Interfacial Energy Balance Governs Initial Cell Spreading DynamicsClick to copy article linkArticle link copied!
- Jifeng RenJifeng RenSchool of Biomedical Engineering, Capital Medical University, Beijing 100069, ChinaBeijing Key Laboratory of Fundamental Research on Biomechanics in Clinical Application, Capital Medical University, Beijing 100069, ChinaDepartment of Biomedical Engineering, College of Biomedicine, City University of Hong Kong, Hong Kong 999077, ChinaMore by Jifeng Ren
- Shuhuan Hu*Shuhuan Hu*Email: [email protected]Department of Biomedical Engineering, College of Biomedicine, City University of Hong Kong, Hong Kong 999077, ChinaBGI-Shenzhen, Shenzhen,Guangdong 518083, ChinaMore by Shuhuan Hu
- Yi LiuYi LiuDepartment of Biomedical Engineering, College of Biomedicine, City University of Hong Kong, Hong Kong 999077, ChinaMore by Yi Liu
- Siping HuangSiping HuangDepartment of Biomedical Engineering, College of Biomedicine, City University of Hong Kong, Hong Kong 999077, ChinaMore by Siping Huang
- Jingqian ZhangJingqian ZhangDepartment of Biomedical Engineering, College of Biomedicine, City University of Hong Kong, Hong Kong 999077, ChinaMore by Jingqian Zhang
- Qi GaoQi GaoDepartment of Biomedical Engineering, College of Biomedicine, City University of Hong Kong, Hong Kong 999077, ChinaMore by Qi Gao
- King Wai Chiu LaiKing Wai Chiu LaiDepartment of Biomedical Engineering, College of Biomedicine, City University of Hong Kong, Hong Kong 999077, ChinaMore by King Wai Chiu Lai
- Raymond H. W. Lam*Raymond H. W. Lam*Email: [email protected]. Phone: +852-3442-8577. Fax: +852-3442-0172.Department of Biomedical Engineering, College of Biomedicine, City University of Hong Kong, Hong Kong 999077, ChinaCity University of Hong Kong Shenzhen Research Institute, Shenzhen Guangdong 518172, ChinaMore by Raymond H. W. Lam
Abstract
Cell spreading is a fundamental process in physiological and pathological contexts, including tissue formation, wound healing, and cancer cell extravasation. Previous studies have examined biophysical mechanisms governing early spreading (around 1–10 min) while biomolecular processes also begin to emerge, yet initial spreading in an even earlier stage (<1 min) remains largely unexplored. Here, we present a deterministic model based on interfacial energy balance─integrating strain energy, surface adhesion energy, and viscous dissipation─to quantitatively describe initial spreading dynamics. Using interference reflection microscopy (IRM), we characterize spreading behaviors of three breast cell lines (MCF-10A, MCF-7, and MDA-MB-231) on extracellular matrix-coated substrates. Model predictions, incorporating biomechanical and biochemical parameters measured through IRM and atomic force microscopy (AFM), show strong agreements with experimental observations. This work provides a universal framework for understanding initial spreading and offers insights into strategies to regulate initial cell spreading, with potential applications in cancer treatment and tissue engineering.
This publication is licensed for personal use by The American Chemical Society.
Note
Made available for a limited time for personal research and study only License.
Introduction
Results and Discussion
Model
Figure 1
Figure 1. Visualization of initial cell spreading. (a) Key parameters involved in the initial cell spreading process. (b) Configuration of IRM using a laser scanning confocal microscope. (c) Glass and PDMS substrates coated with fluorescent fibronectin. (d) IRM images of an initial spreading MCF-10A cell on fibronectin-coated glass (upper) and their cross-sectional views (lower, processed with intensity inversion and thresholding for better visualization), captured as 10s, 60s, and 300s. All other cases are available in Figure S1. (e) Transient spreading area (Ac) of MCF-10A cells spreading on fibronectin-coated glass substrates. Gray lines represent single-cell dynamics; and the black line represents the average Ac (t). Error bars are standard errors of the mean. All cases with N numbers are available in Figure S2. Scale bar: 10 μm.
Experimental Validation
| MCF-10A | MCF-7 | MDA | |
|---|---|---|---|
| E (kPa) | 1.30 ± SE0.06 | 0.74 ± SE0.04 | 0.66 ± SE0.04 |
| η (kPa•s) | 2.62 ± SE0.16 | 2.55 ± SE0.17 | 3.89 ± SE0.25 |
| ζb, Col (μJ/m2) | 1256 ± SE69 | 2218 ± SE462 | 1892 ± SE259 |
| ζb, FN (μJ/m2) | 529 ± SE101 | 1185 ± SE65 | 1974 ± SE153 |
| Rn (μm) | 3.52 ± SE0.08 | 4.01 ± SE0.07 | 4.12 ± SE0.16 |
Figure 2
Figure 2. Prediction and measurement of the cell spreading area (Ac). (a) Scatter plots of predicted and measured α (left) and β (right) for MCF-10A, MCF-7, and MDA-MB-231 cells spreading on different substrate materials coated with collagen type 1 (Col) or fibronectin (FN). N = 112 in total; and N for different cases are mentioned in Figure S2. (b) Power-law fittings of measured cell–substrate contact radius (Rc = √(Ac/π)) against scaled time βt for. The black dash line indicates the expected power–law relation (∼t1/4) predicted by the model.
Parametric Study of Initial Spreading Rate
Figure 3
Figure 3. Roles of key parameters. (a) Scatter plot of αβ against measured initial spreading rate. The dash line indicates proportional linear fitting of all points (legends are shared with the same one in Figure 2a). (b) Scatter plot of measured initial spreading rate against R/Rn. The solid line indicates the prediction based on the model with average cell properties (legends are shared with the same one in Figure 2a). (c) Average initial spreading rates of cells spreading on different substrate conditions. Asterisks represent p < 0.05. (d) Correlations between initial spreading rate and the derived μ/ζb for all cases. The dash line indicates the linear regression of the average values. Error bars are standard errors of the mean. N = 112 in total; and N for different cases are mentioned in Figure S2.
Individual Cell Viscoelasticity
Figure 4
Figure 4. Characterization of single-cell viscoelasticity. (a) IRM (left) and AFM (right) measurements of MDA-MB-231 cells during the initial spreading process. Scale bar: 20 μm. (b) Scatter plots of single-cell E*/μ against R for individual cells. (c) Scatter plots of αβ against single-cell E*. Blue dash lines indicate fitting results of “fast spreading” cells, and red dash lines indicate fitting results of “slow spreading” cells. (d) Correlation of μ and E* for different individual cells. Each dash line indicates the linear fitting of individual cells with the same cell type.
Methods
Substrate Preparation and Cell Seeding
Interference Reflection Microscopy
IRM Image Processing
Cell Properties Quantification
Measurement of In Situ Single-Cell Viscoelasticity
Cell Culture
Statistics
Conclusion
Supporting Information
The Supporting Information is available free of charge at https://pubs.acs.org/doi/10.1021/acs.langmuir.5c03250.
Representative IRM images for selected cell types spreading on different substrate conditions; fitting of cell spreading area; power-law relationship of the spreading radius; and cell properties measurement using an elasticity microcytometer (PDF)
Terms & Conditions
Most electronic Supporting Information files are available without a subscription to ACS Web Editions. Such files may be downloaded by article for research use (if there is a public use license linked to the relevant article, that license may permit other uses). Permission may be obtained from ACS for other uses through requests via the RightsLink permission system: http://pubs.acs.org/page/copyright/permissions.html.
Acknowledgments
We thank financial supports from the Hong Kong Research Grant Council (GRF 11217323 and 11206324) and the National Natural Science Foundation of China (NSFC 12402380).
References
This article references 28 other publications.
- 1Gout, S.; Tremblay, P.-L.; Huot, J. Selectins and Selectin Ligands in Extravasation of Cancer Cells and Organ Selectivity of Metastasis. Clin. Exp. Metastasis 2008, 25 (4), 335– 344, DOI: 10.1007/s10585-007-9096-4Google ScholarThere is no corresponding record for this reference.
- 2Min, B.-M.; Lee, G.; Kim, S. H.; Nam, Y. S.; Lee, T. S.; Park, W. H. Electrospinning of Silk Fibroin Nanofibers and Its Effect on the Adhesion and Spreading of Normal Human Keratinocytes and Fibroblasts in Vitro. Biomaterials 2004, 25 (7–8), 1289– 1297, DOI: 10.1016/j.biomaterials.2003.08.045Google ScholarThere is no corresponding record for this reference.
- 3Grinnell, F. Fibronectin and Wound Healing. J. Cell. Biochem. 1984, 26 (2), 107– 116, DOI: 10.1002/jcb.240260206Google ScholarThere is no corresponding record for this reference.
- 4Babcock, D. T.; Brock, A. R.; Fish, G. S.; Wang, Y.; Perrin, L.; Krasnow, M. A.; Galko, M. J. Circulating Blood Cells Function as a Surveillance System for Damaged Tissue in Drosophila Larvae. Proc. Natl. Acad. Sci. U.S.A. 2008, 105 (29), 10017– 10022, DOI: 10.1073/pnas.0709951105Google ScholarThere is no corresponding record for this reference.
- 5Cuvelier, D.; Théry, M.; Chu, Y.-S.; Dufour, S.; Thiéry, J.-P.; Bornens, M.; Nassoy, P.; Mahadevan, L. The Universal Dynamics of Cell Spreading. Curr. Biol. 2007, 17 (8), 694– 699, DOI: 10.1016/j.cub.2007.02.058Google ScholarThere is no corresponding record for this reference.
- 6Frisch, T.; Thoumine, O. Predicting the Kinetics of Cell Spreading. J. Biomech. 2002, 35 (8), 1137– 1141, DOI: 10.1016/S0021-9290(02)00075-1Google ScholarThere is no corresponding record for this reference.
- 7Ibata, N.; Terentjev, E. M. Development of Nascent Focal Adhesions in Spreading Cells. Biophys. J. 2020, 119 (10), 2063– 2073, DOI: 10.1016/j.bpj.2020.09.037Google ScholarThere is no corresponding record for this reference.
- 8Fouchard, J.; Bimbard, C.; Bufi, N.; Durand-Smet, P.; Proag, A.; Richert, A.; Cardoso, O.; Asnacios, A. Three-Dimensional Cell Body Shape Dictates the Onset of Traction Force Generation and Growth of Focal Adhesions. Proc. Natl. Acad. Sci. U.S.A. 2014, 111 (36), 13075– 13080, DOI: 10.1073/pnas.1411785111Google ScholarThere is no corresponding record for this reference.
- 9Chaudhuri, O.; Gu, L.; Darnell, M.; Klumpers, D.; Bencherif, S. A.; Weaver, J. C.; Huebsch, N.; Mooney, D. J. Substrate Stress Relaxation Regulates Cell Spreading. Nat. Commun. 2015, 6 (1), 6365, DOI: 10.1038/ncomms7365Google ScholarThere is no corresponding record for this reference.
- 10Nisenholz, N.; Rajendran, K.; Dang, Q.; Chen, H.; Kemkemer, R.; Krishnan, R.; Zemel, A. Active Mechanics and Dynamics of Cell Spreading on Elastic Substrates. Soft Matter 2014, 10 (37), 7234– 7246, DOI: 10.1039/C4SM00780HGoogle ScholarThere is no corresponding record for this reference.
- 11Li, J.; Han, D.; Zhao, Y.-P. Kinetic Behaviour of the Cells Touching Substrate: The Interfacial Stiffness Guides Cell Spreading. Sci. Rep. 2014, 4 (1), 3910, DOI: 10.1038/srep03910Google ScholarThere is no corresponding record for this reference.
- 12Bonakdar, N.; Gerum, R.; Kuhn, M.; Spörrer, M.; Lippert, A.; Schneider, W.; Aifantis, K. E.; Fabry, B. Mechanical Plasticity of Cells. Nat. Mater. 2016, 15 (10), 1090– 1094, DOI: 10.1038/nmat4689Google ScholarThere is no corresponding record for this reference.
- 13Sackmann, E.; Smith, A.-S. Physics of Cell Adhesion: Some Lessons from Cell-Mimetic Systems. Soft Matter 2014, 10 (11), 1644– 1659, DOI: 10.1039/c3sm51910dGoogle ScholarThere is no corresponding record for this reference.
- 14Liu, Y.; Ren, J.; Zhang, R.; Hu, S.; Pang, S. W.; Lam, R. H. Spreading and Migration of Nasopharyngeal Normal and Cancer Cells on Microgratings. ACS Appl. Bio Mater. 2021, 4 (4), 3224– 3231, DOI: 10.1021/acsabm.0c01610Google ScholarThere is no corresponding record for this reference.
- 15Hu, S.; Liu, G.; Chen, W.; Li, X.; Lu, W.; Lam, R. H.; Fu, J. Multiparametric Biomechanical and Biochemical Phenotypic Profiling of Single Cancer Cells Using an Elasticity Microcytometer. small 2016, 12 (17), 2300– 2311, DOI: 10.1002/smll.201503620Google ScholarThere is no corresponding record for this reference.
- 16Hu, S.; Lam, R. H. W. Characterization of Viscoelastic Properties of Normal and Cancerous Human Breast Cells Using a Confining Microchannel. Microfluid. Nanofluidics 2017, 21 (4), 68, DOI: 10.1007/s10404-017-1903-xGoogle ScholarThere is no corresponding record for this reference.
- 17Chamaraux, F.; Fache, S.; Bruckert, F.; Fourcade, B. Kinetics of Cell Spreading. Phys. Rev. Lett. 2005, 94 (15), 158102, DOI: 10.1103/PhysRevLett.94.158102Google ScholarThere is no corresponding record for this reference.
- 18Lerche, M.; Elosegui-Artola, A.; Kechagia, J. Z.; Guzmán, C.; Georgiadou, M.; Andreu, I.; Gullberg, D.; Roca-Cusachs, P.; Peuhu, E.; Ivaska, J. Integrin Binding Dynamics Modulate Ligand-Specific Mechanosensing in Mammary Gland Fibroblasts. iScience 2020, 23 (9), 101507, DOI: 10.1016/j.isci.2020.101507Google ScholarThere is no corresponding record for this reference.
- 19Ghaednia, H.; Wang, X.; Saha, S.; Xu, Y.; Sharma, A.; Jackson, R. L. A Review of Elastic–Plastic Contact Mechanics. Appl. Mech. Rev. 2017, 69, 060804, DOI: 10.1115/1.4038187Google ScholarThere is no corresponding record for this reference.
- 20Guilak, F.; Tedrow, J. R.; Burgkart, R. Viscoelastic Properties of the Cell Nucleus. Biochem. Biophys. Res. Commun. 2000, 269 (3), 781– 786, DOI: 10.1006/bbrc.2000.2360Google ScholarThere is no corresponding record for this reference.
- 21Ren, J.; Li, Y.; Hu, S.; Liu, Y.; Tsao, S. W.; Lau, D.; Luo, G.; Tsang, C. M.; Lam, R. H. W. Nondestructive Quantification of Single-Cell Nuclear and Cytoplasmic Mechanical Properties Based on Large Whole-Cell Deformation. Lab Chip 2020, 20 (22), 4175– 4185, DOI: 10.1039/D0LC00725KGoogle ScholarThere is no corresponding record for this reference.
- 22Trickey, W. R.; Baaijens, F. P. T.; Laursen, T. A.; Alexopoulos, L. G.; Guilak, F. Determination of the Poisson’s Ratio of the Cell: Recovery Properties of Chondrocytes after Release from Complete Micropipette Aspiration. J. Biomech. 2006, 39 (1), 78– 87, DOI: 10.1016/j.jbiomech.2004.11.006Google ScholarThere is no corresponding record for this reference.
- 23Lam, R. H. W.; Weng, S.; Lu, W.; Fu, J. Live-Cell Subcellular Measurement of Cell Stiffness Using a Microengineered Stretchable Micropost Array Membrane. Integr. Biol. 2012, 4 (10), 1289– 1298, DOI: 10.1039/c2ib20134hGoogle ScholarThere is no corresponding record for this reference.
- 24Varner, H.; Cohen, T. Explaining the Spread in Measurement of PDMS Elastic Properties: Influence of Test Method and Curing Protocol. Soft Matter 2024, 20 (46), 9174– 9183, DOI: 10.1039/D4SM00573BGoogle ScholarThere is no corresponding record for this reference.
- 25Cao, Y.; Wu, D.; Duan, Y. A New Image Edge Detection Algorithm Based on Improved Canny. J. Comput. Methods Sci. Eng. 2020, 20 (2), 629– 642, DOI: 10.3233/JCM-193963Google ScholarThere is no corresponding record for this reference.
- 26Zhang, Z. L.; Kristiansen, H.; Liu, J. A Method for Determining Elastic Properties of Micron-Sized Polymer Particles by Using Flat Punch Test. Comput. Mater. Sci. 2007, 39 (2), 305– 314, DOI: 10.1016/j.commatsci.2006.06.009Google ScholarThere is no corresponding record for this reference.
- 27Wysotzki, P.; Sancho, A.; Gimsa, J.; Groll, J. A Comparative Analysis of Detachment Forces and Energies in Initial and Mature Cell-Material Interaction. Colloids Surf. B Biointerfaces 2020, 190, 110894, DOI: 10.1016/j.colsurfb.2020.110894Google ScholarThere is no corresponding record for this reference.
- 28Chim, Y. H.; Mason, L. M.; Rath, N.; Olson, M. F.; Tassieri, M.; Yin, H. A One-Step Procedure to Probe the Viscoelastic Properties of Cells by Atomic Force Microscopy. Sci. Rep. 2018, 8 (1), 14462, DOI: 10.1038/s41598-018-32704-8Google ScholarThere is no corresponding record for this reference.
Cited By
This article has not yet been cited by other publications.
Article Views
Altmetric
Citations
Article Views are the COUNTER-compliant sum of full text article downloads since November 2008 (both PDF and HTML) across all institutions and individuals. These metrics are regularly updated to reflect usage leading up to the last few days.
Citations are the number of other articles citing this article, calculated by Crossref and updated daily. Find more information about Crossref citation counts.
The Altmetric Attention Score is a quantitative measure of the attention that a research article has received online. Clicking on the donut icon will load a page at altmetric.com with additional details about the score and the social media presence for the given article. Find more information on the Altmetric Attention Score and how the score is calculated.
Recommended Articles
Abstract
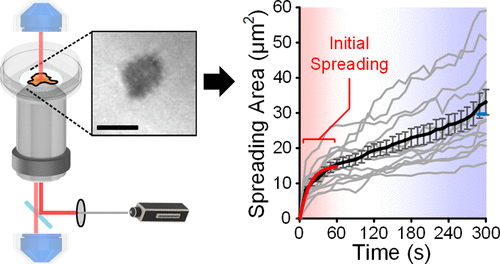
Figure 1
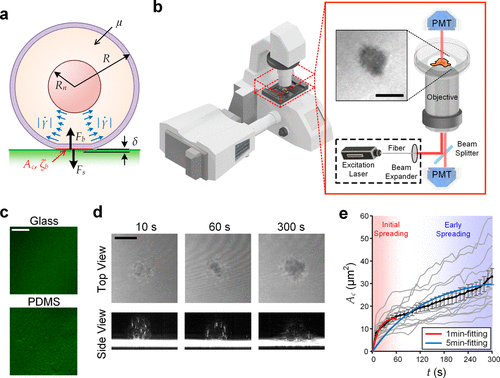
Figure 1. Visualization of initial cell spreading. (a) Key parameters involved in the initial cell spreading process. (b) Configuration of IRM using a laser scanning confocal microscope. (c) Glass and PDMS substrates coated with fluorescent fibronectin. (d) IRM images of an initial spreading MCF-10A cell on fibronectin-coated glass (upper) and their cross-sectional views (lower, processed with intensity inversion and thresholding for better visualization), captured as 10s, 60s, and 300s. All other cases are available in Figure S1. (e) Transient spreading area (Ac) of MCF-10A cells spreading on fibronectin-coated glass substrates. Gray lines represent single-cell dynamics; and the black line represents the average Ac (t). Error bars are standard errors of the mean. All cases with N numbers are available in Figure S2. Scale bar: 10 μm.
Figure 2
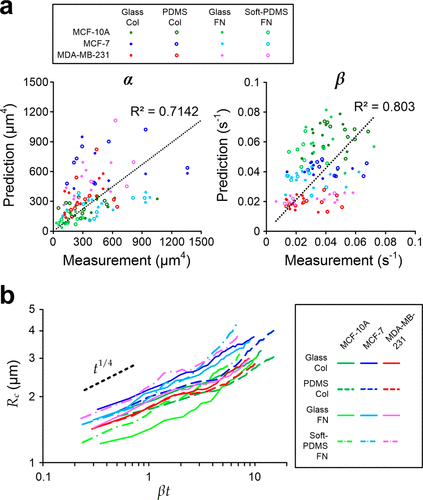
Figure 2. Prediction and measurement of the cell spreading area (Ac). (a) Scatter plots of predicted and measured α (left) and β (right) for MCF-10A, MCF-7, and MDA-MB-231 cells spreading on different substrate materials coated with collagen type 1 (Col) or fibronectin (FN). N = 112 in total; and N for different cases are mentioned in Figure S2. (b) Power-law fittings of measured cell–substrate contact radius (Rc = √(Ac/π)) against scaled time βt for. The black dash line indicates the expected power–law relation (∼t1/4) predicted by the model.
Figure 3
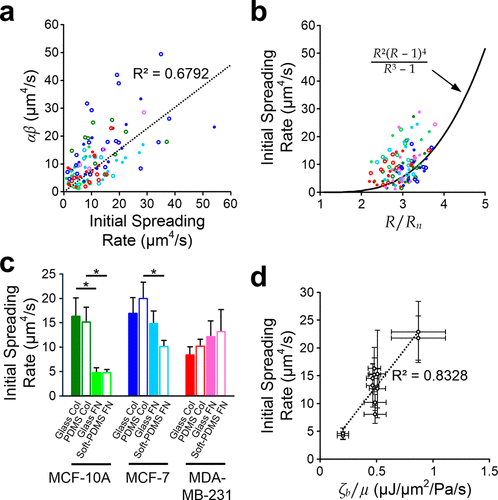
Figure 3. Roles of key parameters. (a) Scatter plot of αβ against measured initial spreading rate. The dash line indicates proportional linear fitting of all points (legends are shared with the same one in Figure 2a). (b) Scatter plot of measured initial spreading rate against R/Rn. The solid line indicates the prediction based on the model with average cell properties (legends are shared with the same one in Figure 2a). (c) Average initial spreading rates of cells spreading on different substrate conditions. Asterisks represent p < 0.05. (d) Correlations between initial spreading rate and the derived μ/ζb for all cases. The dash line indicates the linear regression of the average values. Error bars are standard errors of the mean. N = 112 in total; and N for different cases are mentioned in Figure S2.
Figure 4
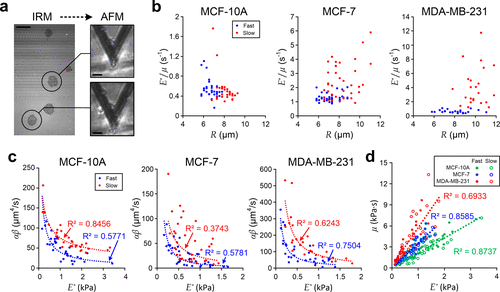
Figure 4. Characterization of single-cell viscoelasticity. (a) IRM (left) and AFM (right) measurements of MDA-MB-231 cells during the initial spreading process. Scale bar: 20 μm. (b) Scatter plots of single-cell E*/μ against R for individual cells. (c) Scatter plots of αβ against single-cell E*. Blue dash lines indicate fitting results of “fast spreading” cells, and red dash lines indicate fitting results of “slow spreading” cells. (d) Correlation of μ and E* for different individual cells. Each dash line indicates the linear fitting of individual cells with the same cell type.
References
This article references 28 other publications.
- 1Gout, S.; Tremblay, P.-L.; Huot, J. Selectins and Selectin Ligands in Extravasation of Cancer Cells and Organ Selectivity of Metastasis. Clin. Exp. Metastasis 2008, 25 (4), 335– 344, DOI: 10.1007/s10585-007-9096-4There is no corresponding record for this reference.
- 2Min, B.-M.; Lee, G.; Kim, S. H.; Nam, Y. S.; Lee, T. S.; Park, W. H. Electrospinning of Silk Fibroin Nanofibers and Its Effect on the Adhesion and Spreading of Normal Human Keratinocytes and Fibroblasts in Vitro. Biomaterials 2004, 25 (7–8), 1289– 1297, DOI: 10.1016/j.biomaterials.2003.08.045There is no corresponding record for this reference.
- 3Grinnell, F. Fibronectin and Wound Healing. J. Cell. Biochem. 1984, 26 (2), 107– 116, DOI: 10.1002/jcb.240260206There is no corresponding record for this reference.
- 4Babcock, D. T.; Brock, A. R.; Fish, G. S.; Wang, Y.; Perrin, L.; Krasnow, M. A.; Galko, M. J. Circulating Blood Cells Function as a Surveillance System for Damaged Tissue in Drosophila Larvae. Proc. Natl. Acad. Sci. U.S.A. 2008, 105 (29), 10017– 10022, DOI: 10.1073/pnas.0709951105There is no corresponding record for this reference.
- 5Cuvelier, D.; Théry, M.; Chu, Y.-S.; Dufour, S.; Thiéry, J.-P.; Bornens, M.; Nassoy, P.; Mahadevan, L. The Universal Dynamics of Cell Spreading. Curr. Biol. 2007, 17 (8), 694– 699, DOI: 10.1016/j.cub.2007.02.058There is no corresponding record for this reference.
- 6Frisch, T.; Thoumine, O. Predicting the Kinetics of Cell Spreading. J. Biomech. 2002, 35 (8), 1137– 1141, DOI: 10.1016/S0021-9290(02)00075-1There is no corresponding record for this reference.
- 7Ibata, N.; Terentjev, E. M. Development of Nascent Focal Adhesions in Spreading Cells. Biophys. J. 2020, 119 (10), 2063– 2073, DOI: 10.1016/j.bpj.2020.09.037There is no corresponding record for this reference.
- 8Fouchard, J.; Bimbard, C.; Bufi, N.; Durand-Smet, P.; Proag, A.; Richert, A.; Cardoso, O.; Asnacios, A. Three-Dimensional Cell Body Shape Dictates the Onset of Traction Force Generation and Growth of Focal Adhesions. Proc. Natl. Acad. Sci. U.S.A. 2014, 111 (36), 13075– 13080, DOI: 10.1073/pnas.1411785111There is no corresponding record for this reference.
- 9Chaudhuri, O.; Gu, L.; Darnell, M.; Klumpers, D.; Bencherif, S. A.; Weaver, J. C.; Huebsch, N.; Mooney, D. J. Substrate Stress Relaxation Regulates Cell Spreading. Nat. Commun. 2015, 6 (1), 6365, DOI: 10.1038/ncomms7365There is no corresponding record for this reference.
- 10Nisenholz, N.; Rajendran, K.; Dang, Q.; Chen, H.; Kemkemer, R.; Krishnan, R.; Zemel, A. Active Mechanics and Dynamics of Cell Spreading on Elastic Substrates. Soft Matter 2014, 10 (37), 7234– 7246, DOI: 10.1039/C4SM00780HThere is no corresponding record for this reference.
- 11Li, J.; Han, D.; Zhao, Y.-P. Kinetic Behaviour of the Cells Touching Substrate: The Interfacial Stiffness Guides Cell Spreading. Sci. Rep. 2014, 4 (1), 3910, DOI: 10.1038/srep03910There is no corresponding record for this reference.
- 12Bonakdar, N.; Gerum, R.; Kuhn, M.; Spörrer, M.; Lippert, A.; Schneider, W.; Aifantis, K. E.; Fabry, B. Mechanical Plasticity of Cells. Nat. Mater. 2016, 15 (10), 1090– 1094, DOI: 10.1038/nmat4689There is no corresponding record for this reference.
- 13Sackmann, E.; Smith, A.-S. Physics of Cell Adhesion: Some Lessons from Cell-Mimetic Systems. Soft Matter 2014, 10 (11), 1644– 1659, DOI: 10.1039/c3sm51910dThere is no corresponding record for this reference.
- 14Liu, Y.; Ren, J.; Zhang, R.; Hu, S.; Pang, S. W.; Lam, R. H. Spreading and Migration of Nasopharyngeal Normal and Cancer Cells on Microgratings. ACS Appl. Bio Mater. 2021, 4 (4), 3224– 3231, DOI: 10.1021/acsabm.0c01610There is no corresponding record for this reference.
- 15Hu, S.; Liu, G.; Chen, W.; Li, X.; Lu, W.; Lam, R. H.; Fu, J. Multiparametric Biomechanical and Biochemical Phenotypic Profiling of Single Cancer Cells Using an Elasticity Microcytometer. small 2016, 12 (17), 2300– 2311, DOI: 10.1002/smll.201503620There is no corresponding record for this reference.
- 16Hu, S.; Lam, R. H. W. Characterization of Viscoelastic Properties of Normal and Cancerous Human Breast Cells Using a Confining Microchannel. Microfluid. Nanofluidics 2017, 21 (4), 68, DOI: 10.1007/s10404-017-1903-xThere is no corresponding record for this reference.
- 17Chamaraux, F.; Fache, S.; Bruckert, F.; Fourcade, B. Kinetics of Cell Spreading. Phys. Rev. Lett. 2005, 94 (15), 158102, DOI: 10.1103/PhysRevLett.94.158102There is no corresponding record for this reference.
- 18Lerche, M.; Elosegui-Artola, A.; Kechagia, J. Z.; Guzmán, C.; Georgiadou, M.; Andreu, I.; Gullberg, D.; Roca-Cusachs, P.; Peuhu, E.; Ivaska, J. Integrin Binding Dynamics Modulate Ligand-Specific Mechanosensing in Mammary Gland Fibroblasts. iScience 2020, 23 (9), 101507, DOI: 10.1016/j.isci.2020.101507There is no corresponding record for this reference.
- 19Ghaednia, H.; Wang, X.; Saha, S.; Xu, Y.; Sharma, A.; Jackson, R. L. A Review of Elastic–Plastic Contact Mechanics. Appl. Mech. Rev. 2017, 69, 060804, DOI: 10.1115/1.4038187There is no corresponding record for this reference.
- 20Guilak, F.; Tedrow, J. R.; Burgkart, R. Viscoelastic Properties of the Cell Nucleus. Biochem. Biophys. Res. Commun. 2000, 269 (3), 781– 786, DOI: 10.1006/bbrc.2000.2360There is no corresponding record for this reference.
- 21Ren, J.; Li, Y.; Hu, S.; Liu, Y.; Tsao, S. W.; Lau, D.; Luo, G.; Tsang, C. M.; Lam, R. H. W. Nondestructive Quantification of Single-Cell Nuclear and Cytoplasmic Mechanical Properties Based on Large Whole-Cell Deformation. Lab Chip 2020, 20 (22), 4175– 4185, DOI: 10.1039/D0LC00725KThere is no corresponding record for this reference.
- 22Trickey, W. R.; Baaijens, F. P. T.; Laursen, T. A.; Alexopoulos, L. G.; Guilak, F. Determination of the Poisson’s Ratio of the Cell: Recovery Properties of Chondrocytes after Release from Complete Micropipette Aspiration. J. Biomech. 2006, 39 (1), 78– 87, DOI: 10.1016/j.jbiomech.2004.11.006There is no corresponding record for this reference.
- 23Lam, R. H. W.; Weng, S.; Lu, W.; Fu, J. Live-Cell Subcellular Measurement of Cell Stiffness Using a Microengineered Stretchable Micropost Array Membrane. Integr. Biol. 2012, 4 (10), 1289– 1298, DOI: 10.1039/c2ib20134hThere is no corresponding record for this reference.
- 24Varner, H.; Cohen, T. Explaining the Spread in Measurement of PDMS Elastic Properties: Influence of Test Method and Curing Protocol. Soft Matter 2024, 20 (46), 9174– 9183, DOI: 10.1039/D4SM00573BThere is no corresponding record for this reference.
- 25Cao, Y.; Wu, D.; Duan, Y. A New Image Edge Detection Algorithm Based on Improved Canny. J. Comput. Methods Sci. Eng. 2020, 20 (2), 629– 642, DOI: 10.3233/JCM-193963There is no corresponding record for this reference.
- 26Zhang, Z. L.; Kristiansen, H.; Liu, J. A Method for Determining Elastic Properties of Micron-Sized Polymer Particles by Using Flat Punch Test. Comput. Mater. Sci. 2007, 39 (2), 305– 314, DOI: 10.1016/j.commatsci.2006.06.009There is no corresponding record for this reference.
- 27Wysotzki, P.; Sancho, A.; Gimsa, J.; Groll, J. A Comparative Analysis of Detachment Forces and Energies in Initial and Mature Cell-Material Interaction. Colloids Surf. B Biointerfaces 2020, 190, 110894, DOI: 10.1016/j.colsurfb.2020.110894There is no corresponding record for this reference.
- 28Chim, Y. H.; Mason, L. M.; Rath, N.; Olson, M. F.; Tassieri, M.; Yin, H. A One-Step Procedure to Probe the Viscoelastic Properties of Cells by Atomic Force Microscopy. Sci. Rep. 2018, 8 (1), 14462, DOI: 10.1038/s41598-018-32704-8There is no corresponding record for this reference.
Supporting Information
Supporting Information
The Supporting Information is available free of charge at https://pubs.acs.org/doi/10.1021/acs.langmuir.5c03250.
Representative IRM images for selected cell types spreading on different substrate conditions; fitting of cell spreading area; power-law relationship of the spreading radius; and cell properties measurement using an elasticity microcytometer (PDF)
Terms & Conditions
Most electronic Supporting Information files are available without a subscription to ACS Web Editions. Such files may be downloaded by article for research use (if there is a public use license linked to the relevant article, that license may permit other uses). Permission may be obtained from ACS for other uses through requests via the RightsLink permission system: http://pubs.acs.org/page/copyright/permissions.html.



