log (N) method for Ewald sums in large systems. J. Chem. Phys. 1993, 98 (12), 10089– 10092, DOI: 10.1063/1.464397There is no corresponding record for this reference.
- 68Fleetwood, O.; Kasimova, M. A.; Westerlund, A. M.; Delemotte, L. Molecular insights from conformational ensembles via machine learning. Biophys. J. 2020, 118 (3), 765– 780, DOI: 10.1016/j.bpj.2019.12.016There is no corresponding record for this reference.
- 69Pan, A. C.; Sezer, D.; Roux, B. Finding transition pathways using the string method with swarms of trajectories. J. Phys. Chem. B 2008, 112 (11), 3432– 3440, DOI: 10.1021/jp0777059There is no corresponding record for this reference.
- 70Hoffmann, M.; Scherer, M.; Hempel, T.; Mardt, A.; de Silva, B.; Husic, B. E.; Klus, S.; Wu, H.; Kutz, N.; Brunton, S. L. Deeptime: A Python library for machine learning dynamical models from time series data. Mach. learn.: Sci. Technol. 2021, 3, 015009, DOI: 10.1088/2632-2153/ac3de0There is no corresponding record for this reference.
- 71Hub, J. S.; de Groot, B. L.; van der Spoel, D. g_wham─A Free Weighted Histogram Analysis Implementation Including Robust Error and Autocorrelation Estimates. J. Chem. Theory Comput. 2010, 6, 3713– 3720, DOI: 10.1021/ct100494zThere is no corresponding record for this reference.
- 72Frisch, M. J.; Trucks, G. W.; Schlegel, H. B.; Scuseria, G. E.; Robb, M. A.; Cheeseman, J. R.; Scalmani, G.; Barone, V.; Petersson, G. A.; Nakatsuji, H., Gaussian 16, Revision C.02; Gaussian Inc.: Wallingford CT, 2016.There is no corresponding record for this reference.
- 73Chai, J.-D.; Head-Gordon, M. Systematic optimization of long-range corrected hybrid density functionals. J. Chem. Phys. 2008, 128, 084106, DOI: 10.1063/1.2834918There is no corresponding record for this reference.
- 74Astore, M. A.; Pradhan, A. S.; Thiede, E. H.; Hanson, S. M. Protein dynamics underlying allosteric regulation. Curr. Opin. Struct. Biol. 2024, 84, 102768, DOI: 10.1016/j.sbi.2023.102768There is no corresponding record for this reference.
- 75Bernetti, M.; Bosio, S.; Bresciani, V.; Falchi, F.; Masetti, M. Probing allosteric communication with combined molecular dynamics simulations and network analysis. Curr. Opin. Struct. Biol. 2024, 86, 102820, DOI: 10.1016/j.sbi.2024.102820There is no corresponding record for this reference.
- 76Stone, J.; Developers, N.T.; Eargle, J.; Sethi, A.; Li, L.; Luthey-Schulten, Z., Dynamical Network Analysis; University of Illinois at Urbana-Champaign, 2012.There is no corresponding record for this reference.
- 77Eargle, J.; Luthey-Schulten, Z. NetworkView: 3D display and analysis of protein· RNA interaction networks. Bioinformatics 2012, 28 (22), 3000– 3001, DOI: 10.1093/bioinformatics/bts546There is no corresponding record for this reference.
- 78Floyd, R. W. Algorithm 97: Shortest path. Commun. ACM 1962, 5 (6), 345– 345, DOI: 10.1145/367766.368168There is no corresponding record for this reference.
- 79Girvan, M.; Newman, M. E. J. Community structure in social and biological networks. Proc. Natl. Acad. Sci. U. S. A. 2002, 99 (12), 7821– 7826, DOI: 10.1073/pnas.122653799There is no corresponding record for this reference.
Supporting Information
Supporting Information
The Supporting Information is available free of charge at https://pubs.acs.org/doi/10.1021/acs.jcim.6c00224.
Additional details on the simulation analysis of structural changes, comparison of ligand stability, transmembrane helix 6 displacement, sampling convergence estimation, energies from quantum chemical calculations, total simulation time for each system, and collective variables for enhanced sampling simulations in this study (PDF)
Terms & Conditions
Most electronic Supporting Information files are available without a subscription to ACS Web Editions. Such files may be downloaded by article for research use (if there is a public use license linked to the relevant article, that license may permit other uses). Permission may be obtained from ACS for other uses through requests via the RightsLink permission system: http://pubs.acs.org/page/copyright/permissions.html.


