Atomic Precision in Personalized Oncology: AI-Designed Nanomedicines Enabling N-of-1 Cancer TherapyClick to copy article linkArticle link copied!
- Miles PourbaghiMiles PourbaghiDepartment of Radiology, Weill Cornell Medicine, New York, New York 10021, United StatesMore by Miles Pourbaghi
- Michelle S. Bradbury*Michelle S. Bradbury*Email: [email protected]Department of Radiology, Weill Cornell Medicine, New York, New York 10021, United StatesSandra and Edward Meyer Cancer Center, Weill Cornell Medicine, New York, New York 10021, United StatesDepartment of Radiation Oncology, Weill Cornell Medicine, New York, New York 10065, United StatesMore by Michelle S. Bradbury
Abstract
Rapid advances in AI-driven molecular design, intelligent nanomaterial engineering, and clinical AI systems are reshaping personalized cancer therapy by enabling coordinated innovation across biological scales, from genome regulation to functional protein interactions. At the molecular level, generative platforms such as AlphaProteo and RFdiffusion now support the rapid design of de novo protein binders with high predicted structural accuracy and low-nanomolar affinities. Acting upstream, AlphaGenome interprets genome-wide regulatory variation to predict how single-base changes in noncoding DNA alter gene control mechanisms, facilitating the prioritization of patient-specific pathways and therapeutic targets. Together, these capabilities are transforming computational design from a speculative approach to a reliable molecular engineering workflow. In parallel, clinically validated nanomedicine platforms provide modular architectures that enhance pharmacokinetics, improve therapeutic indices, and modulate tumor–immune interactions, addressing persistent barriers in solid tumor treatment. Integration with AI-enabled experimental platforms, patient-derived organoid systems, spatial and single-cell profiling, quantitative systems pharmacology models, and microfluidic GMP-compatible manufacturing suggests a feasible path toward accelerated, patient-matched nanomedicine development. While challenges remain─including immunogenicity and manufacturing constraints─the convergence of computation, materials engineering, and regulatory science supports a realistic roadmap toward N-of-1 oncology.
This publication is licensed under
License Summary*
You are free to share(copy and redistribute) this article in any medium or format within the parameters below:
Creative Commons (CC): This is a Creative Commons license.
Attribution (BY): Credit must be given to the creator.
Non-Commercial (NC): Only non-commercial uses of the work are permitted.
No Derivatives (ND): Derivative works may be created for non-commercial purposes, but sharing is prohibited.
*Disclaimer
This summary highlights only some of the key features and terms of the actual license. It is not a license and has no legal value. Carefully review the actual license before using these materials.
License Summary*
You are free to share(copy and redistribute) this article in any medium or format within the parameters below:
Creative Commons (CC): This is a Creative Commons license.
Attribution (BY): Credit must be given to the creator.
Non-Commercial (NC): Only non-commercial uses of the work are permitted.
No Derivatives (ND): Derivative works may be created for non-commercial purposes, but sharing is prohibited.
*Disclaimer
This summary highlights only some of the key features and terms of the actual license. It is not a license and has no legal value. Carefully review the actual license before using these materials.
License Summary*
You are free to share(copy and redistribute) this article in any medium or format within the parameters below:
Creative Commons (CC): This is a Creative Commons license.
Attribution (BY): Credit must be given to the creator.
Non-Commercial (NC): Only non-commercial uses of the work are permitted.
No Derivatives (ND): Derivative works may be created for non-commercial purposes, but sharing is prohibited.
*Disclaimer
This summary highlights only some of the key features and terms of the actual license. It is not a license and has no legal value. Carefully review the actual license before using these materials.
License Summary*
You are free to share(copy and redistribute) this article in any medium or format within the parameters below:
Creative Commons (CC): This is a Creative Commons license.
Attribution (BY): Credit must be given to the creator.
Non-Commercial (NC): Only non-commercial uses of the work are permitted.
No Derivatives (ND): Derivative works may be created for non-commercial purposes, but sharing is prohibited.
*Disclaimer
This summary highlights only some of the key features and terms of the actual license. It is not a license and has no legal value. Carefully review the actual license before using these materials.
License Summary*
You are free to share(copy and redistribute) this article in any medium or format within the parameters below:
Creative Commons (CC): This is a Creative Commons license.
Attribution (BY): Credit must be given to the creator.
Non-Commercial (NC): Only non-commercial uses of the work are permitted.
No Derivatives (ND): Derivative works may be created for non-commercial purposes, but sharing is prohibited.
*Disclaimer
This summary highlights only some of the key features and terms of the actual license. It is not a license and has no legal value. Carefully review the actual license before using these materials.
Figure 1
Figure 1. N-of-1 convergence workflow: From biopsy to bedside. Schematic overview of a rapid-response-engineered therapeutic pipeline enabled by the convergence of AI-driven genomic interpretation and protein design. (A) Patient profiling: A tumor biopsy undergoes whole-genome sequencing and spatial transcriptomic analysis, with AlphaGenome applied to identify patient-specific regulatory variants and splice-derived antigenic peptides through systematic computational variant scanning. (B) AI-driven design: Generative platforms, including AlphaProteo and RFdiffusion, design de novo protein binders targeting the prioritized antigens, while AlphaFold 3 is used to assess predicted structural integrity and surface compatibility. (C) Rapid manufacturing: Automated microfluidic systems conjugate designed binders to an established, clinical-grade nanoparticle backbone (e.g., ultrasmall silica particles or lipid nanoparticles). (D) Ex vivo validation: The resulting patient-matched nanomedicine is evaluated in autologous, immune-competent tumor organoid systems to assess cellular uptake and potential immunogenicity. (E) Clinical administration: The validated N-of-1 therapeutic is administered, with quantitative systems pharmacology (QSP)-based digital twin models guiding dose selection.
The Inflection Point: From Anecdote to Systematic Engineering
Why Nanoscale Architectures Are Uniquely Suited for N-of-1 Oncology
Reimagining the Possible: Four Concepts across the Readiness Spectrum
1. Digital Twin Discovery: From Patient Biopsy to Nanoparticle (Status: Operational in Early Translation)
2. Chameleon Coronas: Programming the “Bug” into a Feature (Status: Emerging Experimental Concept)
3. Multispecific Protein Logic Gates: Boolean Therapeutics (Status: Emerging Experimental Concept)
Figure 2
Figure 2. Boolean therapeutics: Logic-gated nanomedicine design. Conceptual illustration of a multispecific “AND”-gate nanomedicine designed to address tumor heterogeneity while minimizing off-target toxicity. The nanoparticle is functionalized with two distinct AI-designed sensor proteins (Binder A and Binder B) and incorporates a pH-sensitive release mechanism. (Left) Healthy tissue: In the presence of only Target A (e.g., EGFR) and physiological pH, the therapeutic payload remains sequestered, preventing nonspecific release. (Right) Tumor microenvironment: Drug release is triggered only when the Boolean condition (Target A AND Target B) is satisfied in conjunction with low pH. This stringent logic requirement restricts payload delivery to tumor cells coexpressing defined antigens within the acidic tumor niche, thereby increasing the therapeutic index.
4. Self-Assembling Fractal Therapeutics: Maximizing Tumor Penetration (Status: Emerging Experimental Concept)
The Path Forward: Timelines and Implementation Realities
Figure 3
Figure 3. Roadmap to N-of-1 oncology. Projected timeline distinguishing currently operational technologies from emerging experimental concepts. The roadmap illustrates the evolution of AI-designed nanomedicines across three translational phases: (1) Validation and safety (approximately 1–3 Years): Phase I evaluation of single-target nanoparticle conjugates and validation of immunogenicity prediction tools, with AlphaGenome operational for patient-specific target discovery and AlphaProteo supporting rapid binder design. (2) Integration and efficacy (approximately 3–5 Years): Deployment of “Digital Twin” QSP models for dose optimization, initial regulatory approvals of first-in-class AI-designed agents, and adoption of modular, microfluidic manufacturing platforms. (3) Complexity and precision (approximately 5–10 years): Routine clinical use of logic-gated nanotherapeutics and real-time, adaptive formulation cycles. Dashed lines denote technologies that currently require additional in vivo validation (e.g., fractal assemblies and higher-order logic-gated systems).
Acknowledging the Obstacles
Conclusion: A Call to Convergence
References
This article references 34 other publications.
- 1Watson, J. L.; Juergens, D.; Bennett, N. R.; Trippe, B. L.; Yim, J.; Eisenach, H. E.; Ahern, W.; Borst, A. J.; Ragotte, R. J.; Milles, L. F. De novo design of protein structure and function with RFdiffusion. Nature 2023, 620 (7976), 1089– 1100, DOI: 10.1038/s41586-023-06415-8Google ScholarThere is no corresponding record for this reference.
- 2Fox, D. R.; Taveneau, C.; Clement, J.; Grinter, R.; Knott, G. J. Code to complex: AI-driven de novo binder design. Structure 2025, 33 (10), 1631– 1642, DOI: 10.1016/j.str.2025.08.007Google ScholarThere is no corresponding record for this reference.
- 3Dauparas, J.; Anishchenko, I.; Bennett, N.; Bai, H.; Ragotte, R. J.; Milles, L. F.; Wicky, B. I. M.; Courbet, A.; de Haas, R. J.; Bethel, N. Robust deep learning-based protein sequence design using ProteinMPNN. Science 2022, 378 (6615), 49– 56, DOI: 10.1126/science.add2187Google ScholarThere is no corresponding record for this reference.
- 4Abramson, J.; Adler, J.; Dunger, J.; Evans, R.; Green, T.; Pritzel, A.; Ronneberger, O.; Willmore, L.; Ballard, A. J.; Bambrick, J. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature 2024, 630 (8016), 493– 500, DOI: 10.1038/s41586-024-07487-wGoogle ScholarThere is no corresponding record for this reference.
- 5Ma, K.; Mendoza, C.; Hanson, M.; Werner-Zwanziger, U.; Zwanziger, J.; Wiesner, U. Control of Ultrasmall Sub-10 nm Ligand-Functionalized Fluorescent Core-Shell Silica Nanoparticle Growth in Water. Chem. Mater. 2015, 27 (11), 4119– 4133, DOI: 10.1021/acs.chemmater.5b01222Google ScholarThere is no corresponding record for this reference.
- 6Phillips, E.; Penate-Medina, O.; Zanzonico, P. B.; Carvajal, R. D.; Mohan, P.; Ye, Y.; Humm, J.; Gönen, M.; Kalaigian, H.; Schöder, H. Clinical translation of an ultrasmall inorganic optical-PET imaging nanoparticle probe. Sci. Transl Med. 2014, 6 (260), 260ra149, DOI: 10.1126/scitranslmed.3009524Google ScholarThere is no corresponding record for this reference.
- 7De Leon, G.; Zhang, L.; Siddiqui, N. A.; Naguib, N.; Chen, F.; Padmanabhan, R.; Zhang, T.; Monette, S.; Socciarelli, F.; Lee, R. An ultrasmall core-shell silica nanoparticle improves antitumour immunity and survival by remodelling suppressive melanoma microenvironments. Nat. Nanotechnol. 2026, 21 (2), 311– 322, DOI: 10.1038/s41565-025-02083-zGoogle ScholarThere is no corresponding record for this reference.
- 8Chen, F.; Ma, K.; Madajewski, B.; Zhuang, L.; Zhang, L.; Rickert, K.; Marelli, M.; Yoo, B.; Turker, M. Z.; Overholtzer, M. Ultrasmall targeted nanoparticles with engineered antibody fragments for imaging detection of HER2-overexpressing breast cancer. Nat. Commun. 2018, 9 (1), 4141, DOI: 10.1038/s41467-018-06271-5Google ScholarThere is no corresponding record for this reference.
- 9Szymanski, N. J.; Rendy, B.; Fei, Y.; Kumar, R. E.; He, T.; Milsted, D.; McDermott, M. J.; Gallant, M.; Cubuk, E. D.; Merchant, A. An autonomous laboratory for the accelerated synthesis of novel materials. Nature 2023, 624 (7990), 86– 91, DOI: 10.1038/s41586-023-06734-wGoogle ScholarThere is no corresponding record for this reference.
- 10Velapure, P.; Kansal, D.; Bobade, C. Tumor microenvironment-responsive nanoformulations for breast cancer. Discov Nano 2024, 19 (1), 212, DOI: 10.1186/s11671-024-04122-5Google ScholarThere is no corresponding record for this reference.
- 11Wang, J.; Tao, X.; Zhu, J.; Dai, Z.; Du, Y.; Xie, Y.; Chu, X.; Fu, G.; Lei, Z. Tumor organoid-immune co-culture models: exploring a new perspective of tumor immunity. Cell Death Discov 2025, 11 (1), 195, DOI: 10.1038/s41420-025-02407-xGoogle ScholarThere is no corresponding record for this reference.
- 12Helmlinger, G.; Sokolov, V.; Peskov, K.; Hallow, K. M.; Kosinsky, Y.; Voronova, V.; Chu, L.; Yakovleva, T.; Azarov, I.; Kaschek, D. Quantitative Systems Pharmacology: An Exemplar Model-Building Workflow With Applications in Cardiovascular, Metabolic, and Oncology Drug Development. CPT Pharmacometrics Syst. Pharmacol 2019, 8 (6), 380– 395, DOI: 10.1002/psp4.12426Google ScholarThere is no corresponding record for this reference.
- 13Canchola, A.; Li, K.; Chen, K.; Borboa-Pimentel, A.; Chou, C.; Dela Rama, R.; Chen, C. Y.; Chen, X.; Strobel, M.; Riviere, J. E. Meta-Analysis and Machine Learning Prediction of Protein Corona Composition across Nanoparticle Systems in Biological Media. ACS Nano 2025, 19, 37633, DOI: 10.1021/acsnano.5c08608Google ScholarThere is no corresponding record for this reference.
- 14Wang, H.; Arulraj, T.; Ippolito, A.; Popel, A. S. From virtual patients to digital twins in immuno-oncology: lessons learned from mechanistic quantitative systems pharmacology modeling. NPJ. Digit Med. 2024, 7 (1), 189, DOI: 10.1038/s41746-024-01188-4Google ScholarThere is no corresponding record for this reference.
- 15Nobel Prize in Chemistry 2024 , 2024. https://www.nobelprize.org/uploads/2024/10/press-chemistryprize2024-3.pdf.Google ScholarThere is no corresponding record for this reference.
- 16Avsec, Ž.; Latysheva, N.; Cheng, J.; Novati, G.; Taylor, K. R.; Ward, T.; Bycroft, C.; Nicolaisen, L.; Arvaniti, E.; Pan, J. Advancing regulatory variant effect prediction with AlphaGenome. Nature 2026, 649 (8099), 1206– 1218, DOI: 10.1038/s41586-025-10014-0Google ScholarThere is no corresponding record for this reference.
- 17Zambaldi, V.; La, D.; Chu, A. E.; Patani, H.; Danson, A. E.; Kwan, T. O.; Frerix, T.; Schneider, R. G.; Saxton, D.; Thillaisundaram, A. De novo design of high-affinity protein binders with AlphaProteo. arXiv Preprint , arXiv:2409.08022, 2024.Google ScholarThere is no corresponding record for this reference.
- 18Zeni, C.; Pinsler, R.; Zügner, D.; Fowler, A.; Horton, M.; Fu, X.; Wang, Z.; Shysheya, A.; Crabbé, J.; Ueda, S. A generative model for inorganic materials design. Nature 2025, 639 (8055), 624– 632, DOI: 10.1038/s41586-025-08628-5Google ScholarThere is no corresponding record for this reference.
- 19Kp Jayatunga, M.; Ayers, M.; Bruens, L.; Jayanth, D.; Meier, C. How successful are AI-discovered drugs in clinical trials? A first analysis and emerging lessons. Drug Discov Today 2024, 29 (6), 104009, DOI: 10.1016/j.drudis.2024.104009Google ScholarThere is no corresponding record for this reference.
- 20Lee, K. Y.; Lee, G. Y.; Lane, L. A.; Li, B.; Wang, J.; Lu, Q.; Wang, Y.; Nie, S. Functionalized, Long-Circulating, and Ultrasmall Gold Nanocarriers for Overcoming the Barriers of Low Nanoparticle Delivery Efficiency and Poor Tumor Penetration. Bioconjug Chem. 2017, 28 (1), 244– 252, DOI: 10.1021/acs.bioconjchem.6b00224Google ScholarThere is no corresponding record for this reference.
- 21Broske, B.; Binder, S. C.; McEnroe, B. A.; Kempchen, T. N.; Fandrey, C. I.; Messmer, J. M.; Tan, E.; Konopka, P.; Ferber, D.; Yong, M. C. R. Development of AI-designed protein binders for detection and targeting of cancer cell surface proteins. bioRxiv Preprint , 2025. DOI: 10.1101/2025.05.11.652819 .Google ScholarThere is no corresponding record for this reference.
- 22Chen, Z.; Kibler, R. D.; Hunt, A.; Busch, F.; Pearl, J.; Jia, M.; VanAernum, Z. L.; Wicky, B. I. M.; Dods, G.; Liao, H. De novo design of protein logic gates. Science 2020, 368 (6486), 78– 84, DOI: 10.1126/science.aay2790Google ScholarThere is no corresponding record for this reference.
- 23Johnson, A.; Shen, Y.; Zheng, X.; Su, F.; Zeng, J.; Uzunparmak, B.; Breuer, E.-K.; Holla, V.; Kahle, M.; Kim, S.-H. The actionable transcriptome: a framework for incorporating RNA sequencing into precision oncology. Nature Reviews Clinical Oncology 2026, 23, 213, DOI: 10.1038/s41571-025-01110-1Google ScholarThere is no corresponding record for this reference.
- 24Bennett, N. R.; Coventry, B.; Goreshnik, I.; Huang, B.; Allen, A.; Vafeados, D.; Peng, Y. P.; Dauparas, J.; Baek, M.; Stewart, L. Improving de novo protein binder design with deep learning. Nat. Commun. 2023, 14 (1), 2625, DOI: 10.1038/s41467-023-38328-5Google ScholarThere is no corresponding record for this reference.
- 25Valencia, P. M.; Pridgen, E. M.; Rhee, M.; Langer, R.; Farokhzad, O. C.; Karnik, R. Microfluidic platform for combinatorial synthesis and optimization of targeted nanoparticles for cancer therapy. ACS Nano 2013, 7 (12), 10671– 10680, DOI: 10.1021/nn403370eGoogle ScholarThere is no corresponding record for this reference.
- 26Corbo, C.; Molinaro, R.; Tabatabaei, M.; Farokhzad, O. C.; Mahmoudi, M. Personalized protein corona on nanoparticles and its clinical implications. Biomater Sci. 2017, 5 (3), 378– 387, DOI: 10.1039/C6BM00921BGoogle ScholarThere is no corresponding record for this reference.
- 27Praetorius, F.; Leung, P. J. Y.; Tessmer, M. H.; Broerman, A.; Demakis, C.; Dishman, A. F.; Pillai, A.; Idris, A.; Juergens, D.; Dauparas, J. Design of stimulus-responsive two-state hinge proteins. Science 2023, 381 (6659), 754– 760, DOI: 10.1126/science.adg7731Google ScholarThere is no corresponding record for this reference.
- 28Hsia, Y.; Bale, J. B.; Gonen, S.; Shi, D.; Sheffler, W.; Fong, K. K.; Nattermann, U.; Xu, C.; Huang, P. S.; Ravichandran, R. Design of a hyperstable 60-subunit protein dodecahedron. [corrected]. Nature 2016, 535 (7610), 136– 139, DOI: 10.1038/nature18010Google ScholarThere is no corresponding record for this reference.
- 29Ma, T.; Tran, T. B.; Lin, E.; Hunt, S.; Haveman, R.; Castro, K.; Lu, J. Size-transformable nanotherapeutics for cancer therapy. Acta Pharm. Sin B 2025, 15 (2), 834– 851, DOI: 10.1016/j.apsb.2024.11.012Google ScholarThere is no corresponding record for this reference.
- 30Joyce, P.; Allen, C. J.; Alonso, M. J.; Ashford, M.; Bradbury, M. S.; Germain, M.; Kavallaris, M.; Langer, R.; Lammers, T.; Peracchia, M. T. A translational framework to DELIVER nanomedicines to the clinic. Nat. Nanotechnol 2024, 19 (11), 1597– 1611, DOI: 10.1038/s41565-024-01754-7Google ScholarThere is no corresponding record for this reference.
- 31Considerations for the Use of Artificial Intelligence To Support Regulatory Decision-Making for Drug and Biological Products , 2025. https://www.fda.gov/regulatory-information/search-fda-guidance-documents/considerations-use-artificial-intelligence-support-regulatory-decision-making-drug-and-biological.Google ScholarThere is no corresponding record for this reference.
- 32Reflection paper on the use of Artificial Intelligence (AI) in the medicinal product lifecycle , 2024. https://www.ema.europa.eu/en/documents/scientific-guideline/reflection-paper-use-artificial-intelligence-ai-medicinal-product-lifecycle_en.pdf.Google ScholarThere is no corresponding record for this reference.
- 33Sharma, V.; Mottafegh, A.; Joo, J. U.; Kang, J. H.; Wang, L.; Kim, D. P. Toward microfluidic continuous-flow and intelligent downstream processing of biopharmaceuticals. Lab Chip 2024, 24 (11), 2861– 2882, DOI: 10.1039/D3LC01097JGoogle ScholarThere is no corresponding record for this reference.
- 34Carter, P. J.; Quarmby, V. Immunogenicity risk assessment and mitigation for engineered antibody and protein therapeutics. Nat. Rev. Drug Discov 2024, 23 (12), 898– 913, DOI: 10.1038/s41573-024-01051-xGoogle ScholarThere is no corresponding record for this reference.
Cited By
This article has not yet been cited by other publications.
Article Views
Altmetric
Citations
Article Views are the COUNTER-compliant sum of full text article downloads since November 2008 (both PDF and HTML) across all institutions and individuals. These metrics are regularly updated to reflect usage leading up to the last few days.
Citations are the number of other articles citing this article, calculated by Crossref and updated daily. Find more information about Crossref citation counts.
The Altmetric Attention Score is a quantitative measure of the attention that a research article has received online. Clicking on the donut icon will load a page at altmetric.com with additional details about the score and the social media presence for the given article. Find more information on the Altmetric Attention Score and how the score is calculated.
Recommended Articles
Abstract

Figure 1
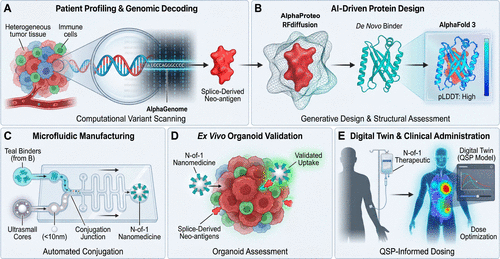
Figure 1. N-of-1 convergence workflow: From biopsy to bedside. Schematic overview of a rapid-response-engineered therapeutic pipeline enabled by the convergence of AI-driven genomic interpretation and protein design. (A) Patient profiling: A tumor biopsy undergoes whole-genome sequencing and spatial transcriptomic analysis, with AlphaGenome applied to identify patient-specific regulatory variants and splice-derived antigenic peptides through systematic computational variant scanning. (B) AI-driven design: Generative platforms, including AlphaProteo and RFdiffusion, design de novo protein binders targeting the prioritized antigens, while AlphaFold 3 is used to assess predicted structural integrity and surface compatibility. (C) Rapid manufacturing: Automated microfluidic systems conjugate designed binders to an established, clinical-grade nanoparticle backbone (e.g., ultrasmall silica particles or lipid nanoparticles). (D) Ex vivo validation: The resulting patient-matched nanomedicine is evaluated in autologous, immune-competent tumor organoid systems to assess cellular uptake and potential immunogenicity. (E) Clinical administration: The validated N-of-1 therapeutic is administered, with quantitative systems pharmacology (QSP)-based digital twin models guiding dose selection.
Figure 2
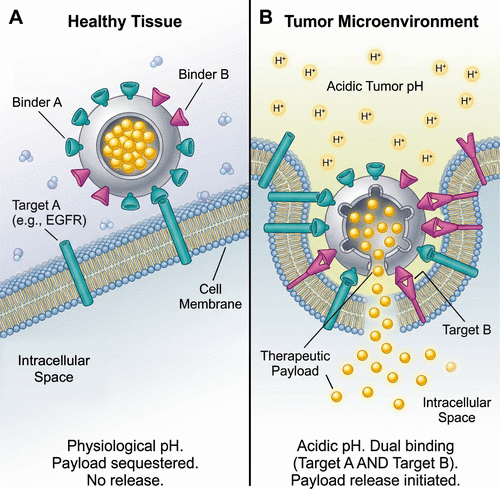
Figure 2. Boolean therapeutics: Logic-gated nanomedicine design. Conceptual illustration of a multispecific “AND”-gate nanomedicine designed to address tumor heterogeneity while minimizing off-target toxicity. The nanoparticle is functionalized with two distinct AI-designed sensor proteins (Binder A and Binder B) and incorporates a pH-sensitive release mechanism. (Left) Healthy tissue: In the presence of only Target A (e.g., EGFR) and physiological pH, the therapeutic payload remains sequestered, preventing nonspecific release. (Right) Tumor microenvironment: Drug release is triggered only when the Boolean condition (Target A AND Target B) is satisfied in conjunction with low pH. This stringent logic requirement restricts payload delivery to tumor cells coexpressing defined antigens within the acidic tumor niche, thereby increasing the therapeutic index.
Figure 3

Figure 3. Roadmap to N-of-1 oncology. Projected timeline distinguishing currently operational technologies from emerging experimental concepts. The roadmap illustrates the evolution of AI-designed nanomedicines across three translational phases: (1) Validation and safety (approximately 1–3 Years): Phase I evaluation of single-target nanoparticle conjugates and validation of immunogenicity prediction tools, with AlphaGenome operational for patient-specific target discovery and AlphaProteo supporting rapid binder design. (2) Integration and efficacy (approximately 3–5 Years): Deployment of “Digital Twin” QSP models for dose optimization, initial regulatory approvals of first-in-class AI-designed agents, and adoption of modular, microfluidic manufacturing platforms. (3) Complexity and precision (approximately 5–10 years): Routine clinical use of logic-gated nanotherapeutics and real-time, adaptive formulation cycles. Dashed lines denote technologies that currently require additional in vivo validation (e.g., fractal assemblies and higher-order logic-gated systems).
References
This article references 34 other publications.
- 1Watson, J. L.; Juergens, D.; Bennett, N. R.; Trippe, B. L.; Yim, J.; Eisenach, H. E.; Ahern, W.; Borst, A. J.; Ragotte, R. J.; Milles, L. F. De novo design of protein structure and function with RFdiffusion. Nature 2023, 620 (7976), 1089– 1100, DOI: 10.1038/s41586-023-06415-8There is no corresponding record for this reference.
- 2Fox, D. R.; Taveneau, C.; Clement, J.; Grinter, R.; Knott, G. J. Code to complex: AI-driven de novo binder design. Structure 2025, 33 (10), 1631– 1642, DOI: 10.1016/j.str.2025.08.007There is no corresponding record for this reference.
- 3Dauparas, J.; Anishchenko, I.; Bennett, N.; Bai, H.; Ragotte, R. J.; Milles, L. F.; Wicky, B. I. M.; Courbet, A.; de Haas, R. J.; Bethel, N. Robust deep learning-based protein sequence design using ProteinMPNN. Science 2022, 378 (6615), 49– 56, DOI: 10.1126/science.add2187There is no corresponding record for this reference.
- 4Abramson, J.; Adler, J.; Dunger, J.; Evans, R.; Green, T.; Pritzel, A.; Ronneberger, O.; Willmore, L.; Ballard, A. J.; Bambrick, J. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature 2024, 630 (8016), 493– 500, DOI: 10.1038/s41586-024-07487-wThere is no corresponding record for this reference.
- 5Ma, K.; Mendoza, C.; Hanson, M.; Werner-Zwanziger, U.; Zwanziger, J.; Wiesner, U. Control of Ultrasmall Sub-10 nm Ligand-Functionalized Fluorescent Core-Shell Silica Nanoparticle Growth in Water. Chem. Mater. 2015, 27 (11), 4119– 4133, DOI: 10.1021/acs.chemmater.5b01222There is no corresponding record for this reference.
- 6Phillips, E.; Penate-Medina, O.; Zanzonico, P. B.; Carvajal, R. D.; Mohan, P.; Ye, Y.; Humm, J.; Gönen, M.; Kalaigian, H.; Schöder, H. Clinical translation of an ultrasmall inorganic optical-PET imaging nanoparticle probe. Sci. Transl Med. 2014, 6 (260), 260ra149, DOI: 10.1126/scitranslmed.3009524There is no corresponding record for this reference.
- 7De Leon, G.; Zhang, L.; Siddiqui, N. A.; Naguib, N.; Chen, F.; Padmanabhan, R.; Zhang, T.; Monette, S.; Socciarelli, F.; Lee, R. An ultrasmall core-shell silica nanoparticle improves antitumour immunity and survival by remodelling suppressive melanoma microenvironments. Nat. Nanotechnol. 2026, 21 (2), 311– 322, DOI: 10.1038/s41565-025-02083-zThere is no corresponding record for this reference.
- 8Chen, F.; Ma, K.; Madajewski, B.; Zhuang, L.; Zhang, L.; Rickert, K.; Marelli, M.; Yoo, B.; Turker, M. Z.; Overholtzer, M. Ultrasmall targeted nanoparticles with engineered antibody fragments for imaging detection of HER2-overexpressing breast cancer. Nat. Commun. 2018, 9 (1), 4141, DOI: 10.1038/s41467-018-06271-5There is no corresponding record for this reference.
- 9Szymanski, N. J.; Rendy, B.; Fei, Y.; Kumar, R. E.; He, T.; Milsted, D.; McDermott, M. J.; Gallant, M.; Cubuk, E. D.; Merchant, A. An autonomous laboratory for the accelerated synthesis of novel materials. Nature 2023, 624 (7990), 86– 91, DOI: 10.1038/s41586-023-06734-wThere is no corresponding record for this reference.
- 10Velapure, P.; Kansal, D.; Bobade, C. Tumor microenvironment-responsive nanoformulations for breast cancer. Discov Nano 2024, 19 (1), 212, DOI: 10.1186/s11671-024-04122-5There is no corresponding record for this reference.
- 11Wang, J.; Tao, X.; Zhu, J.; Dai, Z.; Du, Y.; Xie, Y.; Chu, X.; Fu, G.; Lei, Z. Tumor organoid-immune co-culture models: exploring a new perspective of tumor immunity. Cell Death Discov 2025, 11 (1), 195, DOI: 10.1038/s41420-025-02407-xThere is no corresponding record for this reference.
- 12Helmlinger, G.; Sokolov, V.; Peskov, K.; Hallow, K. M.; Kosinsky, Y.; Voronova, V.; Chu, L.; Yakovleva, T.; Azarov, I.; Kaschek, D. Quantitative Systems Pharmacology: An Exemplar Model-Building Workflow With Applications in Cardiovascular, Metabolic, and Oncology Drug Development. CPT Pharmacometrics Syst. Pharmacol 2019, 8 (6), 380– 395, DOI: 10.1002/psp4.12426There is no corresponding record for this reference.
- 13Canchola, A.; Li, K.; Chen, K.; Borboa-Pimentel, A.; Chou, C.; Dela Rama, R.; Chen, C. Y.; Chen, X.; Strobel, M.; Riviere, J. E. Meta-Analysis and Machine Learning Prediction of Protein Corona Composition across Nanoparticle Systems in Biological Media. ACS Nano 2025, 19, 37633, DOI: 10.1021/acsnano.5c08608There is no corresponding record for this reference.
- 14Wang, H.; Arulraj, T.; Ippolito, A.; Popel, A. S. From virtual patients to digital twins in immuno-oncology: lessons learned from mechanistic quantitative systems pharmacology modeling. NPJ. Digit Med. 2024, 7 (1), 189, DOI: 10.1038/s41746-024-01188-4There is no corresponding record for this reference.
- 15Nobel Prize in Chemistry 2024 , 2024. https://www.nobelprize.org/uploads/2024/10/press-chemistryprize2024-3.pdf.There is no corresponding record for this reference.
- 16Avsec, Ž.; Latysheva, N.; Cheng, J.; Novati, G.; Taylor, K. R.; Ward, T.; Bycroft, C.; Nicolaisen, L.; Arvaniti, E.; Pan, J. Advancing regulatory variant effect prediction with AlphaGenome. Nature 2026, 649 (8099), 1206– 1218, DOI: 10.1038/s41586-025-10014-0There is no corresponding record for this reference.
- 17Zambaldi, V.; La, D.; Chu, A. E.; Patani, H.; Danson, A. E.; Kwan, T. O.; Frerix, T.; Schneider, R. G.; Saxton, D.; Thillaisundaram, A. De novo design of high-affinity protein binders with AlphaProteo. arXiv Preprint , arXiv:2409.08022, 2024.There is no corresponding record for this reference.
- 18Zeni, C.; Pinsler, R.; Zügner, D.; Fowler, A.; Horton, M.; Fu, X.; Wang, Z.; Shysheya, A.; Crabbé, J.; Ueda, S. A generative model for inorganic materials design. Nature 2025, 639 (8055), 624– 632, DOI: 10.1038/s41586-025-08628-5There is no corresponding record for this reference.
- 19Kp Jayatunga, M.; Ayers, M.; Bruens, L.; Jayanth, D.; Meier, C. How successful are AI-discovered drugs in clinical trials? A first analysis and emerging lessons. Drug Discov Today 2024, 29 (6), 104009, DOI: 10.1016/j.drudis.2024.104009There is no corresponding record for this reference.
- 20Lee, K. Y.; Lee, G. Y.; Lane, L. A.; Li, B.; Wang, J.; Lu, Q.; Wang, Y.; Nie, S. Functionalized, Long-Circulating, and Ultrasmall Gold Nanocarriers for Overcoming the Barriers of Low Nanoparticle Delivery Efficiency and Poor Tumor Penetration. Bioconjug Chem. 2017, 28 (1), 244– 252, DOI: 10.1021/acs.bioconjchem.6b00224There is no corresponding record for this reference.
- 21Broske, B.; Binder, S. C.; McEnroe, B. A.; Kempchen, T. N.; Fandrey, C. I.; Messmer, J. M.; Tan, E.; Konopka, P.; Ferber, D.; Yong, M. C. R. Development of AI-designed protein binders for detection and targeting of cancer cell surface proteins. bioRxiv Preprint , 2025. DOI: 10.1101/2025.05.11.652819 .There is no corresponding record for this reference.
- 22Chen, Z.; Kibler, R. D.; Hunt, A.; Busch, F.; Pearl, J.; Jia, M.; VanAernum, Z. L.; Wicky, B. I. M.; Dods, G.; Liao, H. De novo design of protein logic gates. Science 2020, 368 (6486), 78– 84, DOI: 10.1126/science.aay2790There is no corresponding record for this reference.
- 23Johnson, A.; Shen, Y.; Zheng, X.; Su, F.; Zeng, J.; Uzunparmak, B.; Breuer, E.-K.; Holla, V.; Kahle, M.; Kim, S.-H. The actionable transcriptome: a framework for incorporating RNA sequencing into precision oncology. Nature Reviews Clinical Oncology 2026, 23, 213, DOI: 10.1038/s41571-025-01110-1There is no corresponding record for this reference.
- 24Bennett, N. R.; Coventry, B.; Goreshnik, I.; Huang, B.; Allen, A.; Vafeados, D.; Peng, Y. P.; Dauparas, J.; Baek, M.; Stewart, L. Improving de novo protein binder design with deep learning. Nat. Commun. 2023, 14 (1), 2625, DOI: 10.1038/s41467-023-38328-5There is no corresponding record for this reference.
- 25Valencia, P. M.; Pridgen, E. M.; Rhee, M.; Langer, R.; Farokhzad, O. C.; Karnik, R. Microfluidic platform for combinatorial synthesis and optimization of targeted nanoparticles for cancer therapy. ACS Nano 2013, 7 (12), 10671– 10680, DOI: 10.1021/nn403370eThere is no corresponding record for this reference.
- 26Corbo, C.; Molinaro, R.; Tabatabaei, M.; Farokhzad, O. C.; Mahmoudi, M. Personalized protein corona on nanoparticles and its clinical implications. Biomater Sci. 2017, 5 (3), 378– 387, DOI: 10.1039/C6BM00921BThere is no corresponding record for this reference.
- 27Praetorius, F.; Leung, P. J. Y.; Tessmer, M. H.; Broerman, A.; Demakis, C.; Dishman, A. F.; Pillai, A.; Idris, A.; Juergens, D.; Dauparas, J. Design of stimulus-responsive two-state hinge proteins. Science 2023, 381 (6659), 754– 760, DOI: 10.1126/science.adg7731There is no corresponding record for this reference.
- 28Hsia, Y.; Bale, J. B.; Gonen, S.; Shi, D.; Sheffler, W.; Fong, K. K.; Nattermann, U.; Xu, C.; Huang, P. S.; Ravichandran, R. Design of a hyperstable 60-subunit protein dodecahedron. [corrected]. Nature 2016, 535 (7610), 136– 139, DOI: 10.1038/nature18010There is no corresponding record for this reference.
- 29Ma, T.; Tran, T. B.; Lin, E.; Hunt, S.; Haveman, R.; Castro, K.; Lu, J. Size-transformable nanotherapeutics for cancer therapy. Acta Pharm. Sin B 2025, 15 (2), 834– 851, DOI: 10.1016/j.apsb.2024.11.012There is no corresponding record for this reference.
- 30Joyce, P.; Allen, C. J.; Alonso, M. J.; Ashford, M.; Bradbury, M. S.; Germain, M.; Kavallaris, M.; Langer, R.; Lammers, T.; Peracchia, M. T. A translational framework to DELIVER nanomedicines to the clinic. Nat. Nanotechnol 2024, 19 (11), 1597– 1611, DOI: 10.1038/s41565-024-01754-7There is no corresponding record for this reference.
- 31Considerations for the Use of Artificial Intelligence To Support Regulatory Decision-Making for Drug and Biological Products , 2025. https://www.fda.gov/regulatory-information/search-fda-guidance-documents/considerations-use-artificial-intelligence-support-regulatory-decision-making-drug-and-biological.There is no corresponding record for this reference.
- 32Reflection paper on the use of Artificial Intelligence (AI) in the medicinal product lifecycle , 2024. https://www.ema.europa.eu/en/documents/scientific-guideline/reflection-paper-use-artificial-intelligence-ai-medicinal-product-lifecycle_en.pdf.There is no corresponding record for this reference.
- 33Sharma, V.; Mottafegh, A.; Joo, J. U.; Kang, J. H.; Wang, L.; Kim, D. P. Toward microfluidic continuous-flow and intelligent downstream processing of biopharmaceuticals. Lab Chip 2024, 24 (11), 2861– 2882, DOI: 10.1039/D3LC01097JThere is no corresponding record for this reference.
- 34Carter, P. J.; Quarmby, V. Immunogenicity risk assessment and mitigation for engineered antibody and protein therapeutics. Nat. Rev. Drug Discov 2024, 23 (12), 898– 913, DOI: 10.1038/s41573-024-01051-xThere is no corresponding record for this reference.



