Integrated iPRISM Direct-on-Urine Platform for Rapid UTI Diagnosis in a Double-Blind Clinical TrialClick to copy article linkArticle link copied!
- Xin JiangXin JiangDepartment of Biotechnology and Food Engineering, Technion − Israel Institute of Technology, Haifa 3200003, IsraelMore by Xin Jiang
- Ramy FishlerRamy FishlerDepartment of Biotechnology and Food Engineering, Technion − Israel Institute of Technology, Haifa 3200003, IsraelMore by Ramy Fishler
- Gali RonGali RonDepartment of Biotechnology and Food Engineering, Technion − Israel Institute of Technology, Haifa 3200003, IsraelMore by Gali Ron
- Keren BoguslavskyKeren BoguslavskyDepartment of Urology, Bnai Zion Medical Center, Haifa 3104800, IsraelMore by Keren Boguslavsky
- Sarel HalachmiSarel HalachmiDepartment of Urology, Bnai Zion Medical Center, Haifa 3104800, IsraelThe Faculty of Medicine, Technion − Israel Institute of Technology, Haifa 3525433, IsraelMore by Sarel Halachmi
- Ester Segal*Ester Segal*Email: [email protected]Department of Biotechnology and Food Engineering, Technion − Israel Institute of Technology, Haifa 3200003, IsraelMore by Ester Segal
Abstract
Rapid point-of-care (POC) diagnostics for urinary tract infections (UTIs) are critical for targeted therapy and antibiotic stewardship. We report the first double-blind study of a POC diagnostic system for UTI detection and phenotypic antimicrobial susceptibility testing (AST), using the label-free, real-time iPRISM platform (intensity-based phase-shift reflectometric interference spectroscopic measurement), which traps and grows bacteria on photonic silicon chips. In this near-patient study, unprocessed urine samples were tested in a single-use microfluidic device that integrates both infection screening and AST. Infection screening achieved 97% sensitivity and 60% specificity within 90 min; threshold optimization at 75 min improved performance to 81% specificity and 82% sensitivity. For AST, iPRISM correctly classified 100% of gentamicin-exposed samples in just 30 min and achieved 62% sensitivity and 87% specificity for ciprofloxacin within 90 min. Notably, our preliminary data also demonstrate the potential to differentiate between fungal and bacterial infections, thereby broadening its diagnostic applicability. iPRISM delivers clinically actionable results within a relevant time frame, enabling single-visit prescriptions and supporting personalized, data-driven UTI management.
This publication is licensed under
License Summary*
You are free to share(copy and redistribute) this article in any medium or format and to adapt(remix, transform, and build upon) the material for any purpose, even commercially within the parameters below:
Creative Commons (CC): This is a Creative Commons license.
Attribution (BY): Credit must be given to the creator.
*Disclaimer
This summary highlights only some of the key features and terms of the actual license. It is not a license and has no legal value. Carefully review the actual license before using these materials.
License Summary*
You are free to share(copy and redistribute) this article in any medium or format and to adapt(remix, transform, and build upon) the material for any purpose, even commercially within the parameters below:
Creative Commons (CC): This is a Creative Commons license.
Attribution (BY): Credit must be given to the creator.
*Disclaimer
This summary highlights only some of the key features and terms of the actual license. It is not a license and has no legal value. Carefully review the actual license before using these materials.
License Summary*
You are free to share(copy and redistribute) this article in any medium or format and to adapt(remix, transform, and build upon) the material for any purpose, even commercially within the parameters below:
Creative Commons (CC): This is a Creative Commons license.
Attribution (BY): Credit must be given to the creator.
*Disclaimer
This summary highlights only some of the key features and terms of the actual license. It is not a license and has no legal value. Carefully review the actual license before using these materials.
1. Introduction
2. Experimental Section
2.1. Materials
2.2. Preparation of Solutions and Media
2.3. Fabrication of iPRISM Devices
2.4. iPRISM Assay
2.5. Electron Microscopy
2.6. Statistical Analysis
3. Results and Discussion
3.1. Integrated iPRISM Platform and Study Design
Figure 1
Figure 1. Schematic overview of iPRISM for rapid UTI diagnosis and comparative workflow analysis. (a) iPRISM enables simultaneous infection screening and AST directly on clinical urine specimens. Three infection scenarios can be distinguished based on the measured signal: (i) negative infection, (ii) positive infection with an antimicrobial-susceptible uropathogen, and (iii) positive infection with an antimicrobial-resistant uropathogen. (b) Overall flow of iPRISM and clinical routine UTI diagnostics. Current clinical UTI workflow requires sequential steps: (1) standard urine culture (24–48 h for screening), (2) subculture for microbial identification, and (3) AST (e.g., using VITEK 2), with turnaround time from sample to results ranging from 3 to 7 days. The iPRISM system described herein proceeds without culturing, achieving diagnosis within 90 min. (c) Distribution of infection scenario of the collected clinical urine specimens. (d) Isolated causative agents of culture-positive specimens (derived from the red slice in c), categorized by Gram-negative bacteria (blue-purple), Gram-positive bacteria (pink-red), fungal species (yellow-orange), and polymicrobial infections (yellow-green). Created with BioRender.
3.2. Overview of Clinical Urine Specimens and Distribution of Pathogens Detected by the Traditional Culture Approach
3.3. iPRISM Assay for UTI Screening
Figure 2
Figure 2. iPRISM for UTI detection directly on clinical human urine samples. (a–d) Representative real-time characteristic bacterial growth curves as measured by the iPRISM assay for (a) Escherichia coli, (b) Klebsiella spp., (c)Enterococcus faecalis, and (d) Streptococcus agalactiae. (e–h) Corresponding HR-SEM images of the bacterial species in panels a–d, respectively, demonstrating their spatial distribution on the surface of microwells for retrieved silicon chips from the iPRISM device (for each bacterium, the curve corresponding to the image is marked with a star). Scale bars denote 3 μm.
Figure 3
Figure 3. iPRISM for infection screening on fresh untreated clinical urine samples (63 clean samples and 38 bacterial infected samples including the bacterial-fungal coinfection one). (a) Intensity value changes after incubation in iPRISM device for 30, 60, and 90 min. The dashed line represents a double-blind screening threshold of −ΔI (%) = 0. Samples surpassing this threshold were considered positive for UTI infection by the iPRISM assay. (b) Intensity value changes at 75 min. The black dashed line represents the original detection threshold of −ΔI (%) = 0, while the blue dashed line is the new optimized threshold of −ΔI (%) = 2.59. Each circle corresponds to one clinical specimen. Green circles represent specimens identified as noninfected by clinical culture, and red circles represent specimens with positive culture results. (c) ROC analysis at 75 min. The point in the graph indicated by an arrow corresponds to the optimal diagnostic threshold yielding the maximum Youden’s index. (d) Classification agreement at 75 min using the established optimal threshold for data shown in (b). (e) Time-dependent AUC performance. Error bars represent standard error of area under the ROC curve statistics.
3.4. AST Directly from Urine Specimens
Figure 4
Figure 4. Direct iPRISM AST assay performance in clinical human urine samples with exposure to MIC breakpoint concentration of gentamicin (8 μg mL–1). (a, b) Representative gentamicin-resistant (a) and susceptible (b) cases where E. coli was isolated from culture positive urine samples. (i) Real-time iPRISM characteristic curves showing −ΔI (%) values (n = 3). (ii, iii) HR-SEM images of (ii) control and (iii) post-treatment. Scale bars represent 3 μm. (c) iPRISM relative growth (RG) values at 30 min after exposure to a gentamicin breakpoint concentration of 8 μg mL–1 of urine samples from suspected infected UTI patients (n = 23; 2 resistant, 21 susceptible). The dashed line indicates the predefined threshold (RG at 30 min = 0.95) for resistance classification.
4. Conclusions
Supporting Information
The Supporting Information is available free of charge at https://pubs.acs.org/doi/10.1021/acsmeasuresciau.5c00187.
Study participant demographics; iPRISM principles; double-blinded urine test results for microbial infections; differentiation between bacterial and fungal infections using iPRISM; iPRISM-based AST for ciprofloxacin in bacterially infected urine samples (PDF)
Terms & Conditions
Most electronic Supporting Information files are available without a subscription to ACS Web Editions. Such files may be downloaded by article for research use (if there is a public use license linked to the relevant article, that license may permit other uses). Permission may be obtained from ACS for other uses through requests via the RightsLink permission system: http://pubs.acs.org/page/copyright/permissions.html.
Acknowledgments
This work is supported by the Israel Science Foundation (grant no. 2458/21). The authors thank Mrs. Bracha Mendelson and the remainder of the staff at the Department of Urology and Clinical Microbiology of Bnai Zion Medical Center, Haifa, Israel, for the collection of urine specimens, provision of VITEK 2 results, and fruitful discussions. The authors thank Dr. Heidi Leonard, for her valuable insights and discussions on microfluidic design. The authors thank Orna Ternyak and Tatiana Becker at the Micro-Nano-Fabrication and Printing Unit (MNFPU) at the Technion – Israel Institute of Technology for the fabrication of the microstructured silicon diffraction grating chips. Xin Jiang is grateful to the RBNI Scholarships & Prizes for excellence in Nanoscience & Nanotechnology, Technion, Israel. Figure 1 and TOC were created with BioRender.com under a publication license.
References
This article references 87 other publications.
- 1Timm, M. R.; Russell, S. K.; Hultgren, S. J. Urinary tract infections: pathogenesis, host susceptibility and emerging therapeutics. Nat. Rev. Microbiol. 2025, 23 (2), 72– 86, DOI: 10.1038/s41579-024-01092-4Google ScholarThere is no corresponding record for this reference.
- 2Simoni, A.; Schwartz, L.; Junquera, G. Y.; Ching, C. B.; Spencer, J. D. Current and emerging strategies to curb antibiotic-resistant urinary tract infections. Nat. Rev. Urol. 2024, 21 (12), 707– 722, DOI: 10.1038/s41585-024-00877-9Google ScholarThere is no corresponding record for this reference.
- 3Yang, X.; Chen, H.; Zheng, Y.; Qu, S.; Wang, H.; Yi, F. Disease burden and long-term trends of urinary tract infections: A worldwide report. Front. Public Health 2022, 10, 888205, DOI: 10.3389/fpubh.2022.888205Google ScholarThere is no corresponding record for this reference.
- 4Duque-Sanchez, L.; Qu, Y.; Voelcker, N. H.; Thissen, H. Tackling catheter-associated urinary tract infections with next-generation antimicrobial technologies. J. Biomed. Mater. Res., Part A 2024, 112 (3), 312– 335, DOI: 10.1002/jbm.a.37630Google ScholarThere is no corresponding record for this reference.
- 5Bermudez, T.; Schmitz, J. E.; Boswell, M.; Humphries, R. Novel technologies for the diagnosis of urinary tract infections. J. Clin. Microbiol. 2025, 63 (2), e00306–24 DOI: 10.1128/jcm.00306-24Google ScholarThere is no corresponding record for this reference.
- 6Davenport, M.; Mach, K. E.; Shortliffe, L. M. D.; Banaei, N.; Wang, T.-H.; Liao, J. C. New and developing diagnostic technologies for urinary tract infections. Nat. Rev. Urol. 2017, 14 (5), 296– 310, DOI: 10.1038/nrurol.2017.20Google ScholarThere is no corresponding record for this reference.
- 7Singh, S.; Numan, A.; Cinti, S. Point-of-Care for Evaluating Antimicrobial Resistance through the Adoption of Functional Materials. Anal. Chem. 2022, 94 (1), 26– 40, DOI: 10.1021/acs.analchem.1c03856Google ScholarThere is no corresponding record for this reference.
- 8World Health Organization. Global research agenda for antimicrobial resistance in human health; World Health Organization; 2025, https://www.who.int/publications/i/item/9789240102309; Accessed 25.April.02.Google ScholarThere is no corresponding record for this reference.
- 9Tullus, K.; Shaikh, N. Urinary tract infections in children. Lancet 2020, 395 (10237), 1659– 1668, DOI: 10.1016/S0140-6736(20)30676-0Google ScholarThere is no corresponding record for this reference.
- 10Petchakup, C.; Chen, Y. Y. C.; Tay, H. M.; Ong, H. B.; Hon, P. Y.; De, P. P. Rapid Screening of Urinary Tract Infection Using Microfluidic Inertial-Impedance Cytometry. ACS Sens. 2023, 8 (8), 3136– 3145, DOI: 10.1021/acssensors.3c00819Google ScholarThere is no corresponding record for this reference.
- 11Wat, J. K.-H.; Xu, M.; Nan, L.; Lin, H.; To, K. K.-W.; Shum, H. C.; Hassan, S. U. Rapid antimicrobial susceptibility tests performed by self-diluting microfluidic chips for drug resistance studies and point-of-care diagnostics. Microsyst. Nanoeng. 2025, 11 (1), 110, DOI: 10.1038/s41378-025-00938-yGoogle ScholarThere is no corresponding record for this reference.
- 12Yang, J.; Li, G.; Chen, S.; Su, X.; Xu, D.; Zhai, Y. Machine Learning-Assistant Colorimetric Sensor Arrays for Intelligent and Rapid Diagnosis of Urinary Tract Infection. ACS Sens. 2024, 9 (4), 1945– 1956, DOI: 10.1021/acssensors.3c02687Google ScholarThere is no corresponding record for this reference.
- 13Szlachta-McGinn, A.; Douglass, K. M.; Chung, U. Y. R.; Jackson, N. J.; Nickel, J. C.; Ackerman, A. L. Molecular Diagnostic Methods Versus Conventional Urine Culture for Diagnosis and Treatment of Urinary Tract Infection: A Systematic Review and Meta-analysis. Eur. Urol. Open Sci. 2022, 44, 113– 124, DOI: 10.1016/j.euros.2022.08.009Google ScholarThere is no corresponding record for this reference.
- 14Mancuso, G.; Midiri, A.; Gerace, E.; Marra, M.; Zummo, S.; Biondo, C. Urinary Tract Infections: The Current Scenario and Future Prospects. Pathogens 2023, 12 (4), 623, DOI: 10.3390/pathogens12040623Google ScholarThere is no corresponding record for this reference.
- 15Brubaker, L.; Chai, T. C.; Horsley, H.; Khasriya, R.; Moreland, R. B.; Wolfe, A. J. Tarnished gold─the “standard” urine culture: reassessing the characteristics of a criterion standard for detecting urinary microbes. Front. Urol. 2023, 3, 1206046, DOI: 10.3389/fruro.2023.1206046Google ScholarThere is no corresponding record for this reference.
- 16Moreland, R. B.; Brubaker, L.; Tinawi, L.; Wolfe, A. J. Rapid and accurate testing for urinary tract infection: new clothes for the emperor. Clin. Microbiol. Rev. 2025, 38 (1), e00129–24 DOI: 10.1128/cmr.00129-24Google ScholarThere is no corresponding record for this reference.
- 17Zhang, J.; Wang, M.; Xiao, J.; Wang, M.; Liu, Y.; Gao, X. Metabolism-Triggered Plasmonic Nanosensor for Bacterial Detection and Antimicrobial Susceptibility Testing of Clinical Isolates. ACS Sens. 2024, 9 (1), 379– 387, DOI: 10.1021/acssensors.3c02144Google ScholarThere is no corresponding record for this reference.
- 18Gajic, I.; Kabic, J.; Kekic, D.; Jovicevic, M.; Milenkovic, M.; Mitic Culafic, D. Antimicrobial Susceptibility Testing: A Comprehensive Review of Currently Used Methods. Antibiotics 2022, 11 (4), 427, DOI: 10.3390/antibiotics11040427Google ScholarThere is no corresponding record for this reference.
- 19Sturm, A.; Jóźwiak, G.; Verge, M. P.; Munch, L.; Cathomen, G.; Vocat, A.; Luraschi-Eggemann, A.; Orlando, C.; Fromm, K.; Delarze, E. Accurate and rapid antibiotic susceptibility testing using a machine learning-assisted nanomotion technology platform. Nat. Commun. 2024, 15 (1), 2037, DOI: 10.1038/s41467-024-46213-yGoogle ScholarThere is no corresponding record for this reference.
- 20Kandavalli, V.; Karempudi, P.; Larsson, J.; Elf, J. Rapid antibiotic susceptibility testing and species identification for mixed samples. Nat. Commun. 2022, 13 (1), 6215, DOI: 10.1038/s41467-022-33659-1Google ScholarThere is no corresponding record for this reference.
- 21Leonard, H.; Colodner, R.; Halachmi, S.; Segal, E. Recent Advances in the Race to Design a Rapid Diagnostic Test for Antimicrobial Resistance. ACS Sens. 2018, 3 (11), 2202– 2217, DOI: 10.1021/acssensors.8b00900Google ScholarThere is no corresponding record for this reference.
- 22Ho, C. S.; Wong, C. T. H.; Aung, T. T.; Lakshminarayanan, R.; Mehta, J. S.; Rauz, S. Antimicrobial resistance: a concise update. Lancet Microbe 2025, 6 (1), 100947, DOI: 10.1016/j.lanmic.2024.07.010Google ScholarThere is no corresponding record for this reference.
- 23Faigenbaum-Romm, R.; Yedidi, N.; Gefen, O.; Katsowich-Nagar, N.; Aroeti, L.; Ronin, I.; Bar-Meir, M.; Rosenshine, I.; Balaban, N. Q. Uncovering phenotypic inheritance from single cells with Microcolony-seq. Cell 2025, 188 (19), 5313– 5331.e18, DOI: 10.1016/j.cell.2025.08.001Google ScholarThere is no corresponding record for this reference.
- 24David, S. S. B.; Romano, R.; Rahamim-Cohen, D.; Azuri, J.; Greenfeld, S.; Gedassi, B.; Lerner, U. AI driven decision support reduces antibiotic mismatches and inappropriate use in outpatient urinary tract infections. Npj Digital Med. 2025, 8 (1), 61, DOI: 10.1038/s41746-024-01400-5Google ScholarThere is no corresponding record for this reference.
- 25Pew Charitable Trusts. Health Experts Establish Targets to Improve Hospital Antibiotic Prescribing; Pew Charitable Trusts, 2021.https://pew.org/2Qca7pi. Accessed 25.May.26.Google ScholarThere is no corresponding record for this reference.
- 26Huang, R.; Cai, X.; Du, J.; Lian, J.; Hui, P.; Gu, M. Bioinspired Plasmonic Nanosensor for on-Site Antimicrobial Susceptibility Testing in Urine Samples. ACS Nano 2022, 16 (11), 19229– 19239, DOI: 10.1021/acsnano.2c08532Google ScholarThere is no corresponding record for this reference.
- 27Zhang, P.; Kaushik, A. M.; Mach, K. E.; Hsieh, K.; Liao, J. C.; Wang, T.-H. Facile syringe filter-enabled bacteria separation, enrichment, and buffer exchange for clinical isolation-free digital detection and characterization of bacterial pathogens in urine. Analyst 2021, 146 (8), 2475– 2483, DOI: 10.1039/D1AN00039JGoogle ScholarThere is no corresponding record for this reference.
- 28Zhang, F.; Jiang, J.; McBride, M.; Yang, Y.; Mo, M.; Iriya, R.; Peterman, J.; Jing, W.; Grys, T.; Haydel, S. E. Direct Antimicrobial Susceptibility Testing on Clinical Urine Samples by Optical Tracking of Single Cell Division Events. Small 2020, 16 (52), e2004148 DOI: 10.1002/smll.202004148Google ScholarThere is no corresponding record for this reference.
- 29Reszetnik, G.; Hammond, K.; Mahshid, S.; AbdElfatah, T.; Nguyen, D.; Corsini, R.; Caya, C.; Papenburg, J.; Cheng, M. P.; Yansouni, C. P. Next-generation rapid phenotypic antimicrobial susceptibility testing. Nat. Commun. 2024, 15 (1), 9719, DOI: 10.1038/s41467-024-53930-xGoogle ScholarThere is no corresponding record for this reference.
- 30Riester, O.; Laufer, S.; Mertelsmann, R.; Deigner, H.-P. Sensors for Antibiotics Susceptibility Testing-Recent Advances. Adv. Sens. Res. 2024, 3 (10), 2400046, DOI: 10.1002/adsr.202400046Google ScholarThere is no corresponding record for this reference.
- 31Zhang, M.; Hong, W.; Abutaleb, N. S.; Li, J.; Dong, P.-T.; Zong, C.; Wang, P.; Seleem, M. N.; Cheng, J.-X. Rapid Determination of Antimicrobial Susceptibility by Stimulated Raman Scattering Imaging of D2 O Metabolic Incorporation in a Single Bacterium. Adv. Sci. 2020, 7 (19), 2001452, DOI: 10.1002/advs.202001452Google ScholarThere is no corresponding record for this reference.
- 32Michael, I.; Kim, D.; Gulenko, O.; Kumar, S.; Kumar, S.; Clara, J. A fidget spinner for the point-of-care diagnosis of urinary tract infection. Nat. Biomed. Eng. 2020, 4 (6), 591– 600, DOI: 10.1038/s41551-020-0557-2Google ScholarThere is no corresponding record for this reference.
- 33Yang, S.; Berto, F. G.; Denstedt, J.; Tang, H.; Zhang, J. Point-of-Care Urinary Tract Infection (UTI) Diagnosis Enhanced by Nanostructured Biosensors: Review Paper. Adv. Sens. Res. 2024, 3 (10), 2400051, DOI: 10.1002/adsr.202400051Google ScholarThere is no corresponding record for this reference.
- 34Velican, A. M.; Măruţescu, L.; Kamerzan, C.; Cristea, V. C.; Banu, O.; Borcan, E. Rapid Detection and Antibiotic Susceptibility of Uropathogenic Escherichia coli by Flow Cytometry. Microorganisms 2020, 8 (8), 1233, DOI: 10.3390/microorganisms8081233Google ScholarThere is no corresponding record for this reference.
- 35Altobelli, E.; Mohan, R.; Mach, K. E.; Sin, M. L. Y.; Anikst, V.; Buscarini, M. Integrated Biosensor Assay for Rapid Uropathogen Identification and Phenotypic Antimicrobial Susceptibility Testing. Eur. Urol. Focus 2017, 3 (2–3), 293– 299, DOI: 10.1016/j.euf.2015.12.010Google ScholarThere is no corresponding record for this reference.
- 36Avesar, J.; Rosenfeld, D.; Truman-Rosentsvit, M.; Ben-Arye, T.; Geffen, Y.; Bercovici, M.; Levenberg, S. Rapid phenotypic antimicrobial susceptibility testing using nanoliter arrays. Proc. Natl. Acad. Sci. 2017, 114 (29), E5787-E5795 DOI: 10.1073/pnas.1703736114Google ScholarThere is no corresponding record for this reference.
- 37Burg, L.; Crewe, G.; DiMeo, J.; Guo, X.; Li, C. G.; Mayol, M.; Tempesta, A.; Lauzier, W.; Markham, R.; Crissy, K. Rapid pathogen identification and phenotypic antimicrobial susceptibility directly from urine specimens. Sci. Rep. 2022, 12 (1), 18315, DOI: 10.1038/s41598-022-22792-yGoogle ScholarThere is no corresponding record for this reference.
- 38Gopal, A.; Yan, L.; Kashif, S.; Munshi, T.; Roy, V. A. L.; Voelcker, N. H.; Chen, X. Biosensors and Point-of-Care Devices for Bacterial Detection: Rapid Diagnostics Informing Antibiotic Therapy. Adv. Healthcare Mater. 2022, 11 (3), e2101546 DOI: 10.1002/adhm.202101546Google ScholarThere is no corresponding record for this reference.
- 39Gonchar, K. A.; Nazarovskaia, D. A.; Domnin, P. A.; Gyuppenen, O. D.; Erokhina, A. A.; Wang, M. AuAg-Modified Porous Silicon Nanowires for Dual-Mode Interferometric and SERS-Based Bacterial Detection. ACS Appl. Mater. Interfaces 2025, 17 (23), 33691– 33700, DOI: 10.1021/acsami.5c06953Google ScholarThere is no corresponding record for this reference.
- 40Arshavsky-Graham, S.; Massad-Ivanir, N.; Segal, E.; Weiss, S. Porous Silicon-Based Photonic Biosensors: Current Status and Emerging Applications. Anal. Chem. 2019, 91 (1), 441– 467, DOI: 10.1021/acs.analchem.8b05028Google ScholarThere is no corresponding record for this reference.
- 41Qin, N.; Zhao, P.; Ho, E. A.; Xin, G.; Ren, C. L. Microfluidic Technology for Antibacterial Resistance Study and Antibiotic Susceptibility Testing: Review and Perspective. ACS Sens. 2021, 6 (1), 3– 21, DOI: 10.1021/acssensors.0c02175Google ScholarThere is no corresponding record for this reference.
- 42Van Belkum, A.; Burnham, C.-A. D.; Rossen, J. W. A.; Mallard, F.; Rochas, O.; Dunne, W. M. Innovative and rapid antimicrobial susceptibility testing systems. Nat. Rev. Microbiol. 2020, 18 (5), 299– 311, DOI: 10.1038/s41579-020-0327-xGoogle ScholarThere is no corresponding record for this reference.
- 43Myndrul, V.; Arreguin-Campos, R.; Iatsunskyi, I.; Di Scala, F.; Eersels, K.; van Grinsven, B. Photonic sensor based on surface imprinted polymers for enhanced point-of-care diagnosis of bacterial urinary tract infections. Biosens. Bioelectron. 2025, 268, 116916, DOI: 10.1016/j.bios.2024.116916Google ScholarThere is no corresponding record for this reference.
- 44Ganguly, A.; Ebrahimzadeh, T.; Zimmern, P.; De Nisco, N. J.; Prasad, S. L.-F. Novel Electrofluidic Capacitor Biosensor for Prostaglandin E2 Detection toward Early and Rapid Urinary Tract Infection Diagnosis. ACS Sens. 2022, 7 (1), 186– 198, DOI: 10.1021/acssensors.1c01951Google ScholarThere is no corresponding record for this reference.
- 45Zhuang, Q. Q.; Lu, L. Y.; Lin, Y. L.; Yan, X. L.; Chen, Q. Q.; Jiang, Y. C. A Self-Calibrating Chemiluminescence Sensor for Rapid and Precise Antibiotic Prescribing Guidelines on Urinary Tract Infections. ACS Sens. 2025, 10 (3), 2203– 2211, DOI: 10.1021/acssensors.4c03503Google ScholarThere is no corresponding record for this reference.
- 46Kaushik, A. M.; Hsieh, K.; Mach, K. E.; Lewis, S.; Puleo, C. M.; Carroll, K. C.; Liao, J. C.; Wang, T.-Z. Droplet-Based Single-Cell Measurements of 16S rRNA Enable Integrated Bacteria Identification and Pheno-Molecular Antimicrobial Susceptibility Testing from Clinical Samples in 30 min. Adv. Sci. 2021, 8 (6), 2003419, DOI: 10.1002/advs.202003419Google ScholarThere is no corresponding record for this reference.
- 47Zhang, W.; Sun, H.; He, S.; Chen, X.; Yao, L.; Zhou, L. Compound Raman microscopy for rapid diagnosis and antimicrobial susceptibility testing of pathogenic bacteria in urine. Front. Microbiol. 2022, 13, 874966, DOI: 10.3389/fmicb.2022.874966Google ScholarThere is no corresponding record for this reference.
- 48Liao, J. C.; Mastali, M.; Gau, V.; Suchard, M. A.; Møller, A. K.; Bruckner, D. A. Use of Electrochemical DNA Biosensors for Rapid Molecular Identification of Uropathogens in Clinical Urine Specimens. J. Clin. Microbiol. 2006, 44 (2), 561– 570, DOI: 10.1128/JCM.44.2.561-570.2006Google ScholarThere is no corresponding record for this reference.
- 49Hsieh, K.; Mach, K. E.; Zhang, P.; Liao, J. C.; Wang, T.-H. Combating Antimicrobial Resistance via Single-Cell Diagnostic Technologies Powered by Droplet Microfluidics. Acc. Chem. Res. 2022, 55 (2), 123– 133, DOI: 10.1021/acs.accounts.1c00462Google ScholarThere is no corresponding record for this reference.
- 50Jiang, X.; Borkum, T.; Shprits, S.; Boen, J.; Arshavsky-Graham, S.; Rofman, B.; Strauss, M.; Colodner, R.; Sulam, J.; Halachmi, S. Accurate Prediction of Antimicrobial Susceptibility for Point-of-Care Testing of Urine in Less than 90 minutes via iPRISM Cassettes. Adv. Sci. 2023, 10 (31), 2303285, DOI: 10.1002/advs.202303285Google ScholarThere is no corresponding record for this reference.
- 51Leonard, H.; Halachmi, S.; Ben-Dov, N.; Nativ, O.; Segal, E. Unraveling Antimicrobial Susceptibility of Bacterial Networks on Micropillar Architectures Using Intrinsic Phase-Shift Spectroscopy. ACS Nano 2017, 11 (6), 6167– 6177, DOI: 10.1021/acsnano.7b02217Google ScholarThere is no corresponding record for this reference.
- 52Heuer, C.; Leonard, H.; Nitzan, N.; Lavy-Alperovitch, A.; Massad-Ivanir, N.; Scheper, T. Antifungal Susceptibility Testing of Aspergillus niger on Silicon Microwells by Intensity-Based Reflectometric Interference Spectroscopy. ACS Infect. Dis. 2020, 6 (10), 2560– 2566, DOI: 10.1021/acsinfecdis.0c00234Google ScholarThere is no corresponding record for this reference.
- 53Heuer, C.; Jiang, X.; Ron, G.; Ternyak, O.; Scheper, T.; Bahnemann, J. Photonic Si microwell architectures for rapid antifungal susceptibility determination of Candida auris. Chem. Commun. 2024, 60 (10), 1305– 1308, DOI: 10.1039/D3CC04446GGoogle ScholarThere is no corresponding record for this reference.
- 54Heuer, C.; Preuss, J.-A.; Buttkewitz, M.; Scheper, T.; Segal, E.; Bahnemann, J. A 3D-printed microfluidic gradient generator with integrated photonic silicon sensors for rapid antimicrobial susceptibility testing. Lab Chip 2022, 22 (24), 4950– 4961, DOI: 10.1039/D2LC00640EGoogle ScholarThere is no corresponding record for this reference.
- 55Leonard, H.; Jiang, X.; Arshavsky-Graham, S.; Holtzman, L.; Haimov, Y.; Weizman, D. Shining light in blind alleys: deciphering bacterial attachment in silicon microstructures. Nanoscale Horiz. 2022, 7 (7), 729– 742, DOI: 10.1039/D2NH00130FGoogle ScholarThere is no corresponding record for this reference.
- 56Hanley, J. A.; McNeil, B. J. The meaning and use of the area under a receiver operating characteristic (ROC) curve. Radiology 1982, 143 (1), 29– 36, DOI: 10.1148/radiology.143.1.7063747Google ScholarThere is no corresponding record for this reference.
- 57Massad-Ivanir, N.; Mirsky, Y.; Nahor, A.; Edrei, E.; Bonanno-Young, L. M.; Ben Dov, N. Trap and track: designing self-reporting porous Si photonic crystals for rapid bacteria detection. Analyst 2014, 139 (16), 3885– 3894, DOI: 10.1039/C4AN00364KGoogle ScholarThere is no corresponding record for this reference.
- 58Ito, W.; Choi, N.; Letner, G.; Genz, N.; Prokop, D.; Valadon, C.; Sardiu, M. E.; Smith, H.; Whiles, B. B.; Molina, W. R. Preoperative urine culture with contaminants is not associated with increased risk for urinary tract infection after ureteroscopic stone treatment. World J. Urol. 2024, 42 (1), 159, DOI: 10.1007/s00345-024-04793-wGoogle ScholarThere is no corresponding record for this reference.
- 59Nye, T. M.; Zou, Z.; Obernuefemann, C. L. P.; Pinkner, J. S.; Lowry, E.; Kleinschmidt, K.; Bergeron, K.; Klim, A.; Dodson, K. W.; Flores-Mireles, A. L. Microbial co-occurrences on catheters from long-term catheterized patients. Nat. Commun. 2024, 15 (1), 61, DOI: 10.1038/s41467-023-44095-0Google ScholarThere is no corresponding record for this reference.
- 60Muller, A.; Gbaguidi-Haore, H.; Cholley, P.; Hocquet, D.; Sauget, M.; Bertrand, X. Hospital-diagnosed infections with Escherichia coli clonal group ST131 are mostly acquired in the community. Sci. Rep. 2021, 11 (1), 5702, DOI: 10.1038/s41598-021-85116-6Google ScholarThere is no corresponding record for this reference.
- 61Toval, F.; Köhler, C.-D.; Vogel, U.; Wagenlehner, F.; Mellmann, A.; Fruth, A. Characterization of Escherichia coli Isolates from Hospital Inpatients or Outpatients with Urinary Tract Infection. J. Clin. Microbiol. 2014, 52 (2), 407– 418, DOI: 10.1128/JCM.02069-13Google ScholarThere is no corresponding record for this reference.
- 62Gu, W.; Huang, W.; Zhang, J.; Qian, S.; Cao, H.; Ge, L. Evaluation of urinary inflammatory index in rapid screening of urinary tract infection. Sci. Rep. 2020, 10 (1), 19306, DOI: 10.1038/s41598-020-76352-3Google ScholarThere is no corresponding record for this reference.
- 63Gajdacs, M.; Doczi, I.; Abrok, M.; Lazar, A.; Burian, K. Epidemiology of candiduria and Candida urinary tract infections in inpatients and outpatients: results from a 10-year retrospective survey. Cent. Eur. J. Urol. 2019, 72 (2), 209– 214, DOI: 10.5173/ceju.2019.1909Google ScholarThere is no corresponding record for this reference.
- 64Dias, V. Candida Species in the Urinary Tract: Is it a Fungal Infection or Not?. Future Microbiol. 2020, 15 (2), 81– 83, DOI: 10.2217/fmb-2019-0262Google ScholarThere is no corresponding record for this reference.
- 65Behzadi, P.; Behzadi, E.; Ranjbar, R. Urinary tract infections and Candida albicans. Cent. Eur. J. Urol. 2015, 68 (1), 96– 101, DOI: 10.5173/ceju.2015.01.474Google ScholarThere is no corresponding record for this reference.
- 66La Bella, A. A.; Andersen, M. J.; Gervais, N. C.; Molina, J. J.; Molesan, A.; Stuckey, P. V.; Wensing, L.; Nobile, C. J.; Shapiro, R. S.; Santiago-Tirado, F. H. The catheterized bladder environment promotes Efg1- and Als1-dependent Candida albicans infection. Sci. Adv. 2023, 9 (9), eade7689 DOI: 10.1126/sciadv.ade7689Google ScholarThere is no corresponding record for this reference.
- 67Sher, E. K.; Džidić-Krivić, A.; Sesar, A.; Farhat, E. K.; Čeliković, A.; Beća-Zećo, M. Current state and novel outlook on prevention and treatment of rising antibiotic resistance in urinary tract infections. Pharmacol. Ther. 2024, 261, 108688, DOI: 10.1016/j.pharmthera.2024.108688Google ScholarThere is no corresponding record for this reference.
- 68Polo, T. C. F.; Miot, H. A. Use of ROC curves in clinical and experimental studies. J. Vasc. Bras. 2020, 19, e20200186 DOI: 10.1590/1677-5449.200186Google ScholarThere is no corresponding record for this reference.
- 69Silva, S.; Henriques, M.; Oliveira, R.; Williams, D.; Azeredo, J. In Vitro Biofilm Activity of Non-Candida albicansCandida Species. Curr. Microbiol. 2010, 61 (6), 534– 540, DOI: 10.1007/s00284-010-9649-7Google ScholarThere is no corresponding record for this reference.
- 70Lu, X.; Chow, J. J. M.; Koo, S. H.; Tan, T. Y.; Jiang, B.; Ai, Y. Enhanced Molecular Diagnosis of Bloodstream Candida Infection with Size-Based Inertial Sorting at Submicron Resolution. Anal. Chem. 2020, 92 (23), 15579– 15586, DOI: 10.1021/acs.analchem.0c03718Google ScholarThere is no corresponding record for this reference.
- 71Noble, S. M.; Gianetti, B. A.; Witchley, J. N. Candida albicans cell-type switching and functional plasticity in the mammalian host. Nat. Rev. Microbiol. 2017, 15 (2), 96– 108, DOI: 10.1038/nrmicro.2016.157Google ScholarThere is no corresponding record for this reference.
- 72Wijesinghe, G.; Dilhari, A.; Gayani, B.; Kottegoda, N.; Samaranayake, L.; Weerasekera, M. Influence of Laboratory Culture Media on in vitro Growth, Adhesion, and Biofilm Formation of Pseudomonas aeruginosa and Staphylococcus aureus. Med. Princ. Pract. 2019, 28 (1), 28– 35, DOI: 10.1159/000494757Google ScholarThere is no corresponding record for this reference.
- 73Jones-Freeman, B.; Chonwerawong, M.; Marcelino, V. R.; Deshpande, A. V.; Forster, S. C.; Starkey, M. R. The microbiome and host mucosal interactions in urinary tract diseases. Mucosal Immunol. 2021, 14 (4), 779– 792, DOI: 10.1038/s41385-020-00372-5Google ScholarThere is no corresponding record for this reference.
- 74Sujith, S.; Solomon, A. P.; Rayappan, J. B. B. Comprehensive insights into UTIs: from pathophysiology to precision diagnosis and management. Front. Cell. Infect. Microbiol. 2024, 14, 1402941, DOI: 10.3389/fcimb.2024.1402941Google ScholarThere is no corresponding record for this reference.
- 75Brosel-Oliu, S.; Mergel, O.; Uria, N.; Abramova, N.; Van Rijn, P.; Bratov, A. 3D impedimetric sensors as a tool for monitoring bacterial response to antibiotics. Lab Chip 2019, 19 (8), 1436– 1447, DOI: 10.1039/C8LC01220BGoogle ScholarThere is no corresponding record for this reference.
- 76Prinz Setter, O.; Jiang, X.; Segal, E. Rising to the surface: capturing and detecting bacteria by rationally-designed surfaces. Curr. Opin. Biotechnol. 2023, 83, 102969, DOI: 10.1016/j.copbio.2023.102969Google ScholarThere is no corresponding record for this reference.
- 77Soares, A.; Roussel, V.; Pestel-Caron, M.; Barreau, M.; Caron, F.; Bouffartigues, E.; Chevalier, S.; Etienne, M. Understanding Ciprofloxacin Failure in Pseudomonas aeruginosa Biofilm: Persister Cells Survive Matrix Disruption. Front. Microbiol. 2019, 10, 2603, DOI: 10.3389/fmicb.2019.02603Google ScholarThere is no corresponding record for this reference.
- 78Ojkic, N.; Lilja, E.; Direito, S.; Dawson, A.; Allen, R. J.; Waclaw, B. A Roadblock-and-Kill Mechanism of Action Model for the DNA-Targeting Antibiotic Ciprofloxacin. Antimicrob. Agents Chemother. 2020, 64 (9), 10– 1128, DOI: 10.1128/AAC.02487-19Google ScholarThere is no corresponding record for this reference.
- 79CLSI. Performance Standards for Antimicrobial Susceptibility Testing, 35th edition. CLSI supplement M100; Clinical and Laboratory Standards Institute; 2025. http://em100.edaptivedocs.net/dashboard.aspx. Accessed 25.April.03.Google ScholarThere is no corresponding record for this reference.
- 80Needs, S. H.; Donmez, S. I.; Bull, S. P.; McQuaid, C.; Osborn, H. M. I.; Edwards, A. D. Challenges in Microfluidic and Point-of-Care Phenotypic Antimicrobial Resistance Tests. Front. Mech. Eng. 2020, 6, 73, DOI: 10.3389/fmech.2020.00073Google ScholarThere is no corresponding record for this reference.
- 81Wu, W.; Suo, Y.; Zhao, Q.; Cai, G.; Liu, Y.; Jin, W. Inoculum size-insensitive susceptibility determination of urine sample based on in-situ measurement of inducible enzyme activity after 20 min of antibiotic exposure. Anal. Chim. Acta 2023, 1282, 341858, DOI: 10.1016/j.aca.2023.341858Google ScholarThere is no corresponding record for this reference.
- 82Smith, K. P.; Kirby, J. E. The Inoculum Effect in the Era of Multidrug Resistance: Minor Differences in Inoculum Have Dramatic Effect on MIC Determination. Antimicrob. Agents Chemother. 2018, 62 (8), e00433–18 DOI: 10.1128/AAC.00433-18Google ScholarThere is no corresponding record for this reference.
- 83Yi, Q.; Cui, J.; Xiao, M.; Tang, M.-Z.; Zhang, H.-C.; Zhang, G. Rapid Phenotypic Antimicrobial Susceptibility Testing Using a Coulter Counter and Proliferation Rate Discrepancy. ACS Omega 2023, 8 (18), 16298– 16305, DOI: 10.1021/acsomega.3c00947Google ScholarThere is no corresponding record for this reference.
- 84Vasala, A.; Hytönen, V. P.; Laitinen, O. H. Modern Tools for Rapid Diagnostics of Antimicrobial Resistance. Front. Cell. Infect. Microbiol. 2020, 10, 308, DOI: 10.3389/fcimb.2020.00308Google ScholarThere is no corresponding record for this reference.
- 85Alfouzan, W. A.; Dhar, R. Candiduria: Evidence-based approach to management, are we there yet?. J. Mycol. Médicale 2017, 27 (3), 293– 302, DOI: 10.1016/j.mycmed.2017.04.005Google ScholarThere is no corresponding record for this reference.
- 86World Health Organization. Landscape analysis of commercially available and pipeline in vitro diagnostics for fungal priority pathogens; World Health Organization, 2025. https://www.who.int/publications/i/item/9789240105539. Accessed 25.April.12.Google ScholarThere is no corresponding record for this reference.
- 87World Health Organization. WHO issues its first-ever reports on tests and treatments for fungal infections; World Health Organization, 2025. https://www.who.int/news/item/01-04-2025-who-issues-its-first-ever-reports-on-tests-and-treatments-for-fungal-infections. Accessed 25.April.12.Google ScholarThere is no corresponding record for this reference.
Cited By
This article has not yet been cited by other publications.
Article Views
Altmetric
Citations
Article Views are the COUNTER-compliant sum of full text article downloads since November 2008 (both PDF and HTML) across all institutions and individuals. These metrics are regularly updated to reflect usage leading up to the last few days.
Citations are the number of other articles citing this article, calculated by Crossref and updated daily. Find more information about Crossref citation counts.
The Altmetric Attention Score is a quantitative measure of the attention that a research article has received online. Clicking on the donut icon will load a page at altmetric.com with additional details about the score and the social media presence for the given article. Find more information on the Altmetric Attention Score and how the score is calculated.
Recommended Articles
Abstract
Figure 1
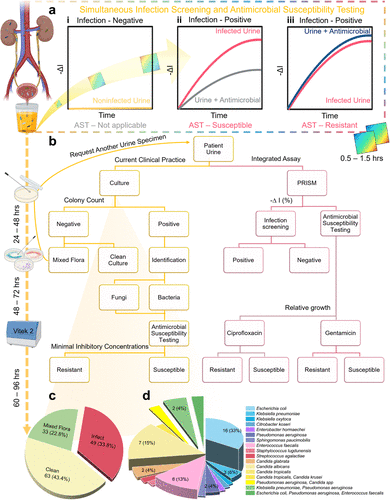
Figure 1. Schematic overview of iPRISM for rapid UTI diagnosis and comparative workflow analysis. (a) iPRISM enables simultaneous infection screening and AST directly on clinical urine specimens. Three infection scenarios can be distinguished based on the measured signal: (i) negative infection, (ii) positive infection with an antimicrobial-susceptible uropathogen, and (iii) positive infection with an antimicrobial-resistant uropathogen. (b) Overall flow of iPRISM and clinical routine UTI diagnostics. Current clinical UTI workflow requires sequential steps: (1) standard urine culture (24–48 h for screening), (2) subculture for microbial identification, and (3) AST (e.g., using VITEK 2), with turnaround time from sample to results ranging from 3 to 7 days. The iPRISM system described herein proceeds without culturing, achieving diagnosis within 90 min. (c) Distribution of infection scenario of the collected clinical urine specimens. (d) Isolated causative agents of culture-positive specimens (derived from the red slice in c), categorized by Gram-negative bacteria (blue-purple), Gram-positive bacteria (pink-red), fungal species (yellow-orange), and polymicrobial infections (yellow-green). Created with BioRender.
Figure 2

Figure 2. iPRISM for UTI detection directly on clinical human urine samples. (a–d) Representative real-time characteristic bacterial growth curves as measured by the iPRISM assay for (a) Escherichia coli, (b) Klebsiella spp., (c)Enterococcus faecalis, and (d) Streptococcus agalactiae. (e–h) Corresponding HR-SEM images of the bacterial species in panels a–d, respectively, demonstrating their spatial distribution on the surface of microwells for retrieved silicon chips from the iPRISM device (for each bacterium, the curve corresponding to the image is marked with a star). Scale bars denote 3 μm.
Figure 3
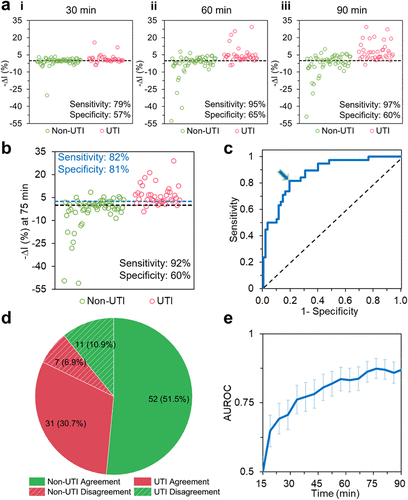
Figure 3. iPRISM for infection screening on fresh untreated clinical urine samples (63 clean samples and 38 bacterial infected samples including the bacterial-fungal coinfection one). (a) Intensity value changes after incubation in iPRISM device for 30, 60, and 90 min. The dashed line represents a double-blind screening threshold of −ΔI (%) = 0. Samples surpassing this threshold were considered positive for UTI infection by the iPRISM assay. (b) Intensity value changes at 75 min. The black dashed line represents the original detection threshold of −ΔI (%) = 0, while the blue dashed line is the new optimized threshold of −ΔI (%) = 2.59. Each circle corresponds to one clinical specimen. Green circles represent specimens identified as noninfected by clinical culture, and red circles represent specimens with positive culture results. (c) ROC analysis at 75 min. The point in the graph indicated by an arrow corresponds to the optimal diagnostic threshold yielding the maximum Youden’s index. (d) Classification agreement at 75 min using the established optimal threshold for data shown in (b). (e) Time-dependent AUC performance. Error bars represent standard error of area under the ROC curve statistics.
Figure 4
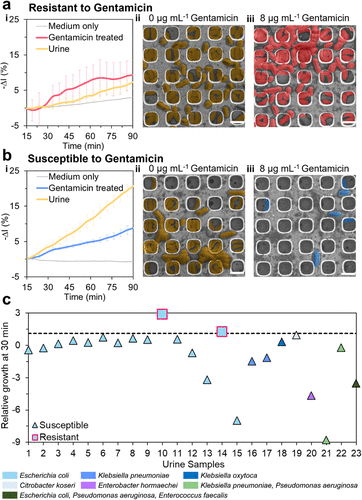
Figure 4. Direct iPRISM AST assay performance in clinical human urine samples with exposure to MIC breakpoint concentration of gentamicin (8 μg mL–1). (a, b) Representative gentamicin-resistant (a) and susceptible (b) cases where E. coli was isolated from culture positive urine samples. (i) Real-time iPRISM characteristic curves showing −ΔI (%) values (n = 3). (ii, iii) HR-SEM images of (ii) control and (iii) post-treatment. Scale bars represent 3 μm. (c) iPRISM relative growth (RG) values at 30 min after exposure to a gentamicin breakpoint concentration of 8 μg mL–1 of urine samples from suspected infected UTI patients (n = 23; 2 resistant, 21 susceptible). The dashed line indicates the predefined threshold (RG at 30 min = 0.95) for resistance classification.
References
This article references 87 other publications.
- 1Timm, M. R.; Russell, S. K.; Hultgren, S. J. Urinary tract infections: pathogenesis, host susceptibility and emerging therapeutics. Nat. Rev. Microbiol. 2025, 23 (2), 72– 86, DOI: 10.1038/s41579-024-01092-4There is no corresponding record for this reference.
- 2Simoni, A.; Schwartz, L.; Junquera, G. Y.; Ching, C. B.; Spencer, J. D. Current and emerging strategies to curb antibiotic-resistant urinary tract infections. Nat. Rev. Urol. 2024, 21 (12), 707– 722, DOI: 10.1038/s41585-024-00877-9There is no corresponding record for this reference.
- 3Yang, X.; Chen, H.; Zheng, Y.; Qu, S.; Wang, H.; Yi, F. Disease burden and long-term trends of urinary tract infections: A worldwide report. Front. Public Health 2022, 10, 888205, DOI: 10.3389/fpubh.2022.888205There is no corresponding record for this reference.
- 4Duque-Sanchez, L.; Qu, Y.; Voelcker, N. H.; Thissen, H. Tackling catheter-associated urinary tract infections with next-generation antimicrobial technologies. J. Biomed. Mater. Res., Part A 2024, 112 (3), 312– 335, DOI: 10.1002/jbm.a.37630There is no corresponding record for this reference.
- 5Bermudez, T.; Schmitz, J. E.; Boswell, M.; Humphries, R. Novel technologies for the diagnosis of urinary tract infections. J. Clin. Microbiol. 2025, 63 (2), e00306–24 DOI: 10.1128/jcm.00306-24There is no corresponding record for this reference.
- 6Davenport, M.; Mach, K. E.; Shortliffe, L. M. D.; Banaei, N.; Wang, T.-H.; Liao, J. C. New and developing diagnostic technologies for urinary tract infections. Nat. Rev. Urol. 2017, 14 (5), 296– 310, DOI: 10.1038/nrurol.2017.20There is no corresponding record for this reference.
- 7Singh, S.; Numan, A.; Cinti, S. Point-of-Care for Evaluating Antimicrobial Resistance through the Adoption of Functional Materials. Anal. Chem. 2022, 94 (1), 26– 40, DOI: 10.1021/acs.analchem.1c03856There is no corresponding record for this reference.
- 8World Health Organization. Global research agenda for antimicrobial resistance in human health; World Health Organization; 2025, https://www.who.int/publications/i/item/9789240102309; Accessed 25.April.02.There is no corresponding record for this reference.
- 9Tullus, K.; Shaikh, N. Urinary tract infections in children. Lancet 2020, 395 (10237), 1659– 1668, DOI: 10.1016/S0140-6736(20)30676-0There is no corresponding record for this reference.
- 10Petchakup, C.; Chen, Y. Y. C.; Tay, H. M.; Ong, H. B.; Hon, P. Y.; De, P. P. Rapid Screening of Urinary Tract Infection Using Microfluidic Inertial-Impedance Cytometry. ACS Sens. 2023, 8 (8), 3136– 3145, DOI: 10.1021/acssensors.3c00819There is no corresponding record for this reference.
- 11Wat, J. K.-H.; Xu, M.; Nan, L.; Lin, H.; To, K. K.-W.; Shum, H. C.; Hassan, S. U. Rapid antimicrobial susceptibility tests performed by self-diluting microfluidic chips for drug resistance studies and point-of-care diagnostics. Microsyst. Nanoeng. 2025, 11 (1), 110, DOI: 10.1038/s41378-025-00938-yThere is no corresponding record for this reference.
- 12Yang, J.; Li, G.; Chen, S.; Su, X.; Xu, D.; Zhai, Y. Machine Learning-Assistant Colorimetric Sensor Arrays for Intelligent and Rapid Diagnosis of Urinary Tract Infection. ACS Sens. 2024, 9 (4), 1945– 1956, DOI: 10.1021/acssensors.3c02687There is no corresponding record for this reference.
- 13Szlachta-McGinn, A.; Douglass, K. M.; Chung, U. Y. R.; Jackson, N. J.; Nickel, J. C.; Ackerman, A. L. Molecular Diagnostic Methods Versus Conventional Urine Culture for Diagnosis and Treatment of Urinary Tract Infection: A Systematic Review and Meta-analysis. Eur. Urol. Open Sci. 2022, 44, 113– 124, DOI: 10.1016/j.euros.2022.08.009There is no corresponding record for this reference.
- 14Mancuso, G.; Midiri, A.; Gerace, E.; Marra, M.; Zummo, S.; Biondo, C. Urinary Tract Infections: The Current Scenario and Future Prospects. Pathogens 2023, 12 (4), 623, DOI: 10.3390/pathogens12040623There is no corresponding record for this reference.
- 15Brubaker, L.; Chai, T. C.; Horsley, H.; Khasriya, R.; Moreland, R. B.; Wolfe, A. J. Tarnished gold─the “standard” urine culture: reassessing the characteristics of a criterion standard for detecting urinary microbes. Front. Urol. 2023, 3, 1206046, DOI: 10.3389/fruro.2023.1206046There is no corresponding record for this reference.
- 16Moreland, R. B.; Brubaker, L.; Tinawi, L.; Wolfe, A. J. Rapid and accurate testing for urinary tract infection: new clothes for the emperor. Clin. Microbiol. Rev. 2025, 38 (1), e00129–24 DOI: 10.1128/cmr.00129-24There is no corresponding record for this reference.
- 17Zhang, J.; Wang, M.; Xiao, J.; Wang, M.; Liu, Y.; Gao, X. Metabolism-Triggered Plasmonic Nanosensor for Bacterial Detection and Antimicrobial Susceptibility Testing of Clinical Isolates. ACS Sens. 2024, 9 (1), 379– 387, DOI: 10.1021/acssensors.3c02144There is no corresponding record for this reference.
- 18Gajic, I.; Kabic, J.; Kekic, D.; Jovicevic, M.; Milenkovic, M.; Mitic Culafic, D. Antimicrobial Susceptibility Testing: A Comprehensive Review of Currently Used Methods. Antibiotics 2022, 11 (4), 427, DOI: 10.3390/antibiotics11040427There is no corresponding record for this reference.
- 19Sturm, A.; Jóźwiak, G.; Verge, M. P.; Munch, L.; Cathomen, G.; Vocat, A.; Luraschi-Eggemann, A.; Orlando, C.; Fromm, K.; Delarze, E. Accurate and rapid antibiotic susceptibility testing using a machine learning-assisted nanomotion technology platform. Nat. Commun. 2024, 15 (1), 2037, DOI: 10.1038/s41467-024-46213-yThere is no corresponding record for this reference.
- 20Kandavalli, V.; Karempudi, P.; Larsson, J.; Elf, J. Rapid antibiotic susceptibility testing and species identification for mixed samples. Nat. Commun. 2022, 13 (1), 6215, DOI: 10.1038/s41467-022-33659-1There is no corresponding record for this reference.
- 21Leonard, H.; Colodner, R.; Halachmi, S.; Segal, E. Recent Advances in the Race to Design a Rapid Diagnostic Test for Antimicrobial Resistance. ACS Sens. 2018, 3 (11), 2202– 2217, DOI: 10.1021/acssensors.8b00900There is no corresponding record for this reference.
- 22Ho, C. S.; Wong, C. T. H.; Aung, T. T.; Lakshminarayanan, R.; Mehta, J. S.; Rauz, S. Antimicrobial resistance: a concise update. Lancet Microbe 2025, 6 (1), 100947, DOI: 10.1016/j.lanmic.2024.07.010There is no corresponding record for this reference.
- 23Faigenbaum-Romm, R.; Yedidi, N.; Gefen, O.; Katsowich-Nagar, N.; Aroeti, L.; Ronin, I.; Bar-Meir, M.; Rosenshine, I.; Balaban, N. Q. Uncovering phenotypic inheritance from single cells with Microcolony-seq. Cell 2025, 188 (19), 5313– 5331.e18, DOI: 10.1016/j.cell.2025.08.001There is no corresponding record for this reference.
- 24David, S. S. B.; Romano, R.; Rahamim-Cohen, D.; Azuri, J.; Greenfeld, S.; Gedassi, B.; Lerner, U. AI driven decision support reduces antibiotic mismatches and inappropriate use in outpatient urinary tract infections. Npj Digital Med. 2025, 8 (1), 61, DOI: 10.1038/s41746-024-01400-5There is no corresponding record for this reference.
- 25Pew Charitable Trusts. Health Experts Establish Targets to Improve Hospital Antibiotic Prescribing; Pew Charitable Trusts, 2021.https://pew.org/2Qca7pi. Accessed 25.May.26.There is no corresponding record for this reference.
- 26Huang, R.; Cai, X.; Du, J.; Lian, J.; Hui, P.; Gu, M. Bioinspired Plasmonic Nanosensor for on-Site Antimicrobial Susceptibility Testing in Urine Samples. ACS Nano 2022, 16 (11), 19229– 19239, DOI: 10.1021/acsnano.2c08532There is no corresponding record for this reference.
- 27Zhang, P.; Kaushik, A. M.; Mach, K. E.; Hsieh, K.; Liao, J. C.; Wang, T.-H. Facile syringe filter-enabled bacteria separation, enrichment, and buffer exchange for clinical isolation-free digital detection and characterization of bacterial pathogens in urine. Analyst 2021, 146 (8), 2475– 2483, DOI: 10.1039/D1AN00039JThere is no corresponding record for this reference.
- 28Zhang, F.; Jiang, J.; McBride, M.; Yang, Y.; Mo, M.; Iriya, R.; Peterman, J.; Jing, W.; Grys, T.; Haydel, S. E. Direct Antimicrobial Susceptibility Testing on Clinical Urine Samples by Optical Tracking of Single Cell Division Events. Small 2020, 16 (52), e2004148 DOI: 10.1002/smll.202004148There is no corresponding record for this reference.
- 29Reszetnik, G.; Hammond, K.; Mahshid, S.; AbdElfatah, T.; Nguyen, D.; Corsini, R.; Caya, C.; Papenburg, J.; Cheng, M. P.; Yansouni, C. P. Next-generation rapid phenotypic antimicrobial susceptibility testing. Nat. Commun. 2024, 15 (1), 9719, DOI: 10.1038/s41467-024-53930-xThere is no corresponding record for this reference.
- 30Riester, O.; Laufer, S.; Mertelsmann, R.; Deigner, H.-P. Sensors for Antibiotics Susceptibility Testing-Recent Advances. Adv. Sens. Res. 2024, 3 (10), 2400046, DOI: 10.1002/adsr.202400046There is no corresponding record for this reference.
- 31Zhang, M.; Hong, W.; Abutaleb, N. S.; Li, J.; Dong, P.-T.; Zong, C.; Wang, P.; Seleem, M. N.; Cheng, J.-X. Rapid Determination of Antimicrobial Susceptibility by Stimulated Raman Scattering Imaging of D2 O Metabolic Incorporation in a Single Bacterium. Adv. Sci. 2020, 7 (19), 2001452, DOI: 10.1002/advs.202001452There is no corresponding record for this reference.
- 32Michael, I.; Kim, D.; Gulenko, O.; Kumar, S.; Kumar, S.; Clara, J. A fidget spinner for the point-of-care diagnosis of urinary tract infection. Nat. Biomed. Eng. 2020, 4 (6), 591– 600, DOI: 10.1038/s41551-020-0557-2There is no corresponding record for this reference.
- 33Yang, S.; Berto, F. G.; Denstedt, J.; Tang, H.; Zhang, J. Point-of-Care Urinary Tract Infection (UTI) Diagnosis Enhanced by Nanostructured Biosensors: Review Paper. Adv. Sens. Res. 2024, 3 (10), 2400051, DOI: 10.1002/adsr.202400051There is no corresponding record for this reference.
- 34Velican, A. M.; Măruţescu, L.; Kamerzan, C.; Cristea, V. C.; Banu, O.; Borcan, E. Rapid Detection and Antibiotic Susceptibility of Uropathogenic Escherichia coli by Flow Cytometry. Microorganisms 2020, 8 (8), 1233, DOI: 10.3390/microorganisms8081233There is no corresponding record for this reference.
- 35Altobelli, E.; Mohan, R.; Mach, K. E.; Sin, M. L. Y.; Anikst, V.; Buscarini, M. Integrated Biosensor Assay for Rapid Uropathogen Identification and Phenotypic Antimicrobial Susceptibility Testing. Eur. Urol. Focus 2017, 3 (2–3), 293– 299, DOI: 10.1016/j.euf.2015.12.010There is no corresponding record for this reference.
- 36Avesar, J.; Rosenfeld, D.; Truman-Rosentsvit, M.; Ben-Arye, T.; Geffen, Y.; Bercovici, M.; Levenberg, S. Rapid phenotypic antimicrobial susceptibility testing using nanoliter arrays. Proc. Natl. Acad. Sci. 2017, 114 (29), E5787-E5795 DOI: 10.1073/pnas.1703736114There is no corresponding record for this reference.
- 37Burg, L.; Crewe, G.; DiMeo, J.; Guo, X.; Li, C. G.; Mayol, M.; Tempesta, A.; Lauzier, W.; Markham, R.; Crissy, K. Rapid pathogen identification and phenotypic antimicrobial susceptibility directly from urine specimens. Sci. Rep. 2022, 12 (1), 18315, DOI: 10.1038/s41598-022-22792-yThere is no corresponding record for this reference.
- 38Gopal, A.; Yan, L.; Kashif, S.; Munshi, T.; Roy, V. A. L.; Voelcker, N. H.; Chen, X. Biosensors and Point-of-Care Devices for Bacterial Detection: Rapid Diagnostics Informing Antibiotic Therapy. Adv. Healthcare Mater. 2022, 11 (3), e2101546 DOI: 10.1002/adhm.202101546There is no corresponding record for this reference.
- 39Gonchar, K. A.; Nazarovskaia, D. A.; Domnin, P. A.; Gyuppenen, O. D.; Erokhina, A. A.; Wang, M. AuAg-Modified Porous Silicon Nanowires for Dual-Mode Interferometric and SERS-Based Bacterial Detection. ACS Appl. Mater. Interfaces 2025, 17 (23), 33691– 33700, DOI: 10.1021/acsami.5c06953There is no corresponding record for this reference.
- 40Arshavsky-Graham, S.; Massad-Ivanir, N.; Segal, E.; Weiss, S. Porous Silicon-Based Photonic Biosensors: Current Status and Emerging Applications. Anal. Chem. 2019, 91 (1), 441– 467, DOI: 10.1021/acs.analchem.8b05028There is no corresponding record for this reference.
- 41Qin, N.; Zhao, P.; Ho, E. A.; Xin, G.; Ren, C. L. Microfluidic Technology for Antibacterial Resistance Study and Antibiotic Susceptibility Testing: Review and Perspective. ACS Sens. 2021, 6 (1), 3– 21, DOI: 10.1021/acssensors.0c02175There is no corresponding record for this reference.
- 42Van Belkum, A.; Burnham, C.-A. D.; Rossen, J. W. A.; Mallard, F.; Rochas, O.; Dunne, W. M. Innovative and rapid antimicrobial susceptibility testing systems. Nat. Rev. Microbiol. 2020, 18 (5), 299– 311, DOI: 10.1038/s41579-020-0327-xThere is no corresponding record for this reference.
- 43Myndrul, V.; Arreguin-Campos, R.; Iatsunskyi, I.; Di Scala, F.; Eersels, K.; van Grinsven, B. Photonic sensor based on surface imprinted polymers for enhanced point-of-care diagnosis of bacterial urinary tract infections. Biosens. Bioelectron. 2025, 268, 116916, DOI: 10.1016/j.bios.2024.116916There is no corresponding record for this reference.
- 44Ganguly, A.; Ebrahimzadeh, T.; Zimmern, P.; De Nisco, N. J.; Prasad, S. L.-F. Novel Electrofluidic Capacitor Biosensor for Prostaglandin E2 Detection toward Early and Rapid Urinary Tract Infection Diagnosis. ACS Sens. 2022, 7 (1), 186– 198, DOI: 10.1021/acssensors.1c01951There is no corresponding record for this reference.
- 45Zhuang, Q. Q.; Lu, L. Y.; Lin, Y. L.; Yan, X. L.; Chen, Q. Q.; Jiang, Y. C. A Self-Calibrating Chemiluminescence Sensor for Rapid and Precise Antibiotic Prescribing Guidelines on Urinary Tract Infections. ACS Sens. 2025, 10 (3), 2203– 2211, DOI: 10.1021/acssensors.4c03503There is no corresponding record for this reference.
- 46Kaushik, A. M.; Hsieh, K.; Mach, K. E.; Lewis, S.; Puleo, C. M.; Carroll, K. C.; Liao, J. C.; Wang, T.-Z. Droplet-Based Single-Cell Measurements of 16S rRNA Enable Integrated Bacteria Identification and Pheno-Molecular Antimicrobial Susceptibility Testing from Clinical Samples in 30 min. Adv. Sci. 2021, 8 (6), 2003419, DOI: 10.1002/advs.202003419There is no corresponding record for this reference.
- 47Zhang, W.; Sun, H.; He, S.; Chen, X.; Yao, L.; Zhou, L. Compound Raman microscopy for rapid diagnosis and antimicrobial susceptibility testing of pathogenic bacteria in urine. Front. Microbiol. 2022, 13, 874966, DOI: 10.3389/fmicb.2022.874966There is no corresponding record for this reference.
- 48Liao, J. C.; Mastali, M.; Gau, V.; Suchard, M. A.; Møller, A. K.; Bruckner, D. A. Use of Electrochemical DNA Biosensors for Rapid Molecular Identification of Uropathogens in Clinical Urine Specimens. J. Clin. Microbiol. 2006, 44 (2), 561– 570, DOI: 10.1128/JCM.44.2.561-570.2006There is no corresponding record for this reference.
- 49Hsieh, K.; Mach, K. E.; Zhang, P.; Liao, J. C.; Wang, T.-H. Combating Antimicrobial Resistance via Single-Cell Diagnostic Technologies Powered by Droplet Microfluidics. Acc. Chem. Res. 2022, 55 (2), 123– 133, DOI: 10.1021/acs.accounts.1c00462There is no corresponding record for this reference.
- 50Jiang, X.; Borkum, T.; Shprits, S.; Boen, J.; Arshavsky-Graham, S.; Rofman, B.; Strauss, M.; Colodner, R.; Sulam, J.; Halachmi, S. Accurate Prediction of Antimicrobial Susceptibility for Point-of-Care Testing of Urine in Less than 90 minutes via iPRISM Cassettes. Adv. Sci. 2023, 10 (31), 2303285, DOI: 10.1002/advs.202303285There is no corresponding record for this reference.
- 51Leonard, H.; Halachmi, S.; Ben-Dov, N.; Nativ, O.; Segal, E. Unraveling Antimicrobial Susceptibility of Bacterial Networks on Micropillar Architectures Using Intrinsic Phase-Shift Spectroscopy. ACS Nano 2017, 11 (6), 6167– 6177, DOI: 10.1021/acsnano.7b02217There is no corresponding record for this reference.
- 52Heuer, C.; Leonard, H.; Nitzan, N.; Lavy-Alperovitch, A.; Massad-Ivanir, N.; Scheper, T. Antifungal Susceptibility Testing of Aspergillus niger on Silicon Microwells by Intensity-Based Reflectometric Interference Spectroscopy. ACS Infect. Dis. 2020, 6 (10), 2560– 2566, DOI: 10.1021/acsinfecdis.0c00234There is no corresponding record for this reference.
- 53Heuer, C.; Jiang, X.; Ron, G.; Ternyak, O.; Scheper, T.; Bahnemann, J. Photonic Si microwell architectures for rapid antifungal susceptibility determination of Candida auris. Chem. Commun. 2024, 60 (10), 1305– 1308, DOI: 10.1039/D3CC04446GThere is no corresponding record for this reference.
- 54Heuer, C.; Preuss, J.-A.; Buttkewitz, M.; Scheper, T.; Segal, E.; Bahnemann, J. A 3D-printed microfluidic gradient generator with integrated photonic silicon sensors for rapid antimicrobial susceptibility testing. Lab Chip 2022, 22 (24), 4950– 4961, DOI: 10.1039/D2LC00640EThere is no corresponding record for this reference.
- 55Leonard, H.; Jiang, X.; Arshavsky-Graham, S.; Holtzman, L.; Haimov, Y.; Weizman, D. Shining light in blind alleys: deciphering bacterial attachment in silicon microstructures. Nanoscale Horiz. 2022, 7 (7), 729– 742, DOI: 10.1039/D2NH00130FThere is no corresponding record for this reference.
- 56Hanley, J. A.; McNeil, B. J. The meaning and use of the area under a receiver operating characteristic (ROC) curve. Radiology 1982, 143 (1), 29– 36, DOI: 10.1148/radiology.143.1.7063747There is no corresponding record for this reference.
- 57Massad-Ivanir, N.; Mirsky, Y.; Nahor, A.; Edrei, E.; Bonanno-Young, L. M.; Ben Dov, N. Trap and track: designing self-reporting porous Si photonic crystals for rapid bacteria detection. Analyst 2014, 139 (16), 3885– 3894, DOI: 10.1039/C4AN00364KThere is no corresponding record for this reference.
- 58Ito, W.; Choi, N.; Letner, G.; Genz, N.; Prokop, D.; Valadon, C.; Sardiu, M. E.; Smith, H.; Whiles, B. B.; Molina, W. R. Preoperative urine culture with contaminants is not associated with increased risk for urinary tract infection after ureteroscopic stone treatment. World J. Urol. 2024, 42 (1), 159, DOI: 10.1007/s00345-024-04793-wThere is no corresponding record for this reference.
- 59Nye, T. M.; Zou, Z.; Obernuefemann, C. L. P.; Pinkner, J. S.; Lowry, E.; Kleinschmidt, K.; Bergeron, K.; Klim, A.; Dodson, K. W.; Flores-Mireles, A. L. Microbial co-occurrences on catheters from long-term catheterized patients. Nat. Commun. 2024, 15 (1), 61, DOI: 10.1038/s41467-023-44095-0There is no corresponding record for this reference.
- 60Muller, A.; Gbaguidi-Haore, H.; Cholley, P.; Hocquet, D.; Sauget, M.; Bertrand, X. Hospital-diagnosed infections with Escherichia coli clonal group ST131 are mostly acquired in the community. Sci. Rep. 2021, 11 (1), 5702, DOI: 10.1038/s41598-021-85116-6There is no corresponding record for this reference.
- 61Toval, F.; Köhler, C.-D.; Vogel, U.; Wagenlehner, F.; Mellmann, A.; Fruth, A. Characterization of Escherichia coli Isolates from Hospital Inpatients or Outpatients with Urinary Tract Infection. J. Clin. Microbiol. 2014, 52 (2), 407– 418, DOI: 10.1128/JCM.02069-13There is no corresponding record for this reference.
- 62Gu, W.; Huang, W.; Zhang, J.; Qian, S.; Cao, H.; Ge, L. Evaluation of urinary inflammatory index in rapid screening of urinary tract infection. Sci. Rep. 2020, 10 (1), 19306, DOI: 10.1038/s41598-020-76352-3There is no corresponding record for this reference.
- 63Gajdacs, M.; Doczi, I.; Abrok, M.; Lazar, A.; Burian, K. Epidemiology of candiduria and Candida urinary tract infections in inpatients and outpatients: results from a 10-year retrospective survey. Cent. Eur. J. Urol. 2019, 72 (2), 209– 214, DOI: 10.5173/ceju.2019.1909There is no corresponding record for this reference.
- 64Dias, V. Candida Species in the Urinary Tract: Is it a Fungal Infection or Not?. Future Microbiol. 2020, 15 (2), 81– 83, DOI: 10.2217/fmb-2019-0262There is no corresponding record for this reference.
- 65Behzadi, P.; Behzadi, E.; Ranjbar, R. Urinary tract infections and Candida albicans. Cent. Eur. J. Urol. 2015, 68 (1), 96– 101, DOI: 10.5173/ceju.2015.01.474There is no corresponding record for this reference.
- 66La Bella, A. A.; Andersen, M. J.; Gervais, N. C.; Molina, J. J.; Molesan, A.; Stuckey, P. V.; Wensing, L.; Nobile, C. J.; Shapiro, R. S.; Santiago-Tirado, F. H. The catheterized bladder environment promotes Efg1- and Als1-dependent Candida albicans infection. Sci. Adv. 2023, 9 (9), eade7689 DOI: 10.1126/sciadv.ade7689There is no corresponding record for this reference.
- 67Sher, E. K.; Džidić-Krivić, A.; Sesar, A.; Farhat, E. K.; Čeliković, A.; Beća-Zećo, M. Current state and novel outlook on prevention and treatment of rising antibiotic resistance in urinary tract infections. Pharmacol. Ther. 2024, 261, 108688, DOI: 10.1016/j.pharmthera.2024.108688There is no corresponding record for this reference.
- 68Polo, T. C. F.; Miot, H. A. Use of ROC curves in clinical and experimental studies. J. Vasc. Bras. 2020, 19, e20200186 DOI: 10.1590/1677-5449.200186There is no corresponding record for this reference.
- 69Silva, S.; Henriques, M.; Oliveira, R.; Williams, D.; Azeredo, J. In Vitro Biofilm Activity of Non-Candida albicansCandida Species. Curr. Microbiol. 2010, 61 (6), 534– 540, DOI: 10.1007/s00284-010-9649-7There is no corresponding record for this reference.
- 70Lu, X.; Chow, J. J. M.; Koo, S. H.; Tan, T. Y.; Jiang, B.; Ai, Y. Enhanced Molecular Diagnosis of Bloodstream Candida Infection with Size-Based Inertial Sorting at Submicron Resolution. Anal. Chem. 2020, 92 (23), 15579– 15586, DOI: 10.1021/acs.analchem.0c03718There is no corresponding record for this reference.
- 71Noble, S. M.; Gianetti, B. A.; Witchley, J. N. Candida albicans cell-type switching and functional plasticity in the mammalian host. Nat. Rev. Microbiol. 2017, 15 (2), 96– 108, DOI: 10.1038/nrmicro.2016.157There is no corresponding record for this reference.
- 72Wijesinghe, G.; Dilhari, A.; Gayani, B.; Kottegoda, N.; Samaranayake, L.; Weerasekera, M. Influence of Laboratory Culture Media on in vitro Growth, Adhesion, and Biofilm Formation of Pseudomonas aeruginosa and Staphylococcus aureus. Med. Princ. Pract. 2019, 28 (1), 28– 35, DOI: 10.1159/000494757There is no corresponding record for this reference.
- 73Jones-Freeman, B.; Chonwerawong, M.; Marcelino, V. R.; Deshpande, A. V.; Forster, S. C.; Starkey, M. R. The microbiome and host mucosal interactions in urinary tract diseases. Mucosal Immunol. 2021, 14 (4), 779– 792, DOI: 10.1038/s41385-020-00372-5There is no corresponding record for this reference.
- 74Sujith, S.; Solomon, A. P.; Rayappan, J. B. B. Comprehensive insights into UTIs: from pathophysiology to precision diagnosis and management. Front. Cell. Infect. Microbiol. 2024, 14, 1402941, DOI: 10.3389/fcimb.2024.1402941There is no corresponding record for this reference.
- 75Brosel-Oliu, S.; Mergel, O.; Uria, N.; Abramova, N.; Van Rijn, P.; Bratov, A. 3D impedimetric sensors as a tool for monitoring bacterial response to antibiotics. Lab Chip 2019, 19 (8), 1436– 1447, DOI: 10.1039/C8LC01220BThere is no corresponding record for this reference.
- 76Prinz Setter, O.; Jiang, X.; Segal, E. Rising to the surface: capturing and detecting bacteria by rationally-designed surfaces. Curr. Opin. Biotechnol. 2023, 83, 102969, DOI: 10.1016/j.copbio.2023.102969There is no corresponding record for this reference.
- 77Soares, A.; Roussel, V.; Pestel-Caron, M.; Barreau, M.; Caron, F.; Bouffartigues, E.; Chevalier, S.; Etienne, M. Understanding Ciprofloxacin Failure in Pseudomonas aeruginosa Biofilm: Persister Cells Survive Matrix Disruption. Front. Microbiol. 2019, 10, 2603, DOI: 10.3389/fmicb.2019.02603There is no corresponding record for this reference.
- 78Ojkic, N.; Lilja, E.; Direito, S.; Dawson, A.; Allen, R. J.; Waclaw, B. A Roadblock-and-Kill Mechanism of Action Model for the DNA-Targeting Antibiotic Ciprofloxacin. Antimicrob. Agents Chemother. 2020, 64 (9), 10– 1128, DOI: 10.1128/AAC.02487-19There is no corresponding record for this reference.
- 79CLSI. Performance Standards for Antimicrobial Susceptibility Testing, 35th edition. CLSI supplement M100; Clinical and Laboratory Standards Institute; 2025. http://em100.edaptivedocs.net/dashboard.aspx. Accessed 25.April.03.There is no corresponding record for this reference.
- 80Needs, S. H.; Donmez, S. I.; Bull, S. P.; McQuaid, C.; Osborn, H. M. I.; Edwards, A. D. Challenges in Microfluidic and Point-of-Care Phenotypic Antimicrobial Resistance Tests. Front. Mech. Eng. 2020, 6, 73, DOI: 10.3389/fmech.2020.00073There is no corresponding record for this reference.
- 81Wu, W.; Suo, Y.; Zhao, Q.; Cai, G.; Liu, Y.; Jin, W. Inoculum size-insensitive susceptibility determination of urine sample based on in-situ measurement of inducible enzyme activity after 20 min of antibiotic exposure. Anal. Chim. Acta 2023, 1282, 341858, DOI: 10.1016/j.aca.2023.341858There is no corresponding record for this reference.
- 82Smith, K. P.; Kirby, J. E. The Inoculum Effect in the Era of Multidrug Resistance: Minor Differences in Inoculum Have Dramatic Effect on MIC Determination. Antimicrob. Agents Chemother. 2018, 62 (8), e00433–18 DOI: 10.1128/AAC.00433-18There is no corresponding record for this reference.
- 83Yi, Q.; Cui, J.; Xiao, M.; Tang, M.-Z.; Zhang, H.-C.; Zhang, G. Rapid Phenotypic Antimicrobial Susceptibility Testing Using a Coulter Counter and Proliferation Rate Discrepancy. ACS Omega 2023, 8 (18), 16298– 16305, DOI: 10.1021/acsomega.3c00947There is no corresponding record for this reference.
- 84Vasala, A.; Hytönen, V. P.; Laitinen, O. H. Modern Tools for Rapid Diagnostics of Antimicrobial Resistance. Front. Cell. Infect. Microbiol. 2020, 10, 308, DOI: 10.3389/fcimb.2020.00308There is no corresponding record for this reference.
- 85Alfouzan, W. A.; Dhar, R. Candiduria: Evidence-based approach to management, are we there yet?. J. Mycol. Médicale 2017, 27 (3), 293– 302, DOI: 10.1016/j.mycmed.2017.04.005There is no corresponding record for this reference.
- 86World Health Organization. Landscape analysis of commercially available and pipeline in vitro diagnostics for fungal priority pathogens; World Health Organization, 2025. https://www.who.int/publications/i/item/9789240105539. Accessed 25.April.12.There is no corresponding record for this reference.
- 87World Health Organization. WHO issues its first-ever reports on tests and treatments for fungal infections; World Health Organization, 2025. https://www.who.int/news/item/01-04-2025-who-issues-its-first-ever-reports-on-tests-and-treatments-for-fungal-infections. Accessed 25.April.12.There is no corresponding record for this reference.
Supporting Information
Supporting Information
The Supporting Information is available free of charge at https://pubs.acs.org/doi/10.1021/acsmeasuresciau.5c00187.
Study participant demographics; iPRISM principles; double-blinded urine test results for microbial infections; differentiation between bacterial and fungal infections using iPRISM; iPRISM-based AST for ciprofloxacin in bacterially infected urine samples (PDF)
Terms & Conditions
Most electronic Supporting Information files are available without a subscription to ACS Web Editions. Such files may be downloaded by article for research use (if there is a public use license linked to the relevant article, that license may permit other uses). Permission may be obtained from ACS for other uses through requests via the RightsLink permission system: http://pubs.acs.org/page/copyright/permissions.html.



